Abstract
Edwardsiella tarda is a fish pathogen that causes systemic infections in fresh water and marine fish. Determining the antigenic proteins is important for the development of an immunodiagnostic tests and a vaccine for effective infection control in fish. In the current study, antigens were detected by immunoblotting and affinity column chromatography using a Japanese flounder (Paralichthys olivaceus) antibody produced by experimental infection with E. tarda. GroEL, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), outer membrane protein A, filament protein, 30S ribosomal protein S6, 50S ribosomal protein L9, cold shock protein, and carbon storage protein were identified as antigens of E. tarda through biochemical analyses of the molecular weights, isoelectric points, and N-terminal amino-acid sequences. These proteins can be easily detected in flounder infected with E. tarda and are potential diagnostic markers.
Edwardsiella tarda is a Gram-negative bacterium of the family Enterobacteriaceae, 30 which is known as a causative agent of edwardsiellosis in fresh water and marine fish. 1,2,22,28 In Japan, even though the Japanese flounder (Paralichthys olivaceus) farming industry has suffered serious damage from the disease, 12 effective infection control has not yet been established. Therefore, information on the virulence, antigenicity, and antibiotic susceptibility is needed for prevention, rapid diagnosis, and therapy of E. tarda infection.
Pathogenesis of E. tarda is multifactorial. Many virulence genes of E. tarda have been identified for clarification of the virulence mechanisms for production of hemolysins, 4,10,31 intracellular replication in macrophage 34,40 and the ability to invade epithelial cells. 11 Recently, in addition to the virulence genes, genes encoding antigenic proteins have been explored for development of a vaccine and immunodiagnosis. Although several genes of E. tarda encoding antigenic proteins have been identified from genomic DNA using artificially prepared immune serum from Japanese flounder, 37 E. tarda antigenic proteins recognized by antibodies of infected fish have not been identified.
In the current study, antigenic proteins of E. tarda were isolated from the cell extracts by immunoblotting and affinity column chromatography under mild conditions using immunoglobulin M (IgM) antibody of Japanese flounder. Furthermore, protein identification was then performed by N-terminal amino-acid sequencing and sequence homology search.
Edwardsiella tarda strain NUF806 was kindly donated by Dr. K. Kanai (Nagasaki University). The E. tarda strain, which was originally isolated in 1997 from Japanese flounder in Nagasaki Prefecture, Japan, has been reported to have high virulence against Japanese flounder using an experimental challenge test. 21 The strain was grown at 25°C for 24 hr on Luria broth agar. a
Two groups (n = 10 each) of Japanese flounder weighing approximately 700 g were intraperitoneally injected with 5.2 × 104 CFU/fish of bacteria and phosphate buffered saline (PBS), respectively. The challenged and control groups were kept in flow-through 500 L-seawater tanks at 23 ± 1°C for 30 days. The serum was sampled from the surviving fish, filtered through a 0.45-μm pore size membrane filter, and stored at −80°C. Then, agglutination antibody titer was measured using microtiter plates against formalin-killed cells (FKC) of the strain NUF 806 according to a previously described method. 6
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was performed according to a previously described method. 15 Two-dimensional electrophoresis (2-DE) was performed as follows. A protein sample for 2-DE was treated with a 2-DE clean-up kit b according to the manufacturer's protocols and solubilized in 7 M urea, 2 M thiourea, 4% 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate (CHAPS), 2 mM tributylphosphine, and 0.2% commercial ampholyte. b The first-dimensional isoelectric focusing was carried out using the 7-cm immobilized pH gradient gel strips (pH range: 3–10 nonliner) b on a commercial isoelectric focusing cell b according to the manufacturer's protocols. The sample was focused at 250 V for 15 min, 4,000 V for 1 hr, and then 4,000 V for a total of 10 kV/hr. The strip was equilibrated b according to the manufacturer's protocols and then subjected to SDS-PAGE. Molecular weight (MW) and isoelectric point (pI) of protein in the gel were examined using a commercial software analyzer. c
Flounder IgM purified from a pool of blood samples of 5 fish previously described 23 was intraperitoneally injected (0.2 mg) into 4-week-old female BALB/c mice on days 0, 21, and 42. Hybridoma cells secreting monoclonal antibodies (mAbs) against the flounder IgM were generated from the spleen cells using the myeloma cell line Sp2/O-Ag14 d according to previously described method. 16 Reaction of mAbs against the flounder IgM was confirmed by immunoblotting and enzyme-linked immunosorbent assay (ELISA). The class and subclass of mAbs were determined using a commercial mAb isotyping kit. e The mAb class IgG was purified from the hybridoma supernatant using a commercial mAb trap kit e according to the manufacturer's protocol.
Edwardsiella tarda antigens were separated from the cell extracts by SDS-PAGE and detected by immunoblotting with mAb. Wet cells (25 μg) of E. tarda NUF806 were lysed with SDS-PAGE sample buffer followed by boiling at 100°C for 3 min. Each sample was subjected to SDS-PAGE and 2-DE, and electroblotted onto a polyvinylidene fluoride (PVDF) membrane b as previously described. 36 After blocking nonspecific binding sites with 3% gelatin, b the membrane was incubated for 1 hr in a 1/50 dilution of flounder antiserum in PBS. Flounder IgM binding to antigens on the membrane was detected with mAbs and goat anti-mouse IgG antibody conjugated with alkaline phosphatase (AP). b The antibody reaction was visualized with an AP conjugate substrate kit. b
Flounder IgM-affinity column was prepared as follows. MAb against flounder IgM was bound to a protein G spin column e according to the manufacturer's protocol. Then, the flounder antiserum (500 μl) was applied to the column, incubated for 60 min at room temperature, and washed 3 times with Tris-buffered saline (TBS). b The flounder IgM, mAb, and protein G were chemically cross-linked to each other with 50 mM dimethyl pimelimidate dihydrochloride f in 200 mM triethanolamine g (pH 8.9) according to the manufacturer's protocol and attached to the protein G spin column. Following the cross-linking reaction, the column was washed 3 times with TBS. Cells of E. tarda NUF806 were mildly lysed using 0.8% n-octyl-β-D-thioglucoside h (NOTG) as previously described. 29 NOTG extracts (400 μl) were applied to the flounder IgM-affinity column equilibrated with TBS containing 0.8% NOTG (NTBS). The column was incubated for 1 hr at room temperature and washed 5 times with NTBS. Edwardsiella tarda antigens captured by flounder IgM were eluted with 0.1 M glycine-HCl buffer (pH 2.6) containing 0.8% NOTG. Each eluted protein was lysed with a 2-DE clean-up kit b according to the manufacturer's protocol and purified by 2-DE.
The antigens purified by SDS-PAGE or 2-DE were electroblotted onto a PVDF membrane b as previously described. 9 The protein bands were cut out of the membrane and sequenced using a gas-phase protein sequencer. i The N-terminal amino-acid sequence was used as a query sequence for the Basic Local Alignment Search Tool (BLAST; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) using the BLASTP program.
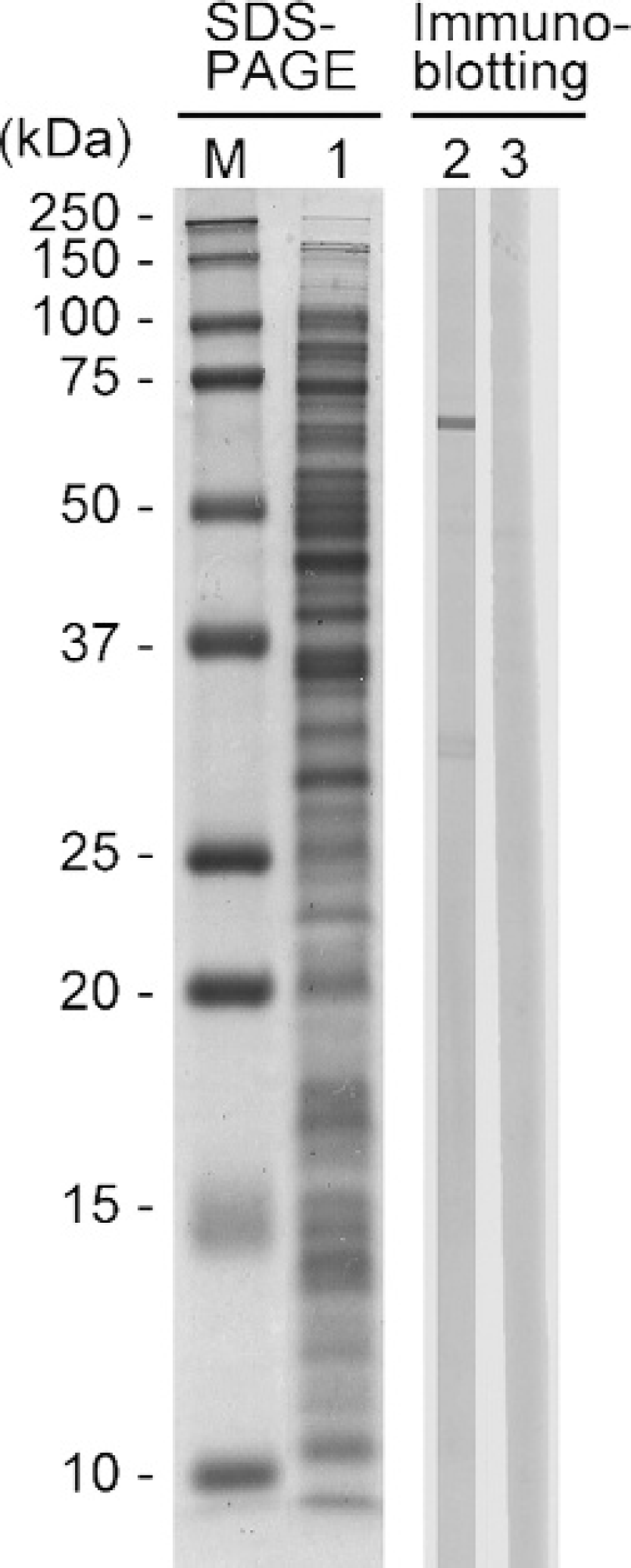
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and immunoblotting of SDS extracts of Edwardsiella tarda NUF806. Lane M: molecular weight marker; lane 1: coomassie brilliant blue stain; lane 2: serum of flounder infected with E. tarda; lane 3: serum of flounder injected with phosphate buffered saline.
Six fish survived within 30 days of experimental infection with E. tarda NUF806. The bacteria were re-isolated from all surviving and dead fish. No fish died in the control group following injection with PBS. From the serum sampled from all surviving fish, the agglutination titer of antiserum were 128–256 for FKC.
The 64.4 kDa band, designated as antigen A, was detected in SDS extracts by immunoblotting (Fig. 1). No band was observed in the immunoblots for the serum of noninfected flounder. The antigen was focused at the position of pH 4.7/64.4 kDa in 2-DE (data not shown).
Many protein bands in the NOTG extracts, which were similar to those in the SDS extracts, were observed by SDS-PAGE (Fig. 2A). In the purification of Edwardsiella tarda antigen from the NOTG extracts using flounder IgM-affinity column, 4 major proteins bands with molecular weight of approximately 64, 40, 35, and 15 kDa were detected in the elution by SDS-PAGE. No protein bands containing the flounder IgM and mAbs were found in the final wash of the IgM nonbinding extracts (data not shown). Approximately 3 μg of antigens were concentrated from the elution fraction and separated by 2-DE (Fig. 3), resolving 14 proteins (spots designated B–O) as the major antigenic proteins.
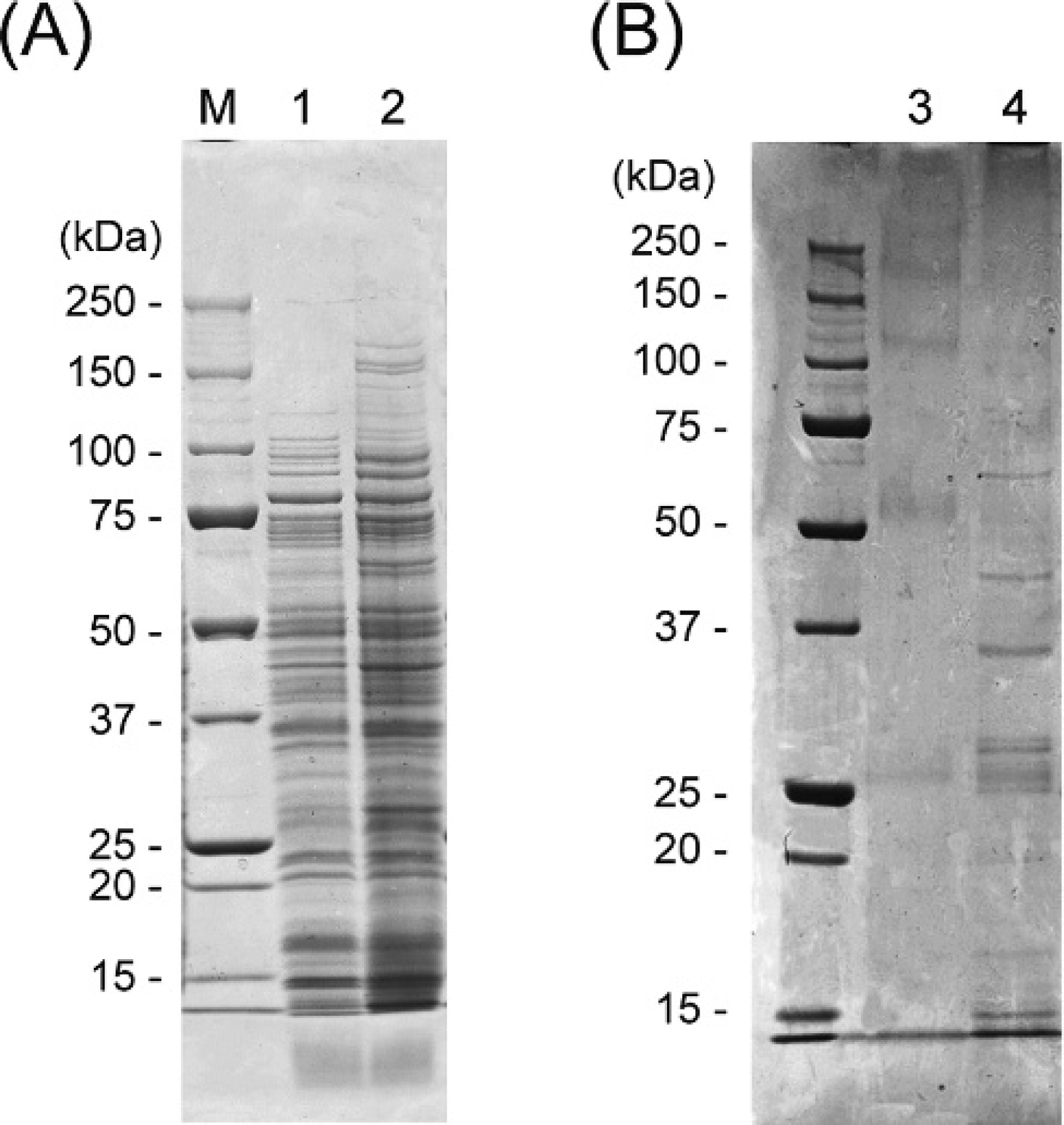
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) of A, Edwardsiella tarda extracts, and B, elution fractions from the flounder immunoglobulin M–affinity column. Lane M: molecular weight markers; lane 1: SDS extracts; lane 2: n-octyl-β-D-thioglucoside (NOTG) extracts; lane 3: elution fraction of phosphate buffered saline (negative control); lane 4: elution fraction of NOTG extracts.
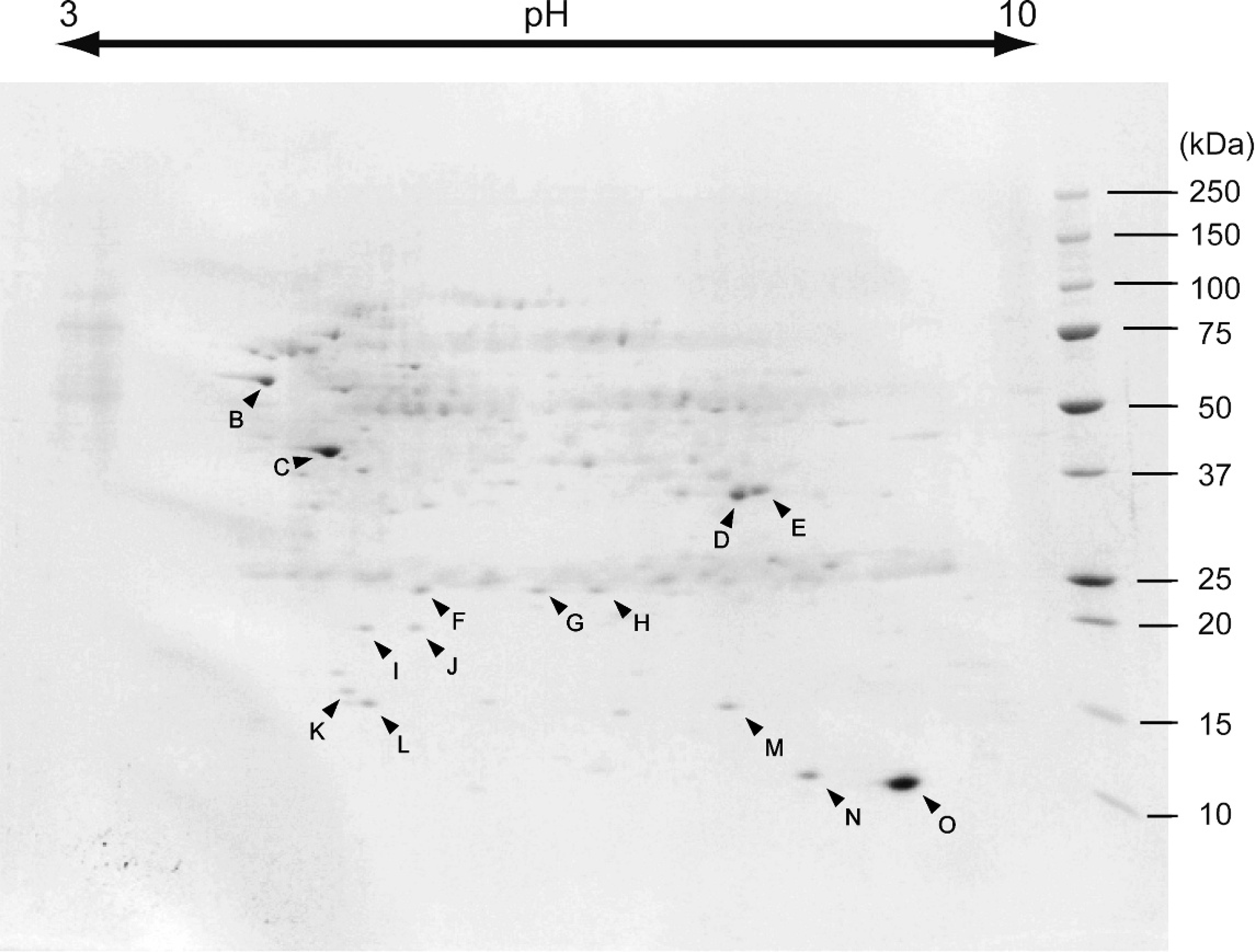
Two-dimensional electrophoresis of Edwardsiella tarda n-octyl-β-D-thioglucoside (NOTG) extract elution fraction from a flounder immunoglobulin M–affinity column.
The 15 antigens, A–O, obtained by immunoblotting or the flounder IgM-affinity column chromatography were subjected to N-terminal amino-acid sequencing (Table 1). The amino-acid sequences of A, B, D, E, and K–O were determined; the sequences of F–J could not be determined as the antigen quantities were below the detection limit or were N-terminally blocked. As the amino-acid sequence, calculated MW, and pI of antigen A matched antigen B, these antigens were determined to be the same. The N-terminal amino-acid sequences of antigens recognized by Japanese flounder IgM were identical to those of enterobacterial proteins, and the predicted MWs and pIs were similar to homologous proteins as listed in Table 1. Therefore, it was determined that the antigens A (B), D, E, and K–O were GroEL, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), outer membrane protein A, filament protein, 30S ribosomal protein S6, 50S ribosomal protein L9, cold shock protein (cspC), and carbon storage regulator (csrA), respectively. Flagellin has been detected as a major antigen of E. tarda using rabbit polyclonal antiserum 37 and observed around 40 kDa at pH 5, which are values similar to the predicted MW and pI (Table 1) on 2-DE in fish pathogenic to E. tarda. 33 Therefore, it was deduced that antigen C was flagellin. The antigen C separated by SDS-PAGE was not detected in immunoblotting analysis. A previous study 37 reported that flagellin separated by SDS-PAGE was not recognized by flounder antibody in immunoblotting analysis. As in antigen C, antigens, except A and B, separated by SDS-PAGE were not detected in the immunoblotting analysis. Conservation of conformational epitope would be important for the recognition of flounder antibody against antigen.
N-terminal amino-acid sequence, molecular weight (MW), and isoelectric point (pI) of Edwardsiella tarda antigens and their homologs.*
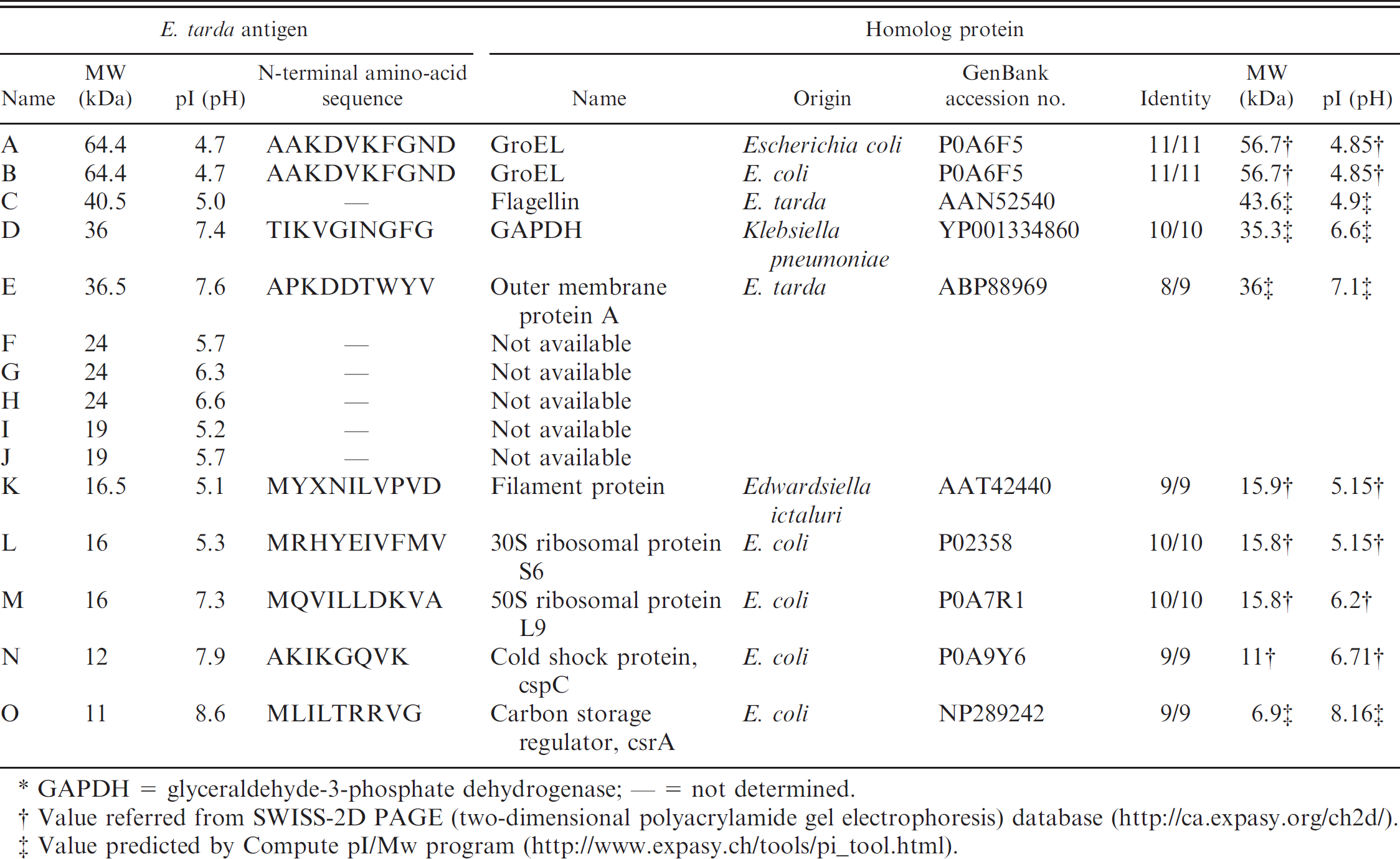
GAPDH = glyceraldehyde-3-phosphate dehydrogenase; — = not determined.
Value referred from SWISS-2D PAGE (two-dimensional polyacrylamide gel electrophoresis) database (http://ca.expasy.org/ch2d/).
Value predicted by Compute pI/Mw program (http://www.expasy.ch/tools/pi_tool.html).
GroEL is known as a 60-kDa heat shock protein (HSP) with a role as a molecular chaperone. 3 The protein is upregulated during infection by most bacteria and is recognized as an immunodominant antigen by the host immune system. 41 With a highly conserved amino-acid sequence among bacteria, 38 GroELs of Flavobacterium psychrophilum and Piscirickettsia salmonis, which are known as fish pathogens, have been recognized by the antiserum of rainbow trout and Atlantic salmon. 32,39 In the current study, the antigen identified as GroEL was clearly detected in immunoblotting analysis and purification with the flounder IgM affinity column. GroEL proteins of fish pathogen will be one of the major antigens recognized by the fish immune system.
Following the first detection of GAPDH of E. tarda by a previous study, 13 a 2005 study 17 reported that the recombinant GAPDH showed vaccine efficacy against E. tarda in experimental infection with Japanese flounder. In the present study, E. tarda GAPDH was recognized by flounder IgM by affinity column chromatography. However, the protein denatured by SDS-PAGE sample buffer was not detected by immunoblotting, suggesting that the native conformation is necessary for antigenicity against fish.
OmpA is a major component of the outer membrane of Escherichia coli, which is widely conserved among Gramnegative bacteria and has been reported to be a contributor to pathogenicity. A previous study 27 reported that the ability to invade brain microvascular endothelial cells was decreased in the E. coli K-1 OmpA gene-deletion mutant. OmpA homologs have been identified in fish pathogens, including A. salmonicida, 7,19 F. psychrophilum, 8 and Yersinia ruckeri. 5 Although the OmpA gene of E. tarda has been registered in GenBank (accession no. ABP88969), the function of OmpA in E. tarda is not well understood.
CsrA belongs to a new class of RNA-binding proteins contributing to translation of target genes or to the stability of target mRNA transcripts. 18 The Csr posttranscriptional system involves switches that control a variety of virulence-linked traits, such as biofilm formation, motility, intracellular replication, antibiotic/stress resistance, and virulence factor production in many pathogenic bacteria. Antibody against E. tarda csrA was detected in serum of the affected flounder. The antibody will assume a role in the prevention of activity of the csrA.
CspC is a homolog of cspA. 26 The cspA family in E. coli consists of 9 homologs, cspA–cspI. However, only cspA, cspB, cspG, and cspI are cold shock inducible. A previous study 25 suggests that cspC is the important element involved in the regulation of a global stress response regulator and is a protein that responds to numerous stresses. Antibody against E. tarda cspC was detected in serum of the affected flounder. Edwardsiella tarda is subjected to stress from the host defense system.
The 30S and 50S ribosomal subunits have 21 proteins and 32 proteins, respectively, 14 including 30S ribosomal protein S6 and 50S ribosomal protein L9. S6 and L9 have been observed on the surface of each ribosomal subunit by immune electron microscopy with an antibody. 35 It was deduced in the current study that S6 protein and L9 protein are antigens easily recognized by the flounder antibody among constitutive proteins of each ribosomal subunit.
Amino-acid sequence of the filament protein of E. ictaluri has been registered in GenBank. However, the function has not yet been elucidated. The amino-acid sequence shares the single conserved domain of universal stress protein Usp (NCBI Conserved Domain Database no. cd00293). The Usp is a small cytoplasmic bacterial protein. The expression is upregulated when the bacterial cell is exposed to stress agents, and Usp enhances the rate of cell survival. 24 It was deduced in the current study that the filament protein of E. ictaluri and antigen K of E. tarda can play such a role in fish.
Development of vaccine for E. tarda infection has never been fully realized. Flounder infected with E. tarda suffer from the disease for prolonged periods and discharge many cells of E. tarda outside of the body. Removal of infected fish and carriers is important for preventing epidemic E. tarda. A previous study 20 reported that flounder discharging E. tarda cells show high antibody titer. Therefore, the detection of antibodies could help identify infected fish and carriers, and the amino-acid sequences of antigens identified in the present study could be useful for developing an immunodiagnostic test and an effective vaccine against E. tarda.
Acknowledgements. This research was supported in part by a grant for research utilizing advanced technologies in agriculture, forestry, and fisheries from the Ministry of Agriculture, Forestry and Fisheries of Japan.
Footnotes
a.
Difco™ LB Agar, Lennox, Becton Dickinson, Tokyo, Japan.
b.
ReadyPrep™ 2-D Cleanup Kit, Bio-Lyte 3/10 Ampholyte, ReadyStrip™ IPG Strip, PROTEAN® IEF cell, ReadyPrep™ Equilibration Buffer I and ReadyPrep™ Equilibration Buffer II; Bio-Rad Laboratories, Tokyo, Japan.
c.
Lane & Spot Analyzer, ATTO Corp., Tokyo, Japan.
d.
Japan Health Sciences Foundation, Tokyo, Japan.
e.
Roche IsoStrip™ Mouse Monoclonal Antibody Isotyping Kit, Protein G HP SpinTrap™ column; GE Healthcare Bioscience, Tokyo, Japan.
f.
Pierce Biotechnology Inc., Rockford, IL.
g.
Wako Pure Chemical Industries Ltd., Osaka, Japan.
h.
Dojindo Laboratories, Kumamoto, Japan.
i.
PPSQ-33A Automated Protein/Peptide Sequencer, Shimadzu Corp., Kyoto, Japan.
