Abstract
Rapid and accurate diagnosis is essential for effective control of foot-and-mouth disease (FMD). The present report describes the practical steps undertaken to deploy a real-time reverse transcription polymerase chain reaction (real-time RT-PCR) to process the samples received during the outbreaks of FMD in the United Kingdom in 2007. Two independent real-time RT-PCR assays targeting different regions (5′UTR and 3D) of the FMD virus (FMDV) genome were used to confirm the presence of FMDV in clinical samples collected from the first infected premises. Once the FMDV strain responsible had been sequenced, a single real-time RT-PCR assay (3D) was selected to test a total of 3,216 samples, including material from all 8 infected premises. Using a 96-well automated system to prepare nucleic acid template, up to 84 samples could be processed within 5 hr of submission, and up to 269 samples were tested per working day. A conservative cut-off was used to designate positive samples, giving rise to an assay specificity of 99.9% or 100% for negative control material or samples collected from negative premises, respectively. For the first time, real-time RT-PCR results were used to recognize preclinical FMD in a cattle herd. Furthermore, during the later stages of the outbreaks, the real-time RT-PCR assay supported an active surveillance program within high-risk cattle herds. To the authors' knowledge, this is the first documented use of real-time RT-PCR as a principal laboratory diagnostic tool following introduction of FMD into a country that was FMD-free (without vaccination) and highlights the advantages of this assay to support control decisions during disease outbreaks.
Introduction
Six years from the previous epidemic of foot-and-mouth disease (FMD) in the United Kingdom, the disease reoccurred during August and September 2007. These FMD outbreaks were in 2 distinct phases affecting 8 separate premises, situated in the west of the county of Surrey, the north of Surrey, and neighboring region of Berkshire in southeast England. 4,14 Rapid and accurate diagnosis is essential for effective control and eradication of FMD, beginning with clinical investigation in the field by veterinary staff of animals suspected of being infected with FMD and followed by rapid confirmation of the presence of FMD virus (FMDV; family Picornaviridae, genus Aphthovirus), antigen, or genome in clinical samples using laboratory methods. In contrast to the 2001 epidemic, diagnostic support for the 2007 outbreaks was characterized by the use of real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assays as a principal tool for decision making.
A number of pan-serotype-specific real-time RT-PCR assays have been developed to target highly conserved regions of the RNA genome of FMDV. 3,10,11 During the past 6 years, these assays have been improved and evaluated for a variety of clinical sample types that might be received for diagnosis. 13,15 In particular, the development and use of automated protocols to prepare nucleic acid template has increased assay throughput and reliability. 12 Parallel testing of samples has shown the sensitivity of these automated real-time RT-PCR assays to be at least equal to that of in vitro virus isolation in cell culture (VI). 9,17 More recently, one-step real-time RT-PCR protocols have further simplified and accelerated the assay for routine use. 16 Together, these improvements have provided data to increase confidence in the use of robust real-time RT-PCR assays for routine diagnostics, justifying their inclusion in laboratory contingency plans for outbreak diagnostics.
The current report describes the role played by automated real-time RT-PCR in laboratory diagnosis of FMD during the U.K. 2007 outbreaks. Clinical samples submitted from the infected farms (and from other report cases representing premises harboring suspect cases of disease) as well as sera collected from high-risk herds were tested using real-time RT-PCR. These and other laboratory data were used for frontline decision making and influenced the actions taken to control and eradicate the outbreaks. To the authors' knowledge, this is the first published use of laboratory-based real-time RT-PCR as a principal diagnostic tool in the face of an FMD introduction into an FMD-free (without vaccination) country.
Materials and methods
Sample collection
During the outbreaks in 2007, a total of 3,246 diagnostic samples were submitted for virologic investigation comprising 50 vesicular epithelia, 1 additional tissue (from a negative herd, sample type not designated), 3,147 blood samples (32 in ethylenediamine tetra-acetic acid [EDTA] and 3,115 clotted serum samples), 2 vesicular fluids, 3 additional fluids (from a negative herd, sample types not designated), 36 esophageal-pharyngeal scrapings (“probangs”), 6 swabs, and 1 fecal sample. Collection of these samples in the field is described elsewhere. 14 Blood samples were collected from suspect cases: initially these were separate EDTA and serum tubes for testing by virologic and serologic assays, respectively, but later in the outbreaks, instructions to veterinarians were modified in accordance with the automated protocols used for nucleic acid extraction (see “Nucleic acid extraction” section) to request duplicate serum samples in plain vacutainer tubes (clotted blood). These samples were collected from a total of 38 separate premises comprising the 8 infected premises (IPs), and an additional 30 farms (report cases) that were subsequently negated based upon laboratory results. Several of the IPs comprised geographically discrete holdings, and these were designated as IP1a, IP1b, etc. During the second phase of outbreaks, clotted bloods (n = 1,861) were collected for active surveillance from 11 farms (14 holdings) containing 490 cattle located within the intensively patrolled area (IPA) in Surrey and 2 neighboring counties. In order to maximize the potential to recognize early infection (or preclinical animals incubating disease), it was agreed to sample and test these high-risk cattle on alternate days (from September 24 through October 16, 2007).
Sample preparation
A 10% (wt/vol) suspension was prepared in 0.04 M phosphate buffer from each vesicular epithelium, other tissue, or from the fecal sample. When testing these suspensions or swab solutions, a 1.4 ml aliquot was taken for the antigen-detection enzyme-linked immunosorbent assay (Ag-ELISA), 6 and further aliquots were used to inoculate cell cultures of bovine thyroid primary cell cultures or IB-RS-2 cells (a permanent line of porcine kidney cells) for VI. 5,6,18 At the same time, epithelial suspensions (ES) and swab samples were prepared for real-time RT-PCR analysis by adding 0.6 ml of the homogenate to 0.9 ml of Lysis/Binding Buffer (supplied with the MagNA Pure LC Total Nucleic Acid Isolation Kit a ). In contrast, blood and probang samples were directly subjected to nucleic acid extraction (see “Nucleic acid extraction” section). Serum samples were aspirated after spinning clotted blood for 10 min at 2,000 × g in a Rotanta 460 R b or Allegra™ 6R c centrifuge.
Nucleic acid extraction
Two separate automated systems were used to generate nucleic acid template from samples for real-time RT-PCR during the outbreaks. These were the BioRobot Universal System, d which can process 96 wells in 120 min and the MagNA Pure LC, a which can generate template from 32 wells in 90 min.
A series of experiments were performed to evaluate the performance of the BioRobot Universal System and to select the most appropriate automated protocol to process samples on this instrument. This work was initially undertaken during the spring and early summer of 2007 in anticipation of a potential bluetongue virus epidemic in the United Kingdom. In terms of sample lysis, the AL buffer supplied with the QIAamp® All Nucleic Acid MDx kit d was not effective with cattle EDTA-blood samples. Improved lysis was achieved by substituting Lysis/Binding Buffer, so this was used for all subsequent testing of material from the outbreaks. Despite this change, intermittent problems with clogging of the filter membranes remained, which did not occur when serum samples were tested in parallel. In light of the demand that was to follow for testing of blood by real-time RT-PCR (active surveillance and preclinical cases), instructions to veterinary field staff were changed to request the collection of 2 clotted blood tubes per animal instead of 1 serum and 1 EDTA-blood per animal.
A strict regime was adopted to process the samples submitted from the outbreaks in order to minimize inter-and intra-assay contamination. For the BioRobot Universal System, 360 μl of Lysis/Binding Buffer was added to each well in an S-Block in a class-II microbiological safety cabinet (located in a room dedicated for RT-PCR sample processing). Where necessary, spaces were left for the addition of 600 μl of prelysed tissue suspension, ES, or swab samples (see “Sample preparation” section). Serum, EDTA-blood, or probang was then carefully added (240 μl) to the appropriate well and then the same volumes of un-infected adult bovine serum mycoplasma (virus screened) e were added as negative control wells. Prepared positive controls (see “Addition of real-time RT-PCR positive and negative controls” section) were also added (600 μl). The total volume of sample (or control material) and buffer in each well was therefore 600 μl. After sample addition, 40 μl of reconstituted Protease d was pipetted into each well and the S-Block securely covered with plastic film. f The S-block was then placed inside the BioRobot Universal System to mix the samples by shaking for 1 min at 600 rpm followed by heating at 56°C for 15 min in a water bath to ensure complete lysis of the samples. Nucleic acid extraction was performed by a customized automated protocol using the reagents from a QIAamp All Nucleic Acid MDx kit. d Total nucleic acids were eluted to a final volume of 50 μl.
To maintain a high daily sample throughput and flexibility in the diagnostic activities, total nucleic acid extraction was also carried out in a MagNA Pure LC as described previously. 16 Samples (and controls) were added to the 32-well cartridge (MagNA Pure LC Sample Cartridge a ) in a dedicated class-II microbiological safety cabinet in a similar manner to the method described for the BioRobot Universal System (see previous paragraph).
Real-time RT-PCR
Two independent real-time RT-PCR assays targeting different regions (5′UTR11 and 3D3) of the FMDV genome are routinely used to test sample material submitted to the World Reference Laboratory for FMD (WRLFMD). The performance of these assays has previously been compared with each other, 2,9 and with the results from conventional diagnostic tests for FMD (VI and Ag-ELISA) 17 demonstrating that these RT-PCR assays, when used together, have a diagnostic sensitivity approaching 100% for diverse field strains of FMDV. 9 This highly sensitive system has also been integrated into the FMD laboratory contingency plan for suspect cases of vesicular disease in the United Kingdom, and therefore both real-time RT-PCR assays were used to test diagnostic material from the first infected premises (IP1a-c). However, performing parallel assays for 2 targets during an outbreak significantly reduced the number of samples that could be processed. Therefore, when sequence data were available for the causative virus strain (within 2 days of the laboratory confirmation of the index case), 4 it was decided to reduce the scope of the real-time RT-PCR to the 3D target alone.
Real-time RT-PCR reactions were prepared manually or by an automated procedure governed by the choice of robot (BioRobot Universal or MagNA Pure LC) for nucleic acid extraction. RT-PCR set up from nucleic acid extracted in the BioRobot Universal was carried out manually (using a multichannel pipette) by adding 5 μl of the template to 20 μl of real-time RT-PCR reaction mix containing 12.5 μl of 2X reaction mix, 0.5 μl of Superscript III/Platinum Taq enzyme mix (both supplied with the Superscript III/Platinum Taq one-step real-time RT-PCR kit e ), 2.0 μl each of the forward and reverse primers, g 1.5 μl TaqMan® probe, g and 1.5 μl nuclease-free water h to a final reaction volume of 25 μl. 15 An automated version of this RT-PCR set up was run in the MagNA Pure LC using the nucleic acid extracted in that robot. ROX reference dye was not included in the reactions.
One-step RT-PCR amplification in Mx4000 or Mx3005P Multiplex Quantitative PCR Systems i with versions 4.2 and 3.2 of the analysis software, respectively, used the following conditions: 60°C for 30 min, 1 cycle; 95°C for 10 min, 1 cycle; 95°C for 15 sec, 60°C for 1 min, 50 cycles. 15 Following amplification, threshold cycle (Ct) values for all reactions were assigned manually as described previously. 11 A negative result was reported if there was no amplification signal in the well (“No Ct” value), while a cut-off Ct value of 32.0 was used to define positive samples. This cut-off was initially based from testing ES using the MagNA Pure LC (as most sample submissions to the WRLFMD from overseas comprise vesicular epithelia; this being the sample of choice for virologic investigation). 16 In order to directly test whether the sample matrix or particular automated nucleic acid extraction system influenced the sensitivity of the real-time RT-PCR, decimal dilution series of FMDV (isolate UKG 34/2001) prepared in uninfected bovine serum e and ES were compared following extraction by the BioRobot Universal and MagNA Pure LC, respectively. Samples producing Ct values between 32.0 and 50.0 were retested (in duplicate) in order to establish a clear positive or negative result, although individual serum samples submitted from premises that had other samples that were clearly FMDV-positive were not typically retested. In the event that retesting was undertaken, primary material was used: the samples were designated negative if both wells yielded a “No Ct” signal. A positive result would have been reported from samples generating Ct values >32.0 on repeat testing if both values were in close agreement.
Addition of real-time RT-PCR positive and negative controls
For nucleic acid extraction and the subsequent steps of the real-time RT-PCR, 6 positive and 6 negative controls spread throughout the plate were included in each batch of 96 samples (BioRobot Universal System), while 2 positive and 6 negative controls were used in each 32-well cartridge processed in the MagNA Pure LC. The purpose of these controls was to monitor the sensitivity, specificity and interplate variation from each assay run. The positive control material (FMDV reference strain: O1 Manisa) was prepared (by dilution in negative bovine ES) to yield a moderately strong Ct value. This level of signal provided a more stringent indicator of the analytical sensitivity of the assay than a strongly positive control and also reduced the risk of intraplate cross contamination of neighboring wells.
Personnel for real-time RT-PCR
Four teams of 2 people each were available as trained real-time RT-PCR testers and had access to 1 BioRobot Universal System and 1 MagNA Pure LC. This arrangement provided the capability to process at least 250 samples within 11 to 12 hr. Most samples arrived at the laboratory in the evening, and those submitted from report cases were set up to run overnight so that results were available first thing the next morning, while the less urgent surveillance samples were processed the next morning.
Results
Validation of BioRobot Universal and comparison of cut-off Ct values used for different sample matrices
The absence of intraplate cross contamination between wells was demonstrated by testing checkerboard plates containing positive and negative samples. For this work, ES containing FMDV type O1 Manisa virus prepared in phosphate buffer (and further diluted in uninfected bovine ES) was mixed with Lysis/Binding Buffer, and 0.5 ml was added to 19 dispersed wells in an S-Block. d The same volume of Lysis/Binding Buffer was added to all remaining wells. All 19 wells containing O1 Manisa virus produced a positive result: mean Ct 18.38 ± 0.50 (coefficient of variation of 2.73%). No signal (“No Ct” value) was obtained from all remaining wells containing only Lysis/Binding Buffer (data not shown).
Results from comparative experiments (Fig. 1) showed the close similarity of the Ct values generated from decimal titrations of FMDV UKG 34/2001 in ES and sera using the MagNA Pure LC and BioRobot Universal instruments, respectively. This experiment, in which the reaction efficiency of the real-time RT-PCR in ES using the MagNA Pure LC was 101.6% and that of the assay in serum using the BioRobot Universal was 100.0%, provided assurance that the same cut-off value (Ct value of 32.0) was applicable during the outbreaks for real-time RT-PCR testing of sera as well as ES.
Analysis of the genome sequence targeted by the diagnostic real-time RT-PCR assay
The sequences of the 3D primers and probe 3 have no degenerate nucleotide positions and were found to have a 100% match with the sequence of the virus that was recovered from samples collected from the first IP (IP1b: GenBank accession number EU448371). In contrast, the primers used for the 5′UTR assay 11 have in-built degenerate bases in order to encompass nucleotide variability of diverse FMDV strains (32-and 8-fold for the forward and reverse primers, respectively). In addition to these positions, there was a further nucleotide substitution in the reverse primer region (position 14) of the virus sequence derived from IP1b that was not accommodated by this assay. As a consequence, the 3D-specific real-time RT-PCR was selected for testing all subsequent samples submitted for laboratory diagnosis in order to increase sample throughput even though the 5′UTR real-time RT-PCR assay was able to detect the U.K. 2007 causative virus strain.
Diagnostic results on samples collected from IPs
Real-time RT-PCR, Ag-ELISA, and VI each confirmed the presence of FMDV on all 8 IPs from respective clinical sample submissions. In addition, 15 preclinically viremic animals were discovered from IP2b. Blood samples had been collected from all 58 cattle in IP2b before the animals were killed and examined postmortem. None had visible lesions. 14 Eleven of the sera were positive by real-time RT-PCR and 11 by VI (7 sera were positive by both assays). A real-time RT-PCR positive result was also obtained from a clinically normal animal in IP1c, and this was the only evidence of infection on this holding.
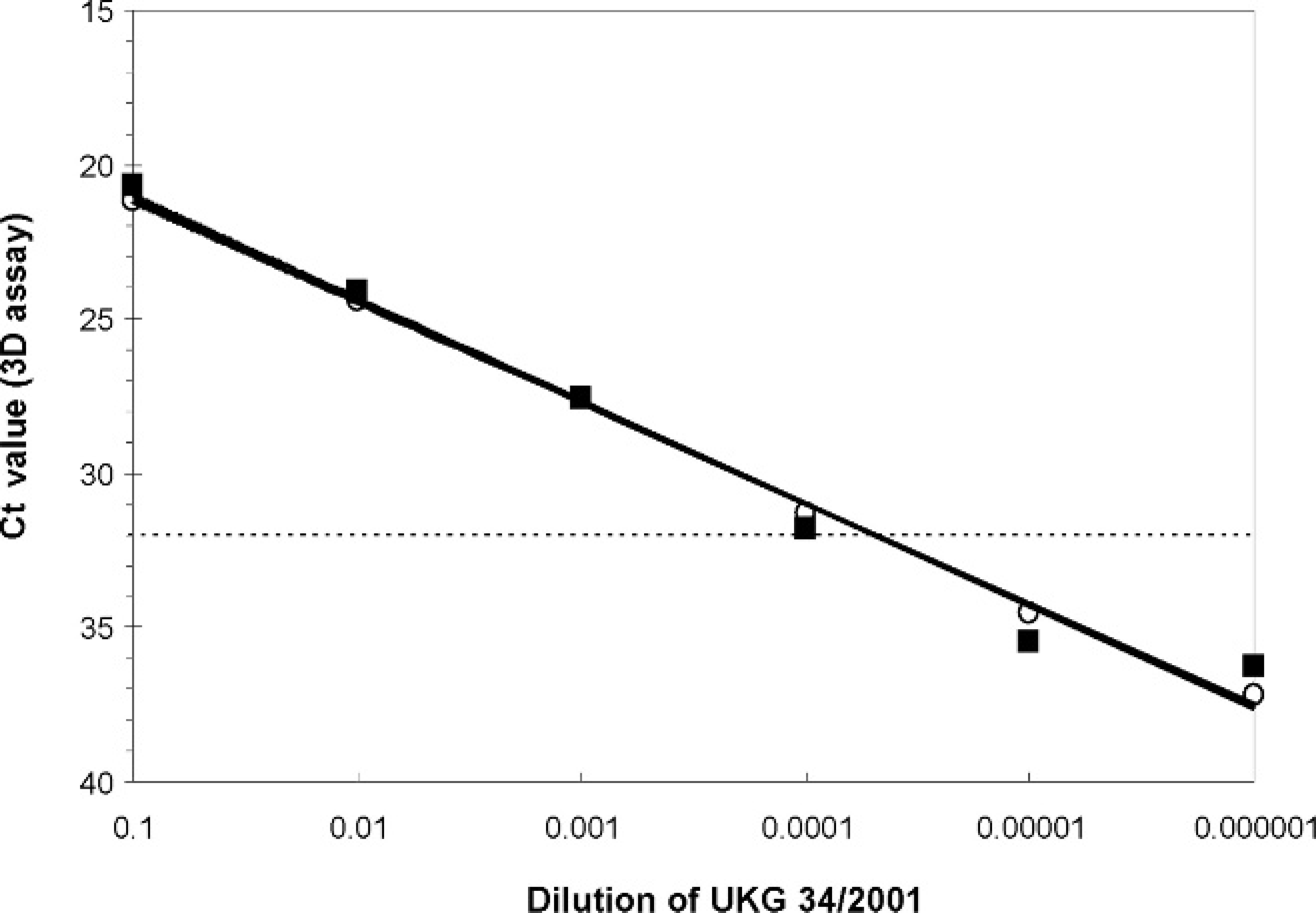
Correlation of the Ct values for the diagnostic real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assay on decimal dilutions of the foot-and-mouth disease (FMD) reference virus UKG 34/2001 in uninfected bovine epithelial suspension (ES) and serum using the MagNA Pure LC and BioRobot Universal for preparation of template, respectively; (ŝ) dilutions prepared in ES followed by extraction in the MagNA Pure LC; (□) dilutions prepared in serum followed by extraction in the BioRobot Universal. Solid lines represent best-fit lines through the points (R 2 values of 0.998 and 0.980 for ES and sera, respectively), and dotted line represents Ct cut-off of 32.0.
No clinical signs were reported by the farmer in either cattle or sheep from IP5 (Virginia Water), but as the bloods collected from the sheep were found to be very strongly seropositive, 14 the stock were inspected, found to have healing FMD lesions, and were quickly slaughtered. Probang samples were collected from all 16 sheep; of these, FMDV was detected in 10 by real-time RT-PCR, 7 of these showed a CPE on VI. One notable discrepancy between the real-time RT-PCR and VI results was evident when sera from IP8b were tested by these methods. Only a single clinically affected animal was found on IP8, and of the 54 submitted sera, 15 were positive by VI but only the one from the clinical case was detected (above the threshold Ct) by real-time RT-PCR both in the initial test and on retesting. Other than this single real-time RT-PCR positive sample (Ct value was 25.73), 4 samples from this IP generated Ct values in the range 32.0–50.0: 3 of these were VI negative, while 1 was VI positive.
Other report cases, and use of real-time RT-PCR for targeted surveillance of FMD
Twenty probangs, 11 ES, and 1 fecal suspension submitted/prepared from other report cases were negative by real-time RT-PCR, VI, and Ag-ELISA. From these cases, 254 sera were also negative both by real-time RT-PCR and VI, and an additional 180 sera were negative by real-time RT-PCR alone. In addition, 3 swab samples were negative by both real-time RT-PCR and VI, while 3 more were negative by real-time RT-PCR alone. For the targeted surveillance program, all 1,861 serum samples submitted from the IPA were negative by real-time RT-PCR (“No Ct” values). Of the 11 farms included in the IPA, 4 holdings were culled as high-risk farms associated with the subsequent occurrence of FMD on IP8. Cattle on the remaining 7 premises (n = 248) tested real-time RT-PCR negative on October 16th and did not have to be culled for disease control.
Compilation of tests performed and comparison of the results obtained by real-time RT-PCR with other laboratory methods
Test selection: compilation of tests performed by real-time RT-PCR, Ag-ELISA, and VI on the diagnostic samples submitted during the U.K. 2007 FMD outbreaks.*
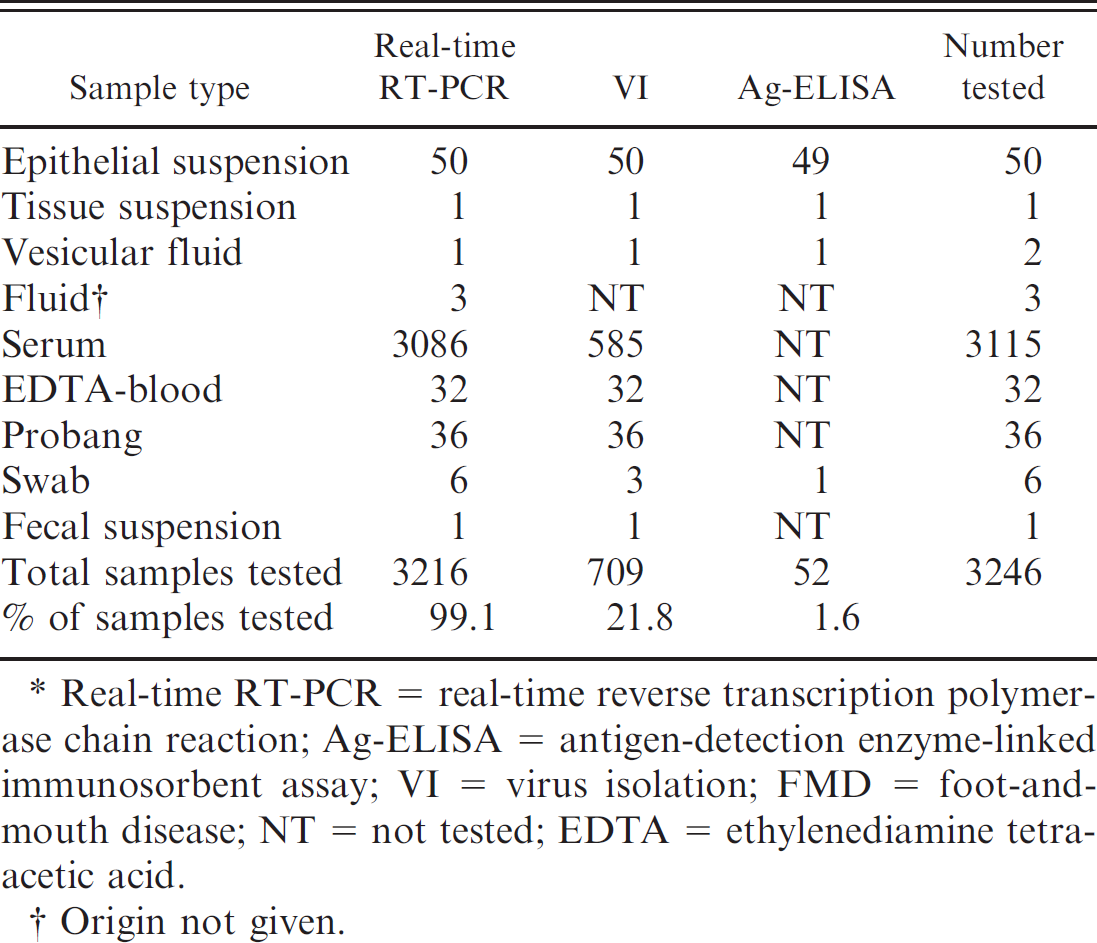
Real-time RT-PCR = real-time reverse transcription polymerase chain reaction; Ag-ELISA = antigen-detection enzyme-linked immunosorbent assay; VI = virus isolation; FMD = foot-and-mouth disease; NT = not tested; EDTA = ethylenediamine tetra-acetic acid.
Origin not given.
Table 1 shows a compilation of the test results achieved by real-time RT-PCR, Ag-ELISA, and VI on the samples received for FMD diagnosis. More serum samples were processed by real-time RT-PCR than by VI, demonstrating the extent of the sample throughput that was achieved with the BioRobot Universal System and MagNA Pure LC automated instruments. Figure 2 shows the numbers of samples tested per day by real-time RT-PCR during the outbreaks. During the second phase of the outbreaks, more than 200 samples per working day were processed by real-time RT-PCR on 6 days, reaching a peak of 269 (all sera) on October 2nd. This peak was associated with the screening of high-risk herds in the IP area north of Egham, United Kingdom.
For some samples, multiple assays were used to detect FMDV allowing the comparative performance of difference assay systems to be assessed. Figure 3 shows the comparison between real-time RT-PCR and VI: there was close agreement between the results on ES, but the methods correlated less well when testing blood samples. Of the 61 sera and 2 EDTA-bloods that tested positive by VI, 40 were also positive by real-time RT-PCR. Conversely, 24 other serum samples from IPs were positive by real-time RT-PCR but failed to show a CPE in cell culture. In total, FMDV was detected in 120 clinical samples using real-time RT-PCR, VI, and Ag-ELISA. One hundred and nineteen of these samples were tested by both real-time RT-PCR and VI (remaining positive sample [single vesicular fluid] was only tested by Ag-ELISA). More positive samples were detected by real-time RT-PCR than VI on all sample types except EDTA-blood (2 detected by VI, 1 by real-time RT-PCR). This was not surprising given the poor performance of the nucleic acid extraction system for this sample type revealed during pilot comparisons between EDTA-blood and serum (see “Nucleic acid extraction” section). Fewer samples were tested by Ag-ELISA (the ELISA only being used to test original tissue suspensions and vesicular fluids): of the 52 samples tested, 21 were FMDV positive, while 31 were FMDV negative. No Ag-ELISA-positive samples tested negative by real-time RT-PCR or VI.
Specificity of real-time RT-PCR
The Ct values from the negative controls (n = 953) and clinical samples from FMD-negative farms (n = 3004) run in the diagnostic assays are shown in Figure 4. The Ct values of 943/953 negative control and 2,972/3,004 clinical samples were ≥50.0 (“No Ct” value). Furthermore, all of the remaining samples produced Ct values above the cut-off value of 32.0 except for 1 negative control. Therefore, the specificity of the assay using the positive/negative acceptance criteria for the negative controls and clinical samples was 99.9% and 100.0%, respectively.
Intra- and interassay reproducibility of real-time RT-PCR
In total, 91 separate nucleic acid plates were prepared during the 2007 U.K. outbreaks: 69 assays were extracted on the BioRobot Universal System, while the other 22 were performed using the MagNA Pure LC. In order to monitor intra-assay variability, the Ct values from the positive controls included in each assay were analyzed (data not shown). Non-parametric tests revealed that the real-time RT-PCR signal generated by positive controls extracted on the BioRobot Universal System (mean ± SD of all controls 20.9 ± 1.5) were significantly stronger than Ct values from the same RNA from ES material using the MagNA Pure LC (mean ± SD of all controls 23.6 ± 1.8: Wilcoxon rank-sum test, P < 0.001). A single positive control well (in run number 79) failed to generate a signal in the assay; in light of these results, the affected plate (containing controls and test samples) was repeated. All other controls were successfully amplified in the assays.
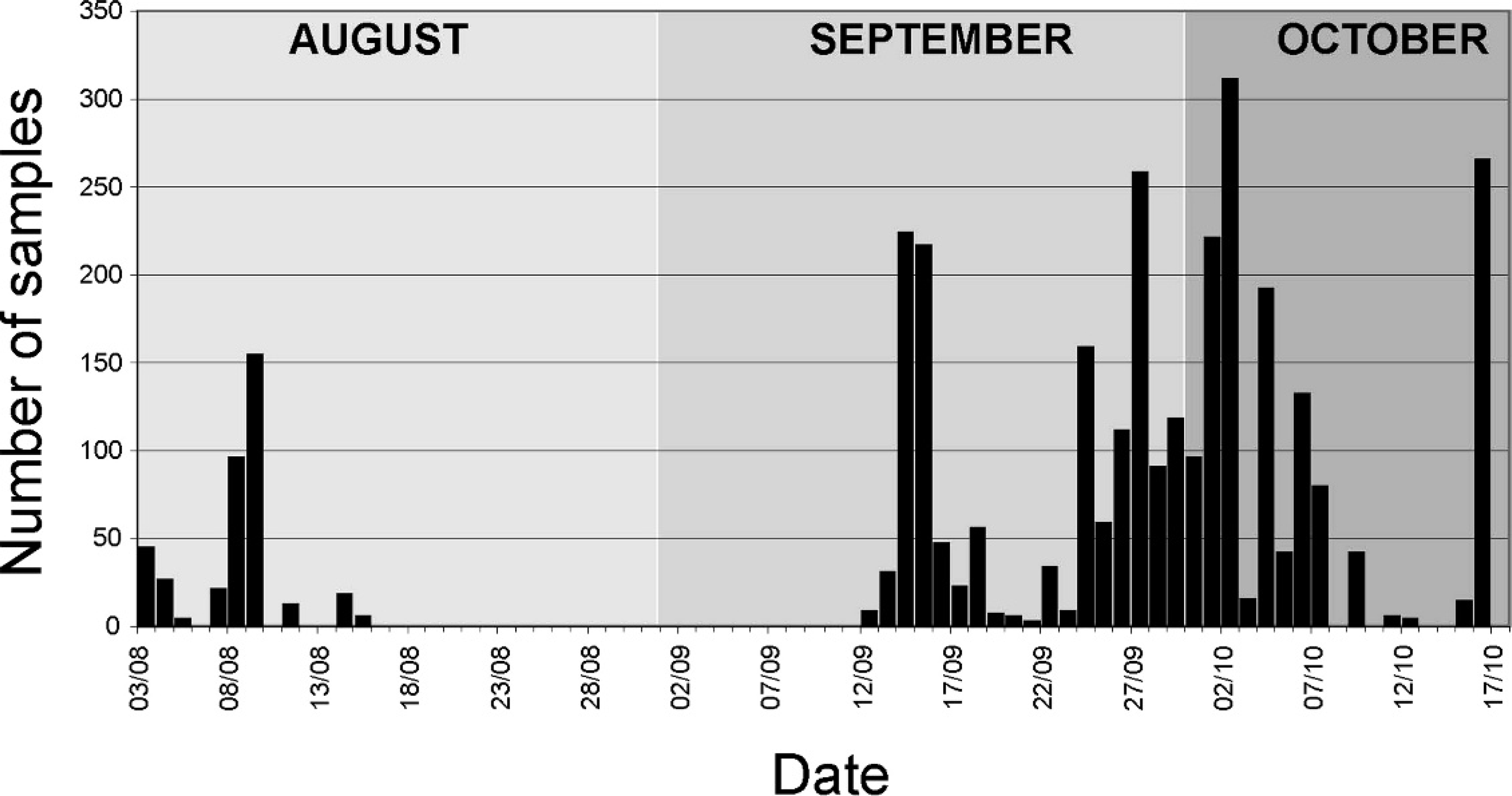
Daily numbers of samples tested by real-time reverse transcription polymerase chain reaction during the United Kingdom 2007 foot-and-mouth disease outbreaks (August 3 through October 17, 2007).
Processing of samples from report cases and from IP areas
Specificity was lower on 2 occasions when strong FMDV-positive clinical samples were included in extractions performed using the BioRobot Universal System. On one of these occasions, 13 negative samples and 9 negative control wells produced Ct values <50.0 when in close proximity to wells producing a strong positive result (the lowest Ct value was 7.9, data not shown). As a consequence of these findings, samples submitted from report cases were generally processed on the MagNA Pure LC and separated from those collected as part of the IPA active surveillance program (processed using the BioRobot Universal System).
Discussion
Real-time RT-PCR assays are widely used for the laboratory diagnosis of FMD and other important human and veterinary diseases. The current report describes the performance of real-time RT-PCR during the outbreaks of FMD that occurred in the United Kingdom during 2007 and outlines some of the key factors that should be considered before adopting these tests for outbreak use.
A highly sensitive parallel-plex real-time RT-PCR system utilizing 2 independent assays 3,11 was initially adopted for testing of samples submitted from the first IP. However, once the full genome of the causative strain had been sequenced, 4 a single assay target was selected, increasing the throughput of samples that could be tested. In addition to supporting high-resolution epidemiologic tracing, 4 sequencing of representative FMDV isolates provided confidence in the continued suitability of the genome region targeted by the real-time RT-PCR. The use of automated nucleic acid extraction equipment alleviated a major bottleneck in the real-time RT-PCR, particularly as the BioRobot Universal System was adopted for use in conjunction with the MagNA Pure LC. This method enabled a high sample throughput to be maintained throughout the duration of the outbreaks. The daily number of sample submissions peaked at 269, which were handled by 4 separate extraction runs on the BioRobot Universal System combined with the use of the Mx4000 and Mx3005P real-time PCR machines. Using a 24-hour rota system with these resources, it would have been possible to have scaled-up the throughput to approximately 500 samples per day.
Testing of samples submitted during these outbreaks produced data to further define the comparative performance of diagnostic tests currently used in the laboratory (real-time RT-PCR, Ag-ELISA, and VI). All 3 assays successfully detected FMDV on all 8 IPs. However, not all serum samples found positive by VI were classified as positive by real-time RT-PCR (Fig. 3). It is possible that the conservative cut-off used in the assay underestimated the number of samples that were positive. However, this only partly explains the discrepancy between real-time RT-PCR and VI, since only 4 VI-positive samples also produced Ct values between 32.0 and 50 in the real-time RT-PCR assay. Of the 23 VI-positive/real-time RT-PCR negative samples, 14 were collected from IP8b. Since these findings were not consistent with other data from the outbreaks, the testing of these samples was repeated using frozen material stored from the original submission. On this occasion, FMDV could only be isolated from a single sample in cell culture. Previous analysis of the 2001 epidemic in the United Kingdom had highlighted samples where real-time RT-PCR assay failure was due to nucleotide substitutions in the genome regions targeted by the primers and probe. 7 Full genome sequencing of representative isolates from all IPs, including IP8b, revealed no genetic differences in the 3D region targeted by the real-time RT-PCR primers/probe set. 4 Furthermore, parallel testing of these samples using the 5′UTR 11 assay failed to detect any additional FMDV positives in the sera from IP8 (data not shown). Therefore, the discrepancy between VI and real-time RT-PCR on IP8 cannot be explained by mismatches in the real-time RT-PCR assay target. Since it is impossible to rule out the effects of a freeze/ thaw cycle, no further conclusions could be made regarding the original results generated for these sera. The possibility that cross contamination had occurred when the VI was originally performed on these samples (as within-batch samples were routinely processed together) could be explored by full-genome sequence analysis: identical sequence data for the 15 isolates would certainly support this hypothesis, while variant sequences would suggest separate sources of FMDV from different animals within the herd.
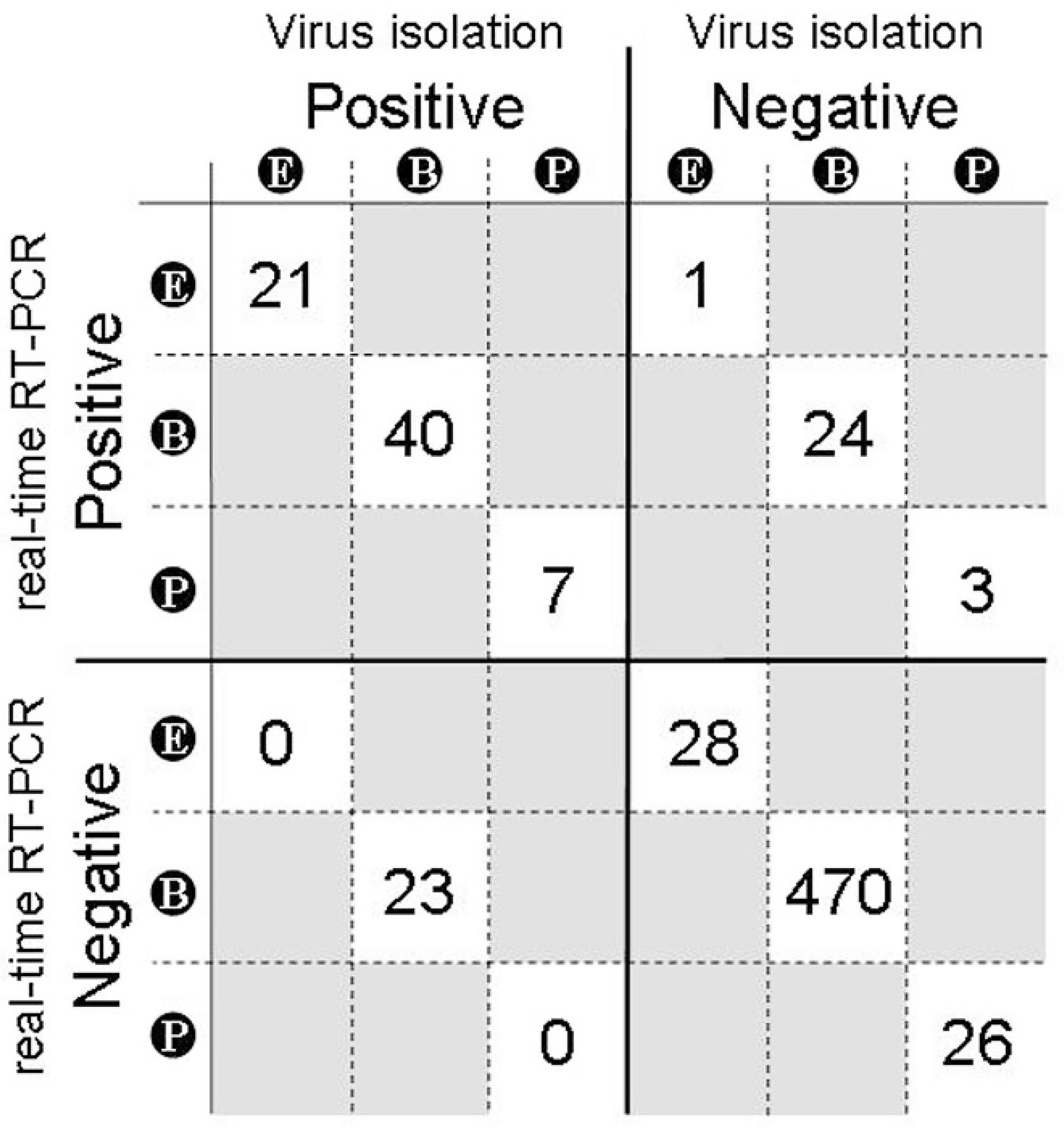
Sensitivity of the diagnostic real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assay in comparison with virus isolation in cell culture on the clinical samples submitted during the U.K. outbreaks for virologic investigation. The numbers of samples of each type testing positive or negative by the respective test are shown in the boxes: E, original suspension of vesicular epithelium; B, blood; P, probang. Results for 4 samples: a single fecal sample, a tissue suspension of unknown origin, a swab, and a vesicular fluid are not shown (first 3 samples were negative by both assay systems, while the vesicular fluid was positive by real-time RT-PCR and virus isolation).
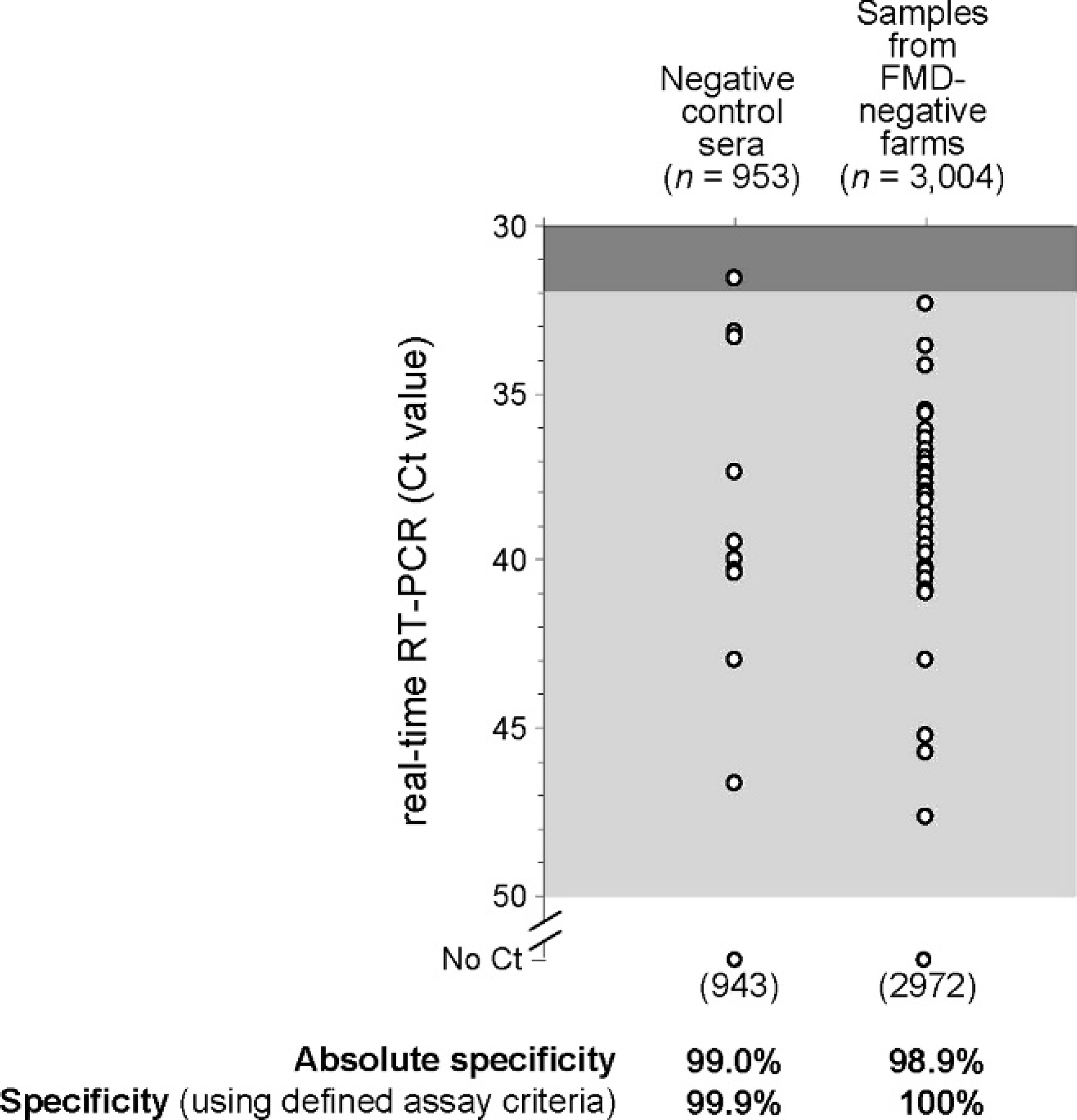
Specificity of the diagnostic real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assay during the testing of samples received during the outbreaks in the United Kingdom. Threshold cycle (Ct) values for negative control wells and clinical samples from foot-and-mouth disease (FMD)-negative farms are shown.
In light of the results above, it is interesting to retrospectively consider the Ct cut-off value used and its effect on the results reported for samples submitted during the outbreaks. Seventy-one additional samples from IPs, which were VI-negative, also gave Ct values in the range between 32.0 and 50.0 (there were also 2 further samples not tested by VI that generated Ct values in this range). None of these samples were retested in order to define their status because they were derived from premises already known to be infected (due to results from other samples). Therefore, the authors can only speculate whether these weak signals were indicative of low-level viremia in these animals, or alternatively had occurred due to within-run cross contamination of the assay yielding false positive results. Clear criteria were established to classify samples as FMDV-positive or -negative and to define those that required a retest. In terms of assay specificity, it was important to adopt a strict regime for sample processing in order to limit the potential for cross contamination that could yield false-positive results. While raising the assay cut-off to a Ct value of 35.0 would have detected a further 22 samples from IPs, this would begin to decrease the assay specificity (Fig. 4), potentially leading to misdiagnosis of FMD in negative herds. This is a particularly important aspect to consider when using real-time RT-PCR to support an active surveillance program that aims to identify preclinical individuals within a herd that are likely to have low-level viremia in the absence of clinical signs. In the event that Ct values between 32.0 and 50.0 were obtained on otherwise FMDV-negative farms, repeat testing would have been undertaken to determine whether these results were indicative of preclinical viremia on a farm at a lower level than observed on IP2b. In fact, during these outbreaks, no individuals from these FMDV-negative farms had repeat Ct values in this range.
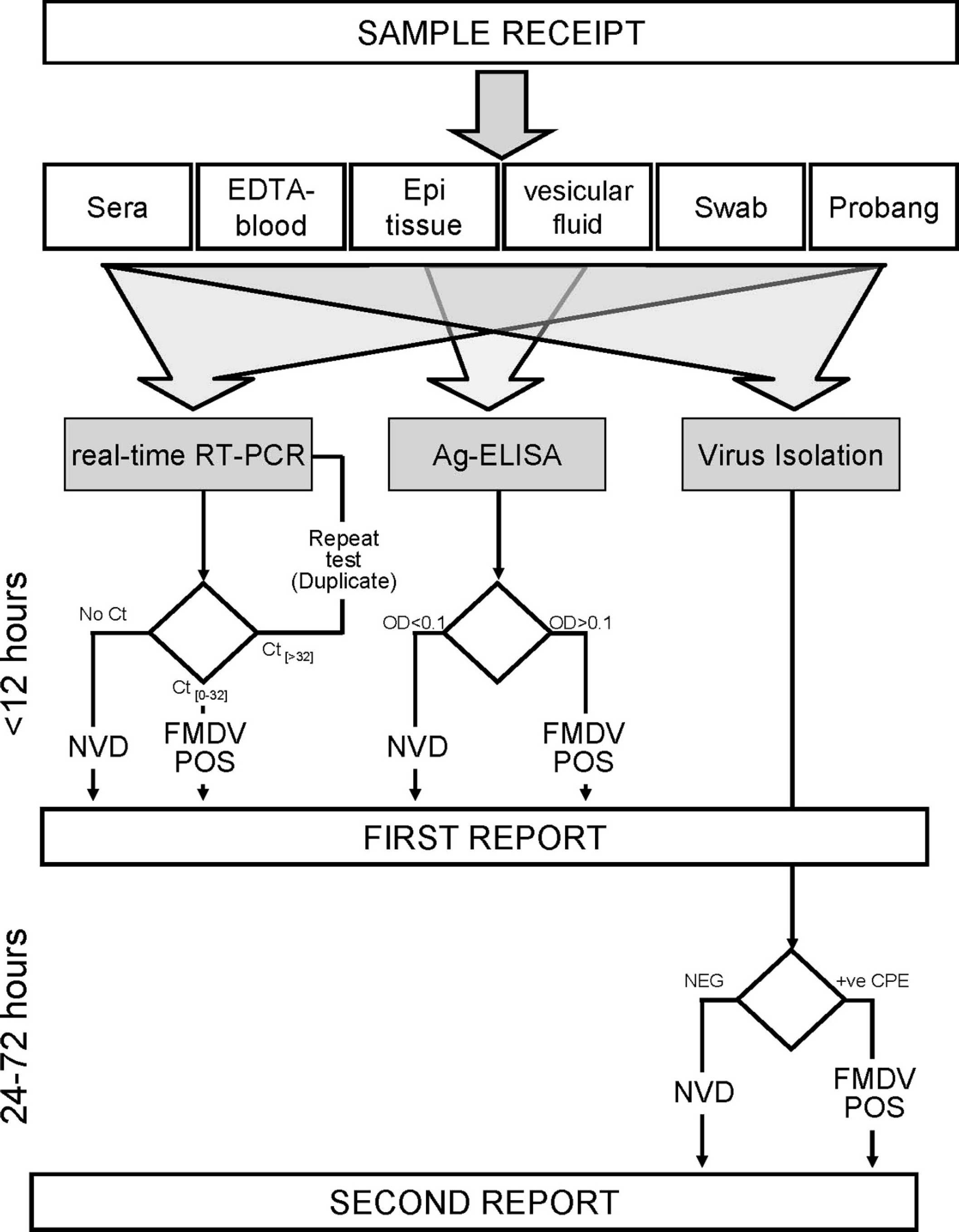
Schematic outline of the Foot-and-mouth disease virus (FMDV) diagnostic procedures used during the 2007 U.K. outbreaks, including timescales and reporting of results. NB Lateral flow devices (Svanodip FMD virus Ag Test) j ; were also used to provide preliminary results within an hour of receipt of tissues from suspect cases. 8 NVD = no virus detected; Ct = threshold cycle; OD = optical density.
In summary, the present report describes the use of real-time RT-PCR during the U.K. FMD outbreaks in 2007. In addition to confirming the presence of FMDV on all the IPs, this assay enabled preclinical infected animals to be identified due to the occurrence of an FMDV viremia in infected animals prior to the generation of macroscopic lesions. Examination of probang samples might enable FMDV to be detected even earlier in the course of infection, 1 but their collection and processing is more cumbersome for use on a large-scale basis compared with sera. Exploiting this ability to accurately detect infected animals in early stages of disease, the real-time RT-PCR assay was used to support an active surveillance program using sera within high-risk cattle herds. An additional advantage of regular sampling in high-risk herds is that it requires close restraint of the cattle, improving the chances of observing clinical signs of early FMD. The results generated by automated real-time RT-PCR were used to support control decisions during the 2007 outbreaks of FMD in the United Kingdom with concomitant reduction in the reliance placed upon VI. An overview of the testing scheme adopted during the outbreaks is shown in Figure 5. Results obtained by real-time RT-PCR and Ag-ELISA were typically obtained at a similar time and were sent to the Department for Environment, Food and Rural Affairs (Defra; as a first report) within 5–6 hr of sample receipt giving the positive/negative status of the suspected premise(s). If required, a second report (24–72 hr after sample receipt) could be produced including VI results. Virus isolation remains a useful laboratory tool for detection of FMDV, not only for propagation of virus for characterization studies but for independent verification of positive real-time RT-PCR results in herds without clinical signs. In comparison with previous FMD outbreaks, the impact of VI was reduced since control decisions were made typically based on the first report: either 1) positive results by real-time RT-PCR alone or with Ag-ELISA confirming the presence of FMD, or 2) negative by real-time RT-PCR alone or with Ag-ELISA for confirmation of negative samples on a farm. To the authors' knowledge, this was the first time that real-time RT-PCR had been used in the face of an FMD outbreak in a previously FMD-free country. With a view to further improving the protocols used for nucleic acid extraction, an alternative protocol has been recently developed for use on the BioRobot Universal System. This protocol requires a smaller sample volume and also performs well on EDTA-blood and could realistically be adopted for routine diagnostic use. Furthermore, the development of mobile RT-PCR platforms for on-site diagnosis of FMD would speed up the issue of test results and enable control procedures to be implemented more rapidly; although the typical throughput for these systems would limit their use to confirmation of clinical cases rather than to the screening of entire herds.
Acknowledgements
The authors thank the following colleagues from the Institute for Animal Health, Pirbright Laboratory: Nick Juleff for providing samples and clinical data used in assay validation, Annette Saunders for invaluable assistance with the laboratory sample management database, and Simon Gubbins for statistical analysis of data. The authors also thank John Buckels of QIAGEN for technical assistance with the BioRobot Universal System. Validation of the real-time RT-PCR assay was funded by Defra projects SE1121 and SE1124. Other colleagues in Defra and the Institute for Animal Health are thanked for their assistance during and after the outbreaks.
Footnotes
a.
Roche Diagnostics Ltd., Lewes, East Sussex, UK.
b.
HETTICH-Zentrifugen GmbH & Co., KG, Tuttlingen, Germany.
c.
Beckman Coulter Ltd., High Wycombe, UK.
d.
Qiagen Ltd., Crawley, West Sussex, UK.
e.
Invitrogen Ltd., Paisley, UK.
f.
BIS Ltd., London, UK.
g.
Sigma-Aldrich Co., Ltd., Gillingham, Dorset, UK.
h.
Ambion Inc., Warrington, UK.
i.
Stratagene, Amsterdam, The Netherlands.
j.
Svanova Biotech AB, Uppsala, Sweden.
