Abstract
Lymphocytosis caused by neoplastic proliferation of small lymphocytes is occasionally difficult to distinguish by morphological examination from nonneoplastic lymphocytosis. To examine the clinical utility of gene rearrangement analysis for demonstrating neoplastic proliferation of small lymphocytes, gene rearrangement analysis was performed in comparison with immunophenotyping using peripheral lymphocytes in dogs with small lymphocytosis. Thirty-one dogs with small-cell lymphocytosis (8,100–884,300
Introduction
Small lymphocytosis in dogs may arise from several causes such as antigenic stimulation, immune-mediated diseases, infectious diseases, and neoplasia. Neoplasia characterized by small lymphocytosis occurs in chronic lymphocytic leukemia (CLL) and in the leukemic phase of low-grade (small-cell) lymphoma. 2 Because neoplastic lymphocytes appear mature, they can be difficult to distinguish from nonneoplastic lymphocytosis by routine morphological evaluation. One of the methods to demonstrate neoplastic expansion of lymphocytes is immunophenotyping by flow cytometry, which characterizes the lineage and diversity of the lymphocytes. Detection of a marked expansion of a phenotypically homogeneous population of lymphocytes or of a population of lymphocytes that expresses an aberrant constellation of lymphocytic antigens is suggestive of neoplasia. 1,12 In addition to immunophenotyping, antigen receptor gene rearrangement analysis, so-called clonality analysis, which detects homogeneity of antigen receptor gene rearrangement, is another method to detect lymphoid neoplasia in cases with small lymphocytosis. Clonality analysis using polymerase chain reaction (PCR) amplification for dogs has been previously reported. 4 More recently, a similar assay was developed based on the canine genome database by targeting the T-cell receptor gamma chain (TCRγ) gene 15 and the immunoglobulin heavy chain (IgH) gene. 9 Although the diagnostic value of PCR analysis has been demonstrated in several types of lymphoproliferative disorders, 7,11,14 only limited studies have been done to identify neoplastic proliferation of lymphocytes in dogs with small-cell lymphocytosis. In the present study, immunophenotypical evaluation was performed on the blood of dogs with small lymphocytosis. The ability to detect cellular clonality by PCR analysis was examined using these lymphocytes.
Materials and methods
Cases
Peripheral blood was obtained from 31 dogs with small-cell lymphocytosis. The number of leukocytes was counted with an automated blood cell counter. a Leukocytic differential and morphological evaluations were performed on Wright-Giemsa–stained smears. The leukocytic differential count was determined by classifying 200 leukocytes on the stained blood smear. Abdominal x-ray and/or ultrasound examination was performed on some dogs.
Flow cytometric analysis
Immunophenotyping of peripheral blood lymphocytes was performed by evaluating cell surface expression of T-cell markers (cluster of differentiation [CD]3, CD4, and CD8) and B-cell markers (CD21 and immunoglobulin G [IgG]) using flow cytometry. b Among the 31 cases enrolled, 5 cases (cases 4, 6, 7, 29, and 31) were not examined for the surface expression of IgG. Peripheral blood was treated with BD Pharm Lyse c to lyse erythrocytes. After washing with phosphate buffered saline, the remaining nucleated cells were labeled with fluorescein-isothiocyanate (FITC)-conjugated mouse monoclonal antibodies d for canine CD3 or CD4 or FITC-conjugated rabbit anti-dog IgG. e Phycoerythrin (PE)-conjugated mouse monoclonal antibodies c for canine CD8 and CD21 were used for double color staining of cells in combinations of antibodies for CD3 and CD21, CD4 and CD8, and CD21 and IgG. FITC- or PE-conjugated mouse IgG was used as a negative control. After staining with the antibodies, the cells were analyzed by a flow cytometry. b Data were analyzed by WinMDI 2.9, f and lymphocytes were identified and gated on a forward-scatter versus side-scatter plot. Regarding dual staining using anti-CD3 and anti-CD21, the company that provided the antibodies recommended that anti-CD3 and anti-CD21 not be used together because of a nonspecific interaction between the antibodies. However, in a preliminary experiment for the current study, in which peripheral blood mononuclear cells (PBMCs) collected from 16 healthy dogs were examined by flow cytometric analysis using a combination of the 2 antibodies, nonspecific interactions of the 2 antibodies were not observed. Therefore, the antibodies were employed for double color staining of immunophenotyping in the current study.
Clonality analysis
Using PBMCs prepared from the 31 cases, PCR-based clonality analysis was performed as described previously. 9,15 Briefly, genomic DNA was extracted from PBMCs using a commercial extraction kit. g For detection of clonal rearrangement of the TCRγ gene, 2 different primers for V segments (Vγa: 5′-CGTGTACTACTGCGCTGCCTGG-3′ and Vγb: 5′-GGCTGTATTACTGTGCCTGCTGG-3′) and 2 different primers for J segments (Jγa: 5′-TACCTTCTG(C/T)AAATATCTTGA-3′ and Jγb: 5′-TGTGCCAGGACCAAGCACTTTGTT-3′) were used. For the analysis of IgH gene rearrangement, the following primers were used: 5′ primer for V segments (IgHV; 5′-ACACGGCC(A/C/G)TGTATTACTGT-3′) and 3′ primer for J segments (IgHJ; 5′-TGAGGAGACGGTGACC-3′). The genomic DNA (250 ng) was independently amplified with Taq DNA polymerase in a 50-μl reaction volume using 4 different combinations of primer sets for the TCRγ gene and a set of primers for the IgH gene. h The PCR cycling protocol consisted of denaturing for 45 sec at 94°C and annealing for 30 sec at 60°C, 69°C, 60°C, 62°C, and 54°C for the primer combinations of Vγa-Jγa, Vγa-Jγb, Vγb-Jγa, Vγb-Jγb, and IgHV-IgHJ, respectively. A 30-cycle extension was performed for 20 sec at 72°C. After PCR amplification, an aliquot (10 μl) of PCR products of the TCRγ gene was size fractionated on a 7.5% polyacrylamide gel, and a similar aliquot of IgH amplicons was size fractionated on a 4.0% agarose gel. Amplification products were visualized in ethidium bromide–stained gels. Clonal proliferation of lymphocytes was demonstrated by observation of discrete bands of PCR products on the gels.
Results
Clinical descriptions
The numbers of lymphocytes in peripheral blood ranged from 8,100 to 884,300/μl (reference range: 1,000–4,800/μl; Table 1). The lymphocytes were small cells with clumped chromatin and inapparent nucleoli. The cytoplasm was frequently abundant and clear to pale blue. Various numbers of fine azurophilic granules were seen in the cytoplasm of lymphocytes in 2 of the 31 cases (cases 3 and 10), indicating large granular lymphocytosis. Seven cases had peripheral lymphadenopathy (cases 1, 11, 24, 25, and 28–30). Among the 17 cases for which abdominal x-ray and/or ultrasound examination were performed, hepatomegaly was detected in 5 cases (cases 1, 5, 7, 12, and 31), and splenomegaly was detected in 6 cases (cases 1, 12, 13, 15, 28, and 31).
Immunophenotyping
Results of immunophenotyping are shown in Table 2. Seventeen of the 31 cases (cases 1–17) had expansion of a homogeneous population of lymphocytes retaining expression of the normal constellation of antigens; 12 cases were of T-cell lineage, including 10 cases of CD8 (cases 1–10) and 2 cases of CD4 T-cell lineage (cases 11 and 12). Five cases were of B-cell lineage (cases 13–17). In the remaining 14 cases (cases 18–31), a population of lymphocytes expressing an aberrant constellation of antigens was detected. The phenotypes included aberrant T-cell antigen expression (CD3+ CD4– CD8–, case 18; CD3– CD4– CD8+, cases 19 and 20; and CD3– CD4+ CD8–, case 21), coexpression of both T- and B-cell markers (CD3+ CD21+, cases 22–30; the scatter plot data of case 28 is shown in Fig. 1 as a representative case), and lack of both T- and B-cell marker expression (case 31). Small-cell lymphocytosis of all 31 cases was considered to be neoplastic based on marked expansion of phenotypically homogeneous lymphocytes or the presence of an aberrant antigen-expressing cell population.
Detection of clonality
Results of the PCR analysis are summarized in Table 2. Clonal proliferation of lymphocytes was detected in 27 cases, including 21 cases involving the TCRγ gene and 6 cases involving the IgH gene. In the analysis of the TCRγ gene, the primer set Vγa-Jγb detected clonality in 18 cases, including 15 cases of monoclonality and 3 cases of biclonality, which is represented as 2 discrete bands of PCR products on the gel. The primer set of Vγb-Jγb detected monoclonality in 3 cases of lymphocytes. Regarding the primer Jγa, clonality was not detectable in any combinations with Vγa and Vγb.
Signalment, liver and splenic involvement, total leukocyte counts, and absolute lymphocyte counts of 31 dogs with small-cell lymphoid neoplasia.*
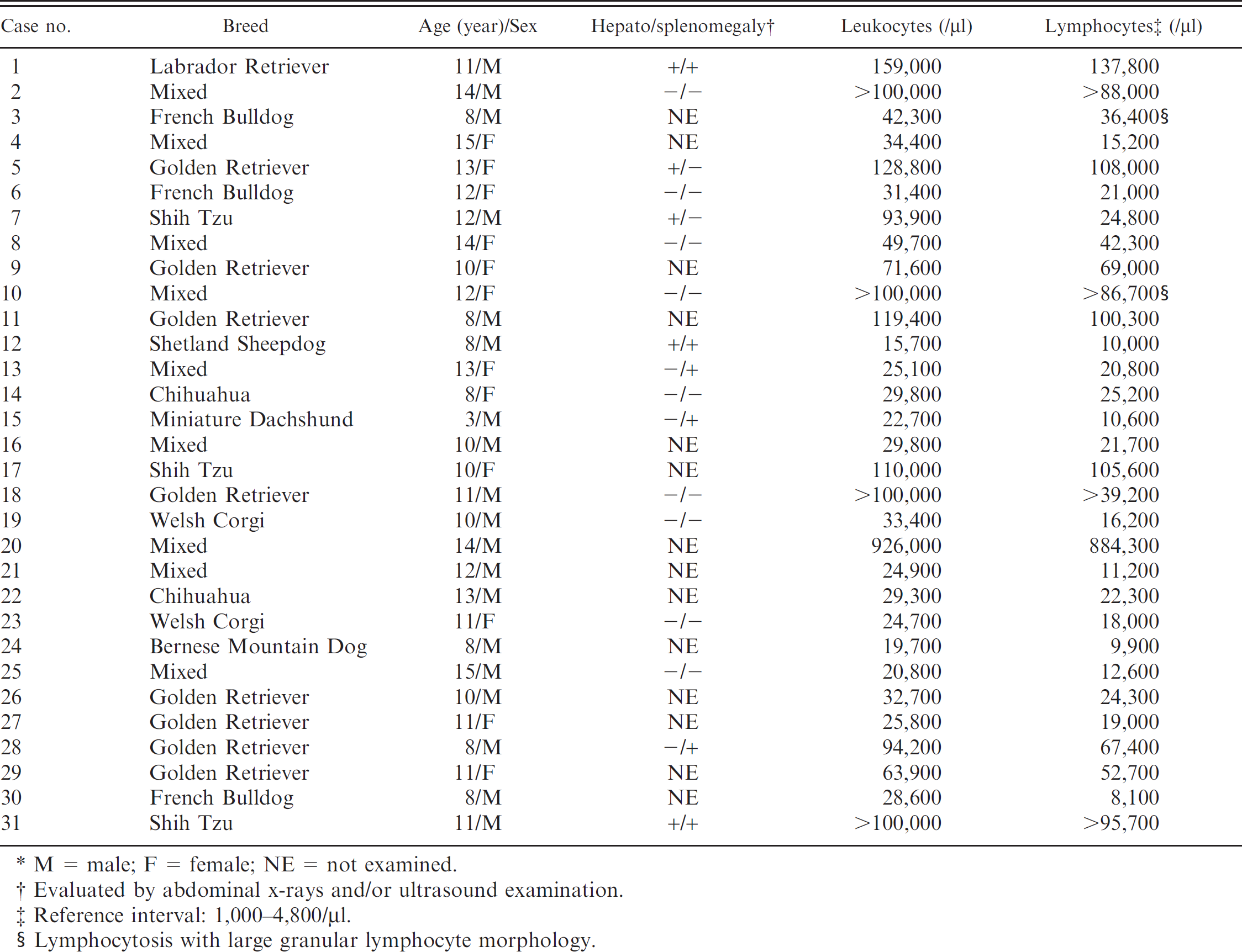
M = male; F = female; NE = not examined.
Evaluated by abdominal x-rays and/or ultrasound examination.
Reference interval: 1,000–4,800/μl.
Lymphocytosis with large granular lymphocyte morphology.
Discussion
Similar to CLL, the leukemic phase of low-grade lymphoma also presents small-cell lymphocytosis. Because CLL may also present with lymphadenopathy, hepatomegaly, and/or splenomegaly due to the infiltration of neoplastic lymphocytes, a distinction between the 2 lymphoproliferative diseases is unclear in veterinary medicine. Therefore, some cases of the leukemic phase of low-grade lymphoma could have been included that were indistinguishable in the present study.
All 31 cases of small-cell lymphocytosis in the present study were confirmed to be neoplastic by immunophenotypic studies, and clonality was detected in 27 cases (87%). In a previous report, 3 clonal rearrangements of antigen-receptor genes with a detection rate of 91% were demonstrated in dogs with lymphoid neoplasia using primer sets that differed from those in the present study. Therefore, 4 cases of lymphocytes in which clonality was not detected were examined by PCR analysis using primer sets previously reported 3 ; however, clonality was not detected by PCR analysis. Although PCR primers for clonality analysis were designed to amplify known V and J segments of TCRγ and IgH canine genes, these primers will not be able to detect clonality in cases that have rearrangement using undefined V and/or J gene segments. Because the number of V and J gene amplicons defined in dogs is still few compared with those in humans and mice, it may be possible that the 4 cases in the present study possessed rearranged genes using undefined V and/or J gene segments. Alternatively, unusual T- and/or B-cell receptor gene rearrangement, such as deletion within genes and incomplete rearrangement, 3,5,8 could have resulted in the failure of the detection of clonality in these 4 cases.
Results of immunophenotyping and clonality analysis for 31 dogs with small-cell lymphoid neoplasia.
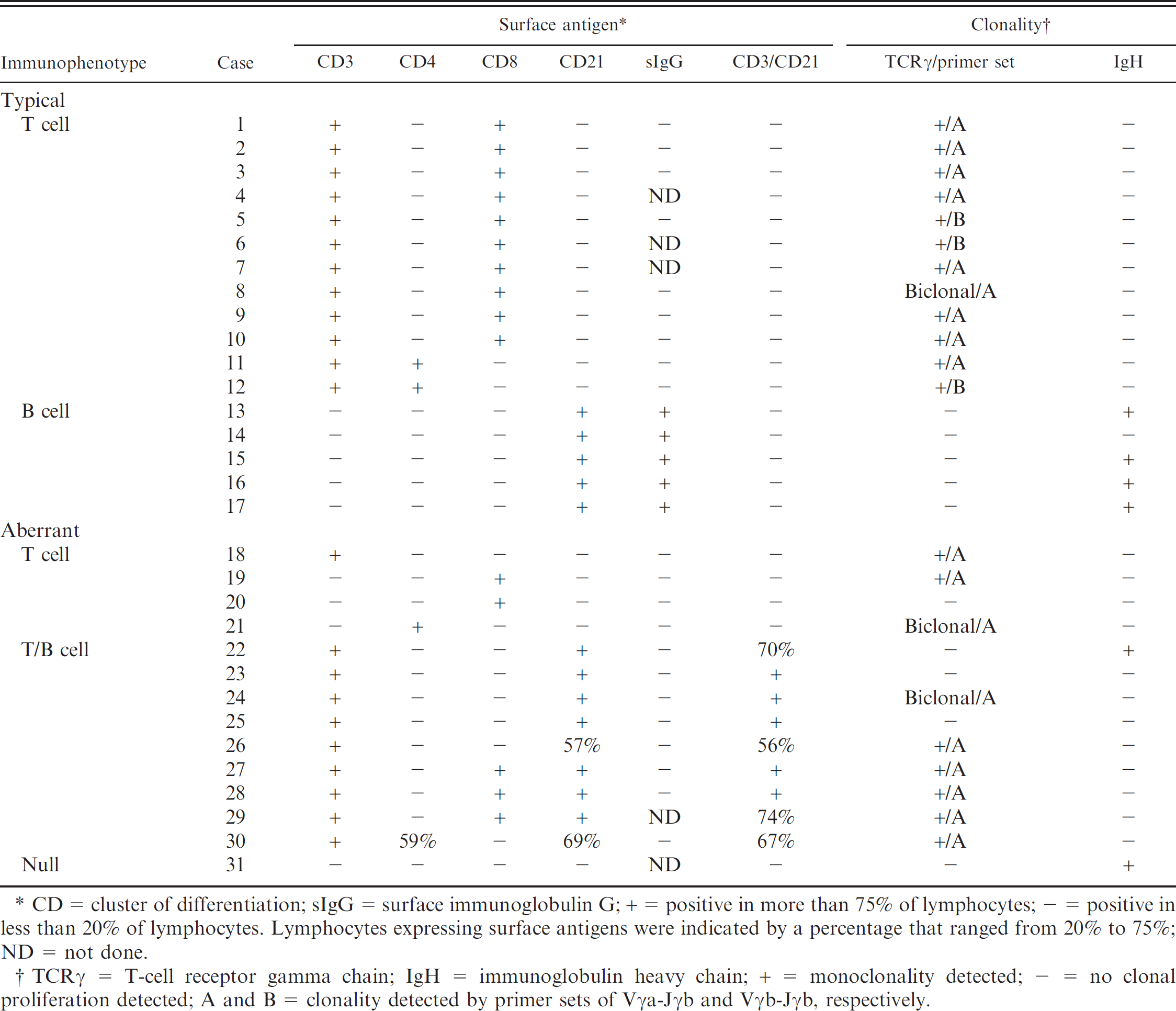
CD = cluster of differentiation; sIgG = surface immunoglobulin G; + = positive in more than 75% of lymphocytes; – =positive in less than 20% of lymphocytes. Lymphocytes expressing surface antigens were indicated by a percentage that ranged from 20% to 75%; ND = not done.
TCRγ = T-cell receptor gamma chain; IgH = immunoglobulin heavy chain; + = monoclonality detected; – = no clonal proliferation detected; A and B = clonality detected by primer sets of Vγa-Jγb and Vγb-Jγb, respectively.
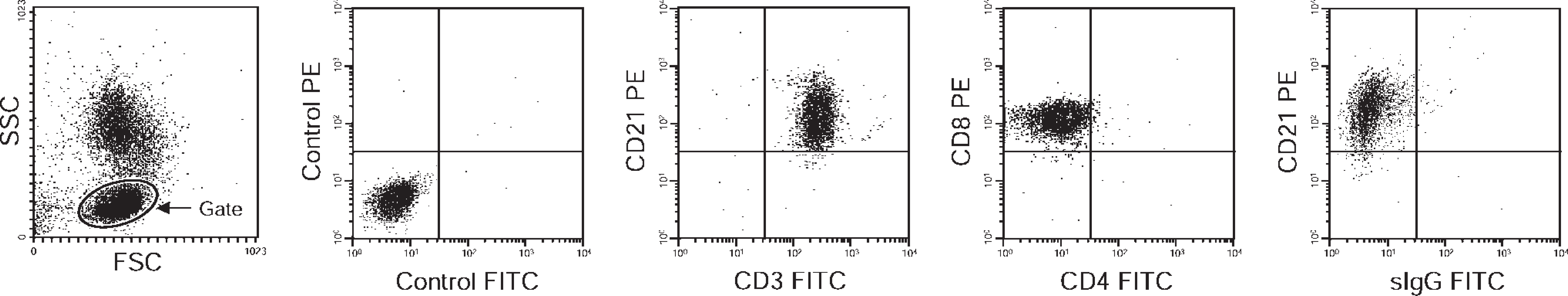
Representative results of flow cytometric analysis in a case with the CD3+ CD21+ phenotype in lymphocytes (case 28). Lymphocytes were gated (indicated by circles) on a forward-scatter (FSC) versus side-scatter (SSC) plot. Cells were simultaneously analyzed with fluorescein-isothiocyanate (FITC)- or phycoerythrin (PE)-labeled monoclonal antibodies against CD3, CD4, CD8, CD21, or surface immunoglobulin G (sIgG), as indicated beside each dot plot. FITC- or PE-labeled mouse IgG was used as a negative control (indicated as control FITC and control PE).
Regarding the correlation of immunophenotyping and clonality analysis, concordance of these analyses was found in cases 1–21. In lymphocytes expressing normal and aberrant T-cell antigens, clonality was detected by the TCRγ gene. In lymphocytes expressing B-cell antigens, clonality was detected by the IgH gene. Two cases were exceptions (cases 14 and 20) because clonality was not detected. Among the cases in which clonality was not detected, lymphocytes from case 20 represented the CD3– CD8+ phenotype. A similar phenotype has been reported in a subset of human natural killer (NK) cells. 10 Although the neoplastic lymphocytes in case 20 did not have large granular lymphocyte morphology, which is typically seen in NK cells, the lymphocytes could have originated from a similar subset of NK cells. In the current study, cases that express both T- and B- cell antigens on the lymphocytes (cases 22–30), namely, coexpression of CD3 and CD21, were frequently found (9 cases; 29% of all cases). The clonality of the lymphocytes was detected in 7 of 9 cases; 6 cases were detected using the TCRγ gene primers, and 1 case was detected using the IgH gene primers. CD3 and CD21 coexpression has been detected in dogs with lymphoma. 13 Similarly, a 2008 study 6 also reported a case of lymphoma with the CD3+ CD21+ phenotype, but a similar finding was not reported in other studies of dogs with lymphoid neoplasia. 1,14 In cases of canine lymphoma reported previously, 13 the coexpression of CD3 and CD21 had been shown in nodal lymphocytes from 4 of 59 dogs. Among these 4 cases, 1 had a TCRγ clonal gene rearrangement and 2 had an IgH clonal gene rearrangement. In the 2008 report, 6 1 case with the CD3+ CD21+ phenotype was found among 16 cases of T-cell lymphoma. This case had a clonal rearrangement in the TCRγ gene but not in the IgH gene. Based on observations in the present study, coexpression of CD3 and CD21 of T- and B-cell lineages would not be an uncommon finding in small-cell lymphoid neoplasia in dogs. However, T-cell lineage appears to be found more frequently. Expression of lymphocyte markers was not detected in 1 case (case 31), although this case was not examined for the surface expression of IgG. Because clonality of the IgH gene was detected in case 31, the lymphocytes might be an aberrant B-cell phenotype that lacked surface expression of CD21. Biclonal patterns in PCR analysis have been reportedly produced by biallelic gene rearrangement of the TCRγ loci in human cases with T-cell neoplasms. 16 The biclonality observed in 3 cases in the present study was assumed to arise by a similar process and reflected the neoplastic nature of the lymphocytes.
In the present study, immunophenotyping was effective in demonstrating neoplastic expansion of small lymphocytes. Moreover, immunophenotyping has recently been advocated as a clinical prognostic indicator in lymphoproliferative disorders. 14 Compared with immunophenotyping, the detection rate of clonality by PCR analysis was unsatisfactory and needs to be improved by identifying more segments encoding the TCRγ and IgH genes. However, from the present study, PCR analysis was considered to be a valuable method to strengthen the evidence of small-cell lymphoid neoplasia in combination with immunophenotyping. Furthermore, PCR analysis of gene rearrangement could be an alternative diagnostic tool when immunophenotyping is not available.
Acknowledgements
This research was supported partially by a Grant-in-Aid for Scientific Research (No. 18580323) and “Academic Frontier” Project for Private Universities: matching fund subsidy (2005–2009) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT).
Footnotes
a.
Celltac MEK-6450, Nihon Kohden, Shinjuku, Tokyo, Japan.
b.
FACSCalibur™, BD, Franklin Lakes, NJ.
c.
BD Biosciences, San Jose, CA.
d.
AbD Serotec, Oxford, UK.
e.
Rockland Immunochemicals Inc., Gilbertsville, PA.
g.
DNeasy® Tissue Kit, Qiagen Inc., Valencia, CA.
h.
Invitrogen Corp., Carlsbad, CA.
