Abstract
Bovine viral diarrhea (BVD) is one of the economically important diseases of cattle. For many years, different types of vaccines have been commercially available, yet this disease is hard to control in high-density population areas. Detection and isolation of bovine viral diarrhea virus (BVDV) from any potential reservoir is vital, especially when considering virus eradication from a herd or locale. One potential source is wild ruminants. Ear notches and lymph nodes were collected from the wild population of white-tailed deer (Odocoileus virginianus) during deer hunting season in Indiana and tested for BVDV with a commercial BVD antigen capture enzyme-linked immunosorbent assay. Two samples out of 745 collected samples were positive, and subsequently cp and ncp BVDV was isolated from 1 ear notch and 1 lymph node. These isolates were genotyped as type 1a and 1b based on sequence analysis of the 5′ untranslated region (UTR). The results of the present study indicate that the prevalence of BVDV in the white-tailed deer population of Indiana is about 0.3%. Wild ruminants infected with BVDV should be taken into consideration during an eradication program of BVDV from the livestock population.
Bovine viral diarrhea virus (BVDV) is a positive, single-strand RNA virus belonging to the family Flaviviridae, genus pestivirus. 11,13,23 Genotyping of pestiviruses can be done using nucleotide analysis of the 5′ untranslated region (UTR) region). 12 There are 3 different known species of viruses within the genus of pestiviruses: BVDV types 1 (BVDV1) and 2 (BVDV2), border disease virus (BDV), and classical swine fever virus (CSFV). BVDV1, BVDV2, and BDV have been shown to infect cattle, swine, sheep, and goat species, while CSFV infects only swine. 20,26 BVDV causes a variety of clinical manifestations in cattle including clinically unapparent to severe acute infections, neonatal diarrhea in calves, 27 and predisposition of calves to respiratory tract infections. 1,4 Both genotypes of BVDV have long been associated with economic losses in the cattle industry. 5 The most important economic consequences of BVDV, however, are reproductive losses. Depending on the time of infection during gestation, BVDV can cause early embryonic death, abortion, congenital defects, or a variety of clinical manifestations including fever, nasal and ocular discharge, profuse watery diarrhea, and mucosal lesions. 10 Most important, fetuses between 18 and 125 days of gestation that survive infection with noncytopathic strains of BVDV invariably develop immune tolerance to the virus and are born persistently infected (PI). 10 PI animals shed large amounts of virus during their lifetime and are the primary spreaders of the virus in the herd. It has been postulated that the key for BVDV control is to interfere with the process by which these PI individuals are generated 15 or to detect and remove these PI animals from herds. BVDV also can be spread to susceptible calves from acutely infected animals. 14
Furthermore, in some areas of the United States, a marked increase in the prevalence of fetal infections was recorded between 1980 and 2000 despite a 60% rate of vaccination. 9 The combined economic impact of BVDV has been estimated at $20 to $57 million loss per million calves in the United States. 8 In 2005, the cost of BVD to U.S. producers was estimated at $760 million to $2.2 billion dollars. 8
Risk factors for BVDV introduction into a noninfected herd have been extensively studied. The introduction of infection through purchased animals appeared to be the most important risk factor. 14 It was concluded from several studies that many wild ruminant species can be infected with BVDV and serve as a potential reservoir for infection to the livestock population. 14 In 2006, the American Association of Bovine Practitioners and National Cattlemen's Beef Association adopted a resolution for eradication of BVDV. 8 In order for the eradication program to be successful, the source of infection needs to be understood in order to prevent reintroduction of BVD into cattle herds that are free of infection. An epidemiologic study was conducted to evaluate BVD infection in wild white-tailed deer (Odocoileus virginianus) in Indiana.
In the fall of 2006, 311 ear notches and 434 lymph node samples were collected across Indiana (Fig. 1) from hunter-harvested deer during the Indiana firearm deer-hunting season. Ear notches were collected in 5-ml snap cap tubes, and lymph nodes were collected fresh in bags and transported to the laboratory. Ear notches, 1 cm in size, were resuspended in 2 ml of phosphate buffer solution (PBS), and 10% of tissue homogenate was made in PBS following procedures recommended by the manufacturer. a A total amount of 4 ml of tissue homogenate was centrifuged at 2,010 × g for 20 minutes. Supernatant from each individual sample of all ear notch and lymph node samples was tested for BVDV antigen using a commercial BVD antigen capture enzyme-linked immunosorbent assay (ELISA) a according to the manufacturer's recommendations.
One positive sample from lymph node and 1 positive ear notch from 2 different white-tailed deer were further confirmed by virus isolation, polymerase chain reaction (PCR), and sequence analysis of positive PCR products. Presence of BVDV in the supernatant from 1 positive ear notch and from 1 lymph node were confirmed directly by reverse transcription (RT)-PCR. 26 Two different genotypes of BVDV were isolated from 1 positive ear notch and 1 lymph node of deer, originating from the locations shown on the map (Fig. 1). In brief, Madin Darby bovine kidney (MDBK) cells were prepared in 48-well plates grown in 5% (vol/vol) horse serum, b 20 mM L-glutamine, c and an antibiotic-antimycotic mixture that consisted of 100 IU/ml penicillin, 10 μg/ml streptomycin, and 50 μg/ml gentamicin in minimum essential media (MEM). b Samples (0.2 ml/well) were inoculated in duplicate or triplicate on cell suspension and left for 24 hours before being removed and replaced with fresh 5% (vol/vol) horse serum MEM. Inoculated cells were observed daily for cytopathic effect (CPE). When CPE was evident in more than 70% of the cell monolayer, cell culture media were harvested and inoculated onto freshly prepared MDBK cells in duplicate (second passage). At day 2 postinoculation, cells on duplicate 48-well plates were fixed after cell culture media were harvested by immersing them in cold 80% aqueous acetone, and subjected to immunofluorescence microscopy using fluorescein isothiocyanate (FITC)-labeled polyclonal antibodies d raised against BVDV. Inoculated duplicate plates with tissue homogenate were stained with BVD-specific monoclonal antibodies #20.10.06 as described 6 at 4 days postinoculation, and a positive signal was observed under a fluorescence microscope; uninoculated wells served as negative control during all procedures. Isolated viruses were plague purified as described earlier. 17
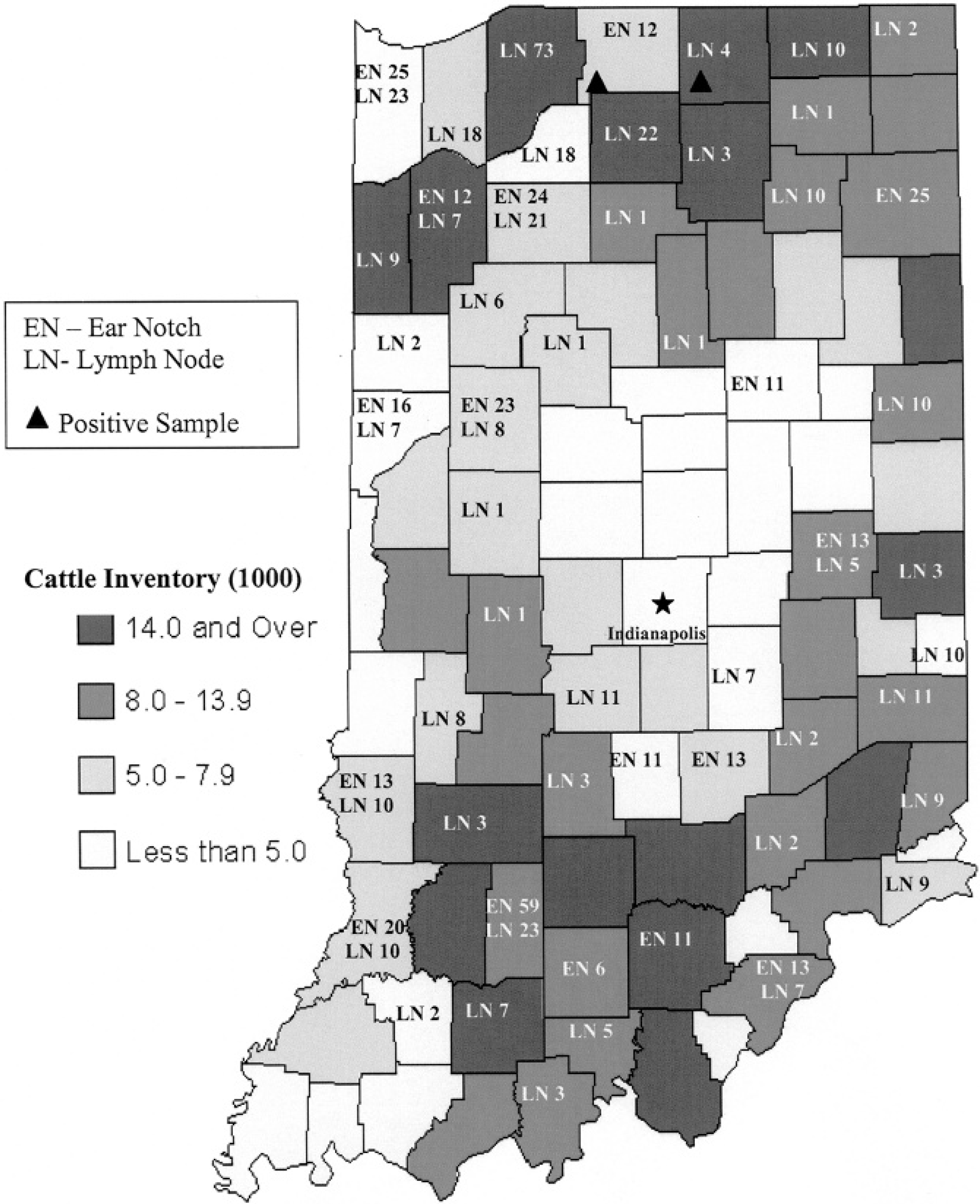
Cattle population inventory and positive and negative wild white-tailed deer samples distribution collected in year 2006 in the state of Indiana.
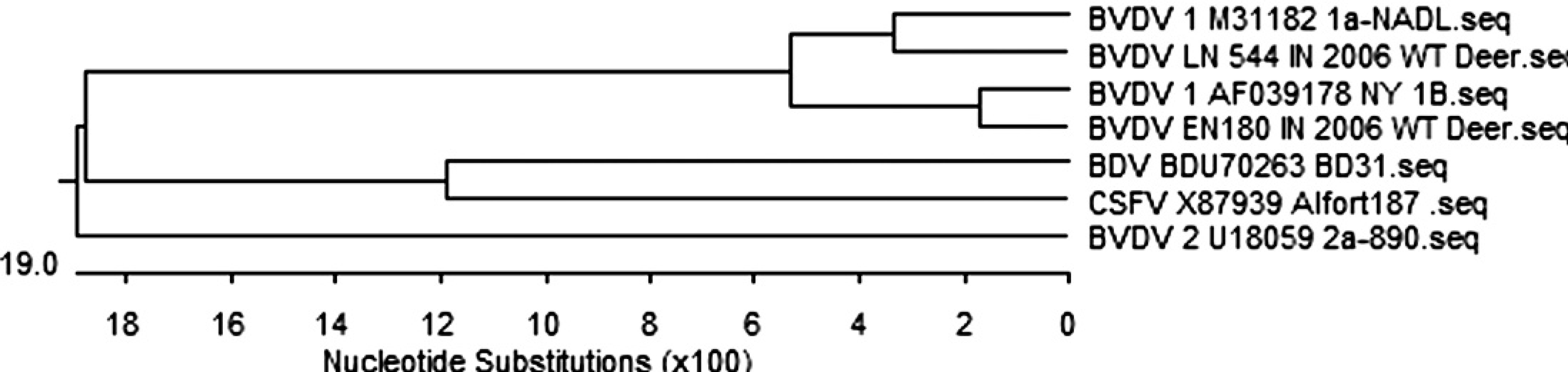
Phylogenetic tree of pestiviruses sequenced at 5′UTR on isolates from white-tailed deer in comparison to reference bovine viral diarrhea virus constructed by DNASTAR program, Clustal W method.
The 2 cytopathic (cp) and 2 noncytopathic (ncp) BVDV isolates from the ear notch and lymph node were amplified and genotyped by RT-PCR as described. 26 In brief, 140 μl of the supernatant were used for RNA extraction using the QIAamp Viral RNA Mini Kit e according to manufacturer's instructions. RT-PCR was carried out with QIAGEN One Step RT-PCR Kit. e Five microliters of the RNA template were added to 45 μl of the RT-PCR master mix. PCR was performed using a set of primers corresponding to the conserved region at 5′UTR and cycling condition as previously described. 21,25
The amplified PCR products with molecular weight 288 base pair (bp) were purified using QIAGEN purification kit c according to the manufacturer's protocol. These products were sequenced using DNASTAR an automated sequencer f at the Purdue genomic core facilities and then analyzed, and their homology to other BVD viruses was determined based on published sequence information and reference control virus. Analysis was done using DNASTAR computer software. g The 2 BVDV isolates from deer were genotyped as type-1a and type-1b BVDV with sequence homology 89.2% to 97% on nucleotide level at 5′UTR in comparison with BVD reference viruses available, and a phylogenetic tree was developed (Fig. 2). There were no differences on nucleotide level at 5′UTR between 2 bio types 1a and 2 biotypes 1b of BVDVs. Based on the results of the samples from hunter-harvested deer, 0.26% of wild deer were estimated to be positive for BVDV in Indiana with 95% confidence level (CI) with range for true population proportion (0.1 < 95%; CI > 0.64). A similar prevalence, speculated to be around 0.6% of BVDV in the Indiana cattle population, was estimated based on samples tested for BVDV in the Indiana Animal Disease Diagnostic Laboratory with 95% CI and range for true population proportion (0.57 < 95%; CI > 0.69; R. Pogranichniy and E. Raizman, personal communication) and other laboratory data. 14
The present report describes isolation and characterization of BVD type 1a and type 1b cp and ncp viruses from the free-range white-tailed deer population in Indiana. It is not possible to determine if the animals were persistently or acutely infected because only limited samples were collected from dead deer, and cp and ncp viruses were isolated. Since BVD isolates were cp and ncp biotypes, it is more likely that the deer were persistently infected animals. The effect of BVDV on the wild white-tailed deer population is not clear, but it is suspected that these animals may develop mild clinical disease such as that described for cattle. 3,22 Since prevalence of BVDV in deer is low, it is likely that cattle serve as the potential reservoir for infection in deer. 25 Pasture fence contact and “other animal traffic” were also identified as risk factors. 24 Direct or indirect contact of infected animals or populations, wild or domestic, with susceptible populations at the interface of their ranges is an important factor responsible for the cause of outbreaks of disease. 2 Evidence for BVD infections of free-ranging populations includes serologic surveys and virus isolations. 16,24 Although serologic surveys in free-ranging and captive populations demonstrate prior infection with BVDV or related pestiviruses in more than 40 species, isolations of BVDV from wild ruminants are few. 7,16,25 BVDV in deer was studied under experimental conditions, and white-tailed deer PI doe was experimentally reproduced with the ncp BVDV 1 strain BJ and the ncp BVDV 2 strain PA131. 18 There is a potential risk of wild animals introducing BVDV infection to livestock. When wildlife species act as disease reservoirs for domestic livestock they can seriously undermine the effectiveness of any disease control strategy in domestic species by escalating the number of susceptible animals in which the disease can persist, especially at the wildlife-livestock interface. 19
White-tailed deer and cattle do share common pastures 19 and might introduce infection to each other. During future eradication programs in the livestock population, it is important to take into consideration the wildlife reservoir for BVDV as one potential source of infection into the domestic ruminant population.
Acknowledgements. The authors thank the Department of Natural Resources of Indiana for cooperation on this project, with special recognition of Dr. Jim Mitchell; students of Purdue University School of Veterinary Medicine, especially Lisa Keefe; and the Animal Disease Diagnostic Laboratory virology personnel for helping with collection of samples and technical assistance.
Footnotes
a.
IDEXX HerdChek ELISA, IDEXX Laboratories, Westbrook, ME.
b.
Sigma Chemical Co., St. Louis, MO.
c.
Invitrogen/Gibco, Carlsbad, CA.
d.
American Bioresearch, Inc., Seymour, TN.
e.
Qiagen Inc., Valencia, CA.
f.
Applied Biosystems, Forest City, CA.
g.
DNASTAR, Madison, WI.
