Abstract
Boar studs are continuously monitored for the presence of porcine reproductive and respiratory syndrome virus (PRRSV) by testing different biological samples by reverse-transcription polymerase chain reaction (RT-PCR). In most cases, samples are run in pools, even though the impact of pooling on the sensitivity of RT-PCR is unknown. The objective of this study was to evaluate the feasibility of using PCR on pooled samples through the estimation of the sensitivity of RT-PCR on different biological samples run individually, in pools of 3 and in pools of 5. Twenty-nine boars were inoculated with a low virulent PRRSV isolate. Serum, blood swab, and semen samples were obtained from each boar every 2 to 3 days for 2 weeks. Each sample was tested by RT-PCR undiluted or diluted 1:3 and 1:5 with negative samples. Eleven of the 29 boars did not appear to get infected from the inoculum, as evidenced by no seroconversion 15 days after inoculation. Data from the other 18 boars showed that serum was the best sample to detect PRRSV during acute infection, with the blood swab sample performing almost as well. Semen samples failed to detect PRRSV infection in most of the cases. Pooling samples at pool sizes of 3 and 5 resulted in a decrease in the sensitivity of RT-PCR. Sensitivity was reduced by 6% and 8%, respectively, when serum or blood swab samples were run in pools of 5. The impact of pooling on the sensitivity of PCR was higher in samples taken during the beginning of the viremic period.
Introduction
Porcine reproductive and respiratory syndrome (PRRS) is one of the most important diseases of swine worldwide. It is caused by the PRRS virus (PRRSV), a positive-sense, single-stranded enveloped RNA virus from the family Arteriviridae. Infection with PRRSV is characterized by reproductive disorders in sows, respiratory distress in young pigs, and no or mild symptoms in boars. 27
Because the virus can be transmitted through artificial insemination with semen from infected boars, 9,26 it is critical that boar studs remain free of PRRSV infection. Since PRRSV surveillance in boar studs cannot rely on clinical signs, a routine sampling protocol is needed to detect any potential introduction of PRRSV in the boar stud. Currently, serum, semen, and/or blood swab 20 samples of most boar studs in the United States are sent to diagnostic laboratories to be tested for the presence of PRRSV by reverse-transcription polymerase chain reaction (RT-PCR). Field experiences suggest that an intensive monitoring protocol, testing a large number of boars by RT-PCR and at a high frequency, is needed to detect a PRRSV introduction before potentially contaminated semen is shipped to PRRS-negative sow farms. Because of the high cost of such a protocol, producers are testing pooled samples. Pooling is a strategy that allows testing a larger number of animals while running the same number of tests, thus keeping the same testing costs. This strategy can successfully be applied if the test has a high analytical sensitivity. 23 However, because of a dilution effect, the sensitivity of the test when run on pooled samples is lower than its sensitivity when run on individual samples. 16 Therefore, an improved understanding of the impact of dilution caused by pooling on the monitoring protocols for boar studs is needed. The objective of this study was to evaluate the feasibility of using RT-PCR on pooled samples to monitor boar studs. This was evaluated through the estimation of the sensitivity of RT-PCR to detect PRRSV in different biological samples tested individually, in pools of 3 and in pools of 5.
Materials and methods
Animals
Twenty-nine boars were obtained from a PRRSV-negative boar stud. They were housed in individual stalls (25 boars) or in individual pens (4 boars) in a mechanically ventilated research barn at the University of Minnesota. The age of the boars ranged from 10 to 42 months, and they were from 7 different genetic lines. After their arrival at the facility, boars were given 9 days to acclimate to the new environment before beginning the study. Animals were housed and cared for according to the guidelines of the University of Minnesota Institutional Animal Care and Use Committee.
Virus inoculation
All boars were inoculated intranasally with 4 ml of an inoculum for a total dose of 104.6 TCID50 of the PRRSV field isolate MN 30–100. This isolate is considered of low virulence and has been found to result in lower amounts of PRRSV genome copies in the serum of infected pigs, compared to a clinically more virulent strain. 2
Sample collection and processing
All boars were sampled (serum only) 3 days before PRRSV inoculation to confirm their PRRSV-negative status. Subsequently, all boars were sampled at 1, 3, 5, 8, 10, 12, and 15 days postinoculation (DPI). At each sampling time, samples of semen, serum, and blood swab were collected from each boar. For semen collection, boars were allowed to mount a dummy, and semen samples were collected with the gloved-hand technique. Immediately after semen collection, 1 aliquot of the semen sample was diluted 1:5 with a commercial semen extender a kept at a constant temperature of 36°C, whereas another aliquot was kept undiluted. Blood for serum samples was collected by venipuncture of the jugular vein. Blood swab samples were collected as previously described 19 with some modifications. To minimize the dilution effect inherent to this technique, blood was collected with a swab b that can absorb 280 μl of liquid and diluted in a tube with 500 μl of saline solution.
All samples were run undiluted and diluted 1:3 and 1:5 to simulate pools of 3 and 5, respectively. All serum samples were diluted with stored serum from a PRRSV-negative animal. Dilution of semen and extended semen samples was performed with semen and extended semen of boars randomly chosen from a PRRSV-negative boar stud. Dilution of blood swab samples was performed with saline solution.
RNA extraction
Viral RNA was extracted from serum and blood samples using a commercial RNA isolation kit c and following the manufacturer's instructions. 25 For semen samples, a commercial RNA isolation column d was used for RNA isolation and purification after a modified cell-lysate homogenization protocol was performed. 4 Specifically, after centrifugation of the raw or extended semen at 660 χ g for 15 min, the seminal cell fraction was obtained by pouring off the supernatant and resuspending the pellet in an equal volume of seminal plasma (extended semen) or phosphate-buffered saline (raw semen). Five hundred microliters of seminal cell fraction was added to 500 μl of guanidium lysis buffer (4 M guanidine thiocyanate, 25 mM sodium citrate [pH 7], 0.5% sarkosyl [N-lauryl sarcosine]), as previously described. 3 Five hundred microliters of this mixture was added to a commercial cell-lysate homogenization column. e The column was centrifuged at a maximum speed of 16,100 χ g for 2 min, and 500 μl of the eluate was removed and combined 1:1 with 70% ethanol. The total 1:1 mixture was then placed on the RNA isolation column, d and the protocol was followed as per the manufacturer's directions with a final elution volume of 30 μl.
Reverse-transcription polymerase chain reaction
Polymerase chain reaction testing was conducted using a commercially available real-time, single-tube, RT-PCR assay f for the detection of US PRRSV. A minor groove binding 5’ nuclease probe and primers were designed from the 3'UTR PRRSV genomic region by alignment of GenBank isolates and based on conserved areas of the 3'UTR primer and probe region. Reverse transcription and PCR were performed in a single tube using a 25 μl reaction volume consisting of 18.9 μl Master mix, 2 μl enzyme mix 1, 0.1 μl enzyme mix 2, and 4 μl of extracted RNA. 25 The reaction tubes were loaded into a real-time PCR instrument, g and software settings of fluorescent detection were set for automatic calculation of the baseline with the background subtraction on. The thermal cycler program consisted of 52°C for 1800 sec, 95°C for 900 sec, and 45 cycles at 94°C for 30 sec, 61°C for 60 sec, and 72°C for 60 sec. The commercial RT-PCR kit f used in this study defines a positive sample as having a cycle threshold (Ct) level ≤37 cycles and a suspect positive sample as having a Ct level between 37 and 39 cycles. For this study, only samples with a Ct level ≤37 were considered positive.
Quantitation of the amount of PRRSV in serum and blood swab samples was performed as previously described. 25 Briefly, known amounts of serially diluted in vitro transcript RNA product (10−1 through 108 copies/μl) were used to generate a standard curve. Copy/ml concentrations of the unknown samples were determined by linear extrapolation of the Ct values plotted against the known concentration of the transcript product.
Enzyme-linked immunosorbent assay
Serum samples were tested for the presence of PRRSV-specific antibodies with a commercial enzyme-linked immunosorbent assay (ELISA) test h following the manufacturer's directions.
Statistical analysis
An estimate of the diagnostic sensitivity was obtained for each combination of sample (serum, blood swab, non-extended semen, or extended semen) and pool size (undiluted, pool of 3, and pool of 5) at each collection time. The sensitivity estimates were calculated as the number of samples that tested positive for each specific test and pool size on a given collection day divided by the number of samples tested.
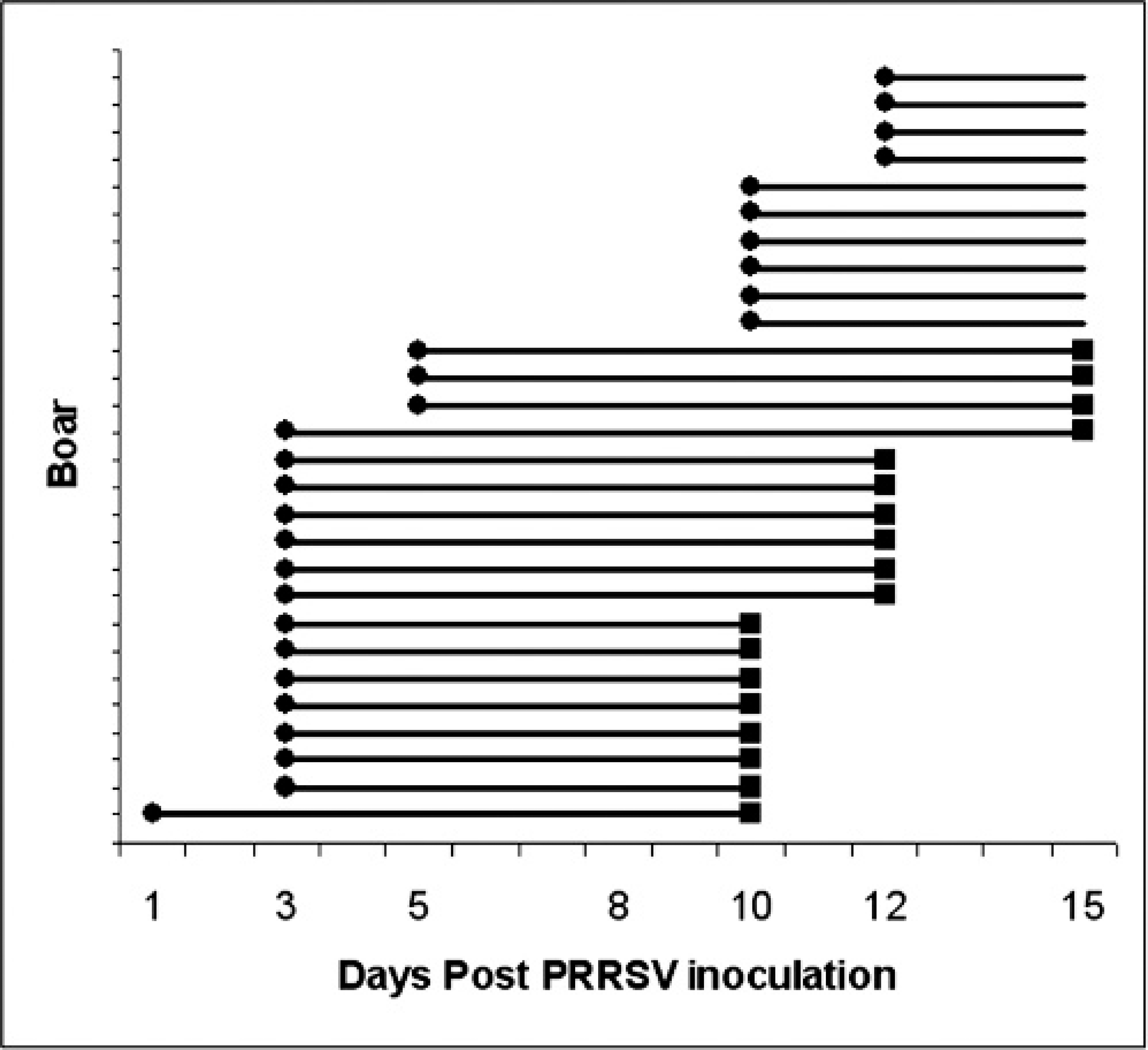
Time line of detection of PRRSV infection in experimentally infected boars by RT-PCR and ELISA on serum samples. The time from the first RT-PCR positive result to the first ELISA positive result is represented by a horizontal line for each boar.
Furthermore, an overall estimate of the sensitivity of pooled serum and blood swab samples was obtained regardless of the collection day. To calculate this estimate, the result of RT-PCR on undiluted serum was used as a gold standard. Therefore, the sensitivity estimates were calculated as the number of samples that tested positive for a given test and pool size divided by the number of samples that tested positive by RT-PCR on serum undiluted. Because samples taken during the first days of viremia might be more likely to result in false-negative results when tested in pools, a second sensitivity estimate was obtained using only the samples from the first collection that tested positive by RT-PCR on serum from each boar. The 95% confidence intervals around sensitivity estimates were calculated using the Jeffreys interval method. 1
A paired t-test was used to compare the average amounts of PRRSV in serum and blood swab samples at each collection day. For the interpretation of the test, the 1-tailed P value was used at a significance level of 0.05, applying the Bonferroni correction for multiple comparisons.
Results
Confirmation of PRRSV infection
Sera from all boars tested negative for the presence of PRRSV and PRRSV-specific antibodies 3 days before inoculation by RT-PCR and ELISA, respectively. Only 18 of the 29 boars had seroconverted by the end of the study (15 DPI). Eight of those boars seroconverted at 10 DPI, 6 seroconverted at 12 DPI, and 4 seroconverted at 15 DPI (Fig. 1). ELISA-positive results were observed in 2 boars before 10 DPI. One boar tested ELISA positive at 3, 5, and 8 DPI, and another boar tested positive at 3 DPI. Because false-positive results are not uncommon with this ELISA test, an indirect fluorescence antibody assay (IFA) was performed on these serum samples as a confirmatory test. 17 The IFA test was negative for all 4 sera; therefore, the sera were considered negative for the presence of PRRSV-specific antibodies.
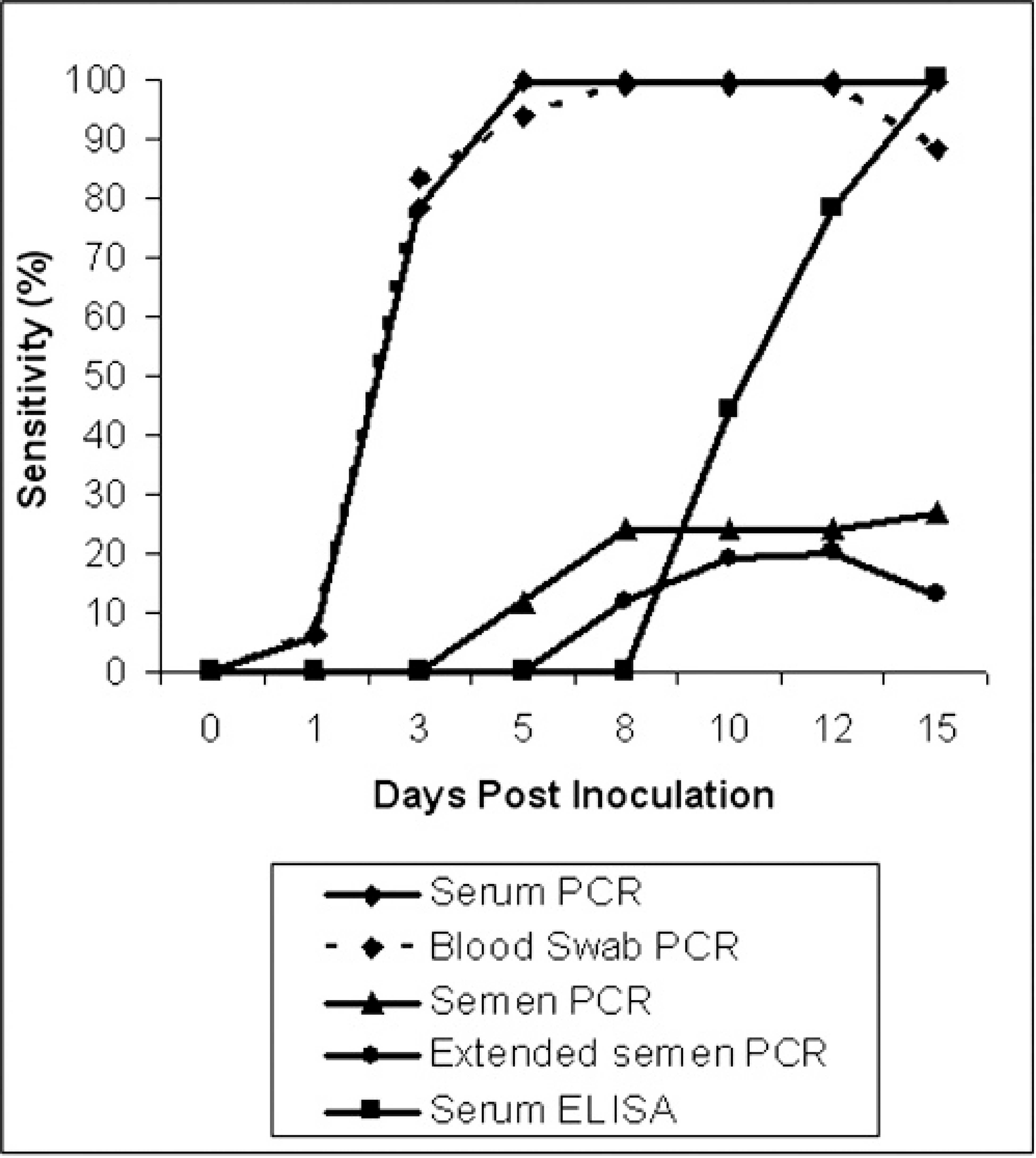
Comparison of the change in sensitivity of RT-PCR to detect PRRSV on different samples from experimentally infected boars over time.
The 18 boars that seroconverted tested positive by RT-PCR on serum for the first time at 1, 3, or 5 DPI (Fig. 1) and were positive thereafter until the end of the study (Fig. 2). Of the 11 boars that did not seroconvert, 10 were found positive by RT-PCR on serum at 10 or 12 DPI and remained positive thereafter. The remaining boar tested negative by PCR on serum through 15 DPI.
PCR sensitivity estimates over time
To obtain sensitivity estimates for specific times post-PRRSV infection, only data from the 18 boars that seroconverted by 15 DPI were used. The sensitivity estimates and their 95% confidence intervals are summarized in Table 1 and Fig. 2.
Serum. Serum was the most sensitive sample in this study. This sample always performed better than any other sample for each boar and sampling time, except in 1 case, in which the serum sample was considered suspect and the blood swab was positive. However, in both samples, the quantity of RNA was very low and near the limit of detection of the assay. The sensitivity estimate for RT-PCR on serum rose from 6% at 1 DPI to 100% at 5 DPI and thereafter (Fig. 2). A decrease in the sensitivity estimate was observed when serum samples were run diluted 1:3 and 1:5 at most of the sampling times (Table 1). The average amount of PRRSV in serum quickly increased during the first 5 days of infection, peaked at 10 DPI, and slightly decreased thereafter (Fig. 3).
PRRSV RT-PCR sensitivity estimates (SE, in %) and 95% CIs for different samples from experimentally inoculated boars, run at different pool sizes and collected at various times after inoculation.
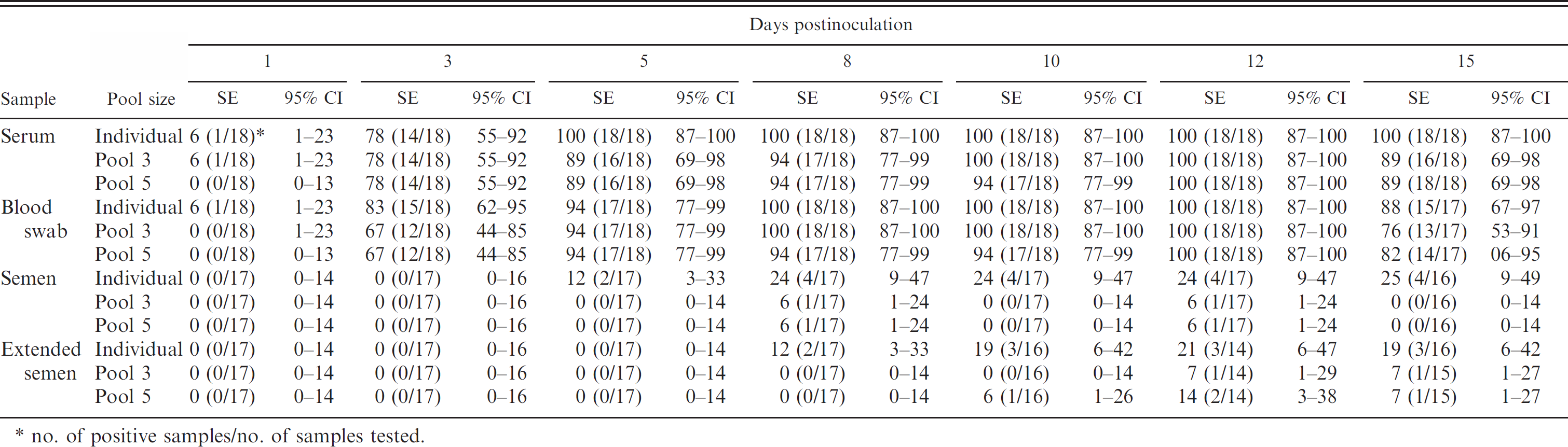
no. of positive samples/no. of samples tested.
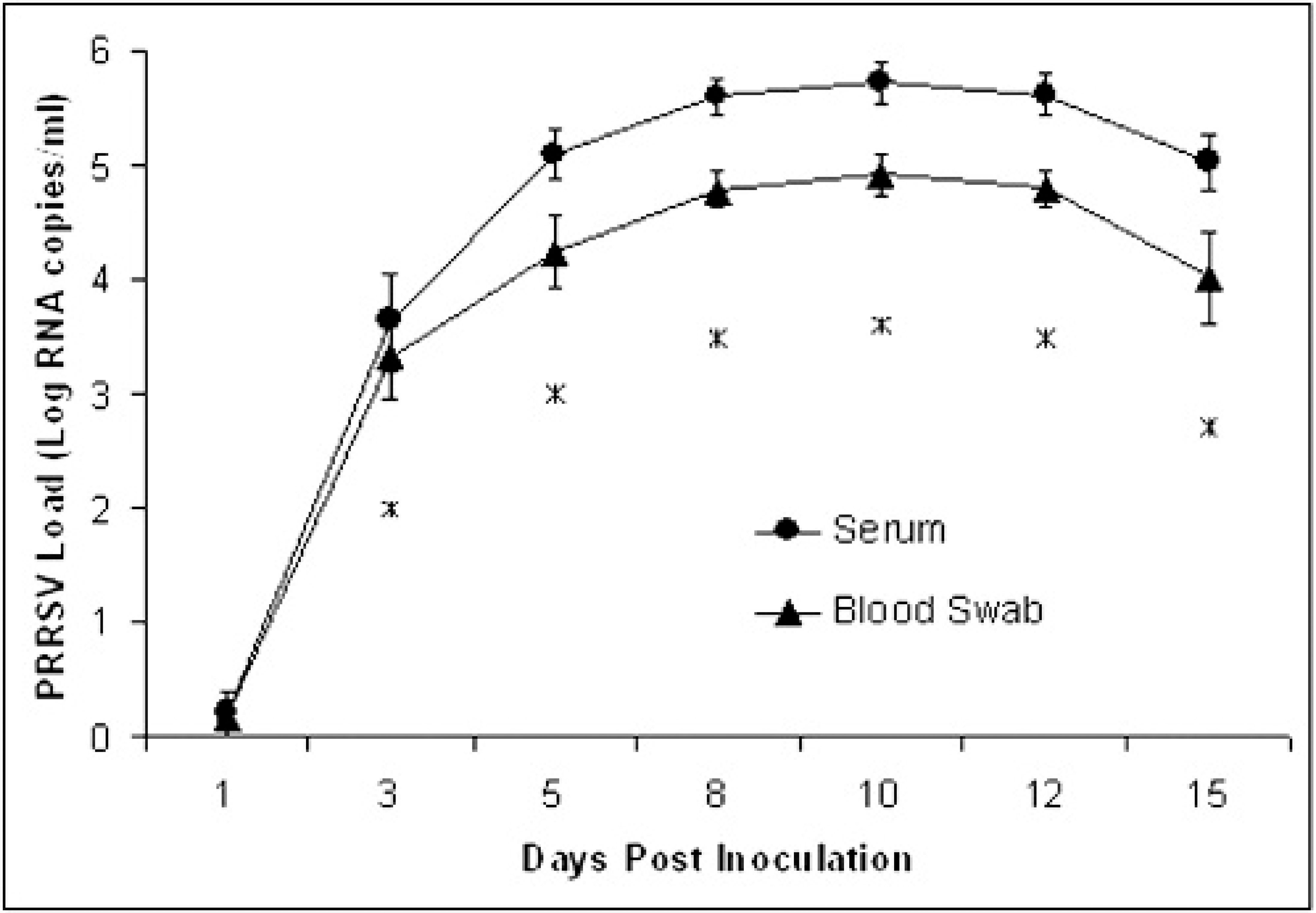
Comparison of the PRRSV load in serum and blood swab samples of boars experimentally infected with PRRSV. The average PRRSV load of 18 boars (Log RNA copies/ml) plus/minus the standard error is represented. The differences that were statistically significant based on a paired t-test are marked with an asterisk.
Blood swab. The blood swab sample could not be collected from 1 animal at 15 DPI. Testing of blood swabs performed similarly to the serum samples. The sensitivity estimate for RT-PCR on blood swab was 6% at 1 DPI, reached 100% at 8 DPI, and dropped to 88% at 15 DPI (Fig. 2). The sensitivity decreased when blood samples were run diluted 1:3 and 1:5 (Table 1). Blood swab samples contained smaller amounts of PRRSV than serum samples at each collection time (Fig. 3). This difference was statistically significant at all collection times except 1 DPI.
Nonextended semen. Semen samples could not be collected from 1 boar, which had to be treated for lameness and refused to mount the dummy throughout the study. Four other samples could not be collected from 4 different boars because of lack of libido, 3 of them at 3 DPI and 1 at 15 DPI. Of the 17 boars from which samples were collected and that were viremic by 5 DPI, PRRSV was detected in nonextended semen from only 7 boars. The first positive samples were detected in 2 boars at 5 DPI, in 2 other boars at 8 DPI, in 1 boar at 10 DPI, and in 2 boars at 15 DPI. Sensitivity estimates went from 12% at 5 DPI to 27% at 15 DPI (Fig. 2). Pooling of positive semen samples with negative semen samples in pools of 3 or 5 resulted in a decrease in sensitivity at each sampling time (Table 1).
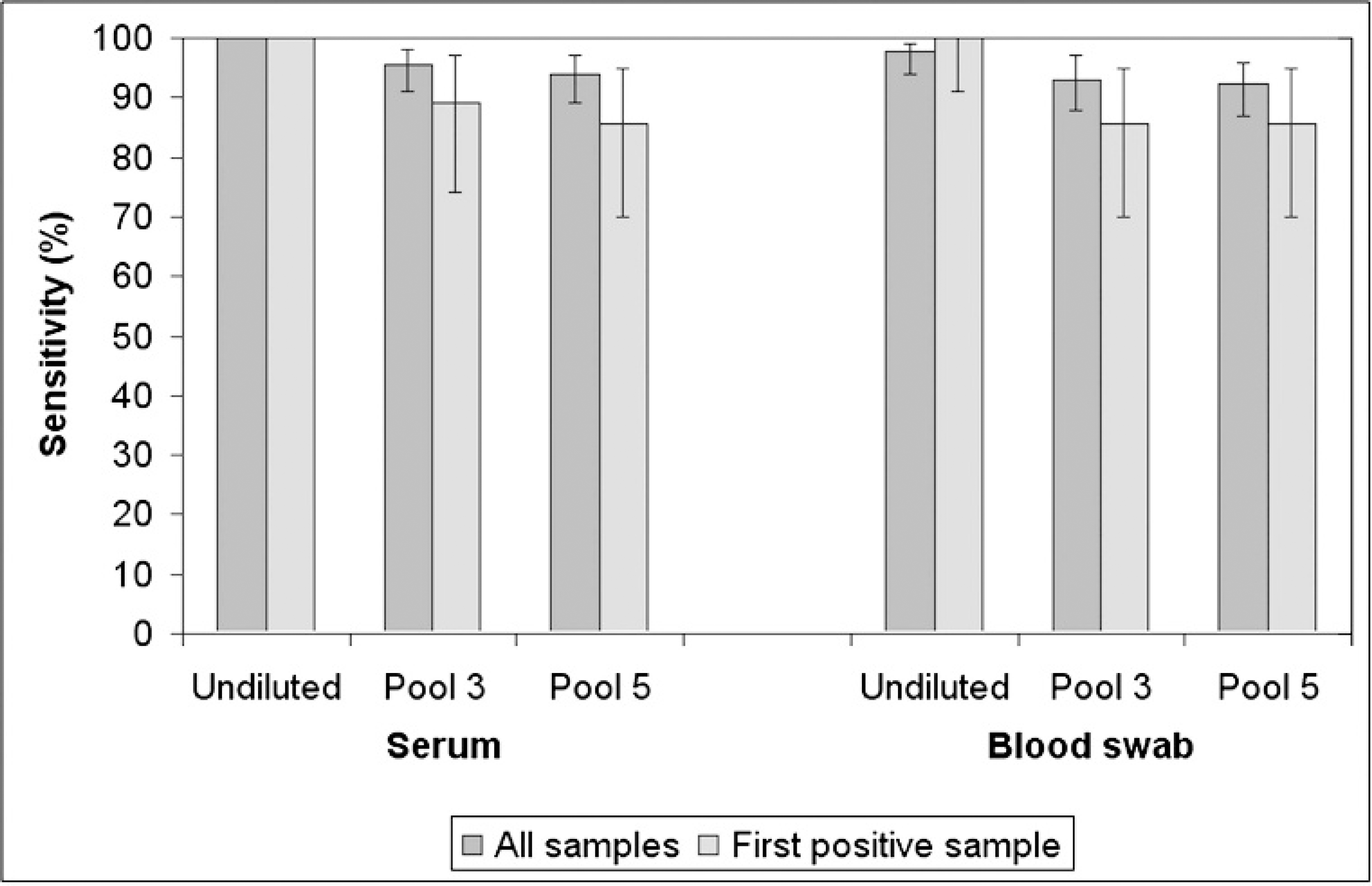
Effect of pooling on the sensitivity of RT-PCR to detect PRRSV on serum and blood swab samples from experimentally infected boars using RT-PCR on serum as the gold standard. The dark bars represent the sensitivity estimates calculated using all the samples that tested positive by RT-PCR on serum (130 samples from 28 boars). The light bars represent the sensitivity estimates including only samples from the first collection that tested positive for each boar by RT-PCR on serum undiluted (28 samples from 28 boars). The error bars represent the 95% confidence interval around each sensitivity estimate.
Extended semen. Extended semen samples proved less sensitive than nonextended semen samples. The first positive samples were detected at 8 DPI, with estimated sensitivity of 12%. After that, sensitivity estimates were around 20% until the end of the study (Table 1; Fig. 2).
Since RT-PCR on serum undiluted was the most sensitive way to detect PRRSV in this study, the data were analyzed using RT-PCR on serum undiluted as a gold standard. For this data analysis, all the samples that were positive by serum RT-PCR when run undiluted were included. Therefore, a total of 130 samples from 28 boars and 7 sampling times were used. Furthermore, a subset of the samples including only the first positive sample for each boar (a total of 28 samples) was analyzed independently, under the assumption that these samples are more likely to result in false-negative results when pooled.
The sensitivity of serum samples in pools of 3 and 5 was 95% (95% confidence interval [CI], 91–98) and 94% (95% CI, 89–97), respectively (Fig. 4). The sensitivity of blood swab when run individually, diluted 1:3, and diluted 1:5 was 98% (95% CI, 94–99), 93% (95% CI, 88–97), and 92% (95% CI, 87–96), respectively (Fig. 4).
When only the first positive sample from each boar was included in the analysis, the sensitivity of serum samples in pools of 3 and 5 dropped to 89% (95% CI, 74–97) and 86% (95% CI, 70–95), respectively (Fig. 4). All the first positive samples by RT-PCR on serum tested positive by RT-PCR on blood swab as well; therefore, the sensitivity estimate for blood swab undiluted was 100% (95% CI, 91–100). When blood swab samples were diluted 1:3 or 1:5, the sensitivity estimates decreased to 86% (95% CI, 70–95; Fig. 4).
Discussion
Pooling is a testing strategy that has been used for a number of purposes including detecting disease 14,15 and estimating the prevalence of disease in a population. 12 It allows testing a larger number of animals faster and at a lower cost than individual testing. However, pooling positive samples with negative samples results in the dilution of the analyte to be detected. 16 Therefore, although pooling can be a valuable tool for monitoring disease, its effect on the sensitivity of the test needs to be evaluated. Pooling of serum, blood swab, and/or semen samples is used routinely by PRRSV-free boar studs to monitor PRRSV status by RT-PCR. In this study, sensitivity estimates have been obtained for RT-PCR on different biological samples run individually, in pools of 3, and in pools of 5.
It has been demonstrated that infection with highly virulent PRRSV isolates results in larger amounts of viral copies in serum than infection with attenuated isolates or isolates associated with low virulence. 2,11,24 Therefore, the decrease in sensitivity due to pooling could be influenced by the PRRSV isolate. Positive samples from animals infected with a low-virulent isolate would be more likely to test negative when pooled than those from animals infected with a high-virulent isolate. Consequently, the use of a highly virulent isolate could result in the overestimation of the sensitivity of RT-PCR for pooled samples. Because of the importance of false-negative results in samples from a boar stud, the study was conducted under a worst-case scenario, using a PRRSV strain of low virulence administered intranasally at an inoculation dose lower than those used in previous boar studies. Besides the choice of inoculation strain, dose, and route, other factors that made this study a worst-case scenario were the way the pools were made and the interpretation of the RT-PCR results. The pools were made by combining each of the study samples with known negative samples, therefore assuming that every pool has only 1 positive sample. The RT-PCR kit used in this study considers as suspect any sample giving a Ct value between 37 and 39. 25 To ensure a worst-case scenario, such results were considered as negative in the data analysis of this study. However, only 9 of 386 positive sera and 5 of 374 positive blood swabs were in the suspect category, indicating a very low number of samples in this range, and most of these were in the diluted samples. Under these conditions, the sensitivity estimates of RT-PCR for pooled samples that were obtained could be underestimated, but they are not likely to be overestimated.
A limitation of this study was that 11 of the 29 inoculated boars did not show results consistent with an infection from the inoculum at day 0 (i.e., detectable viremia by 5 DPI and ELISA-detectable antibodies by 2 weeks postinfection). 5,6,8,18,20,22,25 Furthermore, 1 of the boars did not test positive for serum RT-PCR at any sampling time (although it tested positive in serum and tissues by RT-PCR when it was killed at 17 DPI; data not shown), and the other 10 became viremic as detected by RT-PCR at 10 or 12 DPI and remained positive thereafter. Based on this delayed viremia and the fact that boars were housed in the same barn, it can be speculated that these 10 boars were infected later by direct or indirect contact with infectious boars. It is important to note that boars were housed in the same air space, they were allowed nose-to-nose contact with boars housed in the adjacent stalls or pens, and they had contact with secretions (saliva, semen) from other boars at the collection pen. Another possible explanation would be that these boars did become infected from the inoculum but their levels of PRRSV in serum remained under the detection limit of the RT-PCR until 10 or 12 DPI. This hypothesis is regarded as very unlikely because seroconversion is expected between 10 and 15 DPI, and the boars that became viremic at 10 to 12 DPI did not seroconvert throughout the duration of the follow-up (15 DPI). In addition, the quantitative RT-PCR sensitivity has been established to detect virus concentrations as low as 10 TCID50/ml and corresponds to a nested RT-PCR previously validated with the use of a swine bioassay. 7
While most of the experimental inoculations of boars with PRRSV reported in the literature used a dose of about 106 TCID50, 5,6,8,18,20,22,25 in the present study, the boars were inoculated with a lower dose of 104.6 TCID50. This may be the reason why 38% of the boars did not get infected from the inoculum. This theory is supported by a recent study 10 that showed that the infectious doses needed to infect 60% and 90% of the animals by an intranasal inoculation were 104.4 and 106 TCID50, respectively. Those estimates, however, were obtained using data from experimental inoculations of 2-week-old pigs with the VR-2332 PRRSV isolate. In contrast, another study reported that 29 of 40 boars had seroconverted 13 days after intranasal inoculation with 104 TCID50 of PRRSV isolate MN 30–100 and that 36 of the 40 boars tested positive by PRRSV RT-PCR within the first 6 days after infection. 19
To estimate the diagnostic sensitivity of RT-PCR over time, the 18 boars that had seroconverted by 15 DPI and were positive by RT-PCR on serum within the first 5 days after inoculation were considered infected with the inoculum at day 0. It is interesting to note that 3 of the 18 boars had the first positive RT-PCR result on serum at 5 DPI. This longer time to viremia in a few boars is consistent with data from previously published experiments in which either boars or sows were housed individually. In one study, 1 of 5 boars tested negative to RT-PCR on serum 4 days after vaccination with a PRRSV-modified live virus vaccine but tested positive 1 week later and seroconverted. 6 Another study reported 1 of 5 PRRSV-inoculated individually housed sows negative by virus isolation in serum at 7 DPI and positive at 14 DPI by virus isolation and serology. 13 Finally, in a recent study, 1 of 5 gilts challenged with PRRSV and housed in individual rooms tested negative to virus isolation on serum at 3 and 5 DPI but positive at 9 DPI on serum and 5 DPI on nasal swab. 21 Therefore, it is suggested that viremia may not necessarily be observed within the first 3 days after exposure to PRRSV.
Results from this study suggest that serum is the best sample to detect PRRSV in boars during acute infection. Moreover, under the conditions of this study, the blood swab sample performed almost as well as the serum sample. In contrast, the semen sample failed to detect PRRSV infection in most of the cases. These findings are in agreement with findings of previous studies. 19,25 It is important to note that the blood swab technique used in the present study has been modified from the protocol originally described 20 to increase its sensitivity by increasing the swab absorbance and decreasing the volume of the diluent. Therefore, the estimates of sensitivity of RT-PCR on blood swab samples should be applied only in cases in which the same sampling protocol is used.
Pooling samples, at pool sizes of both 3 and 5, resulted in a decrease in the sensitivity of RT-PCR. About 6% of the samples that would be detected by RT-PCR on individual serum would be missed if they were run in pools of 5. Furthermore, about 8% of those samples would be missed if, instead of individual serum samples, blood swab samples in pools of 5 were used. If one takes into account that using pooling would allow sampling a larger number of animals, this decrease in sensitivity might be compensated for by the benefits of a larger sample, especially in the case of a low prevalence of disease. 16 However, the results of this study also show that the decrease in sensitivity due to pooling would be larger in boars during the first 1 to 5 days of infection. In this stage of infection, up to 14% of the RT-PCR-positive individual serum samples would be missed if pools of 5 were used. Whether the benefits of pooling outweigh its disadvantages is a complicated question. Several factors such as the prevalence of PRRSV-infected boars at the time of sampling, the stage of infection of each infected boar, the sample size, and the pool size play a role. Further research should investigate the interplay of these factors to determine the best combination of sample size and pool size to be used for routine monitoring of PRRSV in boar studs.
Acknowledgements
This project was supported by the National Research Initiative of the USDA Cooperative State Research, Education and Extension Service, grant 2004–35605–14197; the Advanced PRRS Research Award from Boehringer Ingelheim; and PIC. We thank Dr. Mike Murtaugh and Dr. Martha Fuentes from the University of Minnesota for the preparation and titration of the inoculum. We also acknowledge the technical expertise of the Molecular Diagnostic Section at South Dakota State University including Matt Dammen, Gina Steinlicht, Simon Althoff, and Becky Hewer.
Footnotes
a.
Safe Cell Plus Extender, IMV International Corporation, Maple Grove, MN.
b.
Sterile rayon-tipped applicator REF. 25–806 1PRLB QUICKVIEW, Puritan Medical products Company LLC, Guilford, ME.
c.
QIAamp Viral RNA Mini-Kit, Qiagen, Inc., Valencia, CA.
d.
RNeasy column, Qiagen Inc., Valencia, CA.
e.
Qiashredder column, Qiagen Inc., Valencia, CA.
f.
Tetracore U.S. PRRSV Master Mix, Tetracore Inc., Rockville, MD.
g.
Smart Cycler II block, Cepheid, Sunnyvale, CA.
h.
HerdChek PRRS 2xR Antibody Test Kit, IDExx Laboratories, Westbrook, ME.
