Abstract
For the easy survey of Marek's disease virus (MDV), feather tip–derived DNA from MDV-infected chickens can be used because feather tips are easy to collect and feather follicle epithelium is known to be the only site of productive replication of cell-free MDV. To develop a diagnostic method to differentiate highly virulent strains of MDV from the attenuated MDV vaccine strain, CVI988, which is widely used, nested polymerase chain reaction (PCR) was performed to detect a segment of the meq gene in feather tip samples of chickens experimentally infected with MDV. In chickens infected with Md5, a strain of oncogenic MDV, the meq gene was consistently detected, whereas the L-meq gene, in which a 180–base pair (180-bp) sequence is inserted into the meq gene, was detected in CVI988-infected chickens. Moreover, the meq gene was mainly detected even in chickens co-infected with both Md5 and CVI988. These results suggest that this method is appropriate for the surveillance of the highly virulent MDV infection in the field.
Introduction
Marek's disease virus (MDV: family, Herpesviridae; subfamily, α-Herpesvirinae; genus, Mardivirus; species, Gallid herpesvirus 2) is the cause of Marek's disease (MD), which manifests as malignant lymphomas in visceral organs of infected chickens. 10 The feather follicle epithelium (FFE) is the only site where large numbers of infectious viral particles are produced, and chickens are infected by inhalation of feather dust particles containing cell-free virus. 10,27 MDV is divided into 3 antigenically related serotypes named 1, 2, and 3. Serotype 1 MDV strains cause MD in chickens. 10 Although MD had previously caused serious economic losses to the poultry industry, the disease is currently well controlled by vaccination. 33 Strains of attenuated serotype 1 MDV, naturally nononcogenic serotype 2 (Gallid herpesvirus 3), and serotype 3 (herpesvirus of turkey [HVT] or Meleagrid herpesvirus 1) have been used as monovalent or bivalent vaccines; an attenuated serotype 1 MDV strain designated CVI988 is considered to be the most protective vaccine currently available and is used in many countries. 10,39 However, the evolution of MDV in the field has resulted in the generation of more virulent strains such that MD cases have been reported even in vaccinated chickens. 38
The MDV genome consists of a unique long (UL) region and a unique short (US) region, both flanked by a set of inverted repeat regions, located terminally (terminal repeat long [TRL] and terminal repeat short [TRS]) and internally (internal repeat long [IRL] and internal repeat short [IRS]). 16,32 MDV genes consistently expressed in MDV-transformed cell lines and tumor samples have been studied for the search of viral factors related to MDV-induced latency and tumorigenicity. 28–31 Among them, a serotype 1 MDV-specific gene, meq, was identified in the TRL and IRL regions. 28–31 The meq gene encodes a 339-amino acid N-terminal basic region-leucine zipper (bZIP) trans-activator protein, which resembles the Fos/Jun family of oncoproteins, suggesting that the meq gene product, Meq, plays a key role in the cell transformation induced by oncogenic MDV. 21,25,26,30 In addition, it has been previously reported, by Southern blot analysis, a 180-base pair (180-bp) insertion was found in the meq open reading frames (ORF) of CVI988 and CVI988/R6, attenuated vaccine strains of MDV, whereas this meq gene, termed as L-meq, was not detected in Md5 and RB1B, strains of oncogenic MDV. 24 A recent study has shown that highly virulent MDV strains carry the meq gene but not the L-meq gene. 34 Although the transactivation domain of Meq is characterized by 6 proline-rich repeats (PRR), the L-meq gene product (L-Meq) contains 9 PRR in the transactivation domain, suggesting that L-Meq may have a transactivator activity that is different from that of Meq. 13
The traditional diagnosis of MD is based on clinical signs and pathological findings. However, more specific and easier methods for the surveillance of MDV are necessary. Therefore, this study focused on the development of a diagnostic method using feather tips. Methods that have been reported for the detection of MDV from feather tips include virus detection from feather tips by the agar gel precipitation test, enzyme-linked immunosorbent assay (ELISA), and dot blot hybridization. 15,17,22,27 A previous study has shown that the polymerase chain reaction (PCR) is a convenient tool for monitoring MDV in poultry populations. 18 In addition, recently, methods for the detection and quantitation of MDV genomes by real-time PCR have been reported. 1,3,19,20 A real-time PCR assay for the quantitation of the CVI988 genome can be used to monitor virus loads in vaccinated chickens and for research application to improve vaccine strategies for different breeds of chickens. 2,4 However, these PCR protocols are not able to differentiate oncogenic strains of MDV from CVI988, a widely used vaccine; therefore, there is a need for a detection method that is specific for oncogenic MDV strains and does not cross-react with vaccine strains. In this study, a nested PCR, specific for oncogenic MDV strains for surveillance of MDV infections, has been developed.
Sequences of MDV-specific primers used for PCR amplification.
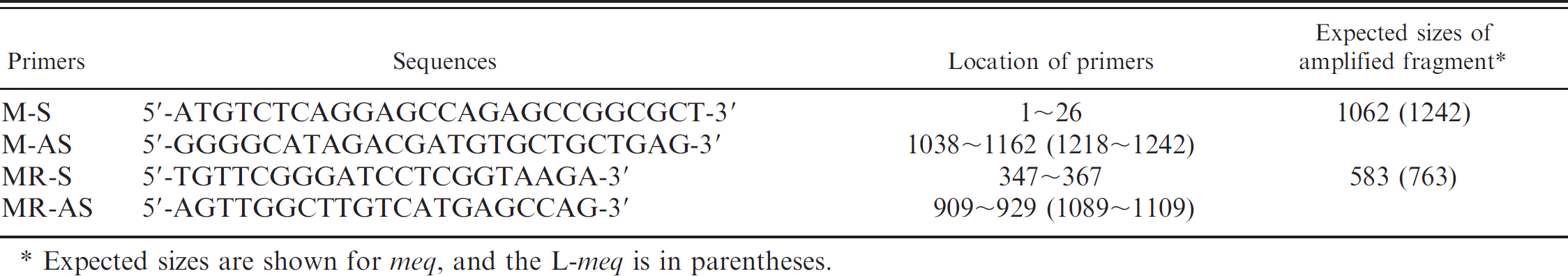
Expected sizes are shown for meq, and the L-meq is in parentheses.
Materials and methods
Experimental chickens. Fertilized eggs from commercial chickens a were hatched in the authors' laboratory, and the chicks were raised in isolators.
Viruses and cells. A strain of oncogenic MDV, Md5, was isolated in chicken kidney cell culture from infected chickens. 41 Vaccine strains used were attenuated serotype 1 MDV strain, CVI988, b and a nonpathogenic serotype 3 strain, FC126. c All viral strains were propagated in chicken embryo fibroblasts (CEF). When cytopathic effects were visible in 100% of the cell monolayer, the infected cells were harvested, and virus titers were measured by plaque assay. 40 These infected CEFs were used as inoculum for experimental infection of chickens.
Inoculation of viruses and sample collection from infected chickens. Three groups of 3-day-old chickens (5 chickens/group) were inoculated intraperitoneally with either 1,000 plaque forming units (PFU)/0.2 ml of Md5; 2,000 PFU/0.2 ml of CVI988; or 2,000 PFU/0.2 ml of FC126; respectively. In addition, 2 groups of chickens (5 chickens/group) were co-infected with 1,000 PFU/0.2 ml of Md5 and either 2,000 PFU/0.2 ml of CVI988 or 2,000 PFU/0.2 ml of FC126, respectively.
Two or 3 feather tips were collected from wings of chickens infected with MDV once per week for 10 weeks after inoculation. Heparinized blood samples were collected from 2 chickens of each group infected with MDV at 5 weeks after inoculation. Peripheral blood mononuclear cells (PBMC) were prepared using Percoll d density gradient.
DNA extraction. DNA extraction from feather tips was carried out as described previously. 6,7 In brief, the feather tips were cut and immersed overnight at 55°C in 1 ml of the lysis buffer (0.5% sodium dodecyl sulfate [SDS], 0.1 M NaCl, 10 mM Tris pH 8.0, 1 mM ethylene diaminetetraacetic [EDTA]) containing proteinase K at a final concentration of 200 μg/ml. Total cellular DNA was extracted with phenol-chloroform-isoamyl alcohol (25:24:1), precipitated with ethanol, and treated with RNase A at the final concentration of 20 μg/ml. Total cellular DNA was also extracted from PBMC by using a commercial kit e following the manufacturer's instructions.
Nested PCR and conventional PCR. Total cellular DNA samples extracted from feather tips or PBMC were used as templates for nested PCR to detect the meq or L-meq gene. The meg-specific primers used in this study are shown in Table 1. The first round of PCR was performed with the primer set M-S and M-AS to amplify a 1,062-bp meq or a 1,242-bp L-meq gene fragment. 24 After the amplification, 1 μl of the reaction was used for the second round of PCR. The second round of PCR was performed in a 20-μl reaction mixture containing 1.5 mM MgCl2, and the primer set 0.5 μM of MR-S and MR-AS to amplify a 583-bp meq or 763-bp L-meq fragment corresponding to the PRR region. 11 Conventional PCR was performed with MR primers and the same templates. Amplification was conducted for 35 cycles of 94°C for 45 sec, 57°C for 45 sec, and 72°C for 90 sec. The fragments amplified by nested PCR or the conventional PCR were separated on agarose gels (1.5%) and visualized under ultraviolet light after staining with ethidium bromide.
DNA sequencing. The nucleotide sequences of the meq or L-meq gene fragments amplified in this study were determined. The nested PCR products were excised from the gel, purified by using the Geneclean kit, f and cloned into the pGEM-T easy vector. g The pGEM-T plasmid containing the insert was purified by a standard mini-prep method and sequenced by using the BigDye terminator cycle sequencing kit h and model-310 genetic analyzer. h The nucleotide sequences of the meq or L-meq gene were compared with the nucleotide sequence of Md5 reported in GenBank (accession numbers: AF243438) or CVI988. 12
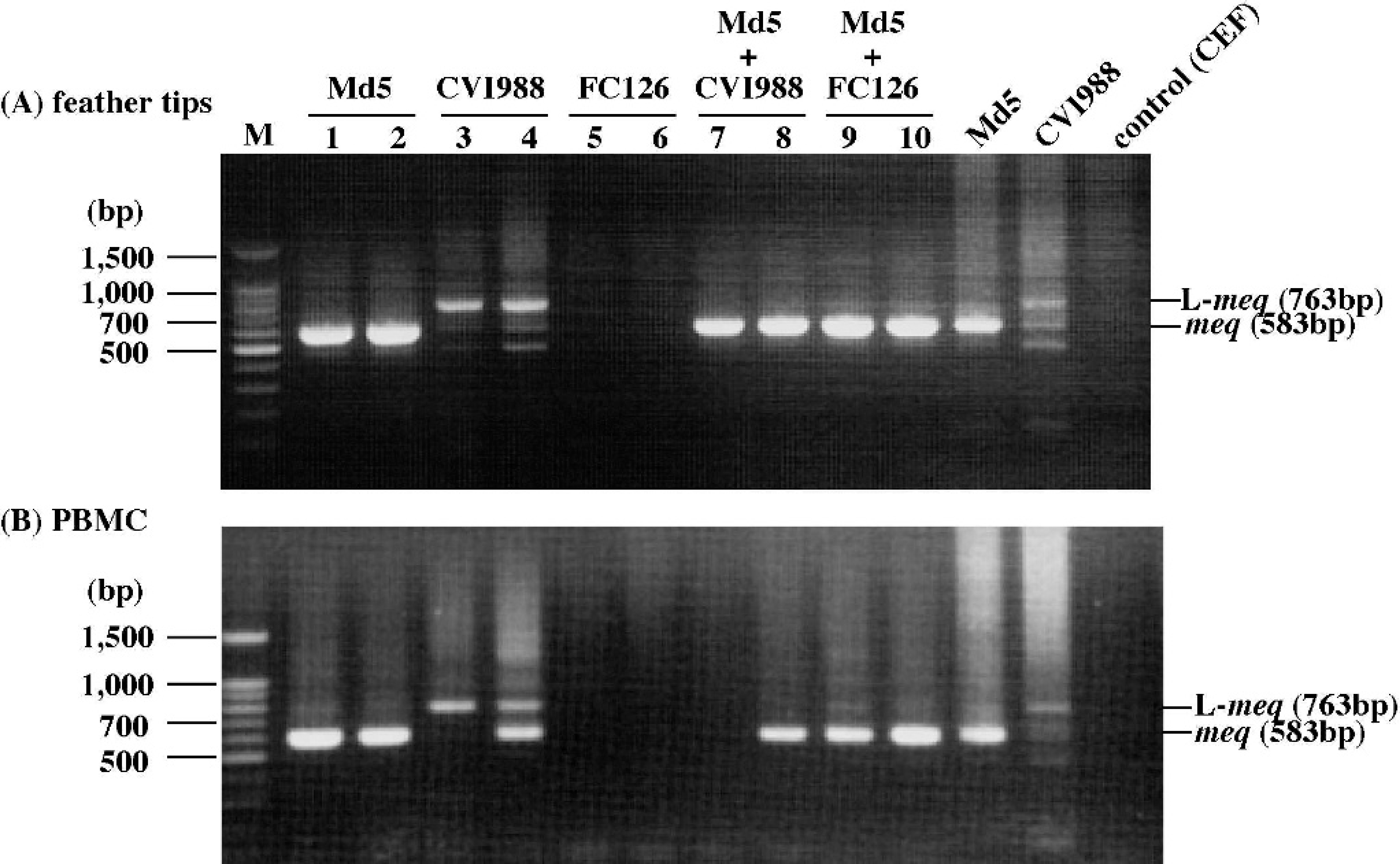
Nested PCR amplification of the meq gene from
Results
Detection of the meq or L-meq gene in feather tips and PBMC by nested PCR. To determine whether the meq or L-meq gene is detected in feather tip DNA with higher sensitivity than in DNA samples of other tissues, chickens were experimentally infected with MDV as described in the Materials and methods section. When feather tips and PBMC collected from infected chickens at 5-weeks postinoculation (pi) were examined by the nested PCR method, the meq and/or L-meq gene was similarly detected in most of the DNA samples of feather tips and PBMC (Fig. 1). In both feather tips and PBMC samples, the meq gene was detected in chickens infected either with Md5 alone or co-infected with Md5 and CVI988 or FC126. On the other hand, the L-meq gene was detected as a major band in DNA samples of chickens infected with CVI988, although the meq gene was also detected as a very faint band. By sequence analysis, it was confirmed that the nucleotide sequences of the meq and L-meq genes amplified from feather tip DNA exactly matched those of the meq gene of Md5 and the L-meq gene of CVI988, respectively (data not shown). In addition, the meq or L-meq gene was detected by the nested PCR using 200 ng, 100 ng, or 50 ng of total cellular DNA from both feather tips and PBMC samples of chickens infected with Md5 or CVI988 alone, indicating that the 2 types of samples yielded similar sensitivities (data not shown). However, neither the meq nor the L-meq gene was detected in chickens infected with FC126, because the meq gene is not present in the genome of FC126. Moreover, the meq gene was not detected in PBMC DNA of 1 chicken co-infected with Md5 and CVI988, although the meq gene was detected in feather tip DNA of the chickens. These results suggest that feather tip DNA samples are available for the nested PCR, similar to PBMC samples.
Detection of the meq gene in feather tip DNA of chickens infected with oncogenic or attenuated MDV. The authors have previously reported that the meq gene is detected in DNA samples extracted from PBMC infected with oncogenic MDV at 1 to 7 weeks pi, while the L-meq gene is detected at 3 to 5 weeks pi. 11 To verify whether this difference in detection periods also occurs in feather tip DNA samples, the meq gene in feather tip DNA of chickens infected with oncogenic MDV, Md5, was amplified from samples collected at 1 to 10 weeks pi. The meq gene was detected in 90.3% of the DNA samples at 1 to 10 weeks pi (Fig. 2A; Table 2). In addition, the L-meq gene was detected as a major band in 100% of the DNA samples of chickens infected with 2,000 PFU of attenuated MDV, CVI988, after 2 weeks pi (Fig. 2B; Table 2), although neither the meq nor L-meq gene was detected in PBMC samples of the same chickens. 11 Neither the meq nor L-meq gene was detected in chickens infected with FC126 throughout the experimental period (data not shown). These results suggest that the detection of the meq or L-meq gene using feather tip samples is an effective method for the differentiation of oncogenic MDV from CVI988.
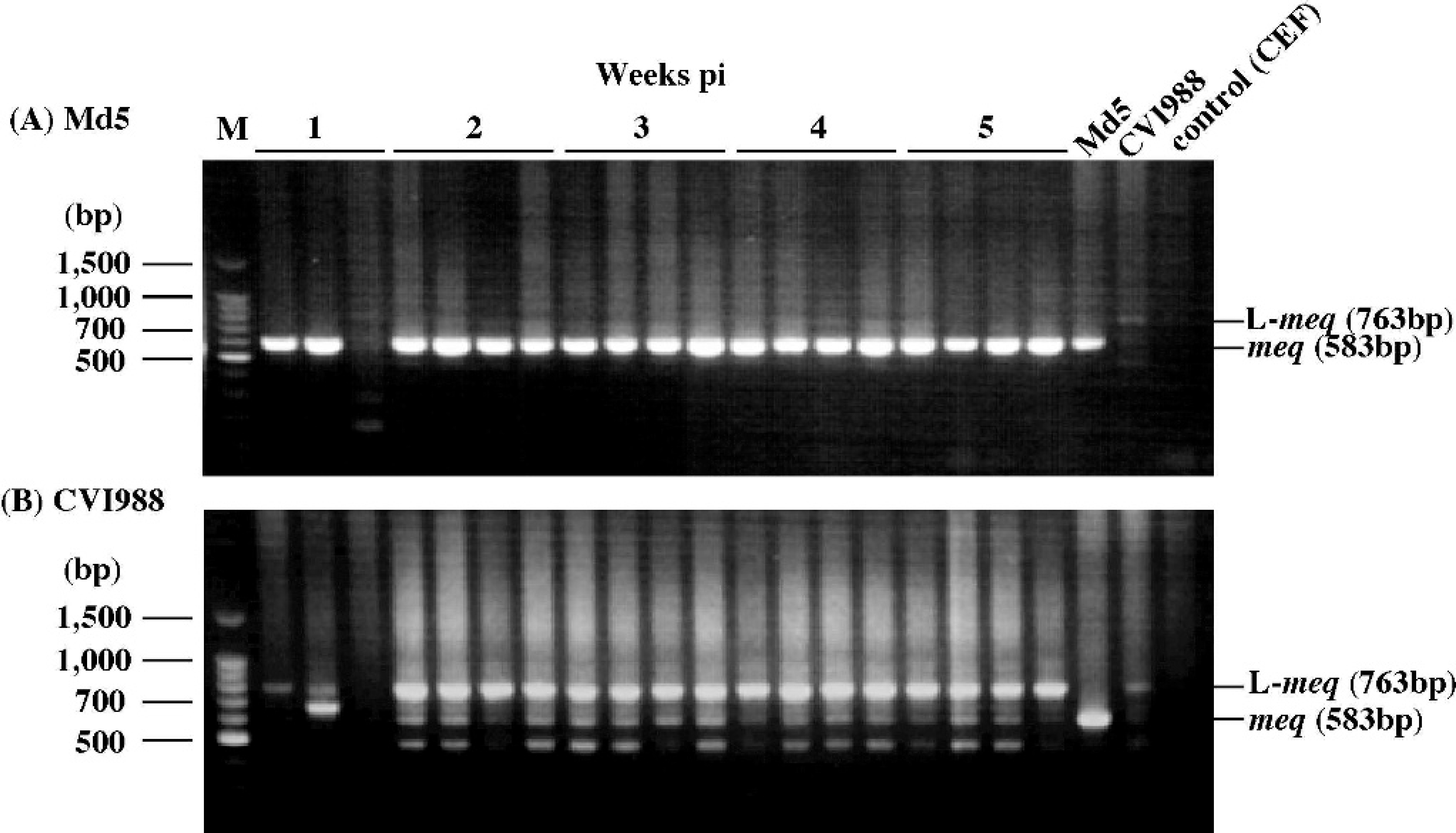
Nested PCR amplification of the meq gene from feather tips of chickens infected with 1,000 or 2,000 PFU/chicken of A, strain Md5, or B, CVI988, respectively. DNA samples were collected at 1, 2, 3, 4, and 5 weeks pi. M = molecular size marker (100-bp DNA ladder g ).
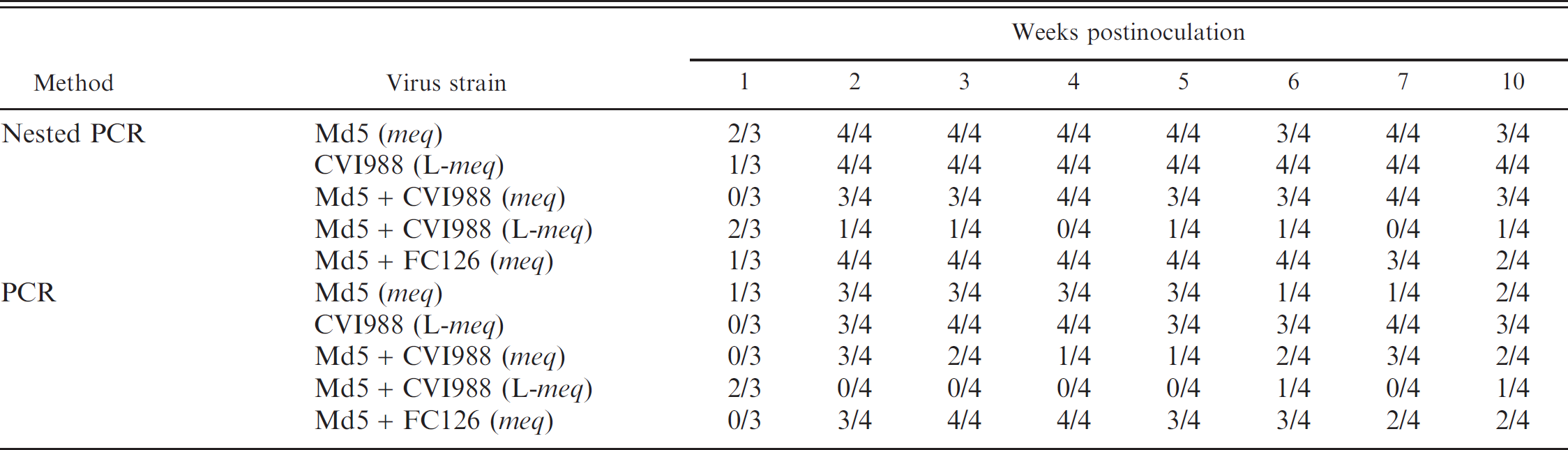
Comparison of the diagnostic sensitivity of nested PCR and conventional PCR for the detection of the meq or L-meq gene in feather tip-DNA samples.
Nested PCR analysis of the meq and L-meq gene in chickens co-infected with oncogenic and vaccine strains. The meq gene was dominantly detected in the mixtures of feather tip DNA of Md5- and CVI988-infected chickens at 5 weeks pi (Fig. 3). To test whether the meq gene is detected by nested PCR using feather tip samples of chickens co-infected with Md5 and either of the vaccine strains, the meq gene was amplified from samples collected from chickens co-infected with Md5 and either CVI988 or FC126. The meq gene was mainly detected in 82.1% (CVI988 and Md5) or 89.3% (FC126 and Md5) of each group of co-infected chickens at 2 to 10 weeks pi (Fig. 4; Table 2). These results show that the meq gene is mainly detected in feather tip DNA of chickens co-infected with oncogenic and vaccine strains, suggesting that oncogenic MDV can be specifically detected in infected chickens by this nested PCR method targeting the meq gene in a flock.
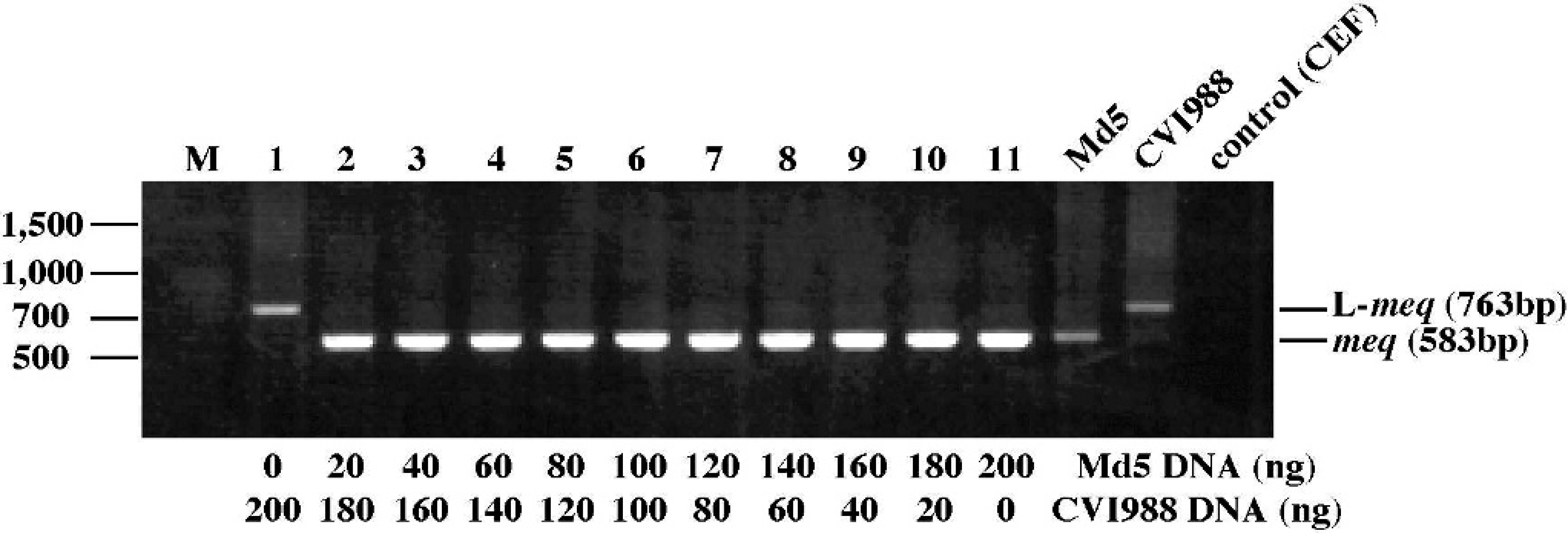
Nested PCR amplification of the meq gene from mixtures of feather tip-DNA of chickens infected with 1,000 PFU/chicken of strain Md5 and 2,000 PFU/chicken of strain CVI988. DNA samples were collected from Md5- and CVI988-infected chickens at 5 weeks pi, and DNA samples from these chickens were mixed at the amounts indicated in the bottom of the figure. M = molecular size marker (100-bp DNA ladder g ).
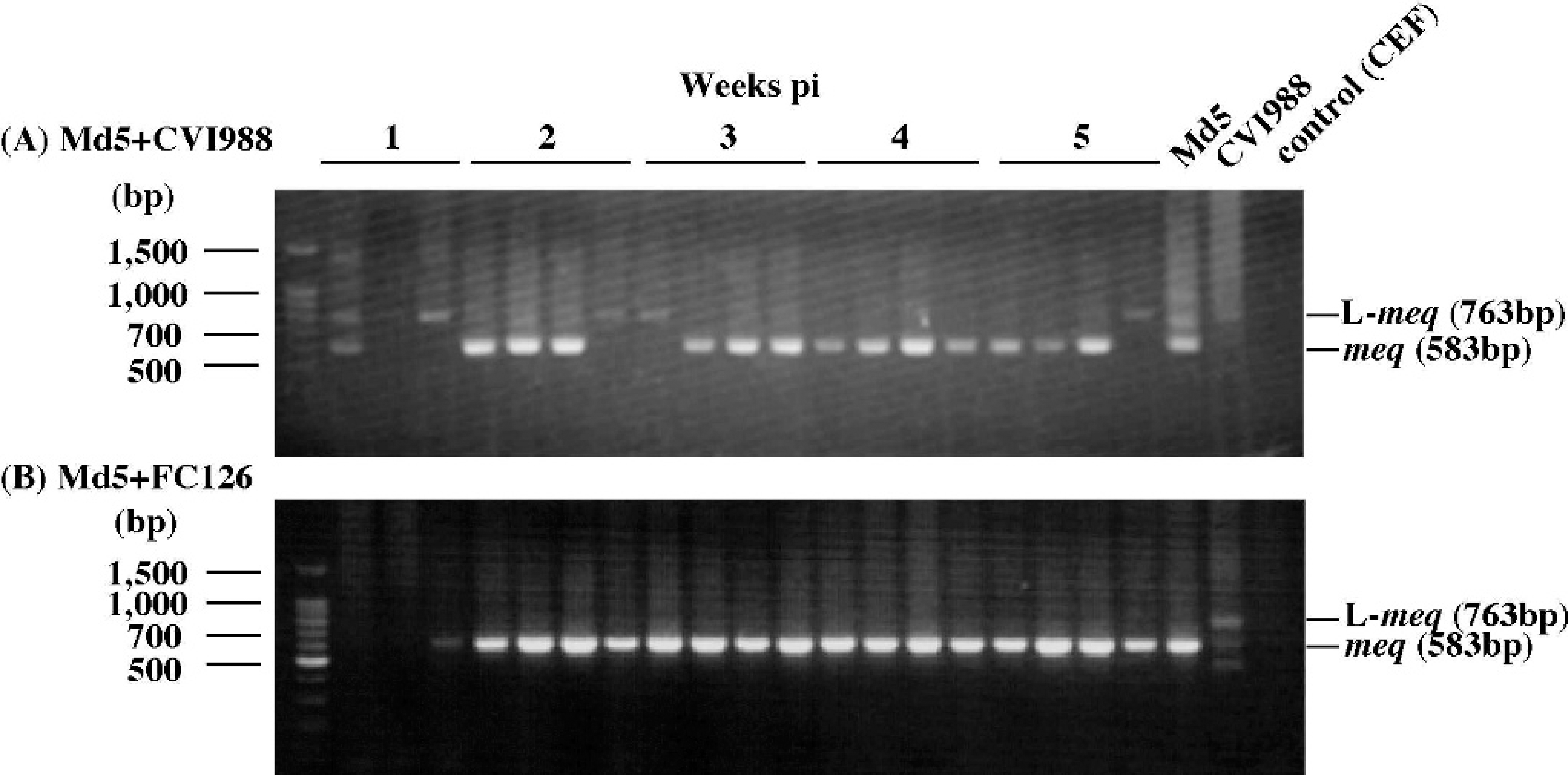
Nested PCR amplification of the meq gene from feather tips of chickens infected with A, 1,000 PFU/chicken of strain Md5 and 2,000 PFU/chicken of strain CVI988, or B, 1,000 PFU/chicken of strain Md5 and 2,000 PFU/chicken of FC-126. DNA samples were collected at 1, 2, 3, 4, and 5 weeks pi. M = molecular size marker (100-bp DNA ladder g ).
Comparison of sensitivity of nested and conventional PCR for the detection of the meq gene in MDV-infected chickens. When the sensitivity of this nested PCR method was compared with the conventional PCR with MR primers and the same volume of templates as nested PCR method, the sensitivity of the nested PCR method was higher than that of the conventional PCR after 3 weeks pi (Table 2). These results suggest that the conventional PCR method is not appropriate for detection of the meq/L-meq gene and that monitoring of oncogenic MDV should continue after the onset of MDV latency. Therefore, this nested PCR method is better than conventional PCR for differentiation of oncogenic MDV from CVI988, since the meq/L-meq gene is constantly detected even at MDV latency.
Discussion
Although the MD is currently well controlled by vaccination, the virulence of field MDV isolates tends to increase. 33,38 Therefore, the development of practical methods to determine the prevalence of oncogenic MDV in the field is necessary. Some PCR-based methods for the detection of MDV in feather tips have been developed for the surveillance of MDV because feather tips are convenient samples for collection in the field and transport to the laboratory. 1,3,7,14,18 20 Since the use of serotype 1 MDV-based vaccines, especially CVI988, has become widespread, specific diagnostic methods to differentiate oncogenic from attenuated MDV using feather tip samples are needed. This study exploited the structural difference in the meq and L-meq genes of oncogenic and attenuated MDV vaccine strain, CVI988, to develop nested PCR for differentiation of oncogenic from attenuated MDV.
In the present study, the feather tip and PBMC samples yielded similar sensitivities (Fig. 1 and data not shown), although it has been reported that the detection of MDV using feather tip samples by PCR had higher sensitivity than those using the buffy-coat and other tissue samples. 18 Moreover, this nested PCR also detected the meq gene of several MDV field isolates, whose nucleotide sequences were different from strain Md5 (Hayashi et al., 2006, unpublished data). Therefore, this method could be applicable for the assay of other MDV strains and for their survey. However, further analysis is needed with regard to the sensitivity.
Previously, the authors reported that both the meq and L-meq genes were detected in DNA samples extracted from PBMC or spleen cells of chickens experimentally infected with oncogenic MDV at 3 to 5 weeks pi, the latent phase of the infection. 11 In the present study, however, only the meq gene was consistently detected in feather tip samples of chickens infected with oncogenic MDV (Md5 strain) (Fig. 2A). The FFE is the site for virus assembly and contains large amounts of cell-free virus, whereas CD4+ T cells in blood are the main target cells for MDV latency. 8–10,35 Because the establishment of MDV latency starts 1 to 2 weeks after infection, the L-meq gene, which was detected in blood samples of chickens infected with oncogenic MDV at 3 to 5 weeks pi, has been suggested as a contributory factor in the maintenance of the MDV latency in CD4+ T cells. 9–11,35 Therefore, these changes (from meq to L-meq during MDV latency) may be specifically observed in blood samples including CD4+ T cells latently infected with MDV. In addition, when the meq and L-meq genes were monitored in chickens infected with attenuated MDV, CVI988, the L-meq gene was constantly detected as a major band after 2 weeks pi (Fig. 2B), suggesting that the use of feather tip samples is more effective than that of blood samples in terms of the differentiation of oncogenic from attenuated MDV.
Since the serotype 1 MDV-based vaccines are widely used, chickens could be co-infected with both oncogenic and attenuated MDV in the field. In chickens infected with CVI988 alone, titers of virus rescued from CD4+ T cells have been shown to be significantly lower than those of chickens infected with Md5 alone. 23 Therefore, it was expected that the meq gene would be predominantly detected even in chickens co-infected with oncogenic MDV and CVI988, because the amount of oncogenic MDV was dominant in feather tips compared with that of CVI988. Indeed, the meq gene was predominantly detected in the mixtures of feather tip DNA of chickens infected with Md5 and CVI988 at 5 weeks pi (Fig. 3). In chickens co-infected with Md5 and CVI988, titers of virus rescued from CD4+ T cells have been shown to be significantly lower than those of chickens infected only with Md5, especially at 1 week pi. 23 In the present study, therefore, the meq gene may not be dominantly detected in co-infected chickens at 1 week pi because the replication of Md5 was suppressed by the vaccination (Fig. 4A). However, the meq gene was mainly detected in co-infected chickens after 2 weeks pi (Fig. 4A). The virus was not rescued from CD4+ T cells of chickens infected only with CVI988, whereas it was rescued from those of chickens co-infected with Md5 and CVI988 at 2 weeks pi, suggesting that the virus isolated from co-infected chickens is mainly Md5. 23 Moreover, CVI988 appears to spread poorly by inhalation of feather dust. 41 Since the amounts of Md5 were more than those of CVI988 in co-infected chickens, the meq gene may be mainly detected in co-infected chickens after 2 weeks pi, although occasionally the L-meq gene was also detected in co-infected chickens. Hence, the nested PCR method is reliable and appropriate for the study of the prevalence of oncogenic MDV in a flock.
In order to differentiate oncogenic from attenuated MDV, this nested PCR method may not be superior to a previously described PCR method that targets a 132-bp segment of the direct repeat region. 5,36 However, the nested PCR method is more advantageous for the survey of highly virulent MDV, compared with 132-bp direct repeat PCR method. Based on their ability to resist immunity induced by different types of vaccines, pathogenic MDV strains are classified as mild (mMDV), virulent (vMDV), very virulent (vvMDV), and very virulent+ (vv+MDV). 38 In addition to CVI988, which is classified as vMDV, MDV strains in which the L-meq gene is present are classified as mMDV or vMDV, whereas the meq gene is present in all strains classified as vvMDV or vv+MDV. 34,39 The target PRR of the meq gene is the site of mutations exhibited by most of the highly virulent MDVs (vvMDV and vv+MDV); this region has been associated with the transcriptional activity of Meq, suggesting that the mutations in this region are associated with MDV virulence. 34 Therefore, it might be possible to speculate on MDV virulence, such as vMDV, vvMDV, vv+MDV, and new virulent MDV strains that could induce future outbreaks, by sequencing the DNA fragments amplified by this nested PCR method.
In 2001, a case of MD was reported in a white-fronted goose (Anser albifrons) in Lake Miyajimanuma of Hokkaido, Japan. 37 Although the roles and significance of wild geese in the epidemiology of MDV remain to be determined, they could play important roles as reservoirs and carriers of MDV. Because some wild birds, such as white-fronted geese, are endangered species and are protected natural resources in Japan, it is almost impossible to collect and analyze samples from their internal organs. However, since feather tips are convenient samples for collection, this nested PCR method can be applied for the surveillance of oncogenic MDV in these wild birds as well as in domestic birds. In conclusion, the detection of the meq gene by nested PCR using feather tip samples is a specific method for monitoring oncogenic MDV in flocks of chickens or wild birds. A study of the prevalence of MDV in wild birds using this nested PCR method is now in progress.
Acknowledgements
This work was supported in part by Grants-in-Aid for Scientific Research (B: 17380203) from the Ministry of Education, Culture, Sport, Science and Technology of Japan, and by the Environmental Technology Development Fund (Kiso 3) from the Ministry of the Environment of Japan.
Footnotes
a.
Hokuren Co., Sapporo, Japan.
b.
Gehn Corp., Inc., Gifu, Japan.
c.
Kyoritushoji Co., Tokyo, Japan.
d.
Amersham Pharmacia Biotech AB, Uppsala, Sweden.
e.
Sankoujunyaku Co., Tokyo, Japan.
f.
Bio101, La Jolla, CA.
g.
Promega, Madison, WI.
h.
Applied Biosystems, Foster City, CA.
