Abstract
This article describes amebic infections in 4 horses: granulomatous amebic encephalitis caused by Balamuthia mandrillaris and Acanthamoeba culbertsoni and systemic infections caused by Acanthamoeba sp. The former infection occurred in 1 of 4 horses spontaneously without any underlying conditions; the latter amebic infection was perhaps “opportunistic” considering the visceral involvement by this protozoan in association with Aspergillus sp. and/or Escherichia coli and Pseudomonas sp. The clinicopathologic findings and demonstration of the amebic organisms using immunohistochemical techniques, culture, polymerase chain reactions, and electron microscopy are presented.
Infections with Acanthamoeba sp. and Balamuthia mandrillaris in 4 horses: clinicopathologic findings.
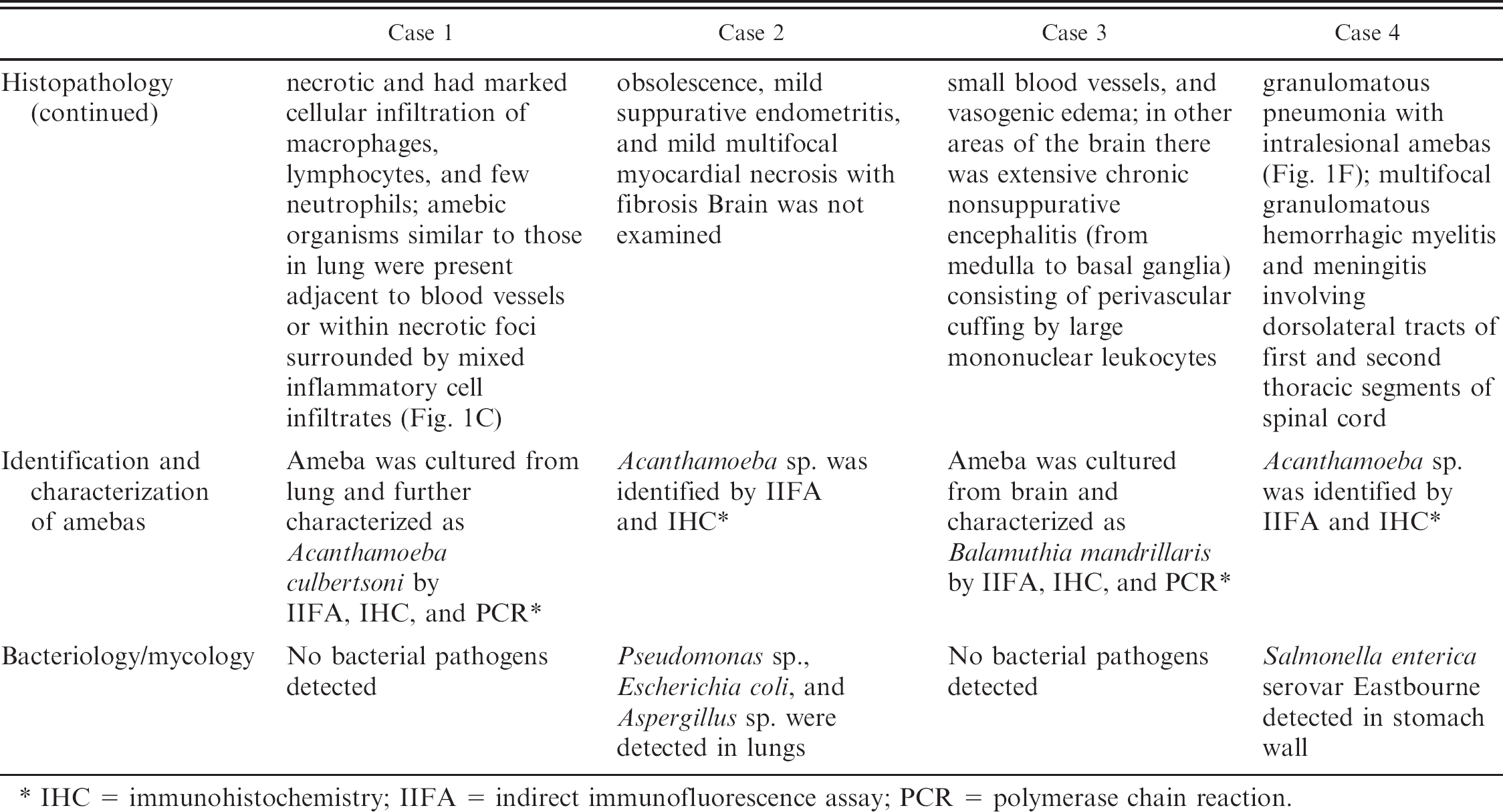
IHC = immunohistochemistry; IIFA = indirect immunofluorescence assay; PCR = polymerase chain reaction.
Amebic encephalitis in humans and animals is caused by several species of free-living amebas belonging to the genera Acanthamoeba, Naegleria, and Balamuthia. 1,5,6,8,12,15,16 In humans free-living amebas cause 3 well-defined diseases: 1) primary amebic meningoencephalitis (PAM) caused by Naegleria fowleri, 2) granulomatous amebic encephalitis (GAE) caused by Acanthamoeba spp. and Balamuthia mandrillaris, and 3) chronic amebic keratitis (AK) caused by species of Acanthamoeba. Both PAM and chronic AK occur in healthy individuals while GAE is often associated with patients having acquired immunodeficiency. 8 , 14 , 15 The occurrence of GAE in animals with immunodeficiency has not been documented. The clinicopathologic findings of the 4 affected horses are described in Table 1.
Formalin-fixed and paraffin-embedded brain (Cases 1 and 3) and lung (Cases 1, 2, and 4) tissue sections were deparaffinized, and indirect immunofluorescence (IIF) and/or immunohistochemistry (IHC) assays were performed as described previously. 6 Acanthamoeba culbertsoni (Fig. 1B), B. mandrillaris (Fig. 1E), and Acanthamoeba sp. (Fig. 1G) were detected.
Attempts were made to isolate the organisms from frozen-thawed unfixed brain (Cases 1 and 3) and lung tissues (Case 1). The brain and lung specimens had been stored at −70°C for 60 days and were transferred on dry ice to the Centers for Disease Control (CDC), Atlanta, Georgia, where cultivation was attempted as previously described. 16 The cultured amebas were further characterized by IIF as A. culbertsoni (from the lung of Case 1) and B. mandrillaris (from the brain of Case 3). In addition, the amebas were processed for transmission electron microscopy and examined in either a Zeiss 10 C or a Zeiss (LEO) model 906E transmission electron microscope at 60 kv accelerating voltage as previously described. 4 Ultrastructural characteristics of the amebas are shown in Fig. 1H, I, and J.
Confirmation of the identity of the amebas and genotyping was performed using polymerase chain reaction (PCR). Centrifuged pellets of amebas (CDC V409C) and CDC V433) were suspended separately in 200 μl of UNSET buffer, and DNA was extracted. 2 , 13 Approximately 100 to 200 ng of genomic DNA from CDC V409C and CDC V433 was used for PCR amplifications. Nuclear 18S rDNA amplification was performed with primers CRN5 (5′-TGGTTGATCCTGCCAGTAG-3′)and SSU2 (5′-CCGCGGCCGCGGATCCTGATCCCTCCGCAGGTTCAC-3′), which span approximately 99% of the gene. PCR products of nuclear 18S rDNA were prepared for automated sequencing using the QIAquick PCR purification kit. a These products were then directly sequenced using automated fluorescent sequencing protocols on an Applied Biosystems automated sequencing system b using primers previously used in Acanthamoeba studies. 14 The 18S rDNA sequence of B. mandrillaris V433 is available under GenBank accession number AF477021, and the 18S rDNA sequence of A. culbertsoni V409C is available under accession number DQ499151. The mitochondrial 16S rRNA gene from A. culbertsoni (CDC V409C) was amplified using primers previously used in one of the coauthor's laboratories (G. B.) to amplify this region in Acanthamoeba sp. 7 The A. culbertsoni (CDC V409C) 16S rDNA sequence is available in GenBank under accession number AF479542. The mitochondrial 16S rRNA gene from B. mandrillaris (CDC V433) was amplified by PCR using primers that specifically amplify this target in B. mandrillaris. 2 The 16S rDNA sequence is available in GenBank under accession number AF477017. The nuclear 18S and mitochondrial 16S rDNA sequences of A. culbertsoni V409C and B. mandrillaris V433 were aligned in the sequence alignment editor ESEE and compared to other sequences in our database to determine their identification. 3 PCR sequence comparison of both the nuclear and mitochondrial sequences from CDC V409C (from Case 1) and CDC V433 (from Case 3) determined the identity of these isolates as A. culbertsoni (Acanthamoeba genotype T10) and B. mandrillaris, respectively.
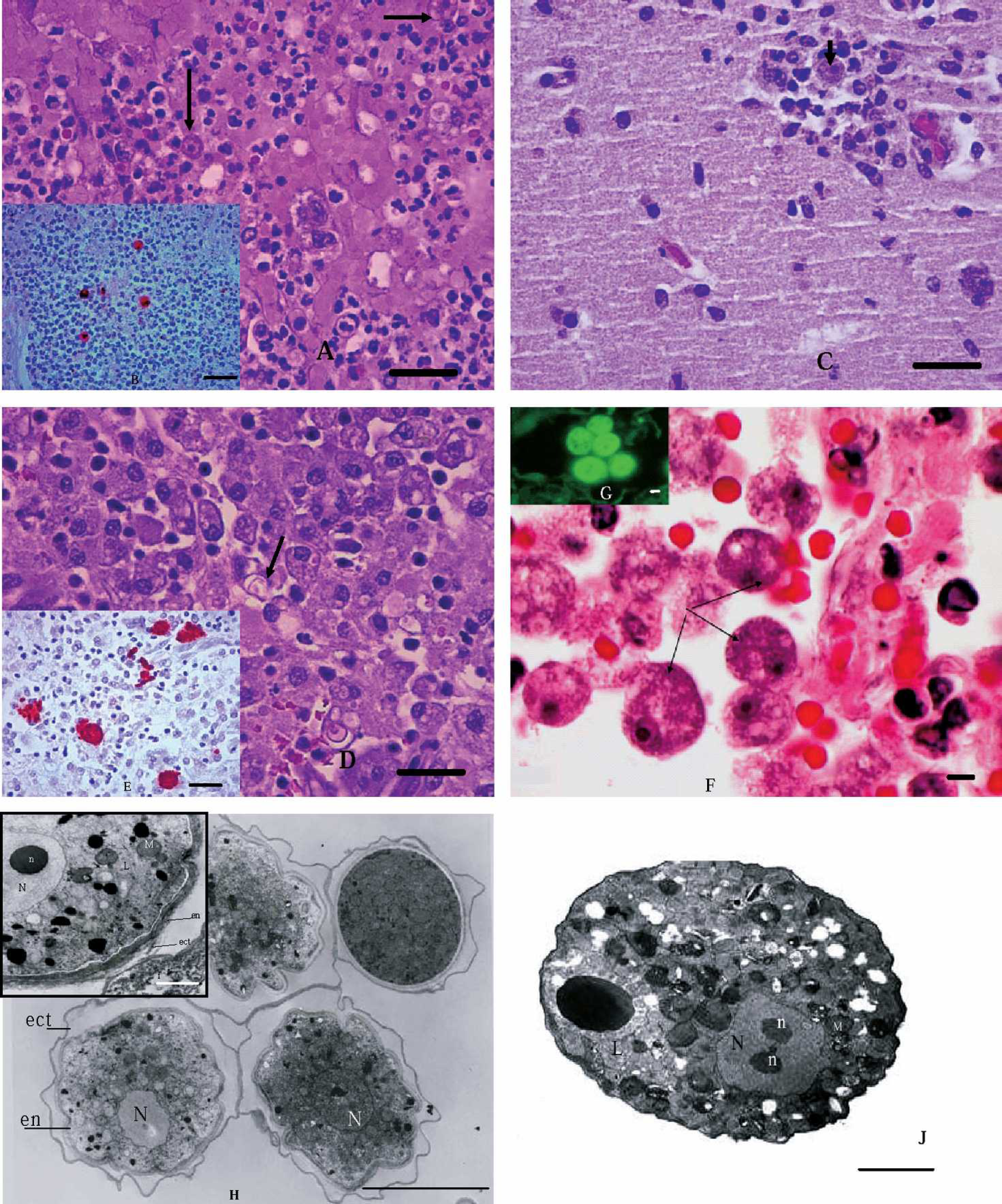
Histologic sections of equine lung and brain and ultrastructures of amebas with amebic infections.
Amebic infection in horses is very rare, and it is noteworthy to mention that all cases have originated in southern California between 1998 and 2001. Kinde and colleagues 6 reported meningoencephalitis in another horse that was also indigenous to southern California. The reason for this cluster of cases in southern California is not clear. Amebas can be easily missed in histologic sections because of their resemblance to macrophages; even when the disease is suspected, confirmation is often not possible because of lack of awareness of the disease and the scarcity of reagents. The authors believe that perhaps there are more cases in horses of amebic infections, “out there,” but not all neurologic cases are submitted to diagnostic laboratories for necropsy examination.
In human patients GAE due to Acanthamoeba species (Acanthamoeba castellanii, A. culbertsoni, and possibly other amebas) has been reported in chronically ill and debilitated or immunologically impaired individuals with severe derangements of host defenses. 8 In Case 4, the horse may have been immunocompromised as it showed thymic atrophy and systemic amebic infection and salmonellosis due to Salmonella enterica serovar Eastbourne. Case 2 had infections with Aspergillus sp., Pseudomonas sp., and Escherichia coli, suggesting that the horse may have already had a weakened immune system and became prone to opportunistic amebic infection. For cases 1 and 3 there was no known predisposing factor or detectable underlying disease and the clinicopathologic findings were due to primary amebic infection. The route of invasion and penetration into the central nervous system in human cases of GAE is hematogenous, probably from a primary site in the lower respiratory tract. 10 GAE has been reported with chronic ulceration of the skin resulting with terminal hematogenous spread to the brain. 8 In all 3 horses with Acanthamoeba infection the lung was highly likely to be the primary site of infection; from there, the organism may have spread hematogenously to other organs including the brain. In Case 3 no evidence of skin lesions, pneumonia, or microscopic lesions in the cribiform plate, olfactory fossae, and ethmoid bones was noted. In general in horses, the pathogenesis of amebic infection is not known but it is speculated that perhaps it is similar to that of humans. Although amebic encephalitis appears to be rare in horses, it should be included in the differential diagnosis with other neurologic conditions.
Acknowledgements. We thank S. Kwiek, E. J. Hurley, and R. Sriram for their technical assistance and Drs. D. Vrono, K. Valko, and T. Brauer (veterinary practitioners) for submitting the cases. Part of the work of P. A. F., D. J. K., and G. C. B. (co-authors) was supported by a grant from the National Institutes of Health (grant EY09073).
Footnotes
a.
Qiagen Inc., Valencia, CA.
b.
Applied Biosystems, Foster City, CA.
