Abstract
The objective of this study was to evaluate the use of real-time TaqMan PCR assays for detection of coinfections with “Candidatus Mycoplasma haemominutum” (Mhm), and Mycoplasma haemofelis (Mhf), in vitro and over time in experimentally infected cats. First, the ability of each real-time PCR assay to detect and quantify mixed infections was determined in vitro by testing mixtures of plasmids containing Mhm and Mhf 16S rDNA with each assay. Subsequently, 4 specific pathogen-free (SPF) cats, 2 of which were splenectomized, were inoculated with blood from a cat infected with both Mhm and Mhf. Sixteen blood samples were then collected from each cat over a 55-day period. Each of the 64 postinoculation samples was tested using both conventional polymerase chain reaction (cPCR) and real-time PCR for the 16S rRNA gene of each organism. When applied to mixtures of plasmid DNA from each species, the results of quantitation with each of the real-time PCR assays approximately reflected the number of plasmid copies present. Forty-nine of 64 post-inoculation samples (77%) were positive using both cPCR and real-time PCR, 4 (6%) were positive using cPCR only, and 3 (5%) were positive using real-time PCR only. Both organisms were detected in 23 samples using real-time PCR. Mixed infections were not detected using cPCR. The size of the corresponding cPCR products suggested infection with Mhm in 4 and Mhf in 18 of these samples. The use of multiple separate real-time PCR assays rather than cPCR alone should thus be considered for epidemiologic studies of hemoplasmosis in cats.
Introduction
Hemotropic mycoplasmas (hemoplasmas) are epierythrocytic mycoplasmas that have never been cultured in vitro. At least 3 species infect cats: Mycoplasma haemofelis (Mhf) and ‘Candidatus Mycoplasma haemominutum’ (Mhm), as well as ‘Candidatus Mycoplasma turicensis’ (Mtc), a new species that was recently described in cats in Switzerland. 5 , 10 , 11 , 18 The latter species appears to be present in cats in North America, as well as an organism related to ‘Candidatus M. haematoparvum’ (Mhp), which was first identified in dogs. 12 , 13 Mhf is capable of causing severe anemia in cats, whereas Mhm has yet to be associated with disease in immunocompetent cats. 4 , 6 , 14 Although uncommon, coinfections with multiple hemoplasma species have been reported. 6 , 14 , 19
Cytologic examination of blood smears has low sensitivity and specificity for diagnosis of hemoplasmosis. 2 , 4 , 6 , 9 The advent of PCR technology has improved our ability to detect hemoplasmas. Conventional PCR (cPCR) is capable of detection and differentiation of Mhm and Mhf based on amplification of 193 and 170 base pair products from the 16S rRNA gene, respectively. 6 However, without post-PCR DNA hybridization or sequence analysis, this does not permit differentiation between Mhm and Mhp or Mhf and Mtc. 12 , 18 Recently, real-time PCR assays have been used for specific detection and quantification of Mhm, Mtc, and Mhf. 3 , 7 , 16 , 18 No studies have specifically studied the sensitivity of PCR assays for detection of coinfections in cats.
During the course of a study aimed to harvest Mhf organisms from splenectomized, experimentally infected cats, it was discovered that the Mhf inoculum used to infect the cats was contaminated with Mhm. Before starting the study, the inoculum tested positive using cPCR and sequencing for Mhf. Before simultaneously analyzing all samples collected over the course of infection using real-time and cPCR, 1 sample was randomly selected from an anemic cat with large numbers of organisms visible on blood smears. The results of cPCR on this sample were positive for a small hemoplasma variant, and the results of a real-time PCR assay for Mhf were negative. Sequencing of the PCR product disclosed the presence of Mhm, yet several cats developed severe anemia. Subsequently, separate real-time PCR assays for Mhf and Mhm were applied to the inoculum and several other samples, which revealed the presence of DNA from both organisms. This afforded an opportunity to compare the results of conventional to those of separate real-time PCR assays for detection of Mhf and Mhm in coinfected cats. The objective of this study was therefore to evaluate the use of real-time TaqMan PCR assays for detection of coinfections with ‘Candidatus Mycoplasma haemominutum’ (Mhm), and Mycoplasma haemofelis (Mhf). The ability of the assays to detect and quantify mixed infections were evaluated 1) in vitro, using plasmid mixtures containing the 16S rRNA gene of each organism, and 2) in vivo, over the course of initial infection, in cats that had been experimentally infected with inocula containing both organisms.
Materials and methods
PCR. Real-time PCR was performed for specific detection of the 16S rRNA gene of Mhm or Mhf as previously described. 13 Briefly, real-time PCR was carried out using a 7700 ABI PRISM SDS instrument and a commercially available Universal Mastermix a containing 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 5 mM MgCl2, 300 μM each of dATP, dCTP, and dGTP, 600 μM dUTP, 0.25 U uracil N-glycosylase (UNG), and 0.625 U of AmpliTaq Gold DNA polymerase per reaction, 400 nM of each primer, 80 nM of fluorogenic TaqMan probe, and 5 μl of diluted template. Amplification was performed using the default cycling parameters of the manufacturer's real-time PCR platform: 2 min at 50°C, 10 min at 95°C, 40 cycles of 15 sec at 95°C and 60 sec at 60°C. Data collection and processing was performed in an attached computer using ABI Sequence Detection System software.
The assay for Mhf used forward and reverse oligonucleotides previously published 6 for conventional hemoplasma PCR, as well as a specific probe (5′-6FAM-TACGAGG-GATAATTATGATAGTACTTCGTGA-TAMRA-3′) that does not bind to Mhm DNA. 13 An additional set of forward (5′-GGCGAACGGGCGAGTAA-3′) and reverse (5′-ACCCCACCAACAACCTAATAGGA-3′) oligonucleotides were used to detect Mhm, together with a specific probe (5′-6FAM-TGGGAAACTAGAGCTTCGCGAGCAG-TAMRA-3′). The sensitivity, specificity, efficiency, and precision of these assays for detection of single infections have been reported elsewhere. 13 A real-time PCR system specific for feline glyceraldehyde-3-phosphate dehydrogenase (GAPDH), as previously described 8 was used as a quality control to test for the integrity of extracted genomic DNA.
Quantification of the number of organisms present in each sample was performed using the standard curve generated by serial dilutions of plasmid DNA template containing known copy numbers of either Mhf or Mhm, and expressed as copy numbers of DNA template per milliliter of blood as previously described (ABI User Bulletin #2, http://docs.appliedbiosystems.com/pebiodocs/04303859.pdf). 15
Conventional PCR was performed using a PCR previously described for detection and differentiation of Mhf and Mhm, which results in amplification of a 193 base pair segment of the 16S rRNA gene sequence of Mhm and a 170 base pair segment of the 16S rRNA gene sequence of Mhf. 6 PCR products were identified following staining with ethidium bromide and ultraviolet transillumination after electrophoresis in 2.4% agarose gels. The results were recorded by one of the authors (ND), who was blinded to the results of real-time PCR analysis.
Plasmid studies. Because the primers used in the Mhf assay could also bind Mhm DNA, and potentially be consumed by an alternative template, the real-time PCR assays were applied separately to mixtures of plasmids in different ratios to validate the performance of each assay in the presence of a mixed infection. Because these organisms cannot be cultured in the laboratory, creation and enumeration of plasmids containing a segment of DNA covering the target region of each assay was performed to demonstrate whether the presence of one organism interferes with detection of the other.
Primers CDsmall84f (5′-GGCGAACGGGCGAGTAA-3′) and 1119r (5′-AAAGAGTAGGGTTGCGCTC-3′) were used to amplify a 1022 base pair fragment of the 16S rRNA gene of Mhm, and a 995 base pair fragment of the 16S rRNA gene of Mhf. Each 50 μl reaction volume contained 1 × reaction buffer, 0.25 U AmpliTaq Gold a , 1.5 mM MgCl2, 0.4 μM each primer, 200 μM of each dNTP and 5 μl DNA template. Cycling conditions were 50°C for 2 min, 95°C for 10 min, followed by 40 cycles of amplification (15 sec at 95°C, 1 min at 60°C) in a thermocycler. b
The resulting products were gel extracted and ligated into commercial plasmids c . Plasmids were transformed using heat-shock into Escherichia coli and isolated using standard methods. DNA sequencing was performed by the dideoxynucleotide chain termination method with an ABI 377 automated sequencer d using M13 primers, to confirm the identity of the cloned product. The partial sequences amplified for Mhm and Mhf included the target sequence for both of the real-time PCR assays. Plasmid copy numbers were determined using spectrophotometry. These plasmids were used to validate single-plex reactions for Mhm and Mhf. 13
To determine how different amounts of DNA for Mhf and Mhm present in the same DNA sample influence detectability by TaqMan PCR, 4 experiments were performed using different ratios of the Mhf and Mhm plasmids. The first experiment contained a fixed amount of Mhf plasmids (103 copies) and decreasing amounts of Mhm plasmids (103, 102, and 10 copies). In the second experiment, 10 copies of Mhf plasmids were mixed with 103, 102, and 10 copies of Mhm plasmids. In the third experiment, 104 copies of Mhf plasmids were mixed with decreasing amounts of Mhm plasmids (106, 104, and 102 copies). Finally, 104 copies of Mhm plasmids were mixed with decreasing amounts of Mhf plasmids (106, 104, and 102 copies). These mixtures were tested in separate reactions with the Mhf and Mhm TaqMan assays, and standard curves were constructed using CT values from the plasmid species present in variable amounts. Plasmid copy number was extrapolated from the single plasmid standard curves for the plasmid species present in constant amounts, and compared with the actual copy number added to each reaction. These experiments were expected to indicate whether quantification of one organism is affected when copy numbers of the other organism are one hundred times more abundant.
Experimental infections. Four specific pathogen-free cats, 2 male neutered and 2 female neutered, ranging from 6 to 12 months of age were used in this study. Conventional PCR for hemoplasma was performed on blood samples from all cats to ensure they were negative before commencing the study (see the following). Two cats (1 male neutered and 1 female; cats 144S and 173S) were subjected to splenectomy 21 days before inoculation. The remaining 2 cats (cats 146 and 177) were not splenectomized. All cats were housed together in the same room. The project was approved by the University of California Davis Animal Use and Care Committee (protocol number 10233).
Each cat was inoculated intraperitoneally with 0.5 ml of EDTA blood from a donor cat infected with both Mhf and Mhm. Aliquots of blood from this donor cat had been treated with glycerol to a concentration of 10%, and stored for several months. Cats received blood from different aliquots of the inoculum, which was initially confirmed to be infected with Mhf alone using cPCR just before inoculation. Subsequent testing using the newly developed real-time PCR assays revealed that the inoculum actually contained both Mhm and Mhf. At the time of inoculation, methylprednisolone acetate (10 mg/kg [4.5 mg/lb], IM) was administered to the splenectomized cats as previously described. 1 Cats were examined daily for 6 wk after inoculation for signs of illness. Rectal temperatures were determined daily for the first week postinoculation (PI), and every other day thereafter. Cats developing severe anemia were treated with doxycycline (10 mg/kg PO, q24h) for 2–5 days, until there was a significant rise in the hematocrit accompanied by clinical improvement.
Sample collection. Blood samples (approximately 0.1–0.2 ml) were collected daily during the 6-wk postinoculation period using jugular venipuncture for determination of PCV, TP, and preparation of peripheral blood smears. Approximately twice weekly (days 3, 5, 7, 10, 13, 17, 20, 22, 27, 31, 38, 41, 45, 48, 52, and 55), 2 ml of blood was collected from each cat into EDTA tubes for conventional and real-time PCR. Thus, a total of 64 samples were analyzed.
Sample analysis. On the day of sample collection, peripheral blood smears were stained with Giemsa and examined for evidence of hemoplasma infection under oil at 100 × magnification. This was performed by one of the authors (JS) without knowledge of PCR results. For PCR analysis, DNA was extracted from 200 μl EDTA blood using a commercially available kit e . DNA was eluted with 200 μl of elution buffer, and 3 separate aliquots were subjected to conventional PCR, real-time PCR for Mhf, and real-time PCR for Mhm. As a measure of the effects of contamination on assay performance, a blood sample from a single client-owned healthy control dog, previously found to be negative for hemoplasmas using conventional PCR, was extracted concurrently. The positive control for detection of Mhf consisted of DNA extracted from an aliquot of the inoculum. For Mhm, the positive control consisted of DNA extracted from a cat that had been experimentally infected solely with Mhm.
Results
Plasmid studies. Mixing the Mhm and Mhf plasmids at different ratios did not affect the amplification efficiencies of the 2 TaqMan systems. The CT values obtained for Mhm indicated that the TaqMan assays detected the Mhm DNA in presence of Mhf DNA without considerable loss of signal; of the 103 molecules of Mhf DNA spiked into varying amounts of Mhm DNA an average of 852 ± 19 molecules were recovered (coefficient of variation 2.18%), and of the 10 molecules of Mhf DNA spiked into varying amounts of Mhm DNA an average of 6 ± 0.6 molecules were recovered (coefficient of variation 9.85%). From the 104 molecules of Mhm DNA spiked into varying amounts of Mhf DNA, an average of 8,950 ± 1,453 molecules were recovered (coefficient of variation 16.23%). From the 104 molecules of Mhf DNA spiked into varying amounts of Mhm DNA, an average of 22,700 ± 1823 molecules were recovered (coefficient of variation 8.05%), which was actually higher than expected but within the same log order of magnitude. All 4 standard curves displayed relatively unaltered amplification efficiencies of 95.6%, 95.3%, 98.8%, and 98.5%, respectively (efficiencies with Mhm and Mhf plasmids when tested separately were 95.9% and 95.3%, respectively).
Results of blood smear evaluation. Organisms were noted on blood smears in all cats at various time points in the study. The sampling times that organisms were seen on blood smears relative to the results of conventional and real-time PCR are shown in Table 1.
Results of real-time and conventional PCR in experimentally infected cats. Feline glyceraldehyde-3-phosphate dehydrogenase (GAPDH) DNA was amplified from all 64 samples. For 49 of the 64 (77%) samples, the results of both conventional and real-time PCR assays were positive for hemoplasma DNA (Table 1). Copy numbers in these 49 samples were calculated to range from 235 to 7.8 × 108 copies of hemoplasma DNA/ml of blood. In 3/64 samples (5%), the results of real-time PCR alone were positive. Copy numbers in these 3 samples were calculated as 553 to 1.8 × 105/ml. In another 4/64 samples (6%), the results of conventional PCR alone were positive. Thus, the number of samples with positive results was similar for both methods (53/64 [83%] for cPCR vs. 52/64 [81%] for real-time PCR).
Comparison of the results of conventional PCR, real-time PCR, and cytologic examination of peripheral blood smears for detection and differentiation of M. haemofelis and ‘Candidatus M. haemominutum’ in cats inoculated simultaneously with both organisms.*
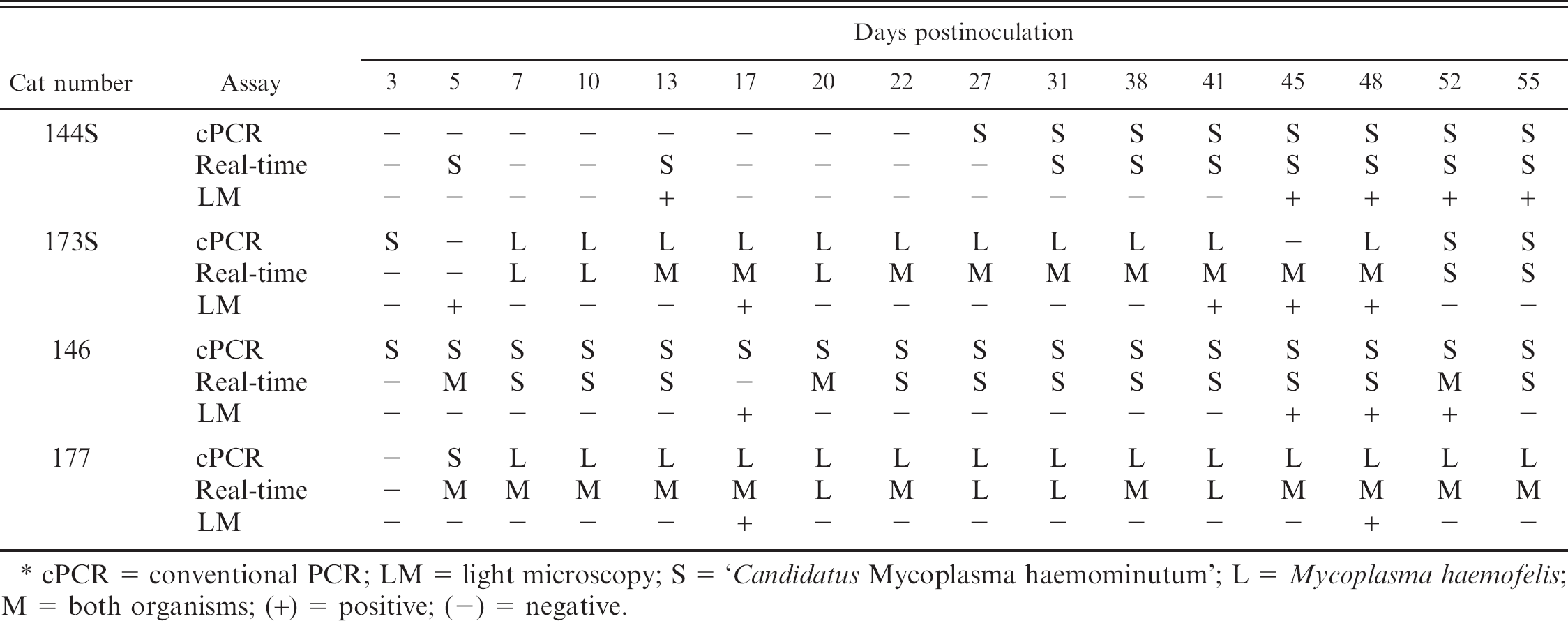
cPCR = conventional PCR; LM = light microscopy; S = ‘Candidatus Mycoplasma haemominutum’; L = Mycoplasma haemofelis:, M = both organisms; (+) = positive; (-) = negative.
Mhm was detected in blood samples from all 4 cats, with 45/64 (70%) of the samples testing positive for this organism using real-time PCR. Blood samples from cats 146, 173S, and 177 also tested positive for Mhf, but most samples collected from cat 146 tested positive only for Mhm as determined using real-time PCR (Table 1). A total of 30/64 (47%) samples tested positive for Mhf using real-time PCR.
The presence of Mhm alone was detected in 22 samples using real-time PCR. The results of cPCR also suggested the presence of this organism alone in 20 of these samples. In the other 2 samples, the results of cPCR were negative. The presence of Mhf alone was detected in 7 samples using real-time PCR. The results of cPCR also suggested the presence of this organism alone in these 7 samples.
The presence of both organisms was identified using real-time PCR in 23 samples, as well as both aliquots of inoculum used to infect the cats. However, the results of cPCR suggested a mixed infection in only 1 aliquot of the inoculum, and not the other aliquot or the 23 postinoculation samples. In the 23 postinoculation samples testing positive for both organisms with realtime PCR, the results of cPCR suggested infection with Mhf alone in 18 samples, Mhm alone in 4 samples, and were negative in 1 sample.
Discussion
In this study, we assessed the ability of multiple separate real-time PCR assays to detect coinfections with Mhm and Mhf, both in vitro and then in experimentally infected cats as compared with conventional PCR. To validate the assays in light of the known pitfalls of competitive inhibition, 2 separate real-time PCR assays for Mhm and Mhf were first applied to mixtures of plasmids in different ratios. The results of these assays were then compared to the results of cPCR when performed on samples collected from cats that had been experimentally infected with both Mhf and Mhm. The in vitro experiments showed that the results of quantitation with each of the 2 realtime PCR assays when applied to mixtures of DNA from each species over the range of concentrations tested approximately (within the same log order of magnitude) reflected the number of genome copies present. Additional work in our laboratory has shown that at plasmid ratios higher than 3 logs, some inhibition of detection of the organism present in lower quantities occurs, possibly because of consumption of primers, polymerase enzyme, and dNTPs as a result of amplification of the 16S rDNA of the species present in larger quantities in the reaction mixture by the Mhf primers, which could also prime PCR for Mhm DNA (data not shown). Further refinement in the ability of real-time PCR to detect coinfections will require development of assays with both primers and probes that target species-specific portions of the genome.
Even with these limitations, the use of multiple real-time PCR assays permitted detection of mixed infections when cPCR did not. This is not surprising given that the primers used for detection of both species in cPCR are the same, and species discrimination is based only on the size of the PCR product. Both species were detected in 25 samples (including the 2 aliquots of inoculum) using real-time PCR, but only 1 of these samples (an aliquot of inoculum) was positive for both species using cPCR. Mixed infections have been documented in naturally infected cats using this cPCR assay. 6 , 14 However, the results of this study suggest that use of cPCR alone to determine the epidemiology of hemoplasma infections may have limitations and possibly underestimate the true prevalence of mixed infections with these organisms.
The results of both real-time and cPCR always agreed on the species present in the 27 samples where only 1 species was detected using real-time PCR. It was sometimes difficult to accurately judge the size of the cPCR product because the difference between the 2 product lengths is small (23 base pairs). However, the results of performing separate real-time PCR assays for Mhf and Mhm allowed unequivocal differentiation between Mhf and Mhm.
Overall, the number of positive samples using the 2 real-time PCR assays and cPCR for detection of hemoplasmas in cats that had been experimentally infected with both Mhm and Mhf were similar, with a total of 49 of 64 samples testing positive using both assays, 4 samples testing positive using cPCR only, and 3 samples testing positive using real-time PCR only. The reasons for these discrepancies are unclear, but possible explanations include low organism loads or inhibitory factors present in the extracted sample. One sample testing negative using cPCR but positive using real-time PCR had a relatively low organism load (<1,000/ml), but this alone could not explain the failure of cPCR to detect hemoplasmas in samples that were positive using real-time PCR, because 4 samples containing approximately the same or lower organism loads tested positive using cPCR. The quantity of feline GAPDH in samples in which only the results of cPCR were positive was not significantly lower than that in samples testing positive using both assays, so inhibitory factors did not seem to be the reason for the reduced sensitivity of real-time PCR in these samples.
All of the samples from 1 cat in this study tested positive only for Mhm whereas multiple samples from the other 3 cats had evidence of infection with both organisms. This may reflect the relative numbers of organisms in the aliquots of inoculum used to infect each cat. Retrospectively, our analysis revealed that one of the aliquots contained 2.6 × 106 copies of Mhf/ml and 1.4 × 104 copies of Mhm/ml, whereas the other contained 3.6 × 104 copies of Mhf and 3.2 × 105 copies of Mhm/ml. The reason for this is unclear, but it is possible that the aliquots had been collected at different time points from the donor cat or sample degradation during storage affected the quantity of DNA in each aliquot.
In some samples, the presence of one organism was sometimes not identified using real-time PCR, possibly because of extremely low organism loads at these time points. Alternatively, inhibition may have occurred if the ratio of organism loads was beyond the range tested using the plasmid dilutions. Failure to detect a coinfection was particularly true for cat 146, where 11/16 postinoculation samples were positive for Mhm and 3/16 samples were positive for both organisms. Thus, a single positive PCR result for 1 organism may not rule out the presence of a coinfection, even when multiple real-time PCR assays are used.
In summary, this study has revealed a similar sensitivity of conventional and real-time PCR for detection of hemoplasma in blood samples collected from experimentally infected cats. However, cPCR was not useful for detection of mixed infections in blood samples collected from cats that were infected with both organisms, probably as a result of competitive inhibition in the assay. In addition, without sequencing of the PCR product, conventional PCR does not differentiate between Mhf and Mtc, 18 nor does it differentiate between Mhm and Mhp. The use of multiple separate real-time PCR assays should thus be considered for studies of the epidemiology of hemoplasmosis in cats, although the ability of the assays reported here to accurately quantify organism numbers in co-infected cats may be limited.
Acknowledgements
The authors would like to acknowledge Claire Allen, and Drs. Catharina Brömel and Elissa Kadar for their assistance with sample collection and care and maintenance of the cats used in this study. We also thank Drs. John Madigan, Niels Pedersen, and Janet Foley for their support and encouragement. This study was supported in part by the Center for Companion Animal Health, School of Veterinary Medicine, at the University of California, Davis.
Footnotes
a.
Applied Biosystems, Inc. Foster City, CA.
b.
Dyad thermocycler, MJ Research Inc., Waltham, MA; Invitrogen Corporation, Carlsbad, CA.
c.
TOPO TA cloning kit, Invitrogen, CA.
d.
Davis sequencing, Davis, CA.
e.
QIAmp blood and tissue kit, Qiagen Inc., Chatsworth, CA.
