Abstract
This communication reports the development and performance assessment of a rapid diagnostic test for identifying horses actively infected with the neurovirulent pathotype of equine herpesvirus-1 (EHV-1). The test is a real-time polymerase chain reaction (PCR)-based assay that uses EHV-1 pathotype-specific TaqMan® reporter probes for discrimination between neuropathogenic and non-neuropathogenic strains of EHV-1 in equine blood or nasal swabs. The diagnostic performance of the new technique was evaluated by testing specimens collected from 234 horses involved in recent outbreaks of EHV-1 myeloencephalopathy at three separate thoroughbred racetracks and one large riding/boarding stable. Side-by-side comparison of the EHV-1 pathotyping results yielded by the new single-step, PCR-based allelic discrimination technique (24-hour turn-around-time) with those generated by a multi-step, conventional nested PCR followed by nucleotide sequencing of the amplified DNA (4-day turn-around-time) revealed complete agreement between the 2 test methods. The ability to rapidly identify horses infected with neuropathogenic strains of EHV-1 using a single-step, PCR-based method has significant implications for future diagnostic evaluation of suspect animals.
The recent upsurge in the number of large, high-mortality outbreaks of equine herpesvirus-1 (EHV-1) neurological disease poses an emerging threat to equine health and to the economic prosperity of horse-related businesses. 2,4,7,9 Rapid, laboratory-based diagnosis of the disease is important for prompt implementation of infection-control measures designed to minimize transmission of the virus to other horses at the outbreak site.
Recent experimental studies have identified a single nucleotide polymorphism (SNP) within the EHV-1 gene encoding the viral DNA polymerase (open reading frame-30 [ORF30]) that is highly associated (p {LT} 0.0001) with the viral attribute of neuropathogenicity for horses. 6 Of the 32 investigated outbreaks of EHV-1 neurological disease that occurred in the United States and the United Kingdom between 2001 and 2006, 30 (94%) were caused by the ORF30 mutant strain of EHV-1 (G. Allen and N. Davis-Poynter, unpublished data). The only method currently available for identification of isolates of EHV-1 as either mutant (ORF30G2254) or wild type (ORF30 A2254) is the specialized, time-consuming technique of nucleotide sequence determination at the identified polymorphic site of the viral genome. 1 Because of multiple obstacles to the adaptability of DNA sequencing to the high throughput environment of veterinary diagnostic laboratories, there remains an urgent need for a more practical test for rapid identification of horses actively infected with mutant, neuropathogenic strains of EHV-1.
The objective of this study was to address this unmet diagnostic need by developing a rapid, inexpensive, and diagnostic laboratory-friendly method for identification of the SNP genetic marker that distinguishes neuropathogenic from nonneuropathogenic strains of EHV-1. The technique for rapid SNP discrimination using real-time PCR and allele-specific TaqMan® detection probes was first described in 1995. 5 The technique is a duplexed, end-point PCR assay that detects sequence variants at a single SNP site in a target DNA fragment. A pair of uniquely labeled TaqMan fluorescent probes are used, one a perfect match to the wild-type DNA sequence and the other a perfect match to the mutated sequence. The assay measures and compares the level of fluorescence associated with hybridization of the 2 TaqMan detection probes to the amplified target DNA at the end of PCR amplification.
Such an allelic discrimination, real-time PCR assay was developed for simultaneous detection and pathotype identification of EHV-1 DNA in nasal secretions or the buffy coat fraction of venous blood collected from horses. Oligonucleotide primers for amplification were designed to be specific for EHV-1 and to amplify a 145-bp ORF30 gene fragment that encompasses the site of the mutation (ORF30 A2254 → G) highly associated with neuropathogenic strains of EHV-1 6 (Table 1). Two EHV-1 pathotype-specific TaqMan detector probes were synthesized and labeled with separate fluorescent dyes, VIC a and 6-FAM a
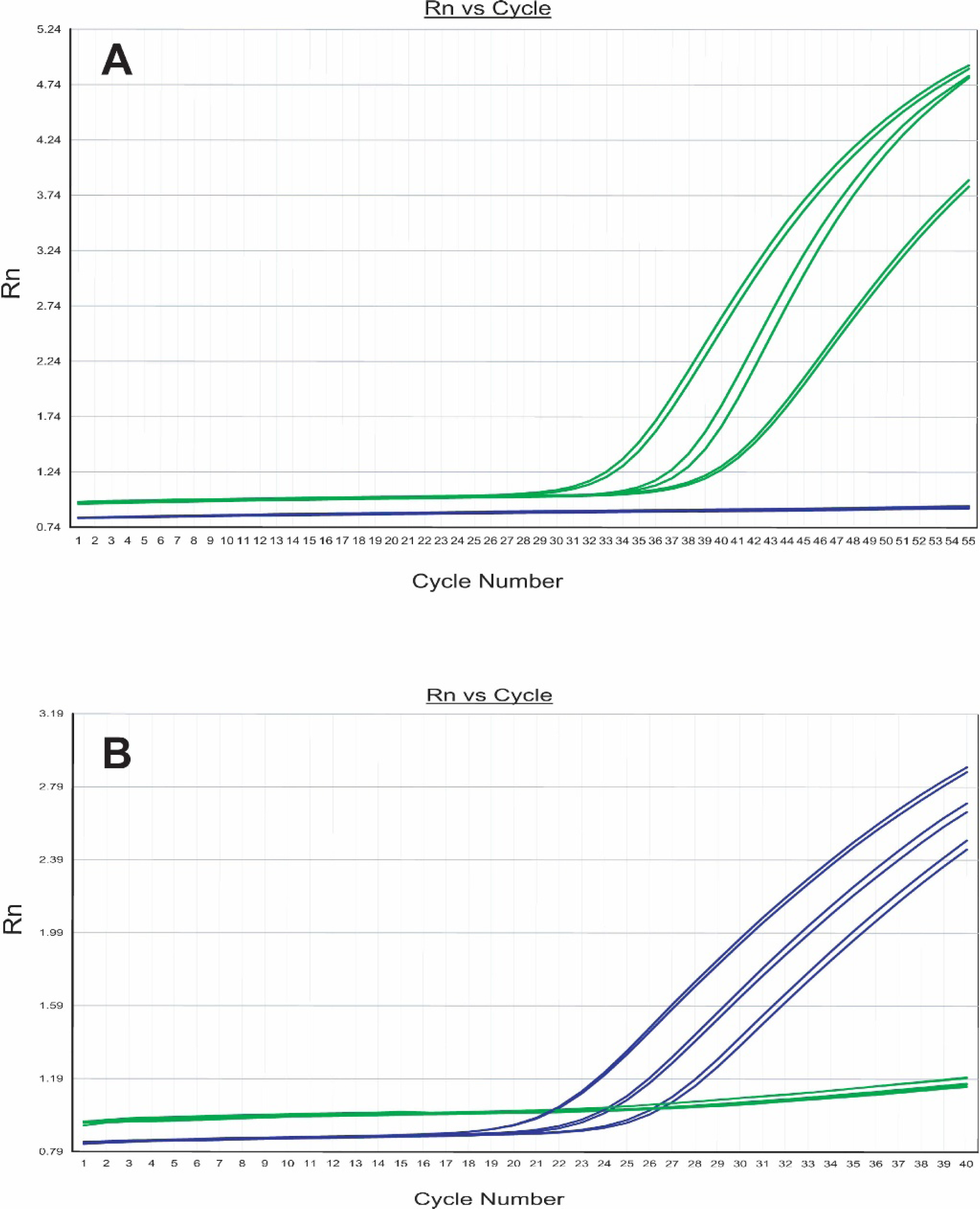
Illustration of ability of described real-time PCR assay to detect EHV-1 DNA in diagnostic equine specimens and to discriminate between ORF30 A2254 wild type and ORF30 G2254 mutant strains of the virus. Shown are real-time PCR amplification plots for EHV-1 DNA isolated from blood leukocytes of 3 horses infected with neuropathotype strains
(Table 1). Placement of the mismatched nucleotide at the center of each TaqMan detector oligonucleotide generated probes possessing 0% binding capacity to the heterologous viral DNA at the annealing temperature (65°C) of the PCR assay (Primer Express Software a ). For each test sample, duplicate real-time PCR reactions were run, each containing 1 μg of test DNA in 25 μl reaction mixture consisting of TaqMan Universal PCR Master Mix a (50 U/ml Taq DNA polymerase; 200 mmol/l each of dATP, dCTP, dGTP, and dTTP; 2.5 mmol/l MgCl2; 50 mmol/l KCl; 1.0 mmol/l EDTA; and 10 mmol/l Tris-HCl, pH 8.3) with 900 nmol/l each of forward and reverse amplification primers and 200 nmol/l each of the 2 pathotype-specific, fluorescent TaqMan probes. Reaction mixtures containing nontemplate water controls, DNA from known neuropathotype or nonneuropathotype EHV-1 isolates (positive controls), and EHV-4 DNA (specificity control) were included in each amplification run. The PCR assay was performed in 96-well plates in an ABI 7500 fast real-time PCR system. a Amplification parameters consisted of an initial denaturation step of 95°C for 10 minutes followed by 55 cycles of 95°C for 15 seconds and 65°C for 1 minute. The time requirement for completing the test, from initial receipt of a diagnostic specimen to the generation of reportable results, was approximately 8 hours.
Oligonucleotide primers and TaqMan® probes used in real-time PCR assay.

Differences in detection probe sequences are shown in bold.
MGB = minor groove binder.
NFQ = nonfluorescent quencher.
Post-run analysis for detection and pathotype identification of EHV-1 DNA present in each test sample was performed by the ABI 7500 SDS allelic discrimination analysis software (version 1.3.1). a Amplification in real time of each genotype of EHV-1 DNA (wild type or neurologic mutant) present in the samples test wells is exhibited by the software as a plot of PCR cycle number versus accumulated level of fluorescence (Rn) from each of the 2 TaqMan reporter probes (Fig. 1). Results of the PCR assay for client reporting and archival storage are outputted by the analysis software as a spreadsheet that provides information, for each sample tested, on test results of the control wells, presence or absence of detectable EHV-1 DNA in the sample, and the neuropathic genotype of any EHV-1 strain detected.
The diagnostic performance (i.e., sensitivity and specificity) of the real-time allelic discrimination PCR test was assessed in regard to both 1) detection of EHV-1 infection in horses, and 2) identification of the infecting EHV-1 strain as possessing either the neuropathogenic or nonneuropathogenic genotype. Evaluation of the assay was performed by the testing of samples submitted from horses involved in 4 recent outbreaks of EHV-1 neurological disease. The study sample of animals consisted of 234 adult horses (196 Thoroughbred and 38 miscellaneous breed horses) known to have been exposed to EHV-1 during large myeloencephalopathy epizootics that occurred between 2003 and 2006 at three thoroughbred race tracks and a large boarding/riding establishment located in the United States.
Total DNA was isolated from submitted equine nasal swabs and/or the leukocyte fraction of submitted EDTA-blood samples by 1) binding to glass fiber-containing spin tubes in the presence of a chaotropic salt, b or 2) the sequential-step isolation technique of cell lysis, salt precipitation of proteins, and isopropanol precipitation of DNA, c respectively. The DNA concentrations in the sample preparations were determined by measuring the spectrophotometric absorbance at 260 nm, and 1-μg DNA samples were tested in duplicate by both 1) the real-time PCR assay described herein, and 2) by conventional nested PCR followed by nucleotide sequence determination of the amplified ORF30 DNA fragment. 1 The test results generated by the 2 diagnostic formats were compared and used to calculate the relative sensitivity and specificity of the new real-time PCR assay, using the conventional nested PCR/DNA sequencing test as the reference test for comparison (Table 2). Relative to the performance of conventional nested PCR, the sensitivity and specificity of the real-time PCR assay for detection of EHV-1 infected horses were 96.3% and 100%, respectively. When compared with DNA sequencing, the sensitivity and specificity of the real-time PCR assay for identification of the neuropathic genotype present in EHV-1 positive horses were both 100%. The assessment verified real-time allelic discrimination PCR as a rapid, sensitive and specific diagnostic tool for use in simultaneously detecting the presence of EHV-1 DNA in equine clinical specimens and, without additional steps, for identifying the neuropathic genotype of the infecting EHV-1 strain. Importantly, the new test allowed detection of simultaneous co-circulation of both EHV-1 pathotypes within the same outbreak (Table 2). In such cases, only horses infected with mutated strains of EHV-1 developed neurological disease.
Evaluation of diagnostic performance of allelic discrimination, real-time PCR assay for detection and pathotype identification of EHV-1 DNA in clinical specimens. *
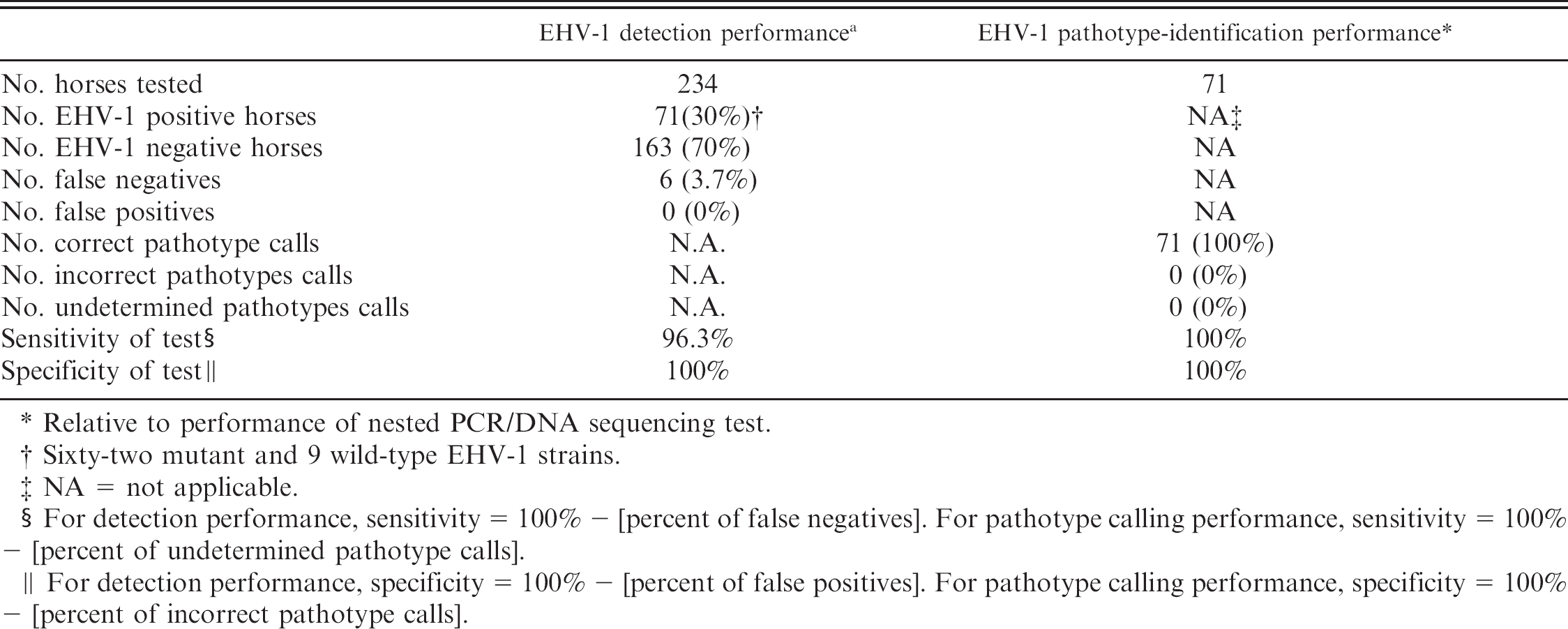

Relative to performance of nested PCR/DNA sequencing test.
Sixty-two mutant and 9 wild-type EHV-1 strains.
NA = not applicable.
For detection performance, sensitivity = 100% -[percent of false negatives]. For pathotype calling performance, sensitivity = 100% — [percent of undetermined pathotype calls].
For detection performance, specificity = 100% — [percent of false positives]. For pathotype calling performance, specificity = 100% — [percent of incorrect pathotype calls].
A carefully worded laboratory reporting of the meaning of the new assay results will be important for avoiding their unjustified over-interpretation by veterinary clientele; e.g., “relative to infection of a horse by ORF30 A2254 wild-type strains of EHV-1, infection by ORF30 G2254 mutant virus strains carries a greater risk for development of neurologic complications.”
Acknowledgements. The author acknowledges with gratitude the receipt of financial support for this study from the Keeneland Association, Grayson-Jockey Club Research Foundation, Columbia Horse Center, and the University of Kentucky Gluck Equine Research Foundation. Published as Paper No. 06-14-093 of the Kentucky Agricultural Experiment Station.
Footnotes
a.
Applied Biosystems, Foster City, CA.
b.
High Pure PCR Template Preparation Kit, Roche, Indianapolis, IA.
c.
Wizard Genomic DNA Purification Kit, Promega, Madison, WI.
