Abstract
Oncogene-induced senescence occurs following oncogene activation in normal cells and is considered as a critical tumor-suppressing mechanism. Ubiquitin-specific protease 10 (USP10) has been reported to play a vital role in oncogene-induced senescence via the deubiquitination-dependent stabilization of p14ARF. However, knowledge of the clinical significance of USP10 and p14ARF expression in patients with small intestinal adenocarcinoma is limited. To study the clinical significance of USP10 and p14ARF expression, we performed immunohistochemistry for USP10 and p14ARF on 195 surgically resected small intestinal adenocarcinoma specimens. Furthermore, we performed methylation analysis on five small intestinal adenocarcinoma samples and matched adjacent normal intestinal tissue samples. UPS10 (
Introduction
Small intestinal adenocarcinoma is a rare tumor and accounts for only 2% of gastrointestinal tract malignancies. 1 There were 10,190 estimated new cases of small intestinal adenocarcinoma in 2017 in the United States. 2 The diagnosis of small intestinal adenocarcinoma can be challenging due to the lack of effective diagnostic tools and non-specific clinical symptoms at advanced stages.1,3 This can negatively affect the survival rate; small intestinal adenocarcinoma remains a global public health problem with a 5-year overall survival rate of 14%–45%. 4
Oncogene-induced senescence, a permanent type of growth arrest, plays a protective role against oncogenic stress-mediated transformation, and it is a mechanism of natural tumor suppression. 5 This has led to increased interest in pathways that mediate oncogene-induced senescence. Oncogene-induced senescence is independent of telomere shortening 6 and requires the involvement of the p19ARF-p53 and p16INK4A-RB tumor-suppressor pathways.7–10 Given that oncogene-induced senescence is composed of multiple mechanisms in response to oncogene activation, the molecular mechanism of oncogene-induced senescence remains unclear. Accordingly, the assessment of oncogene-induced senescence-related proteins in clinical tumor samples will contribute to the understanding of the molecular mechanism of oncogene-induced senescence, and this knowledge could potentially lead to the development of better therapeutic approaches and increased survival of patients.
Ubiquitin-specific protease 10 (USP10) is a member of the ubiquitin-specific protease (USP) family, which specifically cleaves ubiquitin from ubiquitin-conjugated protein substrates, and its related pathways are known to be involved in apoptosis, autophagy, and DNA damage.11,12 Recently, USP10 has been reported to be a novel regulator of p53, providing an alternative mechanism of inhibiting p53 in cancers with wild-type
p14ARF is encoded by an alternative reading frame within the
In this study, we assessed the prognostic significance of USP10 and p14ARF in formalin-fixed paraffin-embedded (FFPE) small intestinal adenocarcinoma specimens by immunohistochemistry and quantitative image analysis. In addition, we investigated the potential association between the loss of USP10 and p14ARF protein expression and methylation.
Materials and methods
Patients and tumor specimens
A total of 195 cases of surgically resected small intestinal adenocarcinoma were retrieved from 22 South Korean hospitals and institutions by the Korean Small Intestinal Cancer Study Group, as previously reported. 1 All procedures were conducted in accordance with the Declaration of Helsinki. Tissue samples were collected from patients who signed the written informed consent form, which was approved by the Regional Institutional Review Board of Asan Medical Center (approval no. S2016-1710-0001). The tumors included primary carcinomas in the duodenum, jejunum, and ileum but excluded carcinomas arising from other gastrointestinal tract sites and the ampulla of Vater, pancreas, cecum, and appendix. Similar to a previous study, 1 clinical and pathological information was recorded, including location, size, histological subtype, differentiation, growth pattern, primary tumor (pT) category, retroperitoneal seeding, other intestinal loop invasions, metastasis to lymph node, and presence of perineural and lymphovascular invasion.
Tissue microarray and immunohistochemistry
Tissue microarray cores were constructed from archival FFPE tissue blocks as previously described.25,26 Briefly, tissue cylinders with a 2-mm diameter were punched from tumor areas of each FFPE tissue block and then brought into a recipient block, including matched normal small intestine. The tissue microarray blocks were cut into serial 5-µm-thick sections, and the sections were deparaffinized in xylene and rehydrated through a graded alcohol series to distilled water. Heat-induced antigen retrieval was performed for 20 min in pH 6 citrate antigen retrieval buffer (Dako, Carpinteria, CA, USA) and in a pH 9.0 buffer for USP10 and p14ARF, respectively. Endogenous peroxidase activity was quenched with 3% H2O2 in water for 10 min. The sections were incubated with anti-USP10 rabbit polyclonal antibody (cat. no. ab72486; Abcam, Cambridge, MA, USA) at 1:1000 dilution for 1 h and anti-p14ARF mouse monoclonal antibody (clone no. 4 C6/4; Cell Signaling Technology, Danvers, MA, USA) at 1:1000 dilution for 1 h. The antigen–antibody reaction was detected with EnVision+ Dual Link System-HRP (Dako) and DAB+ (3, 3′-diaminobenzidine; Dako). The stained sections were lightly counterstained with hematoxylin. The human gastric mucosa was used as a positive control for both USP10 and p14ARF immunoreactivity. Negative controls (IgG and omission of the primary antibody) were concurrently used.
Immunohistochemically stained slides were scanned by NanoZoomer 2.0 HT (Hamamatsu Photonics K.K., Hamamatsu, Japan) at ×20 objective magnification (0.5 μm resolution). The captured images were analyzed using Visiopharm software version 4.5.1.324 (Visiopharm, Hørsholm, Denmark). A brown staining intensity (0 = negative, 1 = weak, 2 = moderate, and 3 = strong) was obtained using a predefined algorithm and optimized settings.27,28 The overall immunohistochemical score (histoscore) was expressed as the percentage of positive cells multiplied by their staining intensity (possible range: 0–300).
DNA methylation analysis
Genomic DNA was isolated from five normal tissue samples and five primary small intestinal adenocarcinoma samples using a standard phenol–chloroform method. The DNA (2 μg) was modified with bisulfite using the EZ DNA Methylation Kit™ (Zymo Research, Orange, CA, USA). Methylation analysis of gene promoters was performed using methylation-specific PCR (MSP) primer pairs located close to the putative transcription start site in the 5′ CpG island with 2 μL of bisulfite-treated DNA as the template and JumpStart REDTaq DNA Polymerase (Sigma–Aldrich, St. Louis, MO, USA) for amplification as previously described.
29
Statistical analysis
The association between protein expression and clinicopathological characteristics was determined using the chi-square test. For survival analysis, expression values were dichotomized (positive vs negative) with the cut-off values showing the most discriminative power (histoscore of 237 for USP10 and 129 for p14ARF). Overall survival according to protein expression was analyzed using the Kaplan–Meier method, and the survival curves were compared with the log-rank test. Subsequently, a Cox multivariate proportional hazards model was used for analysis and adjusted for the following clinical/pathological variables: histologic subtype, pT category, lymph node metastasis, other loop invasions, retroperitoneal seeding, perineural and lymphovascular invasions, sporadic adenoma, and peritumoral dysplasia. Statistical analysis was performed using SPSS version 21.0 (SPSS, Chicago, IL, USA) and R statistical package (version 3.1.2).
Results
Expression of USP10 and p14ARF proteins
Normal epithelial cells in the crypts of the small intestine and epithelial plasma cells were stained by USP10 as a diffuse cytoplasmic pattern. The cytoplasm and nucleus were stained by p14ARF in the normal small intestinal mucosa with an apicoluminal pattern (Figure 1, left column), whereas the tumor cells were stained with diffuse cytoplasmic and nuclear patterns (Figure 1, middle and right columns). Although the TMA contains 195 cases of small intestinal adenocarcinoma and matched normal, 194 out of 195 cases were found suitable for USP10 evaluation due to the complexity of sectioning and heterogeneity. In total, 70 (36.1%) patients showed immunoreactivity for USP10, and 124 (63.9%) patients showed loss of immunoreactivity for USP10. In addition, 120 (61.5%) patients were immunopositive for p14ARF, and 75 (38.5%) patients showed loss of immunoreactivity for p14ARF. USP10 expression status was positively correlated with p14ARF expression status (
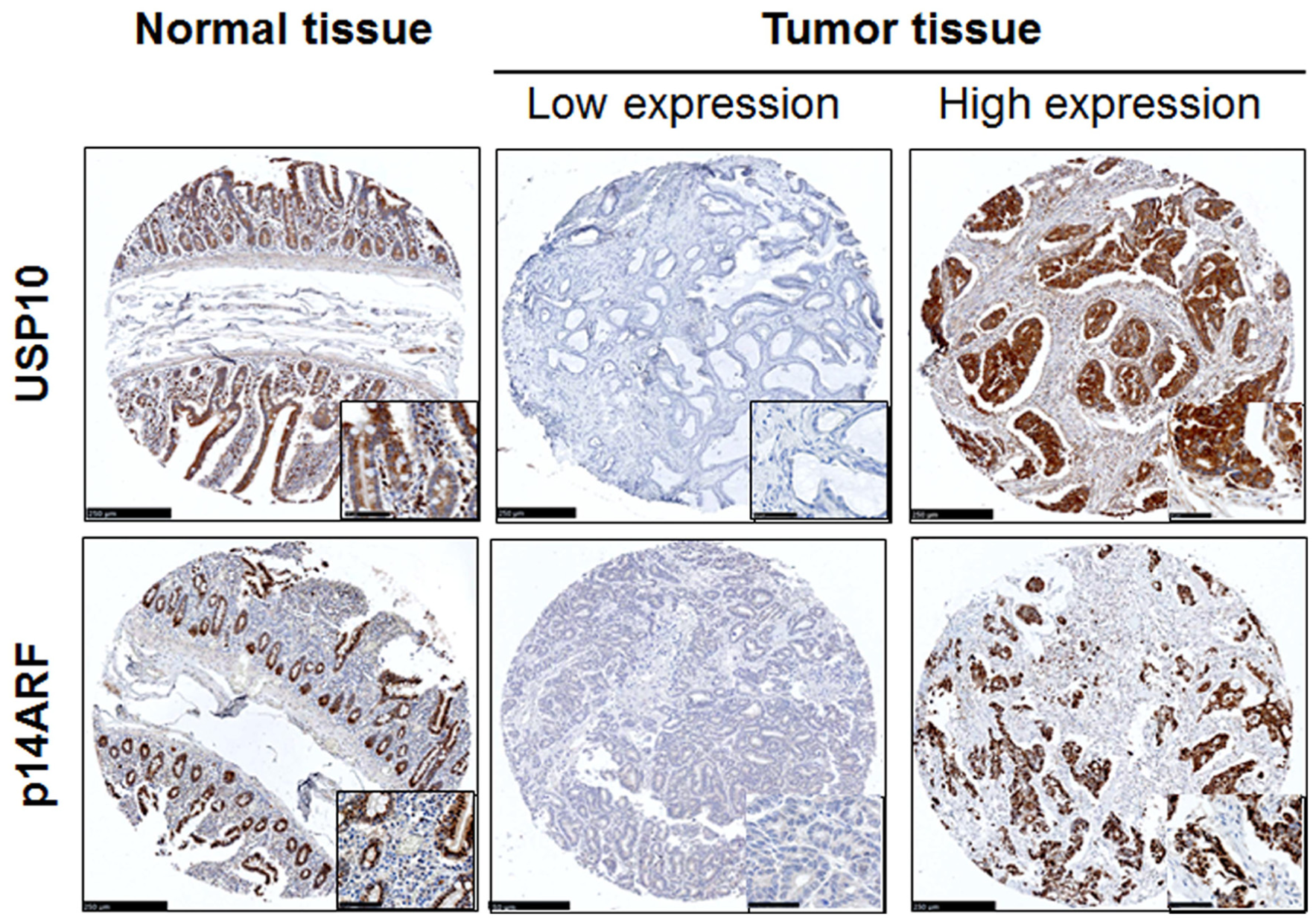
Representative immunohistochemical images of USP10 and p14ARF in formalin-fixed, paraffin-embedded normal and small intestinal adenocarcinoma tissues. Both proteins were expressed in the crypts of the normal small intestine with cytoplasmic and nuclear staining patterns (left column), while both proteins stained with diffuse cytoplasmic and nuclear in tumor cells (middle and right columns). High magnification images are shown in the inset. Scale bar = 250 µm (50 µm in inset).

Correlation between USP10 and p14ARF expression. (a) The expression of USP10 was positively correlated with the expression of p14ARF (
Clinical and pathological characteristics according to USP10 and p14ARF expression
The USP10 and p14ARF expression in small intestinal adenocarcinoma were significantly lower than that in corresponding adjacent normal tissue (

Expression of USP10 and p14ARF in normal and cancerous small intestine tissues. (a) Box plot depiction of IHC data. The histoscores were computed based on intensity and tissue area of positive staining. Small intestinal adenocarcinoma tissue exhibited significantly lower USP10 (
USP10 and p14ARF expression and association with clinicopathological factors in patients with small intestinal adenocarcinoma.

LN: lymph node; pT: primary tumor.
Statistically significant (
Prognostic significance of USP10 and p14ARF protein expression
Patients with the loss of USP10 expression (median survival: 45.1 months) had a significantly worse overall survival compared with those with intact USP10 expression (median survival: 98.4 months, hazard ratio (HR) = 3.02, 95% confidence interval (CI) = 1.91–4.77,
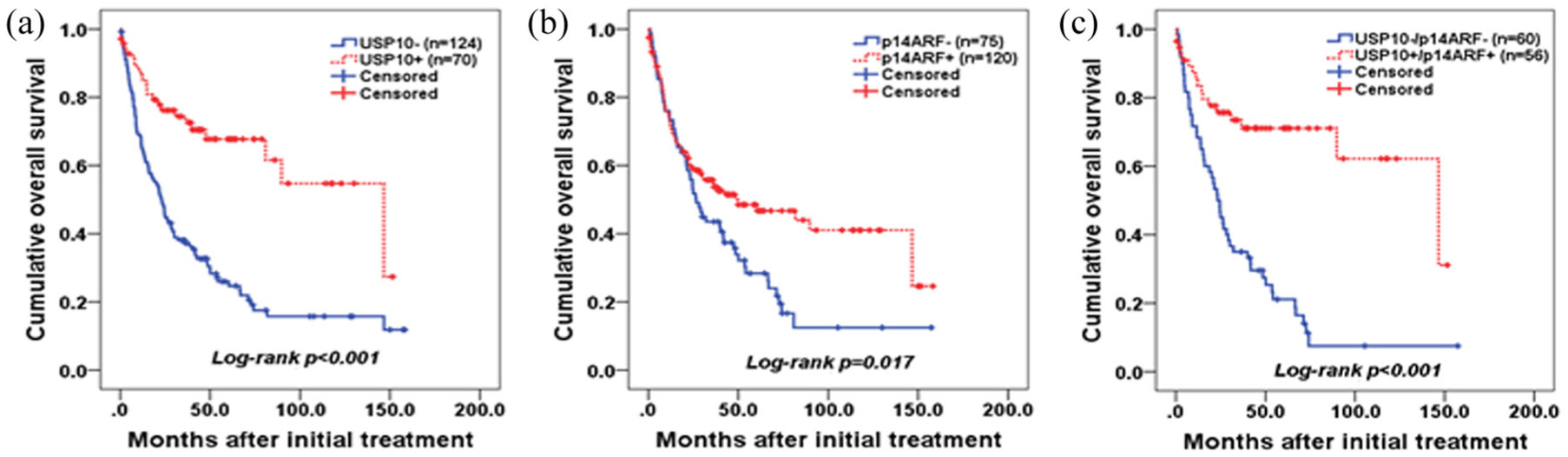
Overall survival analysis. (a, b) Loss of USP10 (
Other parameters including a higher pathological T stage (
Univariate analysis of pathological features affecting patient survival with small intestinal adenocarcinoma.

CI: confidence interval; HR: hazard ratio; pT: primary tumor; LN: lymph node.
Cannot be calculated because more than 50% of patients were alive.
Significant at
Multivariate analysis revealed that the loss of USP10 expression was an independent poor prognostic factor (HR = 3.04, 95% CI = 1.79–5.18,
Multivariate analysis of small intestinal adenocarcinoma.

HR: hazard ratio; CI: confidence interval; pT: primary tumor; LN: lymph node.
Results of exclusion of loss of USP10 expression and loss of p14ARF expression from multivariate analysis.
Significant at
Regulation of USP10 and p14ARF downregulation by promoter hypermethylation
Gene silencing is known to be associated with promoter DNA hypermethylation in human cancers.
31
In addition,

Bisulfite sequencing analysis of the CpG islands in the
Discussion
Due to the rarity of small intestinal adenocarcinoma, there is a lack of sufficient data to adequately characterize the patient population. Moreover, the immunophenotyping and molecular markers that can aid early detection and treatment are limited. Therefore, there is a great need to identify potential molecular markers that will provide prognostic and predictive value for small intestinal adenocarcinoma. In this study, we demonstrated an association between USP10 and p14ARF protein expression and their combined potential as a prognostic marker for small intestinal adenocarcinoma. We also showed that the downregulation of USP10 and p14ARF expression may be associated with promoter hypermethylation.
Two different modes of cellular senescence have been considered. Replicative senescence is observed in aged cells and attributed to short telomeres typically following excessive cell divisions. Oncogene-induced senescence is triggered by oncogenes, resulting in premature senescence or the apoptosis of cells exposed to oncogenic stress including genotoxic and oxidative stress.
24
Both replicative senescence and oncogene-induced senescence share many common changes in gene expression; nevertheless, differences in triggers and effectors have also been characterized.
35
The DNA methylation of promoter CpG islands is strongly associated with specific genes that play a role in the initiation and establishment of senescence.
36
Grasso et al.
37
reported that Nurpr1, a chromatin protein, modified the effects of KrasG12D-induced senescence by regulating Dnmt1 expression and consequently genomic-wide levels of DNA methylation. Promoter methylation is one of the key epigenetic mechanisms that regulate gene expression, and the aberrant DNA hypermethylation of gene promoter regions is an important inactivation mechanism for silencing tumor-suppressor genes in most cancers.
38
Various studies have found that
USPs play important roles in many cellular processes such as the cell cycle, signal transduction, and transcriptional activation, and several USPs have been implicated in human tumorigenesis.
41
USP10 (a member of the USP family) deubiquitinates p53 and MDM2 (a known regulator of cellular p53) are involved in the reverse translocation and degradation of p53.
13
Previous studies have shown that USP10 can suppress tumor formation by enhancing p53 stability.12,13 Recently, Sun et al.
42
suggest that USP10 acts as a tumor suppressor in lung cancer cells by deubiquitinating and stabilizing PTEN. The loss of USP10 expression has been observed in several types of cancers including renal cell,
13
gastric,
14
lung,
42
and pancreatic cancers.
43
In particular, the decreased expression of USP10 has been reported to correlate with poor prognosis in gastric cancer.
14
Conversely, USP10 overexpression has been reported in breast cancer
16
and glioblastoma,
15
and it may be associated with poor outcomes in glioblastoma
15
and prostate cancer.
44
Yuan et al.
13
demonstrated that USP10 overexpression could suppress the growth of tumor cells with wild-type
p14ARF/CDKN2A is well known as a tumor-suppressor protein and an important mediator of oncogenic p53 activation. p14ARF binds to MDM2 and stabilizes p53 in response to DNA damage or cellular stress.47,48 The transcriptional silencing of the
In summary, we demonstrated that the loss of USP10 expression was associated with advanced tumor phenotypes, and the dual loss of USP10 and p14ARF could be a poor prognostic factor for overall survival in small intestinal adenocarcinoma. Furthermore, the loss of USP10 and p14ARF expression may be regulated by promoter hypermethylation in small intestinal adenocarcinoma. In addition, the p14ARF-USP10 axis may play a role in cancer progression. USP10 and p14ARF protein expression could potentially serve as a novel prognostic indicator for predicting survival time in small intestinal adenocarcinoma, which may be helpful in the management of patients with small intestinal adenocarcinoma. Further elucidation of the underlying molecular mechanism of the USP10-p14ARF axis will contribute to the development of novel therapies for small intestinal adenocarcinoma.
Supplemental Material
Supplementary_material__Rv_(TUB-18-0160) – Supplemental material for Dual loss of USP10 and p14ARF protein expression is associated with poor prognosis in patients with small intestinal adenocarcinoma
Supplemental material, Supplementary_material__Rv_(TUB-18-0160) for Dual loss of USP10 and p14ARF protein expression is associated with poor prognosis in patients with small intestinal adenocarcinoma by Joon Seon Song, Joo Mi Yi, Hanbyoul Cho, Chel Hun Choi, Yoonho Park, Eun Joo Chung, Jaewhan Song, Joon-Yong Chung and Seung-Mo Hong in Tumor Biology
Footnotes
Acknowledgements
The authors thank the Korean Small Intestinal Cancer Study Group for providing tissue samples and sample preparations. This study was presented in part at the 24th American Association for Cancer Research Annual Meeting, Washington, DC, USA, 1–5 April 2017.
Author contribution
Joon-Yong Chung and Seung-Mo Hong designed the study; Joon Seon Song, Joo Mi Yi, Hanbyoul Cho, Chel Hun Choi, and Yoonho Park carried out the experiments; Joon Seon Song, Joo Mi Yi, Hanbyoul Cho, Eun Joo Chung, Jaewhan Song, Joon-Yong Chung, and Seung-Mo Hong performed data analysis of the experiments or clinical records; Joon Seon Song, Joo Mi Yi, and Joon-Yong Chung wrote the manuscript with input from all authors; Eun Joo Chung and Jaewhan Song added critical content to the discussion; and Joon-Yong Chung and Seung-Mo Hong were responsible for revising all portions of the manuscript. All contributors met the criteria for authorship and approved the final manuscript. Dr Joon Seon Song and Dr Joo Mi Yi contributed equally to this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
