Abstract
Lymph node metastasis is the most important prognostic characteristic of colorectal cancer. Carcinoembryonic antigen messenger RNA was shown to detect tumor cells that have disseminated to lymph nodes of colorectal cancer patients and to be at least as good as the hematoxylin and eosin method to predict survival in colorectal cancer patients. CXCL17 was recently shown to be ectopically expressed in colon cancer tumors. Therefore, CXCL17 may serve as prognostic marker alone or in combination with carcinoembryonic antigen. CXCL17 and carcinoembryonic antigen messenger RNA levels were determined using quantitative reverse transcription polymerase chain reaction with RNA copy standard in 389 lymph nodes of 120 colon cancer patients (stages I–IV) and 67 lymph nodes of 12 control patients with inflammatory bowel disease as well as in 68 primary tumors and 30 normal colon tissue samples. Lymph nodes of colon cancer patients were analyzed for CXCL17 and carcinoembryonic antigen protein expression by immunohistochemistry. CXCL17 messenger RNA was expressed in primary tumors at high levels, while it was barely detected in normal colon tissue (p < 0.0001). Similarly, CXCL17 messenger RNA levels were significantly higher in hematoxylin- and eosin-positive (hematoxylin and eosin (+)) lymph nodes compared to hematoxylin- and eosin-negative nodes (p < 0.0001). CXCL17 messenger RNA levels were investigated in lymph nodes grouped according to carcinoembryonic antigen messenger RNA levels: low (–), intermediate (int), and high (+). CXCL17 messenger RNA levels were higher in the carcinoembryonic antigen (int) and carcinoembryonic antigen (+) groups compared to the carcinoembryonic antigen (−) group (p = 0.03 and p < 0.0001, respectively). In lymph nodes of stage III and IV patients, CXCL17 messenger RNA levels correlated with carcinoembryonic antigen messenger RNA levels (p < 0.0001, r = 0.56 and p = 0.0002, r = 0.66, respectively). Staining of consecutive lymph node sections for CXCL17 and carcinoembryonic antigen demonstrated that the same cells expressed both proteins. Altogether, these results indicate that CXCL17 in lymph nodes is expressed by tumor cells. Patients were grouped according to the CXCL17 mRNA levels in the highest lymph node with low levels (−) and high levels (+). CXCL17(+) CC patients showed 2.2 fold increased risk for recurrence (P = 0.03) and decreased mean disease-free survival time of 33 months compared to CXCL17(−) CC patients (P = 0.03). CXCL17(+) carcinoembryonic antigen (int) colon cancer patients showed increased risk for recurrence by 8.3 fold (p = 0.04) and decreased mean disease-free survival time of 46 months compared to CXCL17(−) carcinoembryonic antigen (int) colon cancer patient at follow-up after 12 years (p = 0.02). The presence of tumor cells expressing CXCL17 in regional lymph nodes is a sign of poor prognosis. Analysis of CXCL17 messenger RNA is particularly useful to detect less differentiated colon cancer tumors expressing relatively low carcinoembryonic antigen messenger RNA levels. Thus, CXCL17 messenger RNA in combination with carcinoembryonic antigen messenger RNA may be used as a complementary tool to the hematoxylin and eosin method for detection of poorly differentiated, aggressive tumors.
Keywords
Introduction
The most important prognostic characteristic of colorectal cancer (CRC) is the presence of lymph node metastasis.1–3 Currently, approximately 50% of patients with tumor-cell-positive lymph nodes, that is, stage III (any T N1-2 M0), and about 25% of patients with no detected tumor-cell-positive lymph nodes, that is, stage I (T1-2 N0 M0) and stage II (T3-4 N0 M0) patients, will recur. 4 These relatively dismal results suggest that tumor cells in lymph nodes vary in aggressiveness, that presence of tumor cells in lymph nodes in many cases is missed by the present standard method, that is, examination of hematoxylin and eosin (H&E)-stained tissue sections, and that the treatment modalities for lymph node positive (H&E(+)) CRC patients is inadequate at least partly due to lack of ability to subgroup these patients with respect to tumor aggressiveness. Taken together, these factors lead to both under-treatment of stage I and II patients and over-treatment of some stage III patients. Previously, selected biomarker messenger RNAs (mRNAs) were shown to detect tumor cells that have disseminated to lymph nodes of CRC patients. Carcinoembryonic antigen (CEA) mRNA was shown to be at least as good as H&E to predict survival in CRC patients5,6 and kallikrein-related peptidase 6 was shown to be a prognostic maker. 7
In search for additional biomarker mRNAs that could be used to predict survival in CRC, we recently discovered that the chemokine CXCL17 is ectopically expressed in colon cancer (CC) 8 and therefore potentially could predict survival. Levels of CXCL17 in CC tumors were found to correlate with the levels of the antigen-presenting cell marker CD86, suggesting that CXCL17 expression in colon is related to an inflammatory environment. 8 Moreover, CXCL17 mRNA levels correlated with CCL2 mRNA levels. 8 CCL2 affects accumulation and function of polymorphonuclear myeloid-derived suppressor cells (MDSCs) in colonic carcinogenesis. 9
CXCL17 is a chemokine attracting monocytes, macrophages, and mature- and immature dendritic cells.10–13 CXCL17 is classified as a “dual chemokine,” that is, a chemokine with homeostatic expression in some tissues and in other tissues only under inflammatory conditions. 14
The focus of this study is to determine CXCL17 mRNA expression levels in lymph nodes of CC patients and evaluate their prognostic value, alone and in relation to the expression levels of CEA mRNA. In addition, the cellular distribution of the CXCL17 and CEA proteins in lymph nodes was investigated by immunohistochemistry. CXCL17 was found to be a prognostic marker in CC complementary to CEA and especially informative for patients with relatively low CEA levels in their lymph nodes.
Materials and methods
Patients and tissue specimens for mRNA analysis
Primary tumor specimens from 68 CC patients (median age = 73.5 years, range = 41–88 years; 38 women and 30 men) were retrieved after surgery. Lymph nodes were also retrieved from these patients. None of the patients received preoperative treatment. A total of 15 patients were in stage I, 29 in stage II, 19 in stage III, and 5 in stage IV (any T, any N, and M1). The tumor samples, approximately 0.5 × 0.5 × 0.5 cm in size, were collected immediately after resection, snap-frozen, and stored at −70°C until RNA extraction. Normal colon samples retrieved from the proximal or distal resection margin of CC tumors were collected from 30 of the patients (median 72 (57–85) years; 13 women and 17 men).
Lymph nodes were dissected from surgically removed specimens and bisected. One half of each node was fixed for routine H&E staining and the other half was snap-frozen in liquid nitrogen and stored at −70°C until RNA extraction. Lymph nodes were retrieved from a total of 120 CC patients (median 70 (41–88) years; 66 women and 54 men): 76 of the lymph nodes were from 23 patients in stage I, 191 lymph nodes were from 52 patients in stage II, 90 lymph nodes were from 36 patients in stage III and 32 lymph nodes were from 9 patients in stage IV. Control lymph nodes (n = 67) were from 12 patients (median 23 (9–32) years; 2 women and 10 men) with ulcerative colitis (n = 11) and Crohn’s disease (n = 1).
Patients and tissue specimen for immunohistochemistry
Lymph nodes were collected from 10 CC patients (median 80 (71–91) years). Six H&E(−) lymph nodes were obtained from five patients; one in stage I, two in stage II, and two in stage III, and six H&E(+) lymph nodes were obtained from five patients in stage III.
RNA extraction
Total RNA was extracted using the acid guanidinium thiocyanate-phenol-chloroform method as described. 7 Briefly, tissues were exposed to the components mentioned above followed by centrifugation at which step RNA is separated from DNA and protein. The RNA-containing solution was subsequently collected and RNA precipitated by isopropanol.
Real-time quantitative reverse transcription polymerase chain reaction
A real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR) assay for CXCL17 mRNA with RNA copy standard was constructed using specific primers placed in different exons and a reporter dye-labeled probe hybridizing over the exon boundary in the amplicon. The qRT-PCR was performed using the 3′-primer as template for reverse transcription and Tth DNA polymerase (LightCycler 480 RNA master hydrolysis probes, Cat. No. 04991885001; Roche, Mannheim, Germany). Primer and probe sequences were 5′-primer 5′-AAGCAGTGCCCCTGTGATC-3′, 3′-primer 5′-GGAATGCTTGTTTGGCTTTCT-3′ and probe 5′-AATGTGAAGAAAACAAGACACCAAAGGCAC-3′. The RT-PCR profile was 63°C for 15 min and 95°C for 1 min, followed by 45 cycles of 95°C for 15 sec and 60°C for 1 min. The RNA copy standard was prepared as described. 15 Serial dilutions of the RNA copy standard at concentrations from 103 to 108 copies per microliter were included in each qRT-PCR run. Concentrations in unknown samples were determined from the standard curve and expressed as copies of mRNA per microliter. Concentrations of CEA mRNA copies were determined using the real-time qRT-PCR assay with RNA copy standard described in Öberg et al. 16
The concentration of 18S ribosomal RNA (rRNA) was determined in each sample using real-time qRT-PCR (Applied Biosystems, Foster City, CA, USA) and expressed as arbitrary units from a standard curve of serial dilutions of a preparation of total RNA from human peripheral blood mononuclear cells. One unit was defined as the amount of 18 S rRNA in 10 pg RNA. mRNA concentrations were normalized to the 18 S rRNA concentration in the same sample and expressed as mRNA copies per unit of 18 S rRNA. 17 Emission from the released reporter dye in the PCR reaction was monitored by the ABI Prism 7900 HT Sequence Detection System (Applied Biosystems).
Immunohistochemistry reagents
Anti-CXCL17 monoclonal antibody (mAb; IgG2b; clone 422208, Cat. No. MAB4207, R&D Systems, Minneapolis, MN, USA) and anti-CEA mAb (IgG1; clone II-7, Dako, Glostrup, Denmark) were used. Mouse IgG, ready to use (Dako), served as negative control. Anti-mouse Ig ImmPress enhancement kit was used as secondary reagent (Vector Laboratories, Burlingame, CA, USA). The substrate used was 3,3′-diaminobenzidine (DAB; Vector Laboratories).
Immunohistochemistry procedure
Fresh tissue samples were rinsed with cold phosphate-buffered saline (PBS), snap-frozen in iso-pentane pre-cooled in liquid nitrogen, and stored at −70°C. Frozen tissue was cut into 4- to 6-µm-thick sections with a cryo-microtome (MICROM HM505E; Thermo Fisher Scientific, Waltham, MA, USA). Briefly, as previously described, 18 the sections were fixed with 4% paraformaldehyde for 15 min before air-drying, rehydration in PBS, and immersion in PBS containing 0.03% H2O2 and 2 mM NaN3 at 37°C to quench endogenous peroxidase activity. Thereafter, the sections were incubated with 0.2% bovine serum albumin in PBS followed by horse blocking serum (Vector). Subsequently, sections were incubated with the primary mAb followed by ImmPress anti-mouse Ig. Bound peroxidase was revealed by incubation with 0.05% DAB and 0.03% H2O2 in 0.05 M Tris buffer (pH 7.6) at room temperature. Finally, the sections were counterstained with methyl green. Consecutive sections were stained with anti-CXCL17 and anti-CEA mAb, respectively, and examined using light microscope.
Statistical analysis
The GraphPad Prism 6 (GraphPad Software, San Diego, CA, USA) was used for statistical analyses of mRNA expression levels. Statistical analysis of differences in mRNA expression levels between two groups was performed using two-sided Mann–Whitney U-test. Kruskal-Wallis one-way analysis of variance (ANOVA) test with the Dunn’s multiple comparison post-test was used for analysis of three groups. Correlation analysis between mRNA expression levels was performed using two-tailed Spearman rank correlation test. The SPSS (IBM Corporation, Armonk, NY, USA) was used for statistical analyses of differences between patient groups in disease-free survival time and risk for recurrent disease after surgery according to Kaplan–Meier survival model in combination with the log-rank test and univariate Cox regression analysis. Patients who died from causes other than CC were considered as disease-free. Descriptive values of mRNA expression levels are given as median, interquartile range (IQR) from the 25th to the 75th percentile, and range. Descriptive values of risk of recurrence and disease-free survival time are given as mean and 95% confidence interval (CI). A p value ≤0.05 was considered to be statistically significant.
Results
Colon tumors express CXCL17 mRNA
For accurate determination of CXCL17 mRNA levels, a specific qRT-PCR assay with RNA copy standard was constructed. First, we analyzed 68 primary CC tumors and 30 apparently normal colon tissue samples. All tumor samples expressed high levels of CXCL17 mRNA. Two-thirds of the normal colon samples lacked detectable amounts of CXCL17 mRNA. The median expression level of CXCL17 mRNA in tumor tissue was 0.016 mRNA copies/18 S rRNA unit, which was 16000 times higher than in normal colon (median: 0.0000001 mRNA copies/18 S rRNA unit; p < 0.0001; Figure 1(a)). This difference is in accordance with our previous study. 8

(a) CXCL17 mRNA expression levels in colonic tissues. Normal = normal colon tissue retrieved from the proximal or distal resection margins of CC tumors. Tumor = primary tumor tissue of colon cancer, stages I through IV. (b) CXCL17 mRNA expression levels in lymph nodes of CC patients divided to three groups with respect to CEA mRNA level. CEA (−) = low CEA mRNA levels (<0.013 mRNA copies/18S rRNA unit), CEA(int) = intermediate CEA mRNA levels (0.013–3.67 mRNA copies/18S rRNA unit), and CEA (+) = high CEA mRNA levels (>3.67 mRNA copies/18S rRNA unit). Whiskers indicate the ranges, boxes indicate the 25th–75th percentiles, and horizontal bars inside boxes indicate medians (n = number of samples; p values of statistically significant differences are given).
CEA mRNA levels have previously been determined in these samples. 16 No correlation between CXCL17 mRNA and CEA mRNA levels were seen in primary tumors (r = –0.09; p = 0.5).
Metastatic lymph nodes express CXCL17 mRNA
Two criteria for identification of tumor cells in lymph nodes were used: expression levels of CEA mRNA previously shown to correlate to number of tumor cells 16 and tumor cells identified by the H&E method, performed in clinical routine by independent pathologists at Norrland University Hospital and Helsingborg Hospital. To relate CXCL17 mRNA expression levels to CEA mRNA levels, the lymph nodes were divided into three groups with respect to CEA mRNA content: CEA(−), mRNA values at or below the background level (<0.013 copies/unit); CEA(int), intermediate CEA mRNA values (0.013–3.67 copies/unit); and CEA(+) with mRNA values above the clinical cutoff level (>3.67 copies/unit).5,7 Lymph nodes (n = 389) from CC patients (n = 120; stages I–IV) were analyzed for CXCL17 mRNA expression levels (Figure 1(b)). The median CXCL17 mRNA value of the CEA(−) group was ≤10−7 copies/unit, 0.00008 copies/unit for the CEA(int) group, and 0.004 copies/18S rRNA unit for the CEA(+) group. The differences between the CEA(−) group and the CEA(int) and CEA(+) groups were highly significant (p = 0.03 and p < 0.0001 for CEA(int) and CEA(+), respectively). A highly significant difference in CXCL17 mRNA expression levels was also seen between H&E(−) lymph nodes (n = 366) and H&E(+) lymph nodes (n = 23). Median expression level was 0.000009 (IQR = 0.0000001–0.0003) copies/unit for H&E(−) lymph nodes and 0.002 (IQR = 0.0007–0.02) copies/unit for H&E(+) lymph nodes (p < 0.0001). These results clearly indicate that high CXCL17 mRNA expression levels are related to the presence of CC tumor cells in the lymph nodes.
CXCL17 and CEA mRNA levels correlate with lymph nodes of stage III and IV CC patients
We compared CXCL17 mRNA and CEA mRNA levels in lymph nodes of CC patients belonging to tumor–node–metastasis (TNM) stages I–IV. There was a highly significant correlation between CXCL17 and CEA mRNA levels in lymph nodes of stage III and IV patients (Figure 2(a)), while no correlation is seen in lymph nodes of stage I and II patients.
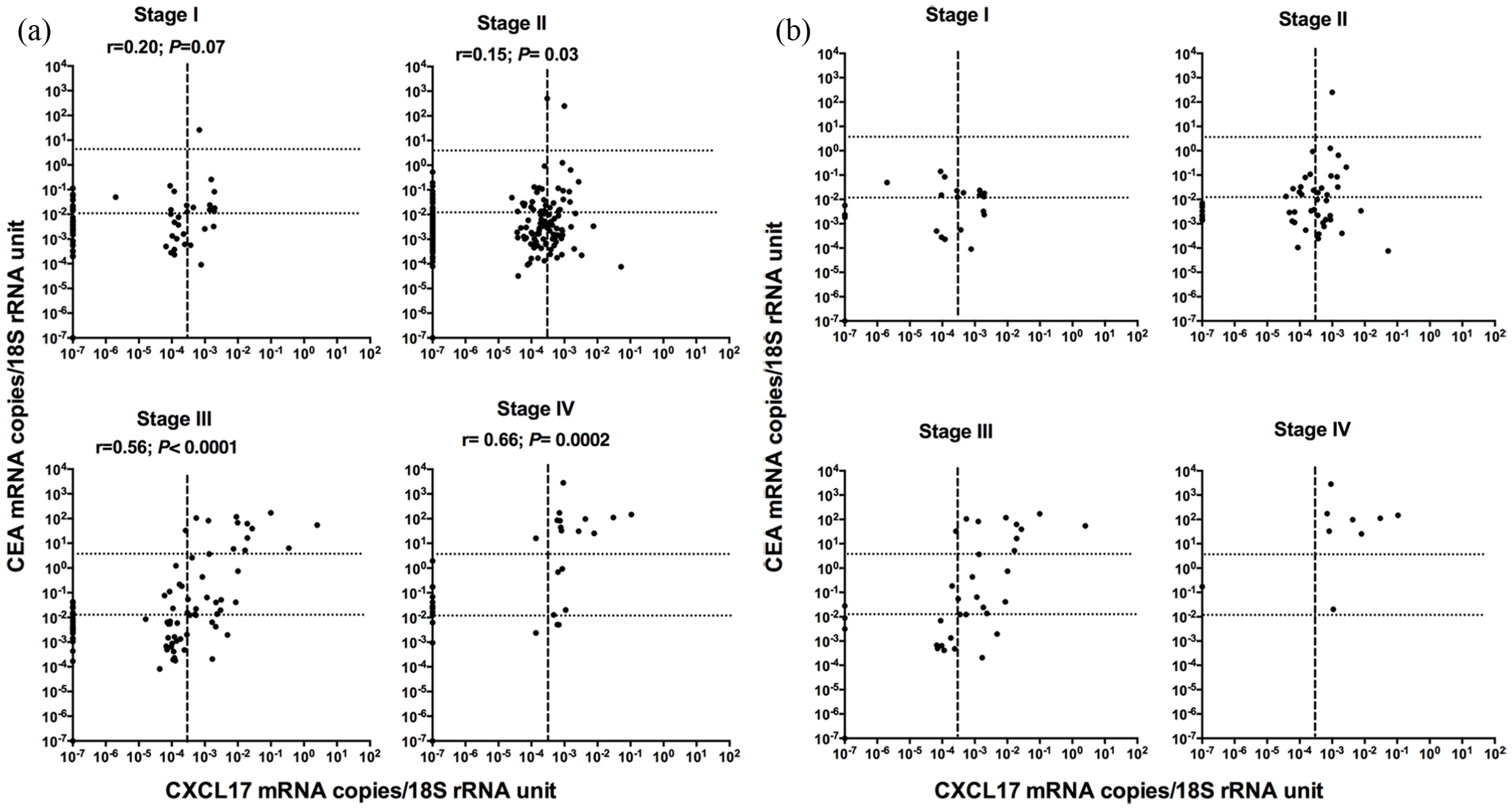
Correlation between CXCL17 mRNA and CEA mRNA levels in lymph nodes of CC patients in stages I through IV. (a) All lymph nodes (n = 389). (b) Lymph node with highest CXCL17 mRNA level for each patient (n = 120). The dashed lines indicate CEA mRNA level 0.013 mRNA copies/18S rRNA unit, that is, the border between CEA(−) and CEA(int) and CEA mRNA level 3.67 mRNA copies/18S rRNA unit, that is, the border between CEA(int) and CEA(+). The hatched line indicates CXCL17 mRNA level 0.0003 mRNA copies/18S rRNA unit, that is, the border between CXCL17(−) and CXCL17(+) (r = correlation coefficient; p values of statistically significant differences are given).
Expression of CXCL17 and CEA proteins in lymph nodes
The immunohistochemistry consecutive section staining method with specific mAbs was used to investigate CXCL17 and CEA protein expression in lymph nodes of CC patients. No positive staining with either mAb was observed in 6 H&E(−) lymph nodes from 5 CC patients (Figure 3(e) and (f)), except for occasional monocyte-/macrophage-like cells stained by anti-CXCL17 mAb only. In six out of six H&E(+) lymph nodes, the anti-CXCL17 and anti-CEA mAbs stained the same cells suggesting that metastatic cancer cells express CXCL17 also at the protein level (Figure 3(a)–(d)). In addition, a few CXCL17-stained macrophage-like cells were seen in H&E(+) nodes. Thus, macrophages could also contribute to the CXCL17 mRNA levels (see above). In two H&E(+) lymph nodes, CXCL17 staining was also observed in some CEA-negative cells with tumor cell morphology, suggesting that CXCL17 may also be expressed in tumor cells with low CEA expression.

Immunoperoxidase staining of tissue sections of lymph nodes with anti-CXCL17 mAb and anti-CEA mAb. (a and b) Consecutive sections of an H&E(+) lymph node stained with (a) anti-CXCL17 mAb and (b) anti-CEA mAb. Original magnification: 32×. (c and d) The same tissue sections at an original magnification of 200×. (c) The area enclosed in the red box in (a; anti-CXCL17 staining) and (d) the area in the red box in (b; anti-CEA staining). (e and f) Consecutive sections of an H&E(−) lymph node stained with (e) anti-CXCL17 mAb and (f) anti-CEA mAb at 32× magnification.
Expression of CXCL17 mRNA in lymph nodes is associated with poor prognosis of CC patients
To investigate the clinical utility of analyzing CXCL17 mRNA expression in lymph nodes for predicting tumor recurrence, hazard risk ratio was calculated using Cox regression analysis. Each patient was represented by the lymph node with the highest level (Figure 2(b)). The patients were divided into four groups according to the 25th, 50th, and 75th percentile of the CXCL17 mRNA values, corresponding to cutoff values of 0.0001, 0.0003, and 0.0014 copies/unit, respectively. The patients were followed for 12 years. The two groups with the highest cutoff values both differed significantly from the group with the lowest expression values but not from each other. Therefore, they were combined and termed CXCL17(+). Similarly, the two groups with the lowest mRNA values did not differ with statistical significance from each other. They were therefore combined and termed CXCL17(−). This procedure yielded a cutoff level of 0.0003 copies/unit.
Patients in the CXCL17(+) group showed a 2.2-fold increased recurrence rate compared to the CXCL17(−) group (p = 0.03; Table 1) indicating that high levels of CXCL17 mRNA is associated with poor prognosis. At 12 years follow-up, the difference in mean survival time between the two groups was 33 months (p = 0.02) according to Kaplan–Meier analysis (Figure 4(a) and Table 1).

(a) Kaplan–Meier cumulative survival curves for CC patients divided into two groups CXCL17(−) and CXCL17(+) according to the CXCL17 mRNA level at the 50th percentile. (b) Kaplan–Meier cumulative survival curves of CXCL17(−) and CXCL17(+) patients for the CEA(int) subgroup of CC patients. The patients were followed for 12 years. Differences in postoperative survival time between the two groups are given as a Δ-value in months and statistical significance as p value (n = number of patients in the respective group).
Comparative analysis of average survival time after surgery and risk for recurrence of disease of CC patients with CXCL17(+) and CXCL17(−) lymph nodes.

mRNA: messenger RNA; CI: confidence interval; rRNA: ribosomal RNA; CC: colon cancer; CEA: carcinoembryonic antigen.
Mean survival time after surgery for patients followed for 12 years as calculated by cumulative survival analysis according to Kaplan–Meier.
Hazard ratio with 95% confidenece interval (CI) for patients followed for 12 years as calculated according to univariate COX regression analysis.
CEA(int): CEA mRNA levels in the highest lymph node between the highest level of control lymph nodes (0.013 mRNA copies/18S rRNA unit) and the clinical cutoff level (3.67 mRNA copies/18S rRNA unit).
Interestingly, patients in the CXCL17(+)CEA(int) group had a hazards ratio of 8.3 (p = 0.04) for recurrence (Table 1). Furthermore, the difference in mean survival time was 46 months (p = 0.02; Figure 4(b) and Table 1). Thus, patients in the CXCL17(−)CEA(int) group survived on average 143 months compared to 97 months for the CXCL17(+)CEA(int) group.
CXCL17(+) patients were of all four TNM stages, but the fraction was highest in stages III and IV (Figure 2(a)). Table 2 shows the number of CXCL17(+)CEA(−), CXCL17(+)CEA(int), and CXCL17(+)CEA(+) patients who have died from CC. It can be seen that CXCL17(+) lymph nodes can also predict cancer death in the CEA(int) and CEA(−) group including stage II patients who had no detected H&E(+) lymph nodes. Supplementary Table S1 summarizes the results for all patients whose lymph nodes were CXCL17(−). It can be seen that five patients, two in stage II and three in stage III, had died from CC during the observation period. However, none of these lymph nodes were H&E(+), CEA(+), or CEA(int), suggesting that the lymph nodes on which staging was based were not available for analysis in this study.
Fraction of CC patients with lymph nodes belonging to any of the three categories CXCL17(+)CEA(+), CXCL17(+)CEA(int), and CXCL17(+)CEA(−) that have died in cancer during the 12-year observation period.

CC: colon cancer; CEA: carcinoembryonic antigen; H&E: hematoxylin and eosin.
H&E status of the lymph node referring the patient to the indicated CXCL17/CEA mRNA category. Data are expressed as H&E(+) lymph nodes/total number of lymph nodes in the respective category.
Note that in several cases, the investigators did not have access to the lymph node(s) that referred patients to stage III by H&E examination.
Expression levels of CXCL17 mRNA in lymph nodes of patients with inflammatory bowel disease are low
In all, 67 lymph nodes from 11 ulcerative colitis patients and 1 Crohn’s disease patient were also analyzed for CXCL17 mRNA expression. All lymph nodes except one showed values below the 50th percentile, that is, the CXCL17(+) cutoff. The value of the positive node was 0.004 copies/unit.
Risk for recurrence and survival time after surgery are not correlated to CXCL17 mRNA levels in primary CC tumors
No difference in recurrence risk or survival time was seen in CC patients divided into CXCL17(+) and CXCL17(−) patients using the median mRNA of primary CC tumors as cutoff (median: 0.016 copies/unit). Moreover, no correlation was found between mRNA levels in the primary tumor and highest lymph node of the patient (r = 0.03; p = 0.8).
Discussion
This study shows that expression of high levels of CXCL17 mRNA in regional lymph nodes is a sign of poor prognosis. Thus, recurrence rate was increased 2.2 fold and mean survival time was decreased by 33 months compared to patients with low levels of CXCL17 mRNA when evaluated 12 years after surgery. Of particular interest is the finding that CXCL17 appears to be most useful as a prognostic marker in the group of CC patients whose lymph nodes express relatively low levels of CEA, that is, the CEA(int) group.5,7 In this group, the risk of recurrence was 8.3 fold higher in the CXCL17(+) group compared to the CXCL17(−) group, and mean survival time was decreased by 46 months after 12 years. Thus, CXCL17 mRNA complements CEA mRNA as a prognostic marker for surgically treated CC patients.
The reason why CXCL17 mRNA complements CEA mRNA as a prognostic marker for CC is probably because CXCL17 is expressed both in mature and immature CC tumor cells while CEA, despite its name, is a normal adult colon tissue component highly expressed in the mature epithelial cells at the free luminal surface, but relatively weakly expressed in the immature epithelial cells in the crypts.6,19 That CXCL17 can be expressed in immature tumor cells is also suggested by the analysis at the protein level where some cells determined to be tumor cells based on morphological criteria expressed CXCL17 but not CEA. This notion is further supported by our previous study in which different CC cell lines were studied. Thus, CXCL17 mRNA was expressed at higher levels in the undifferentiated CC cell line CaCO2 than the more differentiated T84 CC cell line. 8 Thus, it is possible that some of the CEA(int) lymph nodes are populated by CXCL17-producing tumor cells with features of immature epithelial cells, including low CEA mRNA levels per cell. It is interesting to note that CXCL17 expression was correlated to poor prognosis also in hepatic cancer. 20
Here, we confirm that CXCL17 mRNA is ectopically expressed in primary CC tumors. It was not expressed at all in 21 of the 30 normal colon tissue samples collected from the resection margins of operated CC patients, and in the residual 9 normal colon samples, only low levels of CXCL17 mRNA were detected (cf. Ohlsson et al. 8 ). The cellular source of the weak signal found in one-third of the normal colon tissue samples is presently unknown, but two possibilities present themselves: either the signal is due to a few tumor cells dislodged from the main tumor mass or due to macrophage-like cells occurring in inflammatory tissues.
What are the functional consequences of CXCL17 expression in CC tumor cells? Apart from CC tumor cells, anti-CXCL17 mAb stained some cells morphologically identified as macrophage-like in the lymph nodes of both stage I and II patients and stage III patients. CCL2 expression was enhanced in CC tissues. 9 CCL2 attracts MDSCs, which in turn appear to inhibit T-cell effector functions creating a tumor-permissive environment. 9 Since CCL2 mRNA and CXCL17 mRNA expression correlate, 8 it will be of great interest to investigate whether MDSCs also are affected by tumor cell–derived CXCL17 in lymph nodes of CC patients.
In conclusion, expression of CXCL17 by tumor cells in regional lymph nodes of CC patients is a marker for poor prognosis. CXCL17 mRNA seems to be particularly useful to detect less differentiated CC tumors expressing relatively low CEA mRNA levels. In combination with CEA mRNA, CXCL17 mRNA could be used clinically as a complementary biomarker to H&E for identification of patients with H&E(−) lymph nodes at risk of recurrence due to the presence of aggressive, immature tumor cells in the nodes.
Supplemental Material
Supplementary_Table_S1 – Supplemental material for Lymph node CXCL17 messenger RNA: A new prognostic biomarker for colon cancer
Supplemental material, Supplementary_Table_S1 for Lymph node CXCL17 messenger RNA: A new prognostic biomarker for colon cancer by Yomna Rashad, Lina Olsson, Anne Israelsson, Åkeke Öberg, Gudrun Lindmark, Marie-Louise Hammarström, Sten Hammarström and Basel Sitohy in Tumor Biology
Footnotes
Acknowledgements
Basel Sitohy, Yomna Rashad, Marie-Louise Hammarström, and Sten Hammarström conceived and designed the experiments. Gudrun Lindmark and Åke Öberg collected the surgical specimens. Anne Israelsson constructed the CXCL17 and CEA mRNA qRT-PCR assays. Yomna Rashad, Anne Israelsson, and Lina Olsson performed qRT-PCR determination of CXCL17 and CEA mRNA levels. Basel Sitohy performed immunohistochemistry and consecutive staining method. Basel Sitohy, Sten Hammarström, Lina Olsson, and Marie-Louise Hammarström analyzed the data and wrote the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by grants from the Swedish Research Council-Medicine and Health (Basel Sitohy) and the Swedish Research Council-Natural and Engineering Sciences (Marie-Louise Hammarström), the Medical Faculty of Umeå University (Basel Sitohy and Marie-Louise Hammarström), the County Council of Västerbotten (Basel Sitohy), and the Lions Cancer Research Fund (Basel Sitohy) and the Stig & Ragna Gorthon Foundation, Helsingborg (Gudrun Lindmark).
Informed consent
Samples were collected after patients’ written, informed consent. The study was approved by the Local Ethics Research Committee of the Medical Faculty, Umeå University, Umeå, Sweden (registration no. 03-503).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
