Abstract
Outcomes of children with high grade neuroblastoma remain poor despite multi-agent chemotherapy regimens. Rhodiola crenulata extracts display anti-neoplastic properties against several cancers including breast cancer, melanoma, and glioblastoma. In this study, we evaluated the anti-neoplastic potential of Rhodiola crenulata extracts on human neuroblastoma cells. Through this work, cell viability and proliferation were evaluated following treatments with ethanol (vehicle control) or Rhodiola crenulata extract in neuroblastoma, NB-1691 or SK-N-AS cells, in vitro. HIF-1 transcriptional activity was evaluated using a dual luciferase assay. Quantitative real-time polymerase chain reaction was utilized to assess the expression of HIF-1 targets. Selected metabolic intermediates were evaluated for their ability to rescue cells from Rhodiola crenulata extract-induced death. Lactate dehydrogenase, pyruvate kinase, and pyruvate dehydrogenase activities and NAD+/NADH levels were assayed in vehicle and Rhodiola crenulata extract-treated cells. The effects of Rhodiola crenulata extracts on metabolism were assessed by respirometry and metabolic phenotyping/fingerprinting. Our results revealed striking cytotoxic effects upon Rhodiola crenulata extract treatment, especially prominent in NB-1691 cells. As a greater response was observed in NB-1691 cells therefore it was used for remaining experiments. Upon Rhodiola crenulata extract treatment, HIF-1 transcriptional activity was increased. This increase in activity correlated with changes in HIF-1 targets involved in cellular metabolism. Serendipitously, we observed that addition of pyruvate protected against the cytotoxic effects of Rhodiola crenulata extracts. Therefore, we focused on the metabolic effects of Rhodiola crenulata extracts on NB-1691 cells. We observed that while the activities of pyruvate kinase and pyruvate dehydrogenase activities were increased, the activity of lactate dehydrogenase activity was decreased upon Rhodiola crenulata extract treatment. We also noted a decline in the total NAD pool following Rhodiola crenulata extract treatment. This correlated with decreased cellular respiration and suppressed utilization of carbon substrates. Through this work, we observed significant cytotoxic effects of Rhodiola crenulata extract treatment upon treatment on NB-1691 cells, a human neuroblastoma cell line with MYCN amplification. Our studies suggest that these cytotoxic effects could be secondary to metabolic effect induced by treatment with Rhodiola crenulata extract.
Introduction
Neuroblastoma is the most common extra-cranial solid malignancy in infants and children. Occurring in approximately 1 in 100,000 children, neuroblastoma is responsible for 15% of all pediatric cancer deaths.1–4 While these tumors of neurocrest origin may arise anywhere along the sympathetic ganglion chain, they most commonly arise from the adrenal medulla. 1 The clinical course of this highly malignant neoplasm is quite variable depending on the location of the primary tumor, age of the patient, and stage of disease at presentation. The DNA content of cells, amplification status of the proto-oncogene MYCN, the stage of disease, and age of the patient at presentation are important variables for determining the treatment and outcomes of this aggressive disease.1–4 Treatment often involves surgical resection and multi-agent chemotherapy in which the toxicity of the chemotherapeutic agents utilized often limits treatment in this young and fragile population.1–5 Despite intensive research, outcomes of children with aggressive variants of neuroblastoma remain poor.
Rhodiola plants have been utilized medicinally for centuries in Eurasian cultures. The genus Rhodiola is composed of nearly 90 different species of plants, which grow throughout Eastern Europe and Asia. Rhodiola plants grow throughout regions of high altitude, cold climates, and barren soil. 6 Given this, these plants have evolved to produce numerous adaptogenic compounds, which allow the plant to thrive in such harsh environments. Rhodiola plants display adaptogenic and medicinal properties due to the active compounds contained within the plant extracts, such as salidroside, tyrosol, caffeic acid, rosin, and rosavin. 6 Today, Rhodiola extracts are available as natural supplements used for the treatment of fatigue, depression, anemia, impotence, gastrointestinal ailments, infections, and nervous system disorders. These extracts are believed to convey improved physical endurance and boost the immune system. Minimal side effects of Rhodiola supplements have been reported. Extracts from Rhodiola crenulata (RCEs) are bioactive and bioavailable and they are under investigation as a therapy for several types of cancers, including breast cancer, melanoma, and glioblastoma.7–10 RCEs, and the compounds contained within it, have the potential to serve as a novel adjunct for the treatment of a variety of cancers.
Of particular interest is the observation that RCE has greater effects on more aggressive cancer cell types than less aggressive and more differentiated cancer subtypes. In basal-like breast cancer cells, RCE causes an autophagic response suggestive of nutrient deprivation and the cells are less capable of recovering from over time. 8 Therefore, we tested the cytotoxic effects of RCE in two neuroblastoma cell lines, NB-1691 and SKNAS, with and without oncogene MYCN amplification, respectively. We demonstrate that NB-1691 cells with MYCN amplification were more sensitive to RCE than SKNAS with no MYCN amplification. The rapid growth arrest and death was accompanied by cytoplasmic vacuoles similar to what had been previously observed in breast cancer cells. Given the more dramatic cytotoxic effects noted in the MYCN-amplified cell line, further mechanistic evaluation for the effects of RCE treatment was carried out in NB-1691, MYCN-amplified cells.
Cancer cells reprogram their metabolism to survive, grow, and proliferate. The metabolic reprogramming involves alterations in bioenergetics, biosynthesis, and redox balance. 11 Alterations in oncogenes (e.g. MYCN and HIF-1) and tumor suppressors (e.g. TP53) are the underlying cause of metabolic changes in cancer cells. Since more aggressive cancer cell lines tend to develop a dependence on particular pathways for energy, we hypothesized that treatment with the RCE was causing a nutrient deprivation and energetic crisis. In this work, we demonstrate that exposure to RCE decreases both oxygen consumption rates (OCRs) and extracellular acidification rates (ECARs) suggestive of reduced metabolism. Further assays confirmed alteration in pyruvate dehydrogenase (PDH) and glucose utilization following RCE treatment. The effects on viability could be partially rescued by addition of pyruvate confirming a starvation response. This is the first mechanistic study demonstrating a significant impact of RCE on the metabolism of an aggressive neuroblastoma cell line.
Materials and methods
RCE preparation
Rhodiola crenulata root extract (RCE) was obtained in powdered form (Barrington Chemical Corporation, Harrison, NY, USA). It was subjected to high-performance liquid chromatography (HPLC) analysis, as described in Kwon et al., 12 to evaluate the levels of the described phenolic compounds. A total of 0.13 ± 0.08 mg/g dry weight (dw) of tyrosol, 2.15 ± 0.01 mg/g dw of salidroside, and 2.27 ± 0.31 mg/g dw of gallic acid were detected. RCE powder was dissolved in a 10% ethanol (vehicle) solution in distilled water and the solution was filter sterilized prior to use in cell culture
Cell culture
NB-1691 and SK-N-AS human neuroblastoma cells were graciously provided by Dr Andrew Davidoff’s laboratory at St. Jude’s Hospital for Children (Memphis, TN, USA). NB-1691 is an MYCN-amplified neuroblastoma line derived from a recurrent retroperitoneal tumor in a 1.9-year-old male, whereas SKNAS are non-MYCN-amplified cells obtained from the bone marrow of an 8-year-old female with recurrent neuroblastoma. 13 Cells were cultured in either Roswell Park Memorial Institute (RPMI)-1640 medium containing 2 mM of glutamine (Thermo Fisher, Grand Island, NY, USA) or Dulbecco’s Modified Eagle’s Medium (DMEM; Thermo Fisher). RPMI and DEMEM media were supplemented with 10% fetal bovine serum (FBS), 100 U/mL penicillin, and 100 μg/mL streptomycin (Gibco, Grand Island, NY, USA) or 20 µg/mL gentamicin (Thermo Fisher). Cells were cultured at 37°C under 5% CO2 in a humidified incubator.
Morphologic evaluation
The cellular morphologies following treatment with RCE compared to vehicle control (10% ethanol) were evaluated using a Nikon Eclipse TE2000-U inverted microscope using MetaVue software (Universal Imaging Corporation, Downingtown, PA, USA).
Evaluation of viability with trypan blue exclusion
Cell viability was determined by trypan blue exclusion assay using Vi-Cell XR (Beckman Coulter, Brea, CA, USA) following 72 h treatment with RCE or vehicle.
MTS proliferation assay
A CellTiter 96 Aqueous One Solution MTS (Promega, Madison, WI, USA) assay was performed according to the manufacturer’s directions in order to test for viability. A total of 1 × 104 cells were seeded on a 96-well microtiter plate and treated with 0–200 μg/mL RCE or vehicle control. After incubation for 24–96 h as indicated elsewhere, media was removed and replaced with 100 μL of 1× phosphate-buffered saline (PBS) mixed with 20 μL of MTS reagent per well. The plate was then incubated at 37°C for 1–4 h. The absorbance of each well was measured at 490 nm wavelength using an EnSpire® multimode automated plate reader (PerkinElmer, Waltham, MA, USA).
Clonogenic assay
About 300 cells were seeded onto 60-mm tissue culture dishes (three plates per treatment group). After allowing the cells to adhere for 24 h, plates were treated with 50 μg/mL RCE or vehicle control. A smaller dose of RCE was used in clonogenic assays to ensure survival during colony establishment and avoid acute cytotoxic effects associated with higher doses. Plates were also additionally treated with ±exposure to 5 Gy γ-radiation using a 137Cs irradiator (Gammator-B; Radiation Machinery Corporation, Parsippany, NJ, USA) 1 h following RCE treatment as radiation is often utilized as an adjunct to neuroblastoma therapy clinically. The media was replaced with the appropriate treatment every 3–4 days. After approximately 2 weeks, colonies were fixed with methanol and stained with 2% crystal violet in 50% methanol. A colony was defined as a cluster of five or more cells. The number of colonies was quantified visually and compared between the treatment groups (vehicle vs RCE treated). The experiment was performed in triplicate for all treatment groups.
Hypoxia inducible factor transfection and luciferase assay
NB-1691 cells were plated in a 24-well plate at a density of 8 × 104 cells/well. Cells were transfected the following day with a reporter construct containing an inducible hypoxia-inducible factor (HIF) promoter (Cignal HIF Pathway Reporter Assay Kit (LUC) #CCS-007L; Qiagen, Valencia, CA, USA) as well as a non-inducible reporter construct (negative control) and a constitutively active reporter construct (positive control). Vectors were transfected using Lipofectamine® 2000 (Invitrogen, Carlsbad, CA, USA) as per manufacturer’s instructions. After 24 h of transfection, the cells were treated with either ethanol vehicle control or 100 μg/mL of RCE for 24 h. Cells were washed with 1× PBS and lysed using 1× passive lysis buffer (Promega). Luciferase activity was measured using the Dual-Luciferase® reporter assay system (Promega) according to the manufacturer’s instructions. Light output was measured with a TD-20/20 Luminometer (Turner Designs, Sunnyvale, CA, USA), and relative luciferase activity was calculated by firefly luciferase activity/renilla luciferase activity. Luciferase activity of RCE-treated cells was compared with vehicle-treated control cells.
RNA isolation and quantitative real-time polymerase chain reaction
NB-1691 cells were treated with 100 μg/mL RCE or vehicle for 24 h. Here, a relatively lower RCE dose was used to keep cells alive for 24 h to permit analysis of changes in gene expression. RNA was extracted from cells using Trizol (Invitrogen, Carlsbad, CA, USA) per protocol utilizing acid-phenol extraction method. Following RNA extraction, up to 10 μg of RNA samples was then treated with deoxyribonuclease (DNase) (Thermo Fisher) in order to improve purity of samples. RNA levels were quantified utilizing a NanoDrop reader (Thermo Scientific, Wilmington, DE, USA). Relative levels of messenger RNA (mRNA) were determined by quantitative real-time polymerase chain reaction (qRT-PCR) using the Stratagene Mx3005P RT-PCR system (Agilent Technologies, La Jolla, CA, USA). All values were normalized to the amplification of GAPDH transcript. The PCR primer sequences evaluated are listed in Table 1. The assays were performed using the one-step 2× Brilliant SYBR Green qRT-PCR Master Mix Kit (Agilent Technologies, Santa Clara, CA, USA) containing both 200 nM of forward primer and reverse primer and 50 ng of total sample mRNA. Target mRNA amplification protocol included one cycle of 50°C for 30 min; one cycle of 95°C for 10 min; 35 cycles each 95°C for 30 s, 55°C for 1 min, and 72°C for 30 s. All experiments were performed in triplicate.
Primers utilized in qRT-PCR. List of primers utilized for qRT-PCR to evaluate the effects of RC treatment of NB-1691 cells on genetic expression.

Evaluation of pyruvate supplementation on the effects of RCE on NB-1691 viability
In order to evaluate whether media supplementation with pyruvate alters the outcomes of RCE treatment on NB-1691 cells, viability and growth were evaluated utilizing trypan blue exclusion, MTS assays, and clonogenic assays. Prior to performance of these assays, NB-1691 cells were grown in media supplemented with 1 mM of pyruvate. For MTS and viability assays, cells were treated in suspension with 200 μg/mL RCE or vehicle ± 1 mM pyruvate. Cells were then incubated for 24 h and assays were performed. Viability and MTS assays were then carried out as described above. Clonogenic assays were performed in the presence and absence of 1 mM pyruvate as described above.
Evaluation of nutrient supplementation on the effects of RCE on NB-1691 viability
All experiments were performed on 1 × 104 NB-1691 added to DMEM media lacking pyruvate, glucose, and glutamine. Cells were treated with 200 μg/mL RCE or vehicle. Media was then additionally supplemented with 25 mM glucose, 2 mM GlutaMAX, or 0.4 mM non-essential amino acids (NEAAs). Cells were treated with RCE in suspension and then plated on a 96-well microtiter plate and incubated for 24 h. MTS assay was then performed to assess for viability.
Lactate dehydrogenase activity assay
About 1 × 106 NB-1691 cells were treated with 200 μg/mL RCE or vehicle for a total of 2 h in suspension in microcentrifuge tubes (triplicates/group). After 2 h, samples were prepared per lactate dehydrogenase (LDH) Activity Assay kit (cat# MAK066; Sigma Aldrich, St. Louis, MO, USA). Samples were evaluated on an EnSpire multimode automated plate reader (PerkinElmer) to determine the initial absorbance at 450 nm wavelength [(A450)initial]. Repeat measurements were obtained every 5–10 min until the value of the most active samples exceeded that of the highest NADH standard (12.5 nmoles/well). LDH activity was determined by subtracting the blank value from each standard and unknown value. The [(A450)final] was then subtracted from the [(A450)initial] and this value was compared to the standard curve to calculate the amount of NADH generated by LDH over the evaluated time period. Results of samples treated with vehicle were compared to those treated with RCE.
Pyruvate kinase activity assay
About 5 × 105 NB-1691 cells were treated with 200 μg/mL RCE or vehicle for a total of 2 h in microcentrifuge tubes. Samples were evaluated in triplicate. After 2 h, samples were prepared per pyruvate kinase (PK) Activity Assay (cat# MAK072; Sigma Aldrich) protocol. Samples were evaluated on an EnSpire multimode automated plate reader (PerkinElmer) to determine the initial [(A570)initial] absorbance at a wavelength of 570 nm. The plate was then incubated at room temperature in the dark and repeat measurements were obtained every 5–10 min. PK activity was determined by subtracting the blank value from each standard and unknown value. The [(A570)final] was then subtracted from the [(A570)initial] and this value was compared to the standard curve to calculate the amount of pyruvate generated by PK over the evaluated time period. Results of samples treated with vehicle were compared to those treated with RCE.
Nicotinamide adenine dinucleotide pool (NAD+ + NADH) quantification assay
About 8 × 105 NB-1691 cells were treated with 200 μg/mL RCE or vehicle ± 1 mM pyruvate for 2 h in microcentrifuge tubes. Samples were evaluated in triplicate. After 2 h, samples were prepared per nicotinamide adenine dinucleotide (NAD+) quantification protocol (cat. # MAK037; Sigma Aldrich). Samples were allowed to develop at room temperature from 1 to 4 h in which measurements were obtained on an EnSpire multimode automated plate reader (PerkinElmer) to determine the absorbance at a wavelength of 450 nm (A450) to assess for a total level of NADH and NAD+ together. The blank value was subtracted from each standard and unknown value and this value was compared to the standard curve to calculate the amount of NAD pool (NAD+ plus NADH) generated. This was then was compared between samples treated with vehicle and RCE.
Evaluation of PDH activity in neuroblastoma in vitro
About 1 × 106 NB-1691 cells were treated with 200 μg/mL RCE or vehicle for a total of 2 h in microcentrifuge tubes, per protocol recommendations. All treatment samples were prepared in triplicate. After 2 h, samples were prepared per PDH Activity Assay Kit protocol (cat. #MAK183; Sigma Aldrich). Samples were allowed to develop at 37°C for 2–3 min following which absorbance at a wavelength of 450 nm (A450) was measured on an EnSpire multimode automated plate reader (PerkinElmer). Additional measurements were obtained every 5–10 min at 450 nm. A blank value was subtracted from each standard and unknown value and this value was compared to the standard curve to calculate the amount of NADH produced by PDH as a means to evaluate PDH activity. Absorbance of NADH was then was compared between samples treated with vehicle and RCE.
Respirometry
The respiratory activity of cells was measured by microplate-based respirometry as previously described14,15 using XF24-3 Analyzer (Agilent Seahorse, Billerica, MA, USA). NB-1691 cells were cultured in V7 PS plates and 48 h later cells were treated with vehicle or RCE. The respiration rates were measured using the XF24-3 Analyzer from Agilent Seahorse (Agilent, Santa Clara, CA, USA 16 ). The effect of RCE was tested at different cell densities ranging from 20,000 to 100,000 cells/well. After 48 h, cells were then treated with 200 µg/mL RCE or vehicle in the culture medium for a total of 2 h. Then, medium was replaced with low K+ buffer (LKB: 3.5 mM KCl, 10 mM KH2PO4, 1.2 mM Na2SO4, 2.0 mM MgCL2, 1.3 mM CaCL2, 15 mM glucose, and 120 mM NaCl, and 0.4% fat-free bovine serum albumin (BSA), pH 7.4). Cells were incubated in LKB for 1 h prior to analysis. Respectively, the injection ports A–D were loaded with oligomycin (2 μg/mL), FCCP (2 μM), pyruvate (10 mM), and rotenone (1 μM) with antimycin A (1 μg/mL). Calibration of sensor cartridges was performed according to manufacturer’s instructions (Agilent Seahorse). About 2–5 wells/per group were utilized.
Metabolic phenotyping of RCE treatment
Metabolic phenotyping of control and RCE-treated cells was performed using 96-well phenotyping microarrays for mammalian cells (PM-M1 plates; Biolog Inc., Hayward, CA, USA 17 ). The carbon substrates arrayed in PM-M1 plates are listed in Table 2. Plates were run in triplicate for each treatment. About 3 × 104 NB-1691 cells were seeded into each well and allowed to adhere to the bottom of each well overnight in IF-M1 media (1×, Biolog Inc.) containing 0.3 mM gentamicin, 1× penicillin/streptomycin, and 5% dialyzed FBS. The following day, cells were treated with 200 µg/mL RCE or vehicle. About 10 µL of redox dye was added to each well on all plates. Samples were then allowed to run on OmniLog PM instrument (Biolog Inc.). Absorbance at 590 and 750 nm was recorded every 15 min over 24 h for all plates. Kinetic measurements and graphs were obtained from the OmniLog PMM kinetic software.
Metabolic Phenotypic Assay: Carbohydrate and carbohydrate metabolites utilized per well of PM-M1 biology plates for cellular metabolism of NB-1691 cells treated with RC or ETOH.

Statistical analysis
All results were analyzed using either a two-tailed Student’s t test with one-way analysis of variance (ANOVA) or a two-way ANOVA. Statistical outliers were identified utilizing Grubbs’ test. All graphs were made and statistical analysis was performed using GraphPad Prism Software (Prism; GraphPad Software, Inc., San Diego, CA, USA).
Results
Cytotoxic effects of RCE on neuroblastoma cells
To determine the cytotoxic effects of RCE, we used two cell lines, the NB-1691 and SKNAS with and without MYCN amplification, respectively. A cell viability assay (Vi-cell XR) revealed that NB-1691 cells were more sensitive to the treatment with RCE compared to SKNAS cells (Figure 1(a)). These results were particularly striking because MYCN-amplified neuroblastoma typically behaves more aggressively. Therefore, progressing forward NB-1691 cells were utilized for all subsequent studies. Upon microscopic analysis of NB-1691 cells, we noted that RCE-treated group had a significant proportion of dead or abnormal appearing cells. Morphological changes of RCE treatment consisted of more rounded appearing cells, which were less adherent (Figure 1(b)). Furthermore, cells treated with RCE-developed cytoplasmic vacuoles within several hours of treatment.

Treatment of neuroblastoma cells with RCE reduces viability and alters morphology. (a) Trypan blue exclusion as assessed by a Vi-Cell demonstrates that NB-1691 cells were more sensitive to the treatment with RC extract as compared to SKNAS cells. Effects of treatment under anchorage independent conditions were even more striking. Viability graphed as % vehicle treated control. (b) Phase cell contrast microscopic images of live NB-1691 cells treated with either vehicle or 200 μg/mL RCE following 24 h of treatment.
The viability of NB-1691 was reduced by more than 50% upon RCE treatment. The cytotoxic effects were noted to be greater in suspended cells compared to adhered cells (Figure 2(a)). An MTS colorimetric proliferation assay performed utilizing graduated doses of RCE revealed 20%–25% reduction in proliferation compared to vehicle control upon RCE treatment (Figure 2(b)). In addition, to assess the cytotoxic effects of RCE-treated cells, we performed clonogenic assay as described in the “Materials and methods” section. NB-1691 cells exposed to 50 μg/mL RCE for 24 h formed less colonies compared to control cells (Figure 2(c), 24 h EtOH vs 24 h Rhodiola). To determine the relative effect of RCE compared to γ-radiation at a clinically relevant dose, cells were treated with ±5 Gy radiation. The radiation treatment resulted in a significant reduction in the number of established colonies in control cells (p < 0.0001; Figure 2(c), EtOH). Strikingly, upon continuous RCE treatment, no colonies were observed, regardless of radiation exposure (Figure 2(c), Rhodiola).
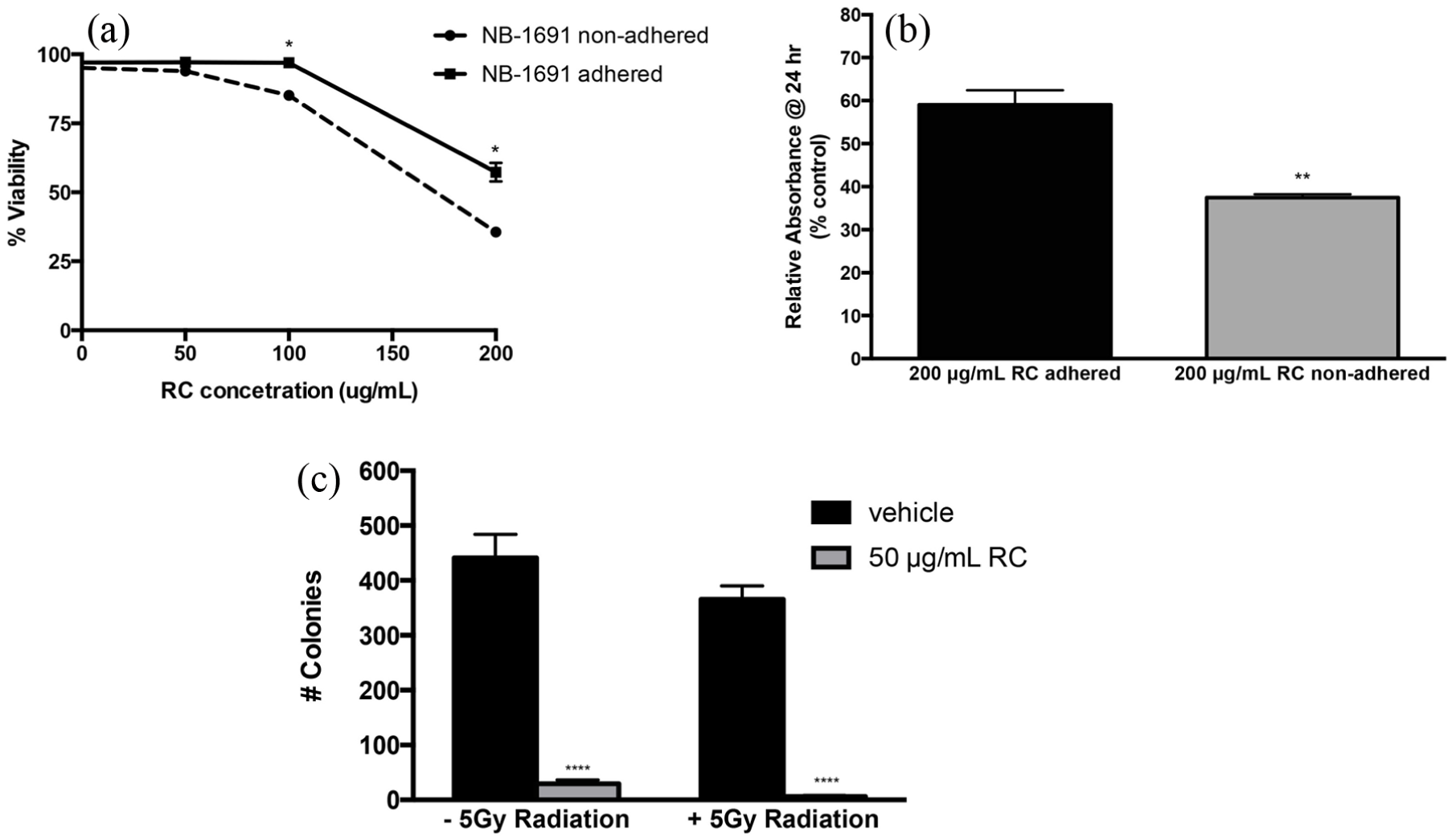
Viability and proliferation assessment of RCE treatment in NB-1691 cells. (a) 72-h viability evaluated with trypan blue exclusion on NB-1691 treated following both adherence to cell plate (black line) and prior to adherence to the place (dashed line). Results presented as viability relative to vehicle-treated cells. (b) MTS assay performed on NB-1691 cells treated with 0 and 200 μg/mL RCE either in suspension or following adherence to the culture plate. Results presented as absorbance relative to vehicle. (c) Clonogenic evaluation of NB-1691 cells treated with 50 μg/mL RCE or vehicle control for 24 h ± 5 Gy radiation. Total colonies containing five or more cells were quantified in each treatment group. Error bars indicate ±standard error of the mean, *p < 0.05; **p < 0.01; ****p < 0.001.
RCE effects on HIF and HIF-related gene expression
Rhodiola species have been used by people to help acclimate to high altitudes and low-oxygen conditions. Salidroside, a key compound in RCE, has been shown to induce HIF-1α accumulation in human fibroblasts to prevent hypoxia. 18 HIF-1α in cancer cells, however, triggers nutrient uptake and cellular metabolism. In order to evaluate potential genes involved with nutrient uptake and cellular metabolism, we monitored mRNA expression of HIF-1α and several HIF-1 targets by qRT-PCR. Selected target genes involved in glycolysis (HK2, PKM2) and nutrient transport (MCT1, GLUT1) were monitored. Our data revealed a significant upregulation of HIF-1α expression upon RCE treatment (p < 0.05, Figure 3(b)). Additional upregulation of PKM2 and MCT1 was also observed (Figure 3(b)). Given the increased expression of HIF-1α, further confirmation of alterations in HIF-1 activity was performed. NB-1691 cells transfected with a HIF-luciferase reporter element exhibited a significant upregulation in HIF promoter activity upon RCE treatment (p = 0.0094) compared to vehicle (Figure 3(a)), which corroborated qRT-PCR findings.
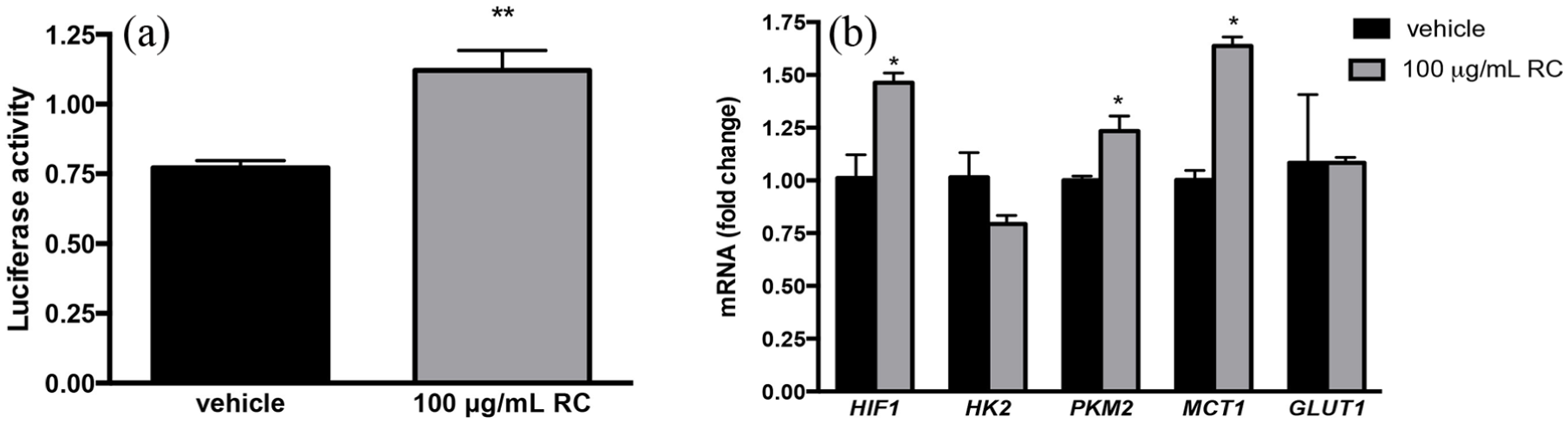
Assessment of HIF-1 and HIF-1 effectors’ transcriptional activity upon RCE treatment in NB-1691 cells. (a) NB-1691 cells transfected with luciferase reporter HIF-1 promoter and treated with 100 μg/mL RCE or vehicle for 24 h. Luciferase activity was evaluated and relative luciferase activity is reported. (b) Quantitative RT-PCR was performed from RNA extracted from NB-1691 cells treated with 100 μg/mL RCE or vehicle for 24 h. Changes in mRNA levels normalized to GAPDH expression of HIF-1α, HK2, PKM2, MCT1, and GLUT1 were evaluated. Error bars indicate ±standard error of the mean, *p < 0.05; **p < 0.01.
Pyruvate rescues cells from cytotoxic effects of RCE
We hypothesized that RCE improved cell death due to alterations in nutrient uptake and/or cell metabolism. In order to evaluate whether nutritional alterations occurred upon RCE treatment, nutritional supplements were added back to the media to determine whether the cell viability could be rescued in conjunction with RCE treatment. Upon the supplementation of media with pyruvate, the viability of NB-1691 was rescued from the cytotoxic effects of RCE. This was confirmed utilizing an MTS assay, trypan blue exclusion, and a clonogenic assay. Upon viability evaluation using trypan blue exclusion, RCE resulted in a 69% reduction in viability without pyruvate, while the addition of pyruvate with RCE resulted in only 16.6% reduction in viability, compared to vehicle (Figure 4(a), ∼52% rescue). There was still a significant reduction in viability of pyruvate with RCE compared to vehicle (p = 0.0361). Critically, an improvement in viability upon pyruvate treatment with RCE compared to treatment of RCE without pyruvate was also significant (p < 0.001, Figure 4(a)).
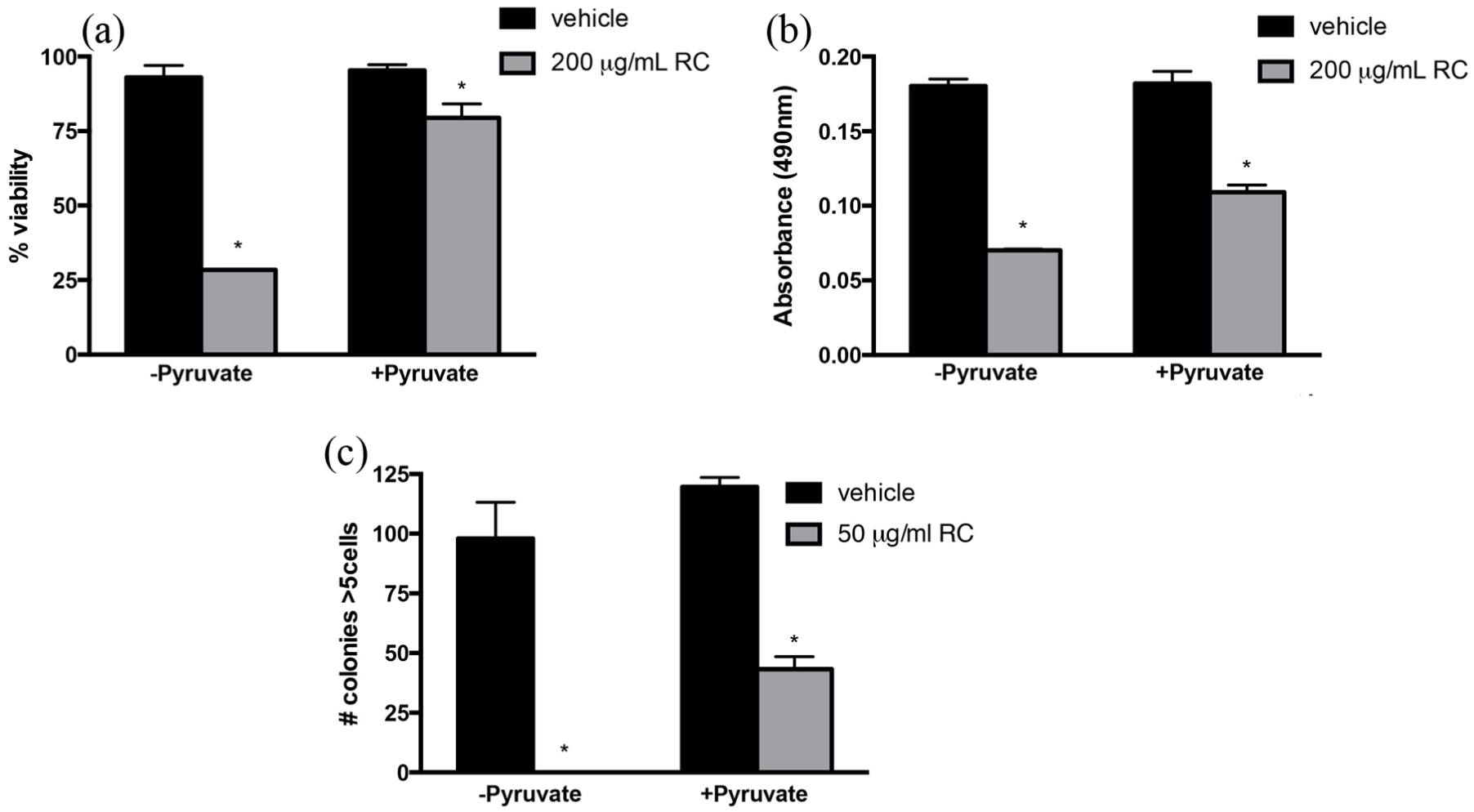
Effect of pyruvate supplementation on RCE treatment in NB-1691 cells. (a) Assessment of viability utilizing trypan blue exclusion on NB-1691 cells treated with 200 μg/mL RCE or vehicle ± pyruvate. (b) MTS assay performed on NB-1691 cells treated with 200 μg/mL RCE or vehicle ± pyruvate (1 mM) following 24 h of treatment. (c) Clonogenic evaluation of NB-1691 treated with 50 μg/mL RCE or vehicle ± 1 mM pyruvate. RCE treatment was applied continuously and plates were evaluated for the development of colony formation over 3 weeks. Following this, cells were fixed and stained with crystal violet and total colonies containing five or more cells were quantified. Error bars indicate ±standard error of the mean, *p < 0.05 comparing RCE-treated cells to vehicle-treated cells.
MTS assay corroborated these results as RCE treatment alone resulted in approximate 60% reduction in absorbance compared to control, while the addition of RCE with pyruvate resulted in a 40% reduction in absorbance. The addition of pyruvate to RCE resulted in a significant improvement in absorbance compared to RCE alone (p < 0.01, Figure 4(b)).
Finally, a clonogenic assay was performed evaluating colony growth and formation following treatment with RCE supplemented with pyruvate. Treatment with vehicle, regardless of the addition of pyruvate, resulted in the development of 70–125 colonies per plate (25%–42% plating efficiency). In contrast, treatment with RCE alone resulted in no colony formation. With pyruvate addition, a significant increase in the number of colonies was observed (an average of 43 colonies per plate, 14% plating efficiency, p < 0.01, Figure 4(c)).
Evaluation of nutrient and metabolic intermediate supplementation on the effects of RCE treatment
While we observed that cell viability was partially rescued by pyruvate, we next wanted to assay whether any other nutrients could reverse the effects of RCE. MTS assay was performed on vehicle and RCE-treated NB-1691 cells in the presence and absence of glucose, NEAA, and GlutaMAX. No significant rescue was observed against RCE toxicity upon the addition of glucose (Figure 5(a)), NEAA (Figure 5(b)), or GlutaMAX (Figure 5(c)).

Effect of alternate nutrient supplementation upon RCE treatment in NB-1691 cells. All experiments shown were performed on 1 × 104 NB-1691 cells treated with RCE or vehicle in suspension ± added nutrient in medium lacking pyruvate, glucose, and glutamine. Cells were incubated for 24 h and then an MTS assay was performed to assess for cell viability. (a) Evaluation of the effects of glucose (25 mM) on RCE treatment. (b) Evaluation of the effects of non-essential amino acids (0.4 mM) on RCE treatment. (c) Evaluation of the effects of GlutaMAX (2 mM) on RCE treatment. Error bars indicate ±standard error of the mean, *p < 0.05.
Evaluation of RCE effect on pyruvate kinase activity
Given that pyruvate was identified to alter the activity of RCE treatment, we then focused our attention on evaluating the enzymes involved in pyruvate metabolism. PK catalyzes the final step in glycolysis forming ATP and pyruvate from phosphoenolpyruvate. PK activity was significantly increased upon treatment with 200 μg/mL RCE compared to vehicle-treated controls following 2 h of treatment (p = 0.0081; Figure 6(a1)). Vehicle-treated cells exhibited an approximate 48 nM/min/mL rate of PK activity, while RCE-treated cells displayed an approximate 65 nM/min/mL rate of PK activity (Figure 6(a2)). Given the degree of upregulation of approximately 35%, we conclude that the cytotoxic effect of RCE was not at the level of PK because its activity increased following treatment and could possibly be upstream within the steps of glycolysis.

Effect of RCE treatment on pyruvate metabolism and total NAD levels on NB-1691 cells. (a) Pyruvate kinase (PK) activity levels evaluated upon NB-1691 treated for 2 h with either RCE 200 μg/mL or vehicle. Results show both (a1) PK activity (nM/min/mL) for each sample as well as (a2) PK activity over time. (b) Lactate dehydrogenase (LDH) activity levels evaluated upon NB-1691 treated for 2 h with either RCE 200 μg/mL or vehicle. Results shown include (b1) LDH activity (mU/mL) for each sample as well as (b2) LDH activity over time. (c) NAD quantification assay performed in NB-1691 cells treated with 200 μg/mL RCE or vehicle ± 1 mM pyruvate for 2 h in which total NAD+ NADH levels were quantified. (d) Pyruvate dehydrogenase (PDH) activity assay performed on NB-1691 cells treated with 200 μg/mL RCE or vehicle for 2 h in which NADH produced by PDH was evaluated. Error bars represent ±standard error of the mean, *p < 0.05; **p < 0.01.
Evaluation of RCE effect on LDH activity
Pyruvate can be converted to lactate through the enzyme LDH, which is a key regulator of cytosolic NADH/NAD+ redox. LDH activity was observed to be significantly reduced upon treatment with 200 μg/mL RCE compared to vehicle-treated NB-1691 cells following 2 h of treatment (p = 0.0196, Figure 6(b1)). On average, vehicle-treated cells exhibited 24 mU/mL rate of LDH activity, while RCE-treated cells had a 16 mU/mL rate of LDH activity (Figure 6(b2), ∼33% reduction). The reduced activity of LDH in the cytosol can affect the regeneration of NAD+ from NADH and thus the NAD+/NADH redox.
Evaluation of RCE effects on total cellular NAD levels
NAD homeostasis plays critical roles in cellular metabolism and growth. LDH and later steps in cellular metabolism provide the cells means to replenish NAD pools. However, should these pools become, cell death could ensue. We observed a significant depleted in the total NAD (NAD+ + NADH) level upon treatment with 200 μg/mL RCE in NB-1691 cells following 2 h of treatment (p < 0.001, Figure 6(c)). RCE-treated cells had about 75% less total NAD than vehicle-treated cells. The decline in NAD pool by RCE could be rescued by the addition of 1 mM pyruvate. This suggests that NAD level is a critical point of interference by RCE treatment in NB-1691 cells and 1 mM pyruvate plays a critical role in NAD homeostasis.
Evaluation of the RCE effects on PDH activity
PDH converts pyruvate to acetyl-coA and generates NADH, which supports respiration. The acetyl-CoA is condensed with oxaloacetate to form citrate, which can be further oxidized inside mitochondria and support respiration or exit mitochondria to support lipid synthesis. The PDH activity was assessed by measuring the rate of NADH production supported by pyruvate as described in the “Materials and methods” section. We observed a significant increase in NADH produced by PDH in 200 μg/mL RCE-treated NB-1691 cell extracts (p = 0.0037, Figure 6(d)). Absorbance at 450 nm was approximately doubled upon treatment with RCE compared to vehicle.
Evaluation of the RCE effects on cellular respiration
To assess the effect of above-noted enzymatic changes on oxygen consumption and extracellular acidification, respirometry was performed using a Agilent Seahorse XF24-3 Analyzer. We assessed the effects of RCE on NB-1691 cells’ respiration at different seeding densities and observed that its effect was more pronounced at lower cell density treated with a 200 μg/mL dose (20K vs 40K; Figure 7(a)). Correspondingly, RCE also reduced extracellular respiration (ECAR) in treated cells (Figure 7(b)). Because respiration was done in glucose-containing medium, this indicates reduced glucose metabolism. However, addition of exogenous 10 mM pyruvate did not increase respiration any further. Inconsistent but reduced effects of RCE at higher cell densities were observed compared to reduced cell densities (Figure 7(c) and (d)). Overall, these data suggest that at least at lower cell density RCE had inhibitory effects on NB-1691 cellular respiration and media acidification.
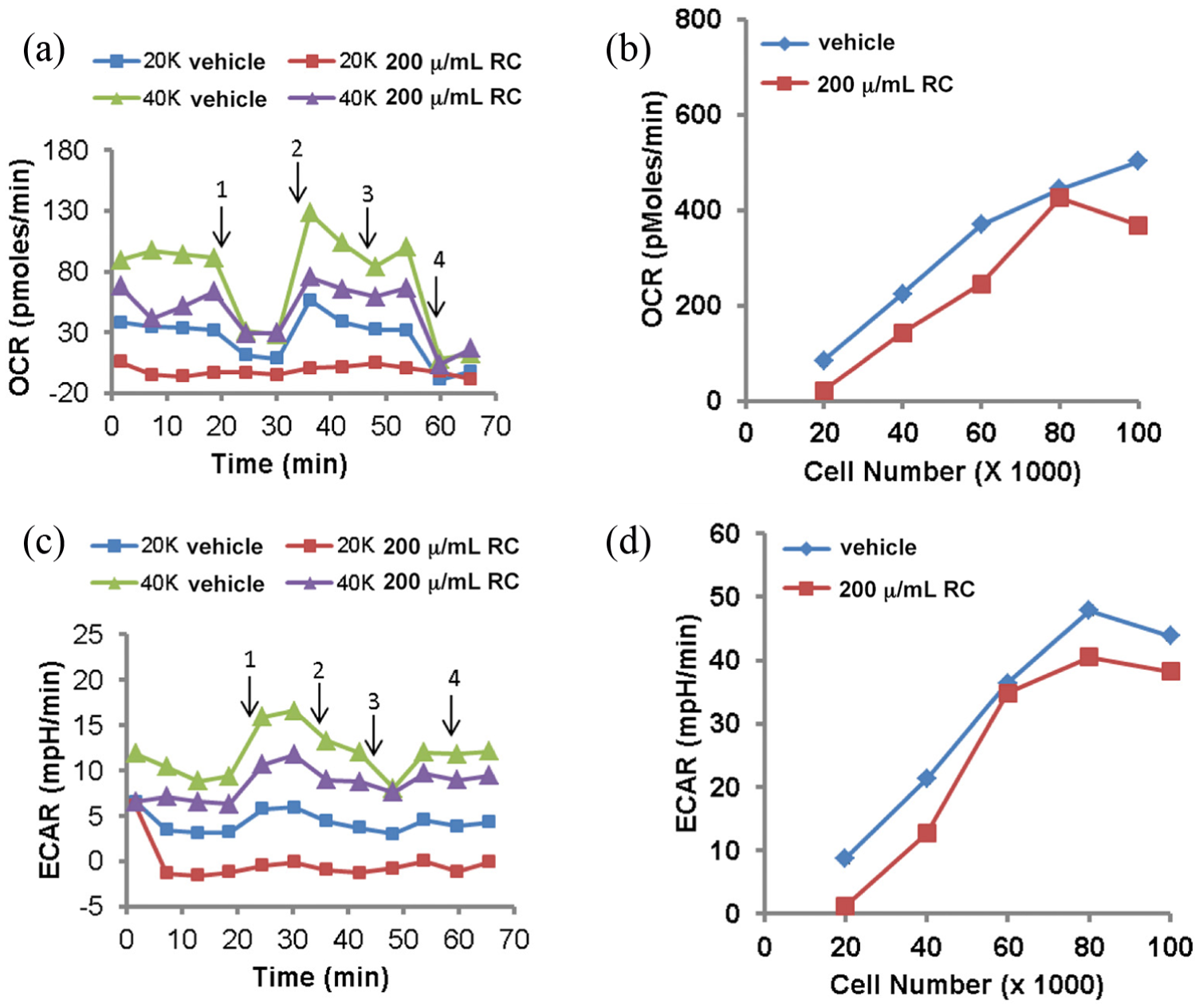
Respirometry analysis of RCE treatment on NB-1691 cells. (a) Oxygen consumption rate (OCR) of neuroblastoma cells upon treatment of RCE over time. Arrows denote serial additions of oligomycin (1), FCCP (2), pyruvate (3), and rotenone with antimycin A (4). (b) Effect of RCE on basal OCR at different cell densities. Rate 4 values plotted. (c) Extracellular acidification rate (ECAR) of neuroblastoma cells upon treatment with RCE over time. ECAR values correspond to OCR values in panel (a). Arrows 1–4 as in panel (a). (d) Effect of RCE on basal ECAR at different cell densities. Rate 4 values plotted. Average values for two wells/group are shown. (a) and (b): RCE dose fixed: 200 μg/mL, (c) and (d): RCE dose varied with cell density 200 μg/mL/20,000 cells.
Evaluation of the RCE effects on overall metabolic phenotype of NB-1691 cells
A phenotypic microarray was performed in order to not only confirm the suppressive effects RCE has upon cellular metabolism but also to investigate additional carbon sources that may play key roles in mechanism of RCE treatment. The PM-M1 microarray analysis of vehicle- and RCE-treated NB-1691 cells showed relatively reduced glucose utilization in the presence of RCE treatment (Figure 8; wells B4–6). Overall, the utilization of glucose and its polymers were reduced in the RCE-treated cells (see Table 2 for details). However, in the RCE-treated cells, the use of other carbon compounds was also increased (Figure 8 and Table 2).
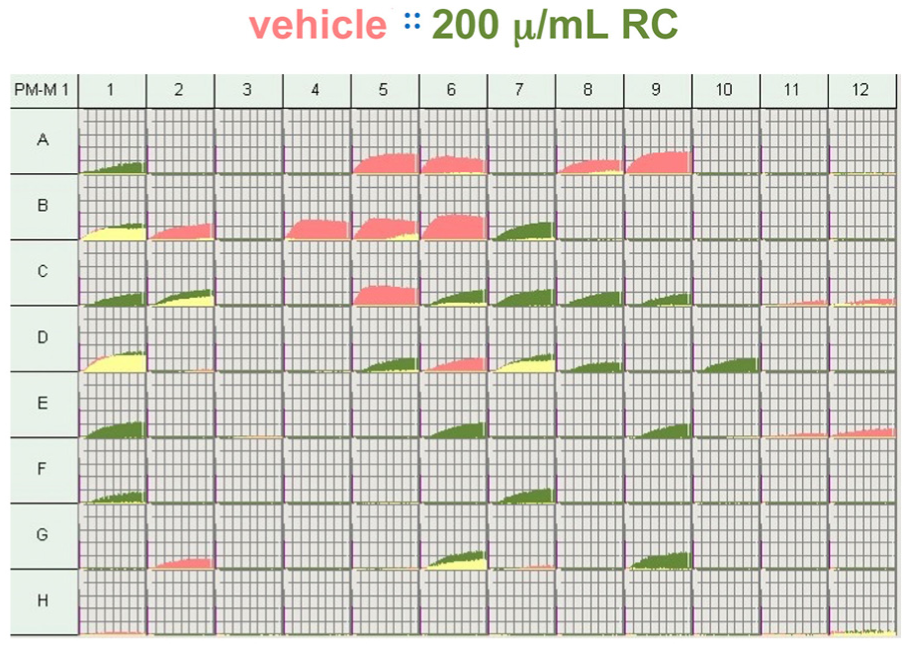
Metabolic phenotyping of RCE treatment on NB-1691 cells. Metabolic phenotyping assay utilizing Biolog PM-M1 phenotyping microarray plates comparing cells treated with RCE and vehicle. See Table 2 for details of carbohydrate and carbohydrate metabolites utilized for metabolism within each well. Kinetic graphs of cellular respiration are represented per treatment well. Red graphs denote wells in which increased respiration was observed in cells treated with vehicle. Green graphs denote wells in which increased respiration was observed in cells treated with RCE. Yellow graphs denote overlapping rates of metabolism by cells treated with either vehicle or RCE.
Discussion
Neuroblastoma remains one of the deadliest childhood cancers. Outcomes of aggressive variants of this disease result in high mortality rates and investigation into novel treatment methods may lead to improved outcomes for this patient population in the future. Through this work, we identified that RCE confers cytotoxic effects upon treatment of neuroblastoma cells in vitro. Furthermore, RCE treatment reduced growth and proliferation of these cells. These cytotoxic effects are likely secondary to metabolic derangements afforded by treatment with this extract. Given the findings observed upon RCE treatment in an aggressive neuroblastoma line, RCE offers therapeutic potential as a possible treatment for neuroblastoma.
RCE has been shown to display anti-neoplastic effects in multiple tumor lines, including breast cancer, melanoma, and glioblastoma.7–10 However, none of these studies evaluated the metabolic mechanism of the effects of RCE treatment. Through this study, we investigated the effects of RCE treatment on MYCN-amplified neuroblastoma cell line NB-1691. MYCN is a critical oncogene for determining the aggressiveness of a patient’s tumor, as amplification of MYCN is associated with more aggressive disease and poorer outcomes.1–4 The cytotoxic effects of RCE were striking upon treatment of this more aggressive neuroblastoma cell line, and the effects were further pronounced upon treatment of cells in suspension. Higher efficacy in suspension suggests that RCE may be able to reduce metastasis as disseminating tumor cells will be more susceptible to RCE. RCE treatment resulted in a rapid cytotoxicity, in which reduced cell growth and increased cell death were observed within 24 h of treatment. Within several hours of RCE treatment on NB-1691 in culture, cells began to develop cytoplasmic vacuoles, suggestive of cellular damage and/or autophagy. Reduced cell proliferation and colony formation were also observed upon RCE treatment. We hypothesized that the causes for these effects could be attributable to nutrient deficiencies or derangements to cellular metabolism, resulting in cell death.
Cancer cells reprogram their metabolism to support survival under stress in order to allow for continued growth and proliferation. This includes diversion of glucose and other carbon sources for biosynthesis apart from oxidation to produce energy.19–22 Very often cancer cells display enhanced glycolysis and suppressed pyruvate oxidation inside mitochondria. The metabolic reprogramming is associated with deregulation of tumor suppressors (e.g. p53) 23 and oncogenes (e.g. MYC and HIF-1α). 23 It is clear that complete oxidation of glucose can release all the carbons as CO2, and therefore, nothing will be available to for macromolecules’ synthesis. Thus, cancer cells have to reprogram their metabolism to balance their anabolic and catabolic needs. This is likely through providing needed intermediates for further biomass development as well as providing cells with energy regardless of oxygen availability. Apart from increased biosynthesis, hypoxia and other conditions that suppress oxidative metabolism can upregulate glycolysis for meeting bioenergetic demands. Therefore, we hypothesized that RCE treatment causes cytotoxic effects on this particularly aggressive neuroblastoma line by interfering with the metabolic pathways required to meet the energy and biomass needs.
HIF-1α plays a key role in cancer cells’ metabolic reprogramming. It promotes glycolysis and suppresses oxidative metabolism.24,25 Because HIF-1α activity is often upregulated in cancer cells to support metabolic adaptations, it serves as target of cancer therapy. In order to determine whether HIF-1α activity was influenced by RCE, we measured HIF-1α gene expression and its activity using a reporter assay. Increases in HIF-1α mRNA expression and activity were observed following RCE treatment. This may be because cells were attempting to respond to the RCE-induced change in the environment through HIF-1α upregulation. The upregulation of mRNAs of glycolytic enzymes controlled by HIF-1α transcription factor suggests that the cells were trying to adapt by altering cellular bioenergetics. However, the early appearance of autophagic vesicles, reminiscent of nutrient deprivation, suggests that the cells were not able to adapt to meet these changes in energetic needs. We speculate that other pathways which RCE treatment will inhibit, such as β-catenin activity, are needed for the efficient reprogramming driven by HIF-1α. Our lab has previously demonstrated in glioblastoma and breast cancer cell lines that RCE inhibits β-catenin activation.9,26 For this reason, we believe that HIF-1α upregulation is likely secondary to the cellular stress imposed by RCE or a partial response due to lack of other transcriptional partners.
In order to evaluate the effect of RCE on bioenergetic pathways in NB-1691 cells, a variety of metabolic nutrients and intermediates were administered in combination with RCE treatment. Interestingly, pyruvate treatment resulted in significant rescue in cell viability in the setting of RCE treatment. The addition of glucose, NEAA, or GlutaMAX, all nutrients that also can be metabolized by cancer cells, failed to result in any appreciable improvement in cell viability upon RCE treatment. Through these experiments, we observed that the effects of RCE were at least partially reversible and dependent upon the availability of important metabolic intermediates, particularly the pyruvate, to prevent cell death and promote optimal growth.
We observed that pyruvate rescued cells from the cytotoxic effects of RC, and hypothesize that this occurs through altering the availability and use of pyruvate for glycolysis, thus negatively impacting cellular respiration. Indeed respiration was reduced in RCE-treated cells. However, the effect was cell-density dependent as was the cytotoxic effects. The activity of PK, the enzyme which converts phosphoenolpyruvate to pyruvate, was not reduced. Rather it was elevated suggesting that impairment in glycolysis could be upstream to PK.
Both glycolysis and the tricarboxylic acid (TCA) cycle require regeneration of the NAD+ from NADH for their sustained operations. One of the NAD+-regenerating pathways operating in cytosol is mediated by LDH, which converts pyruvate to lactate. LDH is often upregulated in cancer cells. 23 It is thought to play key role in cancer cells metabolic reprogramming and their survival. 23 Upon RCE treatment, the activity of LDH activity was decreased. This can negatively impact NB-1691 cell fate, as LDH plays a major role in NAD+ regeneration. While we did not measure the NAD+/NADH ratio, we measured total NAD pool (both NAD+ and NADH). The reduced NAD pool clearly supports our conclusion that glycolysis is likely halted upstream of PK. Therefore, exogenous pyruvate can be expected to play a protective role as we have observed in this study. In addition to protection against RCE-induced death, pyruvate also rescued NAD pool. While the lactate/pyruvate ratio is expected to control NAD+/NADH levels, the observation of reduced overall NAD pool was a surprising and novel finding. A reduction in the total NAD pool is expected to negatively impact both glycolysis and the TCA cycle. If glucose is the only carbon source in respiration assays, it would be reflected in reduced respiration rates and reduced ECARs, which we observed in NB-1691 cells treated with RCE. However, the addition of pyruvate did not increase respiration above the level that was achievable with glucose. Also PDH activity was not suppressed but rather increased by RCE treatments. Thus, pyruvate availability or its oxidation by PDH was not limited for respiratory activity. It is possible that pyruvate in RCE-treated NB-1691 exerts a protective effect by modulating the NAD pool, thus altering the activities of both glycolysis and the TCA cycle. The depletion of NAD pool could be due to a mechanism of cell death induced by RCE, which may involve poly (ADP-ribose) polymerase (PARP-1) activation. The activation of PARP-1 is known to cause NAD depletion, glycolytic arrest, and induced cell death. 27 Cell death involving PARP-1 activation is also rescued by pyruvate supplementation. However, future studies need to be performed to determine whether PARP-1 plays a role in RCE-induced death or if NAD+ depletion can also affect the activity of sirtuins, which play important roles in cell fate determinations. 27
Pyruvate regulates NAD+/NADH redox and aspartate metabolism. Aspartate metabolism is dependent on the respiratory chain activity, impairments of which halt the TCA cycle.28,29 The role of pyruvate metabolism in aspartate biosynthesis in the presence of respiratory chain impairments has been re-emphasized by two recent studies also.30,31 Genetic manipulation of NADH levels in cells also has demonstrated a key role of pyruvate in NAD+ regeneration. 32 Therefore, our findings linking pyruvate and NAD metabolism with RCE effects on NB-1691 cells are not unexpected. However, further studies are required to dissect the exact mechanism of NAD depletion by RCE.
As RCE is a mixture of different compounds, it can affect more than one aspect of cellular physiology including gene expression and enzymes’ activities operating in different pathways. Our biochemical and respirometry-based assays clearly indicated that glycolysis was negatively impacted by RCE treatment. To further validate the negative impact of RCE on glycolysis and assess the overall alterations in carbon metabolism, we performed metabolic phenotyping with the PM-M1 array to examine preferences for different carbon substrates (carbohydrates and carboxylates). As expected, the phenotyping data confirmed downregulation of glycolysis. In general, the utilization of sugars was reduced in RCE-treated cells. However, in this assay, the pyruvate-containing well was not expected to be different from controls. The lactate use was reduced in RCE-treated cells. This agrees with the reduced LDH activity. A difference in lactate uptake can be excluded because MCT1 expression was enhanced by RCE treatment. Apart from the bioenergetics, other compounds whose utilization was increased by RCE may also play some role in cell fate determination, but those will be minor because their production is dependent on sugar metabolism, primarily glucose, unlike in the assay, where they are provided.
The treatment of neuroblastoma typically involves surgical resection and multi-agent chemotherapy. The toxicity of drugs utilized for treatment often limits the therapeutic efficacy of children with neuroblastoma given the side effect profiles. Children who receive chemotherapy at such a young age are susceptible to the lifelong effects of the morbidities associated with these drugs including hepatotoxicity, nephrotoxicity, and cardiotoxicity. 5 Furthermore, children are at increased risk for developing long-term side effects of these agents, such as the development of myelodysplastic syndromes. 5 Any novel agent that can serve as an adjunct to enhance responses or reduce the toxicities associated with current therapeutic options offers the potential for improved outcomes. While this study does not fractionate the extract or examine the in vivo uses in an adjuvant setting, we show that this extract may have the potential to be helpful in treating more aggressive neuroblastoma types through a mechanism of metabolic starvation. Future work will focus on more neuroblastoma lines and in vivo effects.
Conclusion
Through this work, Rhodiola crenulata plant extract was observed to exert dramatic cytotoxic effects on NB-1691 neuroblastoma cells in vitro. These cytotoxic effects were present despite the MYCN-amplified status and aggressive nature of this neuroblastoma cell line. The in vitro effects of RCE appear to result from alterations in metabolic functioning of this aggressive neuroblastoma cell line, resulting in reduced glycolysis and depletion in the NAD+/NADH pool, associated with the metabolism of pyruvate. Because of this, optimal cellular metabolism in these cells cannot be performed, cells are starved of important metabolic intermediates, and cell death ensues. These findings suggest that RCE plant extracts may be a promising novel adjuvant treatment for neuroblastoma. Further work is required to assess RCE treatment in alternate neuroblastoma cell lines, including additional MYCN-amplified and non-MYCN-amplified cell lines as well as evaluate the effects of treatment in an in vivo model, and the potential clinical therapeutic benefits of Rhodiola plant extracts for the treatment of neuroblastoma.
Footnotes
Acknowledgements
The authors would like to thank Baystate Medical Center Department of Surgery for their support of this work as this work was supported by an internal grant through the Department of Surgery and the support of the staff at the Pioneer Valley Life Sciences Institute, including fellow lab members who provided assistance or advice on this work. They would like to specifically acknowledge Kelly Gregory for assistance on the manuscript and Jennifer Ser-Dolansky, Benjamin Schneider, and Maxine Dudek for their assistance with the studies.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
