Abstract
Recent studies have shown that circular ribonucleic acids have differential expression in some diseases. This study compared the expression levels of five circular ribonucleic acids between patients of primary hepatic carcinoma following liver transplantation and healthy individuals for searching a new diagnostic biomarker about primary hepatic carcinoma. We chose differentially expressed targeted circular ribonucleic acids according to fold change ≥2.0 or ≤–2.0 between circular ribonucleic acids microarray of perioperative liver transplantation and normal controls. Then we used the Arraystar home-made micro-ribonucleic acid target prediction software based on TargetScan and miRanda to predict circular ribonucleic acid/micro-ribonucleic acid interactions. And we assess the expression levels of hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 in the peripheral blood of normal controls and liver transplantation patients before transplantation and on the first, third, and seventh days after transplantation by real-time quantitative polymerase chain reaction. We chose five circular ribonucleic acids, two of which have been correlated with micro-ribonucleic acid–related carcinoma recurrence after liver transplantation, hepatocellular carcinoma and analyzed their expression with 2–△△Ct method. The expression level of hsa_circ_100571 and hsa_circ_400031 on day 1 after liver transplantation was higher than pre-transplantation (p < 0.01), and these levels showed a declining trend on post-transplantation. The expression level of hsa_circ_102032 and hsa_circ_103096 on day 1 after liver transplantation was lower than pre-transplantation (p < 0.01) and decreased on post-transplantation. There were the significantly different expressions between the post-transplantation day 7 and normal control (p < 0.01). The expression level of hsa_circ_102347 on day 1 after liver transplantation was lower than pre-transplantation (p < 0.01). This expression showed a declining trend on post-transplantation, and the postoperative day 7 level was similar to normal control (p > 0.05). Five types of circular ribonucleic acid–related micro-ribonucleic acids had varying degrees of upregulation and downregulation between perioperative transplantation of primary hepatic carcinoma patients and normal controls; the hsa_circ_102347 is most likely to have association with primary hepatic carcinoma.
Keywords
Introduction
Circular ribonucleic acid (circRNA) composed of annular arraying exon or intron sequences belongs to a class of endogenous non-coding ribonucleic acids (RNAs) that do not contain protein-coding genes but rather simultaneously regulate multiple genes in vivo. 1 circRNAs have many attributes, such as structural stability, high sequence conservation, abundance, and tissue specificity. 2 Micro-ribonucleic acids (miRNAs) have been found to be associated with various major diseases including cancer, systemic lupus erythematosus, and inflammatory infections. 3 circRNAs play a crucial role in down-regulating the level of miRNA-mediated gene expression by sequestering miRNAs (miRNA sponge). 4 Chen et al. 5 presented that circRNA_100290 may be a competing endogenous RNA to regulate CDK6 expression by sponging up miR-29b and can be used as potential targets in human oral squamous cell carcinomas (OSCC) therapy. Recently, it was reported that circRNAs also regulate or degrade RNA binding proteins and affect the translation of RNA expression. 6 Zhang et al. 7 found that circRNAs, such as the ankyrin repeat domain 52, that are abundant in the nucleus and derived from the gene intron, can positively regulate RNA polymerase II. Therefore, circRNA involved in biological development, growth, and disease processes may function as a novel class of biomarkers for clinical diagnosis and therapeutic effects.
Primary hepatic carcinoma (PHC) is one of the world’s top five most common cancers with 1 million new cases annually. 8 The development and formation of PHC are extremely complicated, which give a huge puzzle to the diagnosis and treatment of PHC. It is a method that some biomarkers of PHC are obtained by screening the differential expression in peripheral blood between patients with PHC and normal people. However, some biomarkers unrelated PHC may be screened without change following liver transplantation. It has some error to choose differential biomakers only according to up-regulated or down-regulated expression between patients with PHC and normal people. Liver transplantation is the most effective treatment for PHC. If we find differential biomarkers by comparing pre-transplantation with normal controls (NCs), and comparing post-transplantation with pre-transplantation, then we would effectively screen biomakers in peripheral blood. For example, the up-regulated/down-regulated biomakers are found through comparing preoperative patients with NCs; the down-regulated/up-regulated biomakers are found through comparing postoperative patients (Post) with preoperative patients. The shared biomakers among them are more likely to be associated with PHC. At present, the methods of clinical monitoring and diagnosis of patients after liver transplantation mainly include imaging, humoral analysis, and liver biopsy, but liver biopsy may bring traumatic pain and complications to patients. Imaging features of cancerous tissue and precancerous lesions often lack specificity. The only means of obtaining circRNA by extracting bodily fluids can reduce the traumatic pain for the patient diagnosis. The diagnosis and treatment of PHC patients would be improved if some novel reliable biomarkers with strong specificity and high sensitivity were identified, such as circRNAs.
Materials and methods
Patients
Peripheral blood samples were collected from four patients with PHC. Patients were recruited from the Nephrology Department at the No. 181 Hospital, Guangxi, China. All of the patients had undergone liver transplantation surgery, and no acute rejection nor recurrence of cancer has been observed after surgery. Peripheral blood samples were also collected from four healthy individuals as NCs from the Physical Center of the Guilin No. 181 Hospital. Peripheral blood samples were collected from patients at various time points: preoperative patients (Pre), 1-day postoperative patients (P1), 3-day postoperative patients (P3), and 7-day postoperative patients (P7). Patient and healthy individuals’ clinical characteristics were extracted from database of the No. 181 hospital and were summarized in Table 1. The aspartate transaminase (AST) and alanine aminotransferase (ALT) <50 IU/L were considered liver function normal. Therefore, the liver function of patients gradually recovered to normal. All of the peripheral blood samples were obtained after receiving informed consent from the participating subject. This study was performed according to the guidelines of the No. 181 Hospital, which abided by the Helsinki Declaration on ethical principles for medical research involving human subjects.
Characteristics of the patients and the normal controls.

NC: normal control; ALT: alanine aminotransferase; AST: aspartate transaminase.
Peripheral blood collection and storage
Whole-blood samples were drawn from a peripheral vein between 7:00 and 9:00 a.m. Peripheral blood samples were collected from four patients and four healthy individuals at the Guilin No. 181 Hospital between 2014 and 2015. Peripheral blood samples used for circRNA detection were collected from patients either at the emergency department or during the physical examination. The plasma was transferred to RNase/DNase-free tubes and stored at −80°C.
circRNAs microarray
Total RNA was extracted from the blood using TRIzol (Invitrogen, Carlsbad, CA, USA). Then, total RNA was quantified using a NanoDrop ND-2000 (Thermo Fisher Scientific, Wilmington, DE, USA). RNA integrity was evaluated by denaturing gel electrophoresis standards. Total RNAs were digested with RNase R (Epicentre, Inc.) to remove linear RNAs and enrich circRNAs. Then, the enriched circRNAs were amplified and transcribed into fluorescent cRNA utilizing Arraystar Super RNA Labeling Kit. The labeled cRNAs were hybridized onto the Arraystar Human circRNA Array (8x15 K, Arraystar). After the slides were washed, the arrays were scanned by the Agilent Scanner G2505C.
circRNA chip data acquisition and analysis
The Agilent Feature Extraction software (version 11.0.1.1) was used to analyze the acquired array images. Quantile normalization and subsequent data processing were performed using the R software package. The differentially expressed circRNAs were screened according to fold change (FC) filtering.
Candidate circRNA selection
Recent evidence has demonstrated that circRNAs play a crucial role in fine tuning the level of miRNA-mediated regulation of gene expression by sequestering miRNAs. The interaction of circRNAs with disease-associated miRNAs indicates that circRNAs are important for disease regulation. 9 miR-203 of low level has been reported to have an association with metastasis of hepatocellular carcinoma after liver transplantation, 10 and miR-483-5p of high level is more specifically expressed in hepatocellular carcinoma. 11 We used the Arraystar home-made miRNA target prediction software based on TargetScan 12 and miRanda 13 to predict the circRNA/miRNA interaction. We chose targeted circRNAs according to FC ≥2.0 or ≤–2.0 in circRNAs microarray of perioperative liver transplantation and NCs, and circRNA-related above-mentioned miRNAs.
Quantitative reverse transcription-polymerase chain reaction
Real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was performed using 2X PCR master mix (Arraystar) on a ViiA 7 Real-time PCR System (Applied Biosystems) following the manufacturer’s instructions. The qRT-PCR was used to quantify the expression of candidate circRNA. The data were analyzed using the ΔCt method.14–16 RNA was reverse-transcribed to cDNA with gene-specific primers. The RT and qRT-PCR primers are listed in Table 2.
Reverse-transcribed real-time quantitative polymerase chain reaction primers.

Statistical analysis
Quantitative data are expressed as the mean ± standard deviation. The relative expression content of circRNAs was analyzed by means of the 2–△△Ct method. All statistical data were analyzed using SPSS software 19.0. The differences in the levels of hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 between samples from liver transplantation patients and healthy individuals were assessed using a t-test with the △Ct data. p-Value of <0.05 was considered significant difference; p < 0.01 was considered extremely significant difference.
Results
RNA analysis with the use of ultraviolet absorption
Quantification and quality assurance of the RNA samples from liver transplantation patients and healthy individuals were determined by measuring the OD260. The OD A260/A280 ratio should be approximately 2.0 for pure RNA (ratios between 1.8 and 2.1 are acceptable). The OD A260/A230 ratio should be greater than 1.8.
Identification of differentially expressed circRNAs in the microarray analysis
To screen differentially expressed circRNA-related PHC, we performed peripheral blood circRNA profiling to compare the Pre pool with the NC pool and the Post pool with the Pre pool. After normalization of the raw data, microarray detected 302 circRNAs had differential expression by comparing Pre with NC. There were 182 up-regulated (FC ≥ 2.0) and 120 down-regulated (FC ≤ –2.0) circRNAs. We compared Post which is the mean of P1, P3, and P7 with Pre. We found that a total of 925 were significant expression. Among them, there were 402 (FC ≥ 2.0) up-regulated and 523 down-regulated (FC ≤ –2.0) circRNAs. Then, we found that 165 circRNAs were up-regulated in the peripheral blood of the Pre patients compared with NC (FC ≥ 2.0) and down-regulated in the blood of the Post patients compared with Pre (FC ≤ –2.0) (Figure 1(a)), and 80 circRNAs were down-regulated in the peripheral blood of the Pre patients compared with NC(FC ≤–2.0) and up-regulated in the peripheral blood of the Post patients compared with Pre (FC ≥ 2.0) (Figure 1(b)).

Differentially expressed common circRNAs of Pre, Post, and NC.
circRNA/miRNA interaction and candidate circRNA
TargetScan and miRanda showed that circRNAs have targeted sites on the following miRNAs. In hsa_circ_400031, the 134th–159rd nucleotides starting from the 5′ terminus were not completely complementary to the miR-483-5p seed region. However, the 305th–331rd nucleotides starting from the 5′ terminus were completely complementary to the miR-483-5p seed region in the 7mer-m8 (Figure 2(a)). In hsa_circ_102347, the 286th–307rd nucleotides starting from the 5′ terminus were not completely complementary to the miR-203 seed region (Figure 2(b)).

The detailed annotation for circRNA–miRNA interactions: (a) complementary situations of hsa_circRNA_400031 and hsa-miR-483-5p and (b) complementary situations of hsa_circRNA_102347 and hsa-miR-203a-5p.
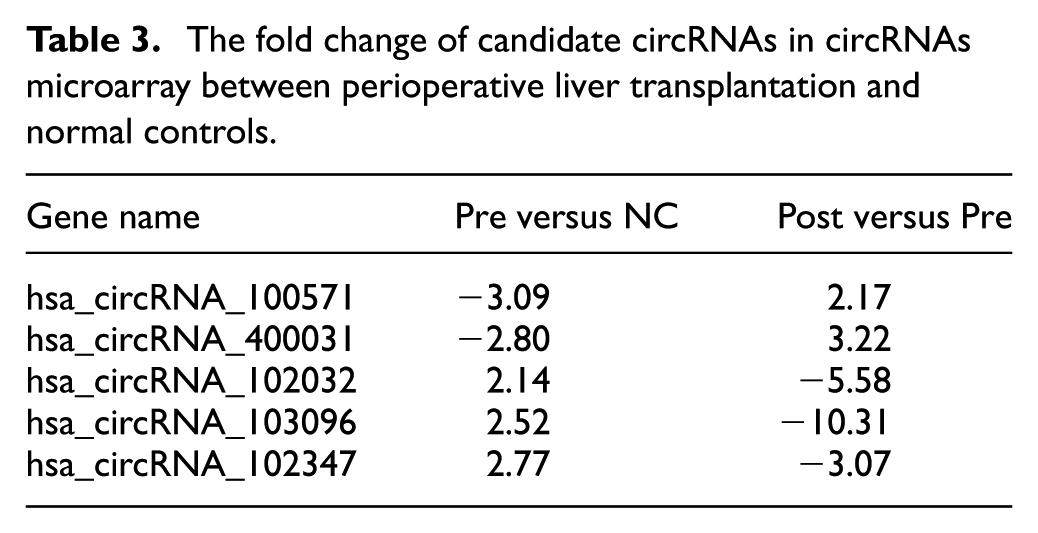
Therefore, we chose differentially expressed hsa_circ_400031 and hsa_circ_102347 from circRNAs microarray analysis according FC more than two times. Among them, hsa_circ_400031 and hsa_circ_102347 interacted with the above-mentioned two miRNAs. We also chose the hsa_circ_100571, hsa_circ_102032, hsa_circ_103096. Although they had no connection with the reported miRNAs, their FC was the most obvious in circRNAs microarray between perioperative liver transplantation and NCs (FC ≥ 2.0 or FC ≤ –2.0). The FC values of five circRNAs in circRNAs microarray between perioperative liver transplantation and NCs are listed in Table 3.
The fold change of candidate circRNAs in circRNAs microarray between perioperative liver transplantation and normal controls.
Test results of quantitative RT-PCR
The Ct value is the number of cycles at which the fluorescence signal passes the appropriate threshold. △Ct was the difference in Ct values between the controls and hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347. △Ct data are listed in Table 4. β-Actin was used as a normalized internal control. –△△Ct was the difference in △Ct values between the experimental samples and the paired control samples. 2–△△Ct represents the relative expression content of hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 (Table 5).
circRNAs △Ct values of the liver transplantation and normal groups.

p-Value and 2–△△Ct of circRNAs expression between the liver transplantation group and normal control group.

p < 0.05 represents the difference between the two groups was significant;**p < 0.01 represents the difference between the two groups was extremely significant.
Analysis of candidate circRNA expression levels
In our study, hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 displayed relative expression levels and p-values between groups of four liver transplantation patients and NCs (Figure 3 and Table 5). The results showed that the expression levels of hsa_circ_100571 and hsa_circ_400031 pre-transplantation were lower than those of NCs, and the relative expression contents (2–△△Ct) were 0.77 and 0.78, respectively (Table 5). However, there were no significantly differential p-values (0.102 and 0.087, respectively). The expression levels of hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 pre-transplantation were significantly higher than those of NCs (p = 6.92E–05, 1.30E–05, 3.82E–05, respectively), and the relative expression contents (2–△△Ct) were 1.39, 1.99 and 1.79, respectively (Table 5). The hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 may have relationship with PHC.

circRNAs expression levels of different groups corrected with the internal reference (β-actin).
The expression level of hsa_circ_100571 on day 1 after liver transplantation was significantly higher than the pre-transplantation levels (p = 0.008), and these levels showed a declining trend on postoperative days 1, 3, and 7. The abnormal postoperative expression could be associated with the surgical trauma rather than PHC (Figure 3(a)). The expression level of hsa_circ_400031 on day 1 after liver transplantation was significantly higher than that pre-transplantation (p = 0.007). Postoperative days 1, 3, and 7 showed reduced overall expression, and the postoperative day 7 level was close to NC (p = 0.809) (Figure 3(e)). Although this result indicated that postoperative days 1 compared pre-transplantation with significant difference, the p-value between the pre-transplantation and normal did not support that the hsa_circ_400031 associated with hepatocellular carcinoma.
The expression levels of hsa_circ_102032 (Figure 3(b)) and hsa_circ_103096 (Figure 3(d)) on day 1 after liver transplantation were significantly lower than those pre-transplantation (p = 2.34E–04 and p = 5.37E–04, respectively) and decreased on postoperative days 1, 3, and 7. The levels of hsa_circ_102032 and hsa_circ_103096 of transplantation patients were significantly differentially expressed on comparing postoperative day 7 with NC (p = 2.87E–07 and p = 3.96E–04, respectively). Decreasing levels of hsa_circ_102032 and hsa_circ_103096 may predict to many hsa_circ_102032 and hsa_circ_103096 of PHC be removed after transplantation. The expression level of hsa_circ_102347 on day 1 after liver transplantation was lower than that pre-transplantation (p = 5.50E–04) (Figure 3(c)). This expression showed a declining trend on postoperative days 1, 3, and 7, and the postoperative day 7 level was similar to NC (p = 0.064). High expression levels of miR-203 indicated that patients had a good recovery without carcinoma recurrence. The postoperative declining trend may indicate that hsa_circ_102347 may be associated with a transplanted liver without carcinoma recurrence.
Discussion
Since circRNA was discovered,17,18 research into the function of circRNAs has increased. Previous reports have shown that CDR1as is a cyclic inhibitor of miR-7 and also has a potential value in monitoring Parkinson’s disease. 19 Although circRNA has been the focus research recently,20–23 it is largely unknown that circRNA-related PHC is screened by monitoring perioperative period of liver transplantation. In the present study, we isolated and analyzed peripheral blood circRNAs from liver transplantation patients of PHC at time points that included pre- and post-transplantation at the first, third, and seventh days after the transplantation. We also chose NCs. We found that a total of 245 had upregulation or downregulation by comparing the Pre pool with the NC pool and the Post pool with the Pre pool. We chose hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 by screening circRNAs. We analyzed the expression levels of five circRNAs and found that had significant differences between perioperative period and NC by p-value and 2–△△Ct. The 245 circRNAs may serve as a novel tool for clinical diagnosis of PHC.
We chose hsa_circ_100571, hsa_circ_400031, hsa_circ_102032, hsa_circ_103096, and hsa_circ_102347 circRNAs that were differentially expressed in NC, Pre, and Post of microarray to analyze the expression levels by p-value and 2–△△Ct. The hsa_circ_400031 and hsa_circ_102347 were associated with miRNAs. According to our results, the hsa_circ_102347 that regulates miRNA-related carcinoma recurrence after liver transplantation is most likely to be a biomarker-associated PHC for adsorption mechanism. And the expression levels of hsa_circ_102032, hsa_circ_103096 may prove that they are associated with PHC. However, the circRNAs half-life is very short; the time of preserving sample was limited. Therefore, we only chose four samples. This may have introduced variability and room for error in analyzing our results, such as the hsa_circ_400031 and hsa_circ_100571. However, a large number of differentially expressed circRNAs can also provide a data basis for subsequent experiments. Absolutely, more samples will be analyzed in subsequent experiments. The adsorption mechanism between the circRNAs and miRNAs, and the regulation networks of circRNAs, miRNAs, proteins, and gene expression in perioperative period of liver transplantation will follow this study. Our work provides the basis of research for mechanism of circRNAs sponge and may also provide a novel method for researching biomarkers of PHC.
Footnotes
Acknowledgements
The authors thank the patients and healthy volunteers who participated in this study. Weiguo Sui and Qing Gan contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by Guangxi Natural Science Foundation (no. 2011GXNSFB018105), Guilin Scientific Research and Technology Development Program (no. 2016012702-1), and no. 2010gxjs037.
