Abstract
Primary human mammary epithelial cells have a limited life span which makes it difficult to study them in vitro for most purposes. To overcome this problem, we have developed a cell line that was immortalized using defined genetic elements, and we have characterized this immortalized non-tumorigenic human mammary epithelial cell line to establish it as a potential model system. human mammary epithelial cells were obtained from a healthy individual undergoing reduction mammoplasty at SIU School of Medicine. The cells were transduced with CDK4R24C followed by transduction with human telomerase reverse transcriptase. Post all manipulation, the cells displayed a normal cell cycle phase distribution and were near diploid in nature, which was confirmed by flow cytometry and karyotyping. In vitro studies showed that the cells were anchorage dependent and were non-invasive in nature. The cell line expressed basal epithelial markers such as cytokeratin 7, CD10, and p63 and was negative for the expression of estrogen receptor and progesterone receptor. Upon G-band karyotyping, the cell line displayed the presence of a few cytogenic abnormalities, including trisomy 20 and trisomy 7, which are also commonly present in other immortalized mammary cell lines. Furthermore, the benign nature of these cells was confirmed by multiple in vitro and in vivo experiments. Therefore, we think that this cell line could serve as a good model to understand the molecular mechanisms involved in the development and progression of breast cancer and to also assess the effect of novel therapeutics on human mammary epithelial cells.
Introduction
Breast cancer is the most common cancer reported in North American women. Primary epithelial cell lines that are used to carry out the research have a limited capacity to proliferate, 1 and this hinders the use of non-malignant cells to understand the molecular mechanism and interactions involved in the initiation, metastasis, and signaling pathways during breast cancer development.2,3 One of the ways to successfully deal with this problem is by using immortalized non-transformed cell lines.4,5 The primary objective of this article is to characterize6,7 the human mammary epithelial cell (HMEC) 2.6 line and establish it as a model for future studies in breast cancer research.
HMEC line the ducts and lobules of the mammary gland, and most of the breast cancers arise from this epithelial cell layer. HMEC have a limited life span. 4 These cells undergo senescence eventually thereby suppressing tumorigenesis and also promoting tissue repair.8,9 HMEC immortalization 10 is one of the key steps in malignant transformation and involves the alterations of a number of genes that are involved in the checkpoints of the cell cycle.1,11
HMEC were obtained from a healthy individual undergoing reduction mammoplasty at SIU School of Medicine. The primary epithelial cell lines were immortalized by transducing them with LXSN encoding CDK4R24C, a mutant form that cannot be inhibited by p16INK4A. This was followed by transduction with human telomerase reverse transcriptase (hTERT),12,13 the catalytic subunit of the telomerase enzyme.
The presence of cyclin-dependent kinase 4 (CDK4), p16INK4A, and telomerase have been verified in the HMEC 2.6 cells,14,15 and they have been passaged for over 20 times to ensure immortality. This study details the establishment of an immortalized non-tumorigenic cell line produced with defined genetic elements.
Materials and methods
Isolation and culture of HMEC
The HMEC 2.6 were derived from an individual undergoing reduction mammoplasty with no known breast pathology, as confirmed by histopathology of the post-surgical specimens. Specimens were obtained from the Simmons Cancer Institute tissue bank under the institutional review board (IRB) approved protocol number: 08-112/12-177. HMEC were prepared by the method of Smith et al. 16 The primary HMEC were derived from mammary tissues that were obtained from mammoplasty. Mammary tissue was then cut into pieces and digested in a mixture of 14 U/mL of hyaluronidase, 15 U/mL of collagenase, 10 µg/mL of insulin, 1× penicillin/streptomycin, 10% fetal calf serum (FCS) in DFCI-1 medium at 37°C for overnight. Cells were sieved through a sterile strainer, washed with phosphate-buffered saline (PBS), and spun down at 1500 r/min for 5 min and plated on 10-cm plate. Fibroblasts were removed by trypsin/ethylenediaminetetraacetic acid (EDTA) treatment. HMEC were immortalized by transducing with Zeocin-based vector carrying hTERT and neomycin-based construct that contained a mutant CDK4(R24C). The antibiotic concentrations for selection are as follows—G418: 100 µg/mL, Zeocin: 20 µg/mL, Puromycin: 2.5 µg/mL, and Hygromycin: 20 µg/mL. The cell line is sensitive to Puromycin and Hygromycin. HMEC was grown at 37°C and 5% CO2 17 in DFCI-1 media 18 supplemented with 10% fetal bovine serum (FBS), 100 units/mL of penicillin, and 100 µg/mL of streptomycin according to the standard protocol. 19 The cells have been passaged around 20 to 23 times to ensure immortality. RAO-1 is a HMEC cell line obtained from healthy individuals undergoing reduction mammoplasty and was immortalized by transduction with catalytic subunit of telomerase (hTERT) after passage through stasis. RAO-4 is derived from RAO-1 by transduction with Q61L mutant H-Ras gene and RAO-4 forms human mammary epithelial carcinoma when injected into nude mice.10,20
Flow cytometry
The cell cycle analysis was carried out to analyze the cellular DNA content, and this was done by lifting the cells from the culture plate with trypsin/EDTA (0.5%/0.1%) and fixing overnight with 80% ethanol at 4°C. The following day, cells were washed with PBS and stained with 10 µg/mL of propidium iodide in PBS for 20 min at room temperature.
Growth kinetics
To determine the growth kinetics of the HMEC 2.6 cell line and compare it with RAO-1 cell line, 20 cells from both lines were harvested as a single cell suspension and 50,000 cells were seeded in a 12-well plate containing 1 mL of the DFCI medium. Every 24 h, the cells were counted by trypsinizing the cultures in triplicate and resuspending the cells in the medium. The readings were taken over a period of 6 days, and, also, trypan blue was used to monitor the viability of the cells.
Western blotting
Cells were lysed on ice using radioimmunoprecipitation assay (RIPA) buffer (Cell Signaling Technology, Danvers, MA, USA). Around 20 µg of protein from each population was run through a 12% polyacrylamide gel under reducing conditions. Proteins were transferred onto nitrocellulose blotting membrane (Amersham Protran 0.2 µm; GE Healthcare Life Sciences, Pittsburgh, PA, USA) and were blocked for an hour at room temperature using Tris-buffered saline with Tween 20 (TBST; 10 mM of Tris-HCl, 150 mM of NaCl, 0.1% Triton X-100, and 0.05% Tween20, pH 8.0) buffer containing 5% skimmed milk. The membranes were incubated with the following antibodies—p53 (mouse monoclonal; 1:1000; Santa Cruz Biotechnology, Dallas, TX, USA), beta-actin (rabbit monoclonal; 1:1000; Cell Signaling Technology, Danvers, MA, USA), CDK4 (mouse monoclonal; 1:1000; Cell Signaling Technology, Danvers, MA, USA), and p16INK4A (1:1000; Cell Signaling Technology, Danvers, MA, USA).
Preparation of RNA, complementary DNA, and reverse transcription polymerase chain reaction
RNA was isolated from the cell lines using RNeasy Qiagen Kit. The complementary DNA (cDNA) synthesis was carried out using RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Waltham, MA, USA), and the cDNA was used as a template for reverse transcription polymerase chain reaction (RT-PCR; GoTaq qPCR Promega kit). The relative messenger RNA (mRNA) expression level was determined by comparative CT method and was normalized to relative β-actin expression.
HMEC 2.6 cells were characterized by IHC—The pattern of expression of different markers in HMEC 2.6 are summarized here.
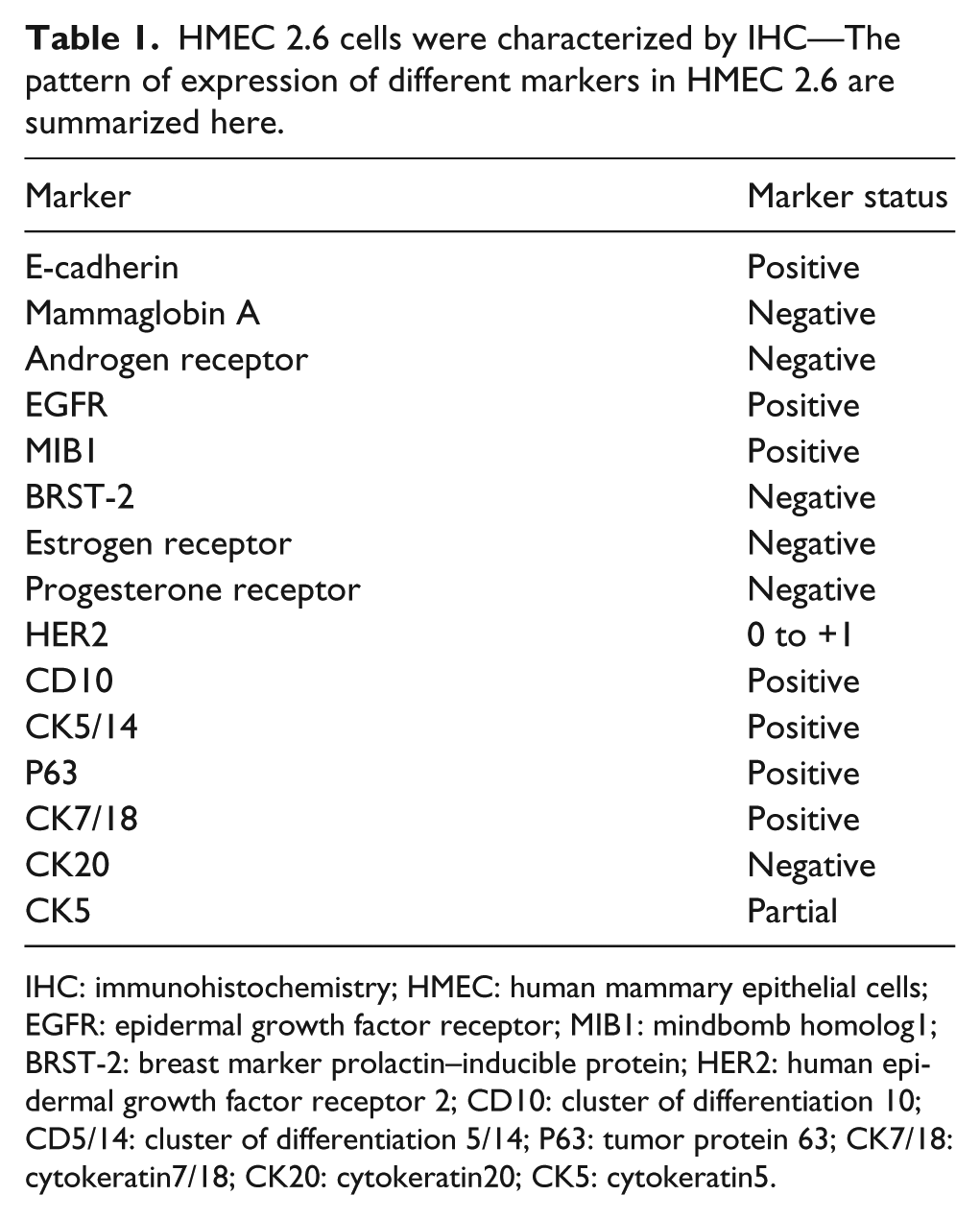
IHC: immunohistochemistry; HMEC: human mammary epithelial cells; EGFR: epidermal growth factor receptor; MIB1: mindbomb homolog1; BRST-2: breast marker prolactin–inducible protein; HER2: human epidermal growth factor receptor 2; CD10: cluster of differentiation 10; CD5/14: cluster of differentiation 5/14; P63: tumor protein 63; CK7/18: cytokeratin7/18; CK20: cytokeratin20; CK5: cytokeratin5.
The RT-PCR primers used for determining the expression level were—hTERT: 5′-AAT CCG TCG AGC AGA GTT-3′ (forward) and 5′-GCG CGG CTT ACC CTT ACC CTT ACC CT-3′ (reverse); β-actin: 5′-AGA AAA TCT GGC ACC ACA CC-3′ (forward) and 5′-AGG AAG GAA GGC TGG AAG AG-3′ (reverse).
Soft agar assay
In total, six-well plates filled with 2 mL of 0.6% noble agar in DFCI-1 media as a bottom layer were allowed to solidify at 4°C for overnight. Around 105 cells were suspended in 1 mL of 0.3% agar in DFCI-1 media and plated on the wells. Each well was fed once a week with 1 mL of 0.3% agar in DFCI-1 media. Wells were scored for colony formation over the next 4 weeks by staining them with 0.005% crystal violet for more than an hour. Cell clumps greater than 60 µm were scored as colonies.
Migration assay
Cell culture inserts were placed on 24 well plates, and 100 µL of serum-free media was added to the chamber along with 200 µL of cells resuspended in serum-free media after trypsinization. Around 700 µL of culture media with serum was added to the lower chambers and allowed to incubate at 37°C for 12–16 h. After incubation, media was removed, and chambers were washed twice with PBS. The cells were fixed by formaldehyde (3.7% in PBS) for 2 min at room temperature. The cells were made permeable by adding 100% methanol. Cells were counted by staining them with Giemsa for 15 min at room temperature.
Immunohistochemistry
Immunohistochemical analysis of HMEC 2.6 was carried out with the help of PhenoPath Laboratories (Seattle, WA) for characterizing the cell line for different cytoplasmic and nuclear molecular markers. Immunohistochemical studies were performed on formalin fixed, paraffin-embedded 4 µm tissue sections. Primary antibodies used in this study to stain different proteins included anti-cytokeratin-7 (OV-TL 12/30; 1:1000; Dako, Carpinteria, CA), anti-estrogen receptor (ID5; 1:200; Zymed Technologies, South San Francisco, CA), anti-progesterone receptor (IA6; 1:50; Lab Vision, Fremont, CA, USA), anti-mammoglobin (304-1A5 1;2000, 31A5 1:400; Corixa, Seattle, WA), anti-cytokeratin 5/6 (5/16 B4; 1:1000; Chemicon, Temecula, CA), anti-cytokeratin 20 (KS 20.8; 1:250; Dako), anti-E-cadherin (HECD-1; 1:300; Zymed Technologies), and Brst-2 (D6; 1:200; Signet, Dedham, MA). After overnight incubation, slides were incubated with secondary antibody. For polyclonal antibodies raised in rabbit, biotinylated goat anti-rabbit immunoglobulin (1:200) was used. For murine monoclonal antibodies, we used biotinylated goat anti-mouse immunoglobulin (1:200). Sections were incubated with secondary antibody for 30 min. The peroxidase staining procedure was done using the ABC Elite kit (Vector Laboratories, Burlingame, CA). The immunostaining reaction was visualized using 0.05% 3,3′-diaminobenzidine in Tris-HCl buffer containing 0.01% hydrogen peroxide, pH 7.6.
Cytogenetics
Metaphase chromosomes from HMEC were prepared using standard cytogenetic techniques. The cells were incubated with 0.06 µg/µL of colcemid at 37°C for 2.5 h and trypsinized with trypsin/EDTA (0.5%/0.1%). The cells were then incubated in 2 mL of 0.075 M potassium chloride for 30 min at 37°C. This was followed by the addition of 1 mL of Carnoy’s fixative (3:1 by volume; methanol: acetic acid) and incubation at room temperature for 10 min. The fixed cells were then spun down and resuspended in 2 mL of Carnoy’s fixative. The prepared metaphase chromosomes were sent to Cell Guidance Systems for G-banding (Wright’s staining) and analysis (St. Louis, MO, USA). Overall, 20 cells were analyzed after G-banding, and the karyotypes were described as per the International System for Human Cytogenetic Nomenclature (ICSN). 21
Nude mice injection
Female BALB/c nude mice (4-week-old) were gamma irradiated to 300 rads. The cell pellet was suspended in Matrigel diluted in unsupplemented Dulbecco’s Modified Eagle’s Medium (DMEM), and 5 million HMEC 2.6 cells were implanted orthotopically in the mammary fat pad.22,23 The positive control cell line used for this assay was RAO-4, a tumorigenic cell line, while the RAO-1 cell line served as the negative control. All cell lines were tested for the presence of any mycoplasma contamination by PCR-based technique using PCR Mycoplasma Detection Kit (Applied Biological Materials, Richmond, British Columbia, Canada). The mice were observed for tumor formation over the following 60 days.24,25
Statistical analysis
Statistical analysis was performed using Student’s t test (two-tailed), and values were expressed as the mean ± standard deviation. p value of less than 0.05 was considered to be statistically significant.
Results
HMEC 2.6 was subjected to cell cycle analysis to look for the DNA content and to enumerate the cells in different stages of cell cycle. As shown in the figures (Figure 1(a) and (b)), the majority of the cells (81%) are in the G1 phase, a minority of the cells are in the S phase (9%), and the remaining (10%) population of cells are in G2/M phase. HMEC 2.6 shows a normal cell cycle phase distribution and confirms a near-diploid population.
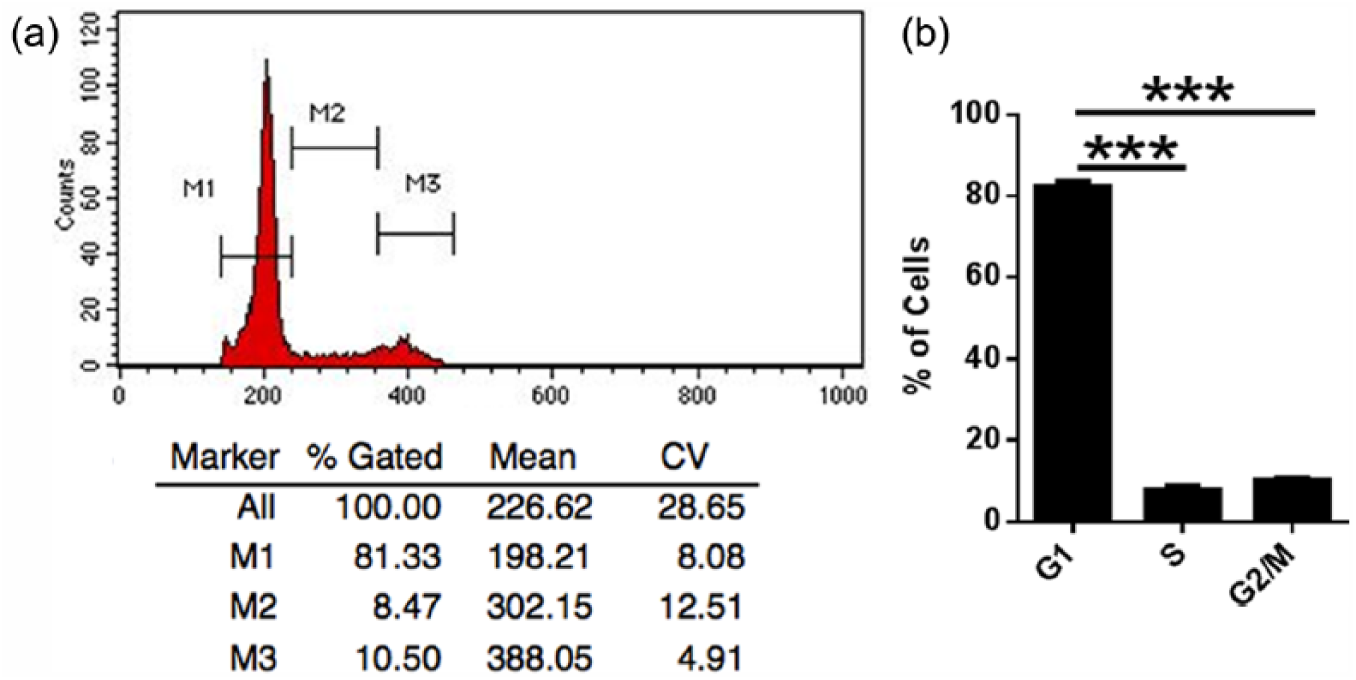
HMEC 2.6 cells were analyzed by flow cytometry. (a) Immortalized HMEC 2.6 line shows normal distribution of cells in each of the four phases of cell cycle. (b) Numerical analysis reveals a statistically significant accumulation of HMEC 2.6 cells in the G1 phase of the cell cycle.
The growth of HMEC 2.6 following inoculation of 50,000 cells is shown in Figure 2 along with RAO-1 cell line that was used as a control for this assay. Cell numbers increased exponentially with RAO-1 having a mean doubling time of 36 h, while HMEC 2.6 grew slowly with a mean doubling time of around 3 days. After Day 4, the growth rate declined in both HMEC 2.6 and RAO-1 cell lines (Figure 2).
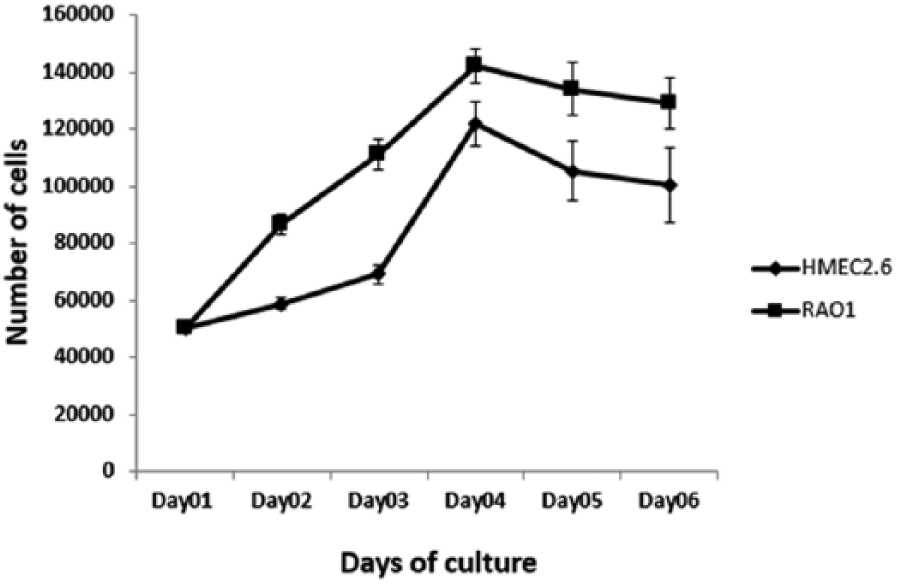
Growth kinetics of HMEC 2.6
HMEC 2.6 cells express high levels of CDK and p16INK4A (Figure 3a) as they were immortalized by introducing a mutant form of CDK4 and transduction of hTERT. However, RAO-1 underwent genome-wide methylation of several genes including p16INK4a as it was allowed to pass through stasis naturally followed by transduction of hTERT. By contrast, HMEC 2.6 does not show altered expression of p16INK4A (Figure 3a). We also see that HMEC 2.6 and RAO-1 express equivalent levels of p53 (Figure 3b). HMEC 2.6 expresses a high level of hTERT due to transduction of hTERT, a key component of immortality when compared to primary HMEC (Figure 3c). Our previous work revealed that RAO-1 cells expressed significantly higher levels of p53 than primary HMEC, 10 and it is likely that p53 levels in HMEC 2.6 are also substantially higher than in primary HMEC.

(a) HMEC 2.6 cells express high levels of CDK4 and maintained expression of p16INK4a, while RAO-1 served as a negative control for p16INK4a expression. (b) HMEC 2.6 cells expressed p53 similar to the positive control RAO-1, while PC3 (prostrate cancer cell line) served as a negative control. (c) HMEC 2.6 cells expressed high levels of hTERT due to the introduction of the catalytic subunit of telomerase enzyme, and primary HMEC served as a negative control.
The anchorage-dependent growth of HMEC 2.6 cells was studied by soft agar assay. The results show that HMEC 2.6 cells do not have the ability to grow in an anchorage-independent manner (Figure 4). This behavior is similar to the negative control cell line, RAO-1 (non-tumorigenic cell line). However, the tumorigenic cell line RAO-4 (positive control) formed visibly larger colonies compared to other cell lines and at least 40 colonies were generated that were scored by using crystal violet stain. The invasive nature of HMEC 2.6 cells was studied by transwell migration assay. The results show that HMEC 2.6 cells are not invasive in nature (Figure 5). These cells behave comparable to the negative control, RAO-1 (non-tumorigenic cell line). The tumorigenic RAO-4 cells which serve as a positive control for this assay are invasive and migrate faster which was confirmed by counting the stained cells under a microscope.

HMEC 2.6 shows no anchorage-independent growth ability

HMEC 2.6 shows less ability to migrate—RAO-4 (positive control) migrates faster, HMEC 2.6 behaves like the negative control for RAO-1 and shows less invasive nature.
Immunohistochemical analysis of HMEC 2.6 confirms that it is a breast epithelial cell line. Figure 6 gives a list of epithelial markers that were tested. The cell line was characterized for various cytoplasmic and nuclear molecular markers as seen in Figure 6. The staining for pan-cytokeratin (CK) was strongly positive confirming the epithelial origin of this cell line. 5 The cells were negative for the presence of estrogen receptor (ER) and progesterone receptor (PR). 6 The cells were positive for cytokeratin 7 and negative for cytokeratin 20 and displayed partial expression for the basal marker, cytokeratin 5. The line is negative for mammoglobin and BRST-2 antigen. The cells are strongly positive for E-cadherin staining but negative for the androgen receptor. The cells are positive for MIB1 and epidermal growth factor expression. The line is positive for other markers such as CD10, CD5, and p63.
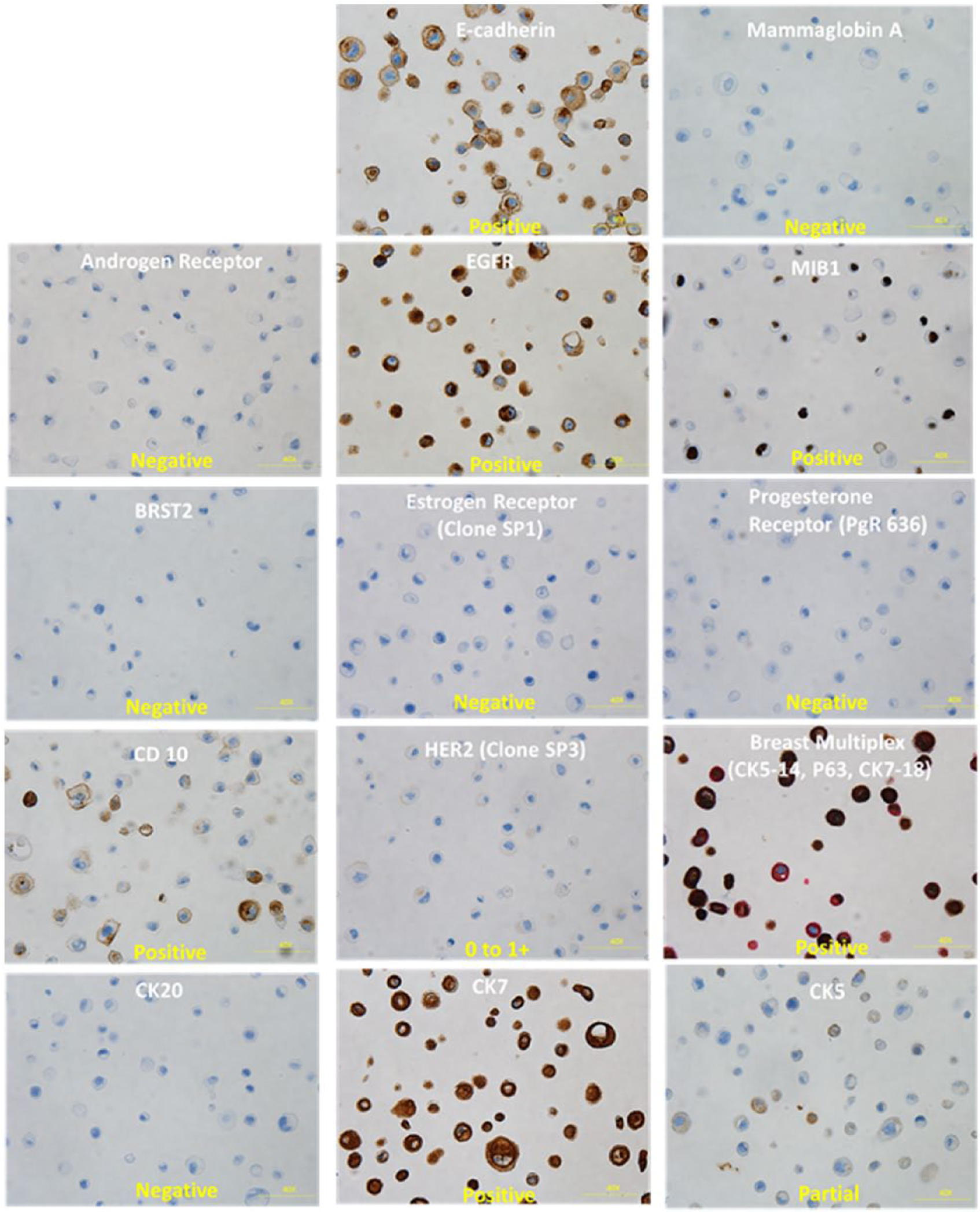
HMEC 2.6 cells were characterized by IHC—Brown stain indicates the positivity of the marker being probed. DAPI (purple) stains the nucleus. The pattern of expression suggests that HMEC 2.6 represents a basal mammary epithelial cell.
Karyotyping results showed a derivative chromosome 1 (48, XX, +1, der(1;21)(q:10;q:10)), in addition to complete trisomy of chromosomes 18 and 20 in 13 of 20 metaphase cells that were analyzed by G-banding. Of the 20 cells analyzed, all the cells showed a derivative chromosome with an entire arm translocation between the long arm of one of the chromosome 1 (breakpoint q10) and the long arm of one chromosome 21(breakpoint q10). We also observed the duplication of chromosome 1 following translocation of the long arm to chromosome 21. The parental karyotype of this cell line was found to be 48, XX, +1, der(1;21)(q:10;q:10),+18,+20 (Figure 7). The summary of the karyotypic changes acquired by HMEC 2.6 during immortalization and further passaging and resulting clonal evolution is shown in Figure 8.

A representative G-banded karyotype of HMEC 2.6. The parental karyotype is 48, XX, +1, der(1;21)(q:10;q:10),+18,+20.

Summary of the karyotypic changes acquired by HMEC 2.6 during immortalization and further passaging and resulting clonal evolution.
The mycoplasma test confirmed that HMEC 2.6 cell line was negative for any mycoplasma contamination (Figure 9) All the mice injected with HMEC 2.6 cells did not have any tumors at the site of injection for up to 6 weeks, thus confirming the benign nature of this cell line. However, we can infer from Figure 10 that RAO-4 cell line which served as a positive control for this assay formed tumors having an average volume of 120 mm3 at the end of 6 weeks in the control group. The mice injected with the negative control cell line RAO-1 did not form any tumor.

HMEC 2.6 cells are mycoplasma negative

HMEC 2.6 shows no tumor formation
Discussion
A variety of methods have been used to immortalize HMEC. It is well known that DNA tumor viruses immortalize cells and have provided a useful model for studying neoplasia for many years. Human papillomavirus (HPV) E6 protein binds p53 and targets it for ubiquitin-mediated degradation, while the E7 protein binds to pRb and inactivates it. 26 HMEC may be immortalized by the combination of HPV 16 E6 and E7 proteins, E7 and hTERT, or by hTERT alone in conjunction with the spontaneous loss of p16 expression. HMECs actively proliferate for several passages in vitro and then become static. 27 The majority of the cells in this stage appear large, flattened, and are less proliferative. Some of the HMEC cells can escape this stasis phase and continue to proliferate, and the number of cells that can escape this depends on the individual donor and the culture conditions. 27 Wong et al. 28 showed that cells were escaping stasis-inactivated p16INK4A through progressive promoter methylation of the CpG islands. Immortalization, however, does not occur till the activation of telomerase occurs. 29 Toouli and colleagues found that Simian virus (SV)40 large T-antigen efficiently immortalized HMEC that were pre-stasis, but when post-stasis cells were used, expression of large T-antigen led to a changed cell morphology, growth arrest, and apoptosis. Exposure to chemical carcinogens has also led to mammary cell immortalization. In 1984, Stampfer and Bartley, 19 exposed rapidly growing primary cultures of HMECs to benzo[a]pyrene for 2–3 days. Benzo[a]pyrene is an environmental carcinogen pollutant that is the result of incomplete combustion of fossil fuel. Upon exposure, DNA adduct formation is noted—particularly between the epoxide metabolite and the deoxyguanosine of DNA in cells susceptible to immortalization/transformation with this agent. Stampfer and Bartley produced two cell lines—184 A1 and 184 B5—with exposure, partial trypsinization, and lengthy culture. The cells still retained expression of keratin, fibronectin, and thioesterase II but displayed an increased nuclear/cytoplasmic ratio, X-ray survival curves with split dose repair, and chromosomal abnormalities. However, the lines were only transformed with the addition of a potent oncogene such as H-Ras. Soule et al. 30 described a spontaneously immortalized normal human breast epithelial cell line and named it as MCF-10. The cells were obtained from mastectomy tissue from a 36-year-old pre-menopausal woman with fibrocystic disease. After 849 days in culture, a population designated MCF-10A was established. The cells exhibited a stable t(3;9)(p13;p22). The cells were non-tumorigenic and non-anchorage independent.
We have attempted to add to the body of knowledge by developing a cell line that was immortalized using defined human genetic elements and that did not rely on passage through stasis as stasis results in the silencing of a wide variety of genes, including E-cadherin, ER, and p16INK4A. Based on our culture conditions, adipocytes and fibroblasts were either removed or could not grow in the culture media. As HMEC 2.6 cells were initially extracted from mammary tissue, we have obtained a population of purified HMECs. The line is diploid as evidenced by flow cytometry and cytogenetics. Consistent with most other mammary cell lines grown as a monolayer, the line does not express ER or PR. The cell line still continues to retain its epithelial nature as it expresses cytokeratin. Cytokeratin 7 expression is present while cytokeratin 20 is absent, conforming to the pattern seen in mammary epithelial tissue. The cell line consistently expressed basal cell markers such as CD10, p63, and cytokeratin 5, suggesting that we have immortalized a basal mammary epithelial cell. 31 However, unlike basal/triple-negative breast cancer, 32 the cells still strongly expressed E-cadherin, suggesting their benign nature.
Haga et al. 33 have shown that HMEC can be immortalized by overexpressing Bmi-1 or using p16INK4a-specific short hairpin that suppresses the expression of p16INK4a along with the introduction of hTERT. They found that HMEC immortalized by p16 shRNA and hTERT or by passage through stasis and introduction of hTERT have a normal or near normal diploid karyotype in contrast to HMEC immortalized by viral oncogenes which showed a tendency for higher chromosomal instability.34,35 Some of the abnormalities seen by Haga et al. on HMEC immortalized p16 shRNA and hTERT included trisomy of chromosomes 7 and 20, additional isochromosome 1, isochromosome 8, inversion 1, and derivative (5; 6). Many of these abnormalities have also been noted in the RAO-1 and HMEC 2.6 cell lines. Although these abnormalities do not lead to transformation, they appear to be acquired rather easily in vitro and may enhance the growth of immortalized cells in vitro and perhaps improve the survival of immortal cells in vivo until they fully transform.
Molecular studies confirmed our previous hypothesis that the cell line retains expression of p16INK4A, as it overexpresses a mutant CDK4. Its immortality was confirmed by the expression of telomerase and repeated passaging. TP53 levels were high, and this is consistent with other immortalized primary human mammary cell lines. 29 TP53 has no direct bearing on the immortalization process in HMECs, although it can certainly impact transformation. 36 Although the cells are negative for BRST-2 and mammoglobin, this is not inconsistent with the basal phenotype, and most basal cell breast cancers are negative for these two antigens. 37 This line does not express androgen receptor, which is found in a specific subtype (LAR) of triple negative tumors. 38 HMEC2.6 stained positively for CD10 (acute lymphoblastic leukemia–associated antigen) which is strongly expressed in normal mammary myoepithelial cells. 39 Cytokeratin 5 is highly expressed in mammary progenitor cells of both glandular and myoepithelial lineages of mammary epithelium.40,41 However, it is interesting to note that CK5 levels are reduced in immortalized HMECs 41 which is also seen with our results as HMEC2.6 cells show partial expression for CK5. HMEC2.6 cells do not show detectable levels of ER and PR, and this lack of expression is commonly observed in cells grown in monolayer. 42 HMEC2.6 expresses high levels of epidermal growth factor receptor (EGFR), consistent with a cell of basal mammary origin. Normal mammary epithelial cells also express CK6, CK14, and CK17 while tumor cells mainly express CK8, CK19, and CK19. 41 HMEC 2.6 cells stain positively for the nuclear protein p6339 that is exclusively expressed in mammary myoepithelial cells.43,44 HMEC2.6 cells are positive for CK5, CD10, and p63. This pattern of staining suggests that HMEC2.6 is a HMEC line of basal or myoepithelial origin.
Trisomy 20 and trisomy 7 are common cytogenetic abnormalities seen in immortalized cell lines. 45 Somatic abnormalities in chromosome 20 have been identified in various types of cancer. Studies suggest that some of the genes found on the long arm of this chromosome may play pivotal roles in controlling the division and growth of the cells. Two of the metaphase spreads show a derivative on chromosome 20 that could have possibly occurred due to translocation from chromosome 7 p13 to chromosome 20p13. Some of the genes found on chromosome 7p13 include CDK13, a regulator of cell cycle and is one of the genes that amplifies and transforms cells, 46 ZMIZ2, a zinc finger motif that plays a novel role in regulating the Wnt/β-catenin pathway, 47 and H2AFV, a member of H2A histone family that is involved in the epigenetic modification of regulating the expression of various genes. 48 Chromosome 20p13 comprised genes such as CDC25B—a cell cycle regulator that also has an important role in the transformation of breast cells, 49 LZTS3—a leucine zipper tumor suppressor that regulates Wnt/β-catenin pathway during cell transformation followed by tumorigenesis, 50 and CENPB—a centromere protein that serves as a prognostic marker for breast cancer patients. 51 Immortalized HMEC gave rise to another chromosomal abnormality, trisomy 7, an aneuploidy that is seen in a number of human cancers. 52 Research studies also suggest that a few of the genes found on chromosome 7 may play important roles in the control of growth and division of cells.53,54 Chromosome 1 is commonly altered in a number of breast cancer cases, 55 and der(1;21)(q:10;q:10) is a common aberration seen in multiple myelomas as well.
A summary of the karyotypic changes involved in the immortalization of HMEC 2.6 is shown in Figure 8. Normal HMEC was passaged, and CDK4R24C, a constitutive mutant of CDK4, and hTERT were introduced into these cells to immortalize them. Cells likely first acquired the karyotype of 48, XX, +1, der(1;21)(q:10;q:10), trisomy 18, and trisomy 20. A fraction of the cells underwent further chromosomal rearrangements to give rise to two related cell populations: one having an additional trisomy 7 and the other with a +der20 t(?7:20)(p13;p13).
Finally, the functional assays (migration, cell growth, and soft agar assays) and nude mouse injection experiments taken together confirmed the non-tumorigenic nature of this cell line. Thus, although these cells have cytogenetic abnormalities, they are still non-invasive and non-tumorigenic when injected into gamma irradiated nude mice. We think, therefore, that this novel cell line can be used as a model to study the impact of a variety of oncogenes on mammary epithelium and to assess the differential toxicity of novel chemotherapeutics on benign and malignant HMECs.
Footnotes
Acknowledgements
The authors gratefully thank the assistance provided by Mr. John Toricelli at the Simmons Cancer Institute Tissue Bank. P.S.J., V.M., and J.C. contributed equally to the manuscript. The cell line is available for purchase through Applied Biological Materials (ABM, Richmond, British Columbia, Canada; #T0454).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by a Concept Development Award grant from the SIU School of Medicine.
