Abstract
The mouse Shb gene coding for the Src Homology 2-domain containing adapter protein B has recently been placed in context of BCRABL1-induced myeloid leukemia in mice and the current study was performed in order to relate SHB to human acute myeloid leukemia (AML). Publicly available AML databases were mined for SHB gene expression and patient survival. SHB gene expression was determined in the Uppsala cohort of AML patients by qPCR. Cell proliferation was determined after SHB gene knockdown in leukemic cell lines. Despite a low frequency of SHB gene mutations, many tumors overexpressed SHB mRNA compared with normal myeloid blood cells. AML patients with tumors expressing low SHB mRNA displayed longer survival times. A subgroup of AML exhibiting a favorable prognosis, acute promyelocytic leukemia (APL) with a PMLRARA translocation, expressed less SHB mRNA than AML tumors in general. When examining genes co-expressed with SHB in AML tumors, four other genes (PAX5, HDAC7, BCORL1, TET1) related to leukemia were identified. A network consisting of these genes plus SHB was identified that relates to certain phenotypic characteristics, such as immune cell, vascular and apoptotic features. SHB knockdown in the APL PMLRARA cell line NB4 and the monocyte/macrophage cell line MM6 adversely affected proliferation, linking SHB gene expression to tumor cell expansion and consequently to patient survival. It is concluded that tumor SHB gene expression relates to AML survival and its subgroup APL. Moreover, this gene is included in a network of genes that plays a role for an AML phenotype exhibiting certain immune cell, vascular and apoptotic characteristics.
Keywords
Introduction
Human acute myeloid leukemia (AML) is initiated by a variety of genetic aberrations and among these, chromosomal translocations, such as t(1;:22), inv(3), t(8;21), t(6;9), t(15;17), t(9;11), and inv(16)/t(16;16), have been identified generating different subclasses of disease.1,2 In addition, mutations that are not the consequences of cytogenetic changes are frequent (FLT3 (fms-related tyrosine kinase 3), NPM1 (nucleophosmin), KIT (v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog), CEBPA (CAAT/enhancer binding protein alpha), TET2 (tet oncogene family member 2), DNMT3A (DNA (cytosine-5)-methyltransferase 3 alpha), IDH1/2 (isocitrate dehydrogenase 1 and 2), NRAS (neuroblastoma rat sarcoma viral oncogene homolog)/KRAS (Kirsten RAS), TP53 (tumor protein p53), BCOR (BCL6 co-repressor), and RUNX1 (runt-related transcription factor 1)). 3 The diversity of these genetic events suggests that the ensuing cellular responses to the initial genetic insult differ between different cases and that each lesion will present a unique signature that will have consequences for disease progression, treatment, and prognosis. In such a scenario, gene mutations will cooperate with the gene and protein expression profile to present a cellular phenotype that depends on both components. For example, acute promyelocytic leukemia (APL), which in most cases results from a chromosomal t(15;17) translocation resulting in a PMLRARA (promyelocytic leukemia retinoic acid receptor A) fusion gene, has distinct clinicopathological features and a favorable prognosis provided treatment based on all-trans retinoic acid or arsenic trioxide. 4 Although numerous alterations in gene expression pathways have been described in different subsets of AML, the complexity of the disease makes it reasonable to assume that many of the genetic or signaling changes that may have an impact on disease characteristics remain unnoticed.
The SHB gene codes for Src homology 2 domain–containing adaptor protein B that operates downstream of tyrosine kinase receptors exerting pleiotropic effects on the cell, such as apoptosis, proliferation, and differentiation, and the cytoskeleton. 5 The Shb knockout mouse shows impaired reproduction, angiogenesis, leukocyte extravasation, hematopoietic stem cell activity, and glucose tolerance. 6 In addition, the T cell response is skewed toward a Th2 phenotype. 7 In all instances studied, Shb gene deficiency was associated with elevated signaling activity in the absence of ligand stimulation of several signaling intermediates including that of focal adhesion kinase (FAK). 6 Noteworthy was the fact that many of the phenotypic effects were observed in Shb+/− mice,8,9 suggesting that minor changes in SHB gene expression may have effects on cellular responses. We recently observed that absence of Shb resulted in shorter latency in a mouse model of BCRABL1 (breakpoint cluster region Abl tyrosine kinase 1)-induced myeloid leukemia because of activation of FAK, increased tumorigenic stem/progenitor cell proliferation, and production of granulocyte colony-stimulating factor (G-CSF) and interleukin-6 (IL-6). 10 SHB gene coding mutations are infrequent in cancer and particularly in AML, have so far none been observed (The Cancer Genome Atlas (TCGA): http://www.cbioportal.org). However, two SHB gene translocations have been observed in acute lymphoblastic leukemia (ALL).11,12 Despite the fact that SHB gene mutations are infrequent in human AML, the results from the experimental mouse model suggest that the SHB gene could have implications for blood neoplasms and that altered expression of this gene may change the cellular phenotype and thus the disease characteristics. This study was conducted in order to address whether human AML patients show differences in their SHB gene expression and whether such alterations have an impact on disease outcome.
Materials and methods
Gene expression by quantitative real-time reverse transcription polymerase chain reaction
RNAs from patients (n = 34) and cell lines were subjected to one-step quantitative real-time reverse transcription polymerase chain reaction (RT-PCR) performed with QuantiTect™ SYBR®Green RT-PCR Kit (QIAGEN, Hilden, Germany) on a LightCycler™ real-time PCR machine (LightCycler 2.0; Roche, Mannheim, Germany). Gene expression was normalized for differences in RNA by subtracting the corresponding GUSB (beta-glucuronidase) Ct value. Statistical comparisons were made on normalized Ct values. Further experimental details can be found in Supplemental materials and methods.
Analysis of data on TCGA website
Patient data and SHB messenger RNA (mRNA) were downloaded from the website13,14 corresponding to the AML study. 3 Data for each patient ID number were matched and the sets were divided in quartiles, and these were subjected to further analysis as indicated in the figures. Gene co-expression data with SHB, PAX5, HDAC7 (histone deacetylase 7), BCORL1 (BCL6 co-repressor-like 1), or TET1 as reference were also downloaded and compared. Data on SHB gene expression in normal blood cells and AML were downloaded from the BloodSpot website15,16 and shown in Supplemental Figure 1.
SHB gene knockdown
NB4 cells were spin infected and green fluorescent protein (GFP)-positive cells were determined from day 0 until day 6 post-infection. The values were normalized by subtracting non-transfected cell values and related to the values on day 2 (set as normalization values for viral transduction efficiency). On day 4, puromycin (4 μg/mL) was added to the cells. Alternatively, MM6 cells were spin infected and cultured for 2 days before adding puromycin to the cultures (2 μg/mL). After 7 days of selection, the cells were used for proliferation experiments in which 105 cells of each virus (control, SHB-A, SHB-B) were placed in two 1 mL tissue culture wells, one with phosphate-buffered saline (PBS) only and the other with PBS + 1 mM isopropyl beta-D-1 thiogalactopyranoside (IPTG). Media were replaced daily and the cells were counted in a Bürker chamber. For more experimental details, see Supplemental materials and methods.
Uppsala cohort of AML RNA samples
Biobanked tumor cell RNAs from 17 patients with newly diagnosed APL (median age: 53 years old, range: 23–77) and from 17 patients with non-APL AML (median age: 58 years old, range: 18–81) were used in the experiments. All patients received remission induction treatment and had given consent to the study and consent to publish. Survival data were obtained from the medical record. The study was approved by the Ethics Committee of Uppsala University (Dnr 2014/233) and was in accordance with the Helsinki Declaration of 1975 as revised in 2008.
Statistical analysis
Mean ± standard error of the mean (SEM) is given for the number of observations indicated. Statistically significant differences were tested by Student’s t test, one-way analysis of variance (ANOVA; Tukey’s or ANOVA on ranks), or log-rank test as indicated. Survival plots were according to Kaplan–Meier analysis.
Results
In silico analysis of SHB gene expression in AML
Since absence of Shb influenced the disease characteristics of BCRABL1-induced myeloid leukemia in mice and since no SHB gene mutations have been observed in AML, we first explored whether human AML showed altered SHB gene expression. The BloodSpot 15 website shows gene expression for any given gene in various AML cases and in normal hematopoietic cell types. In this cohort, all AML cases express SHB mRNA and most express higher mRNA levels than normal hematopoietic cells (Supplemental Figure 1).
SHB gene expression in the TCGA AML database was related to overall survival, age, white blood cells (WBCs), and percent blasts in blood (Supplemental Figure 2). Kaplan–Meier analysis of this cohort showed increased overall survival in patients with tumors expressing low SHB mRNA (Figure 1(a)). This effect was also apparent when examining survival among deceased patients (Figure 1(b)). Subjecting the data in Supplemental Figure 2 to multivariable Cox regression analysis revealed that continuous SHB mRNA and age were significantly related to survival (Table 1). Assessment of SHB gene expression in relation to cytogenetic aberrations (Figure 1(c)) showed high SHB mRNA in patients with t(8;21), RUNX1RUNX1T1, translocations whereas patients with the t(15;17), PMLRARA, translocation exhibited less SHB mRNA. FLT3-mutated tumors expressed less SHB mRNA (Figure 1(d)).
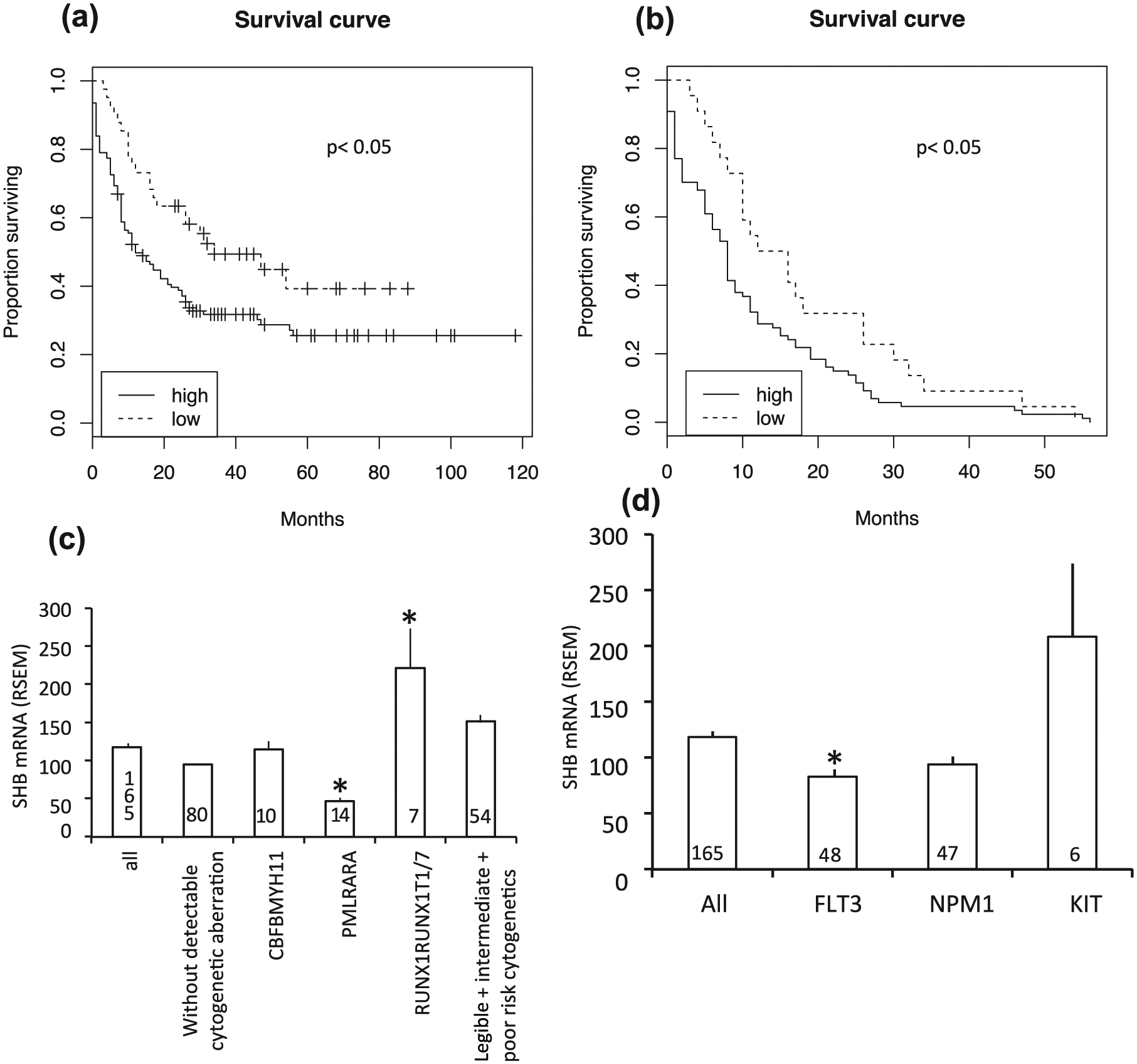
AML survival and cytogenetics/mutations in relation to tumor SHB gene expression in the TCGA cohort. (a) Kaplan–Meier overall survival (in months) comparing the lowest quartile SHB mRNA with patients expressing more SHB mRNA (n = 165). (b) Kaplan–Meier survival curve for deceased patients comparing low SHB mRNA versus rest (n = 109). Log-rank test p values have been indicated in (a) and (b). (c) SHB gene expression in tumors with cytogenetic aberrations. (d) Tumor SHB mRNA among patients carrying FLT3, NPM1, and KIT mutations. Mean ± SEM for the number of observations indicated is given in (c) and (d) (* indicates p < 0.05 when compared with “all” group using one-way ANOVA on ranks).
Multivariable Cox regression analysis of AML patient survival in relation to SHB mRNA, age, white blood cells, and blasts in blood.
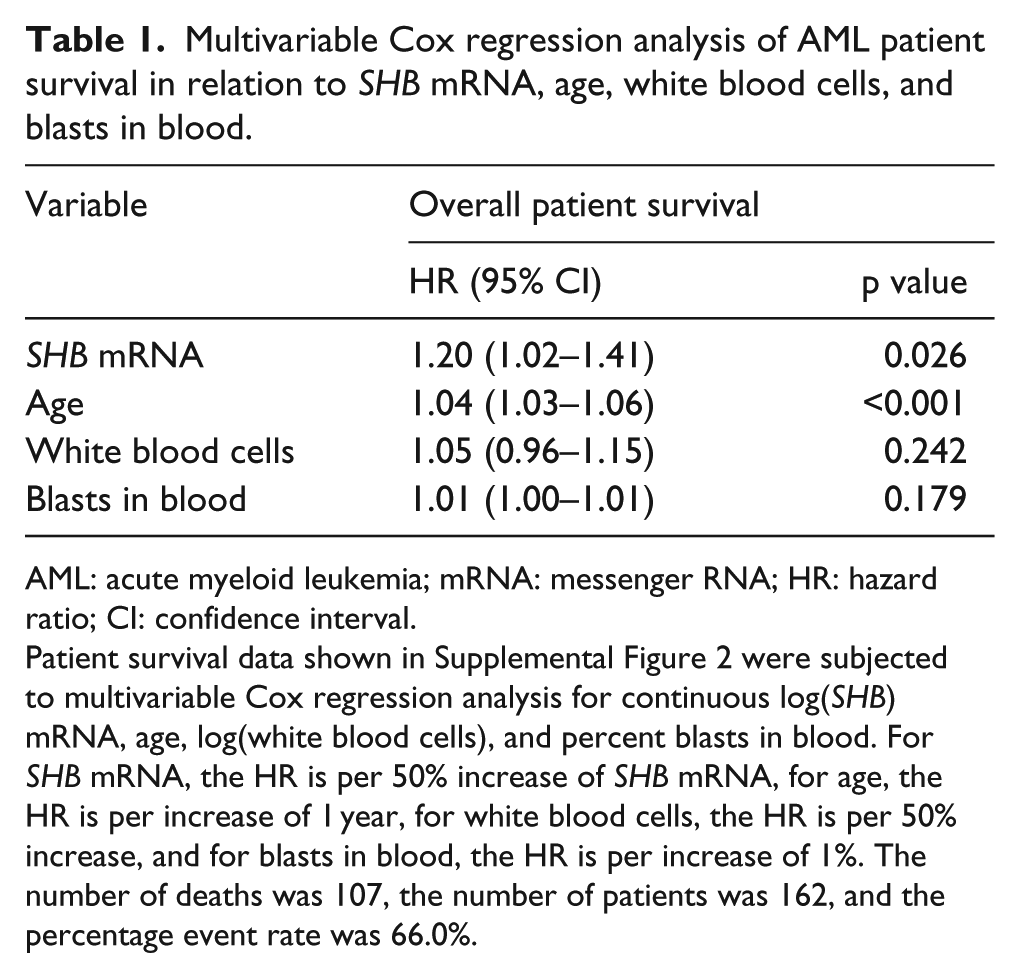
AML: acute myeloid leukemia; mRNA: messenger RNA; HR: hazard ratio; CI: confidence interval.
Patient survival data shown in Supplemental Figure 2 were subjected to multivariable Cox regression analysis for continuous log(SHB) mRNA, age, log(white blood cells), and percent blasts in blood. For SHB mRNA, the HR is per 50% increase of SHB mRNA, for age, the HR is per increase of 1 year, for white blood cells, the HR is per 50% increase, and for blasts in blood, the HR is per increase of 1%. The number of deaths was 107, the number of patients was 162, and the percentage event rate was 66.0%.
SHB gene co-expression in AML patients
The TCGA AML database contains information on gene co-expression of 20,420 genes in 173 patients. The 242 genes co-expressed with SHB using a Pearson score of 0.35 or higher are shown in Supplemental Table 1. Several members of the human leukocyte antigen (HLA) family are present in addition to phospholipase C-γ (PLC-γ) and the leukocyte antigens CD34 (cluster of differentiation 34), CD52, CD74, and CD79B. SHB has been shown to regulate PLC-γ activation in lymphoid cells7,17,18 and hematopoietic stem cell properties10,19 (CD34 is a marker for this feature in leukemia 20 ), whereas other co-expressed genes relate to the lymphoid lineage. Subjecting this set of genes to ToppGene analysis (https://toppgene.cchmc.org) revealed significant enrichment of the following biological functions: T cell selection, development of the vasculature and the circulatory/cardiovascular system, consequences of major histocompatibility complex (MHC) peptide binding, and PLC-γ activity (Supplemental Table 1). Also, notable was the presence of the co-expressed genes of PAX5, HDAC7, BCORL1, and TET1. These are transcriptional regulators implicated in various aspects of leukemia.21–24 Performing the corresponding co-expression analysis of these using the TCGA AML website revealed a signature of co-expressed genes (Supplemental Table 2) that largely overlapped between these four transcriptional regulators and SHB as demonstrated by a Venn diagram showing the number of shared co-expressed genes (Figure 2(a)). Since the SHB and PAX5 co-expressed genes showed the percentagewise largest degree of overlap, expression of SHB and PAX5 was assessed in seven human myeloid leukemic cell lines (Figure 2(b)). Regression analysis revealed a significant correlation between SHB and PAX5 in these cell lines, reinforcing the notion of PAX5 being co-expressed with SHB in patient samples. Table 2 identifies genes that were co-expressed among three or more of the five members of this network (SHB, PAX5, HDAC7, BCORL1, and TET1), whereas Supplemental Table 3 shows genes that were selectively co-expressed between the two listed partners. It should be noted that this network shows some resemblance with the previously identified RNA-sequencing group 6 network 3 with the presence of all genes of that group, except C5ORF23 and MOSC1, in the SHB/PAX5/HDAC7/BCORL1/TET1 network. The genes co-expressed between each pair of the five partners were also subjected to ToppGene analysis (Supplemental Table 4). The SHB/PAX5 co-expressed genes were strongly enriched in numerous functions relating to the immune system, whereas the SHB/BCORL1 co-expressed genes showed a profile of biological functions mainly relating to the development of the vasculature. The PAX5/HDAC7 co-expressed genes also related in many instances to the immune system although their functions were more diverse than those of the SHB/PAX5 co-expressed genes. The HDAC7/TET1 co-expressed genes showed significant clustering around regulation of cell death and immune cell responses. Biological functions related to various aspects of cell adhesion also were detected in several of the comparisons. In conclusion, there appears to be a network consisting of SHB, PAX5, HDAC7, BCORL1, and TET1 that has phenotypic consequences for leukemic cells. These are accentuated lymphoid properties, properties relating to vascular development and cell survival. Since the lowest SHB expressing quartile lived longer, it is assumed that these characteristics confer unfavorable properties to the leukemic cells with respect to patient survival.
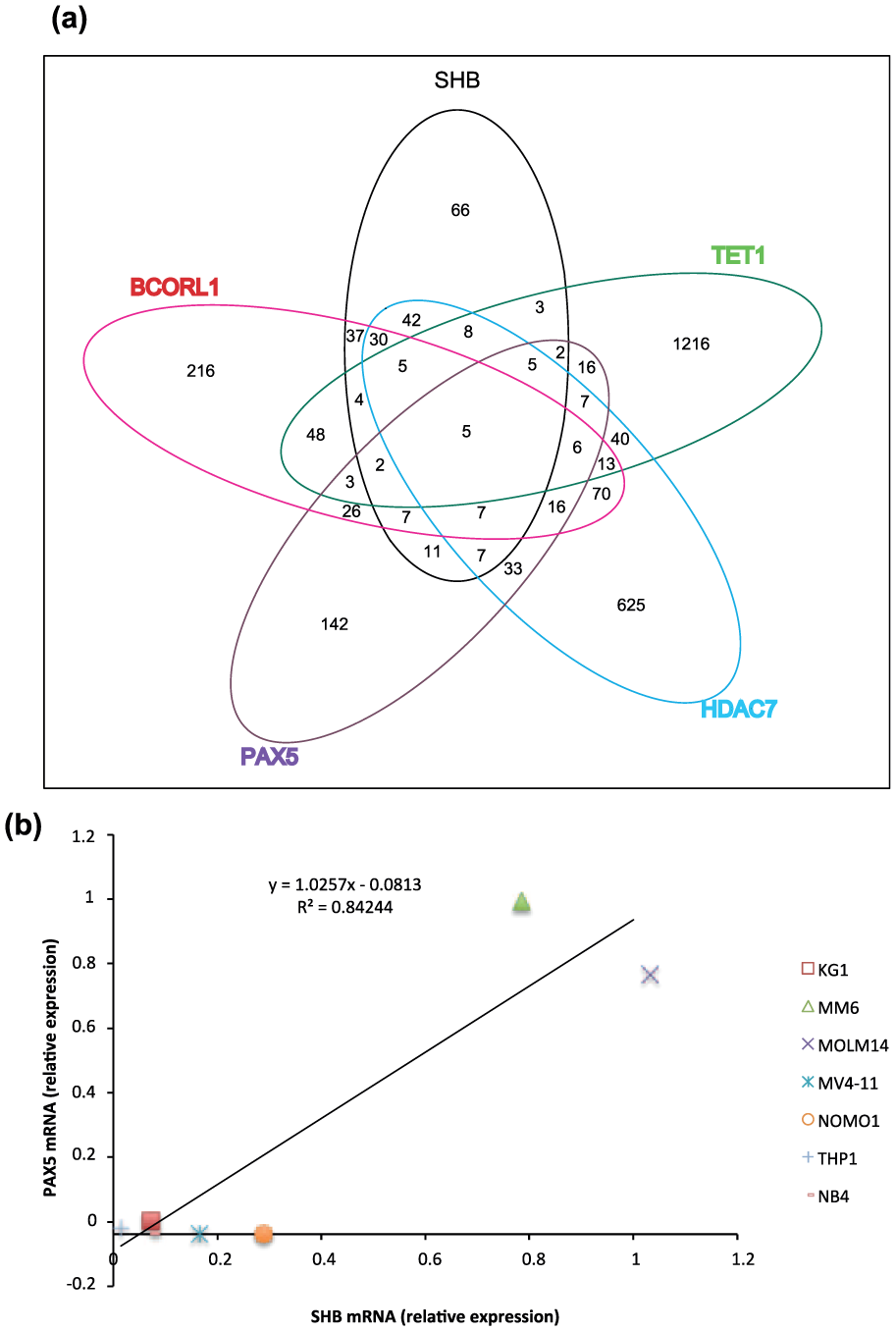
(a) Venn diagram showing overlap in gene co-expression with SHB, PAX5, HDAC7, BCORL1, and TET1.(b) SHB and PAX5 mRNA expression in seven human myeloid leukemic cell lines, as well as trend line and Pearson R2 values.
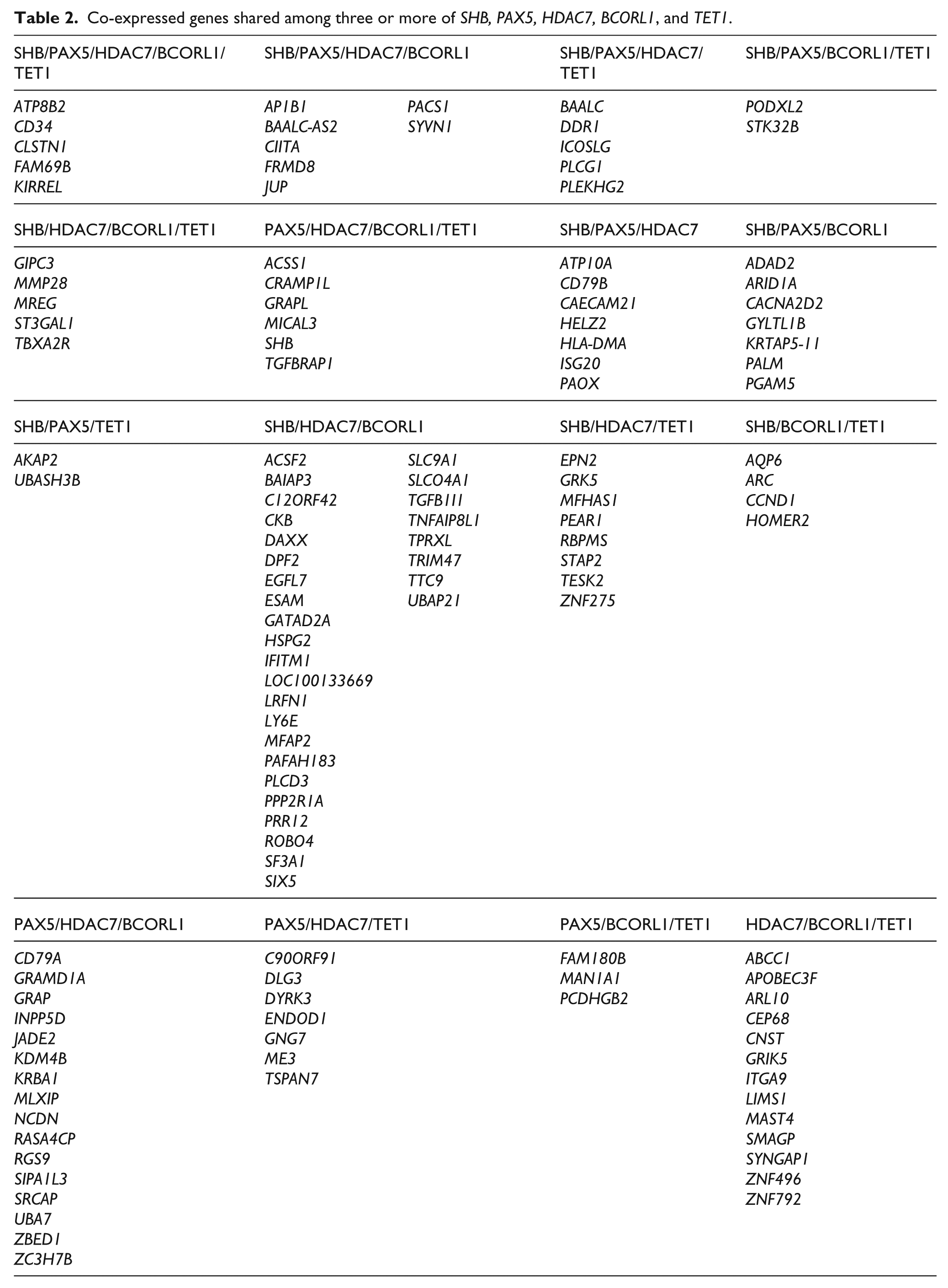
Co-expressed genes shared among three or more of SHB, PAX5, HDAC7, BCORL1, and TET1.
Analysis of APL patients with PMLRARA translocations in the Uppsala AML repository
Since the PMLRARA translocation was found to be associated with a lower tumor SHB mRNA content in the TCGA study, we decided to investigate this further in the AML RNA repository at the Department of Clinical Genetics at Uppsala University by semi-quantitative real-time RT-PCR (qPCR) of SHB mRNA. Patient characteristics and tumor cytogenetics are shown in Supplemental Table 5. Values were normalized for the house-keeping gene GUSB. As was the case in the TCGA AML study, acute promyeolocytic leukemia (APL) tumors expressing the t(15;17) PMLRARA fusion gene had a more than 40% reduction of SHB mRNA (Figure 3(a)) as compared with AML tumors without this chromosomal translocation. PMLRARA APL has a favorable prognosis, and in the Uppsala cohort, 5 out of 17 patients were deceased of which most deaths were attributed to initial complications of the disease. Among the non-PMLRARA patients, deceased patients had higher tumor SHB mRNA content than those in complete remission-1 (Figure 3(b)). Taken as a whole, the Uppsala and TCGA data show a high degree of concordance and suggest that low SHB mRNA correlates with a favorable prognosis and PMLRARA APL.

SHB mRNA expression in Uppsala cohort. (a) AML tumors without the PMLRARA translocation were compared with PMLRARA APL tumors. (b) AML patients (without PMLRARA) alive with complete remission-1 (CR1) were compared with deceased patients with respect to SHB mRNA. SHB qPCR values were normalized for the house-keeping gene GUSB by subtraction of the latter Ct value. Values are relative expression (in arbitrary units). In (a), the median was set at 1, whereas in (b), the lowest value was set as 1. Ct values were converted by the formula 2−Ct. Mean ± SEM is given (* indicates p < 0.05 with a Student’s t test). The numbers of observations are indicated. (c). Correlations between SHB and PAX5, HDAC7, BCORL1, and TET1 mRNAs in the Uppsala cohort by qPCR. Trend lines and Pearson R2 values are shown. All correlations were significant (p < 0.05).
Significant correlations between SHB and its other four network partners (PAX5, HDAC7, BCORL1, and TET1) were also established in the Uppsala AML cohort (Figure 3(c)).
SHB knockdown in a PMLRARA leukemic cell line reduces proliferation
To understand the role of SHB in leukemic cell function, the PMLRARA leukemic cell line NB4 25 was used for SHB knockdown. For SHB, short hairpin RNA (shRNA) knockdown sequences in lentiviruses were used (denoted shSHB -1, -2, -3 and -4) and compared with a control knockdown sequence (control). Figure 4(a) shows a successive decrease in the percentage of GFP-positive cells (virus-infected) during days 3 to 6 using the shSHB -2 and -3 sequences compared with the control, whereas the -1 and -4 sequences showed a trend toward a decrease on day 6 that failed to reach statistical significance, suggesting a difference in knockdown efficacy between the four different target sequences. In four attempts, none of the shSHB sequences generated stable clones after puromycin selection, whereas the control shRNA sequence readily generated long-term expanding clones. Since no surviving knockdown cells could be obtained, an inducible knockdown system was also tested for investigating the role of the SHB mRNA content for leukemic cell proliferation. MM6 cells (unlike THP-1 and NB4 cells) showed significantly decreased SHB mRNA using this system displaying 55% ± 12% and 15% ± 3% reductions in SHB mRNA, respectively, using the SHB-A and SHB-B shRNA sequences (p < 0.05 each). The control sequence did not affect SHB mRNA. Both sequences decreased cell proliferation on days 4 and 5 after IPTG induction in a dose-dependent manner, further implicating SHB gene expression in myeloid leukemic cell proliferation (Figure 4(b)). Consequently, the data support the notion that low SHB gene expression promotes a favorable prognosis due to reduced tumor cell proliferation.
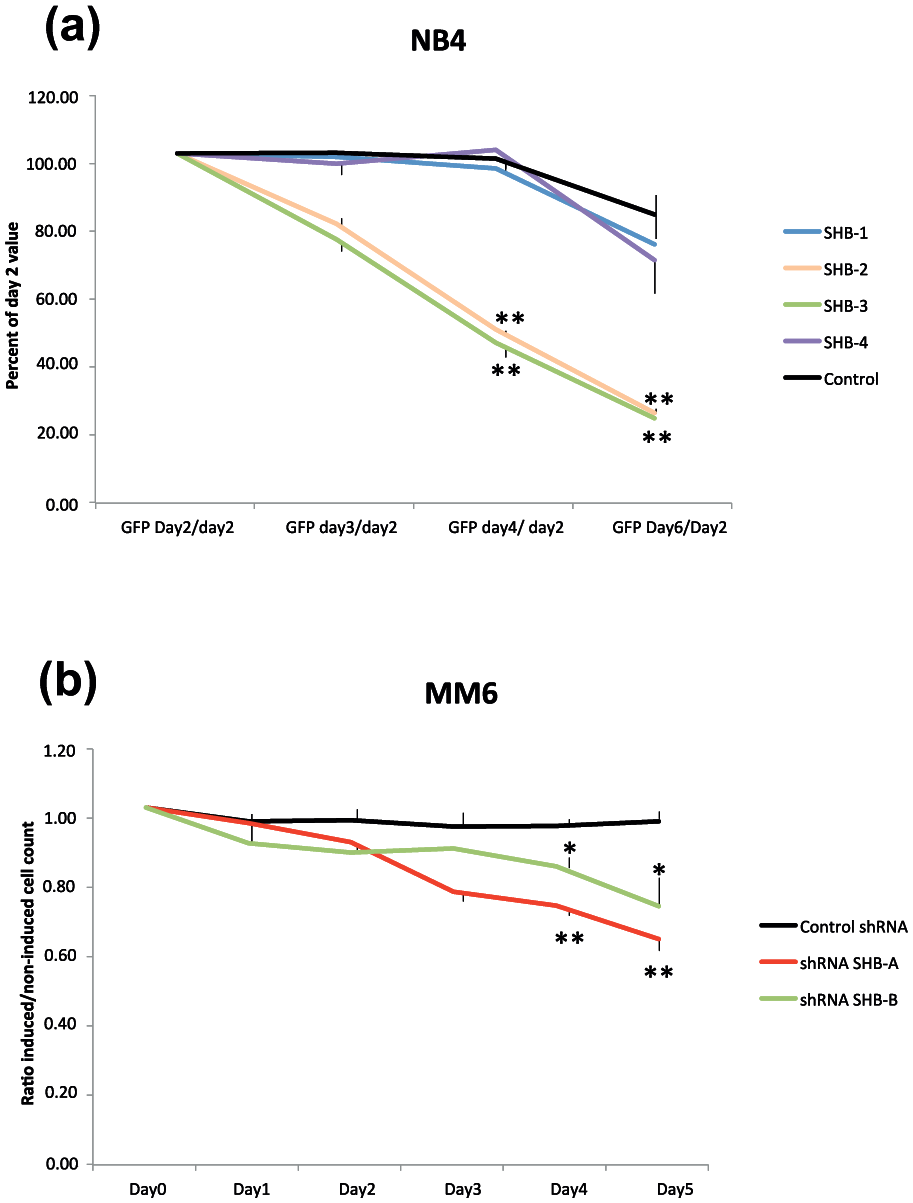
SHB knockdown decreases NB4 proliferation (a) NB4 cells were infected with lentiviruses expressing shSHB-1, -2, -3, -4, and control. Cells were maintained in culture and counted every day. The percentage of GFP-positive cells (percentage of cells infected) was determined on day 2, 3, 4, and 6. Values are percentages of GFP-positive cells on day 2. Values were determined in duplicates for each virus infection, and three separate infections were done. Mean ± SEM is given. Values of significance were tested by ANOVA and compared by Tukey’s test with control on days 4 and 6 (** indicates p < 0.01). (b) Inducible SHB knockdown in MM6 cells inhibits proliferation. Cells were counted every day after IPTG addition, and values are ratios of IPTG cells:vehicle cells. Mean ± SEM for four independent observations (performed at different times) is shown (* and ** indicate p < 0.05 and 0.01, respectively, when compared with control by ANOVA (Tukey’s test)).
Discussion
The current study places the SHB gene in a context of human leukemia. We have previously shown that absence of Shb promotes BCRABL1-induced myeloid leukemia in mice. 10 In contrast, the SHB knockdown experiments in this study show reduced proliferation and failure to select for expanding clones, suggesting that SHB is important for human myeloid leukemic cell proliferation and consequently that there is selective pressure operating against a reduction of SHB gene expression in this setting. This is further supported by the absence of reported SHB gene mutations in human AML, that all patients in the BloodSpot cohort express tumor SHB mRNA and that absence of Shb in mice conferred reduced proliferation of hematopoietic stem cells. 10 We also currently show that SHB gene expression relates to phenotypic properties of leukemic cells, such as overall survival and chromosomal aberrations. Two well characterized chromosomal translocations, PMLRARA: t(15;17)(q22;q21) and RUNX1/RUNX1T1: t(8;21)(q22;q22), were found to exhibit differences in SHB mRNA, the former with reduced mRNA levels and the latter with increased mRNA. Since both have a favorable prognosis, 4 SHB mRNA does not strictly correlate with survival but probably with several phenotypic characteristics, among which survival is one variable, in a heterogeneous disease. SHB knockdown reduced MM6 and PMLRARA NB4 cell proliferation and thus the favorable prognosis of APL and other AML tumors with low SHB mRNA probably relates to decreased leukemic cell proliferation.
The “SHB dependent phenotype” in AML remains elusive but reveals certain features based on the gene co-expression characteristics. The expression of PAX5, HDAC7, BCORL1, and TET1 was correlated with that of SHB. The HDAC7 gene product is a transcriptional repressor involved in lymphoid cell development 23 and angiogenesis 26 and has been shown to play a role for pro-B-ALL 23 or B-ALL. 27 The BCORL1 gene gives rise to a co-repressor of transcription, and HDAC7 is one of its interacting partners. 21 BCORL1 is frequently mutated in AML. 21 The PAX5 protein is a transcription factor necessary for B cell maturation, and the gene is frequently mutated in B cell leukemia. Indeed, one SHB/PAX5 translocation has been described in B-ALL. 11 In AML, PAX5 is commonly overexpressed in tumors exhibiting the RUNX1/RUNX1T1 translocation, and it is associated with a lymphoid cell surface phenotype with increased expression of CD19 and CD79A. 24 However, other AML cases without that chromosomal aberration may also show the “lymphoid” phenotype. 28 The TET1 gene is linked to both T-ALL 29 and AML, the latter as a TET1/MLL translocation or as a product of the MLL/AF9 fusion protein.22,30 The SHB network showed a considerable overlap with the RNA-sequencing group 6 which has an unfavorable prognosis. 3 None of the 24 genes identified as a prognostic signature for AML was found to be co-expressed with SHB, 31 whereas 22 genes (BAALC, BAIAP3, CACNA2D2, CD34, DDR1, FAM69B, GIPC3, GRK5, HSPG2, ICOSLG, IFITM1, ISG20, MREG, PALM3, PEAR1, PLCG1, PLEKHG2, SLCO4A1, STK32B, TBXA2R, TGFB1I1, and ZNF275) profiled in different molecular subtypes of AML 32 were present among the SHB-co-expressed genes listed in Table 1. However, the SHB/PAX5/HDAC/BCORL1/TET1 network shows only limited resemblance to the poor prognosis of 17-gene stemness signature. 33
Currently, when addressing the functional significance of the overlap in co-expression between SHB/PAX5/HDAC7/BCORL1/TET1 by ToppGene analysis, certain phenotypic characteristics can be observed. The most extensive list of biological functions appeared among SHB/PAX5 co-expressed genes in which functions relating to immune cell activation dominated. Likewise were immune cell functions observed in the analysis of PAX5 and HDAC7 co-expressed genes, albeit to a lower degree. The link between PAX5 and immune cell function is well established as described above. Since SHB has been shown to interact with the T cell receptor18,34 and absence of Shb in mice confers a Th2 skewing of the immune response,7,35 it thus seems plausible that SHB is an important player in this context. The role of the “immune” phenotype for AML remains an enigma but should be further investigated. Tentative explanations for the poor prognosis of the high tumor SHB mRNA “immune phenotype” could be activation of various immune reactions that could be deleterious to the patient or it could cause tumor cell evasion from immune cells combating the tumor. 36 Another aspect of the currently observed immune phenotype could be autoimmunity. Patients suffering from autoimmune conditions are more likely to develop myeloid neoplasia such as AML and myelodysplastic syndrome, 37 and the T cell signaling gene expression profile may play a role in that context. The other group of biological functions identified showed a relationship to angiogenesis and the vasculature and these were seen when comparing SHB/BCORL1 co-expressed genes. These features are supported by the well-documented importance of Shb for angiogenesis in the mouse,8,9,38–40 although the relevance of these biological functions for AML is currently unclear. However, there is a considerable overlap in gene expression between vascular and hematopoietic cells, and classical angiogenesis markers, such as CD31, may also play a role for hematopoietic cell function as recently illustrated by the immunosuppressive function of CD31 in T cells. 41 Finally, the TET1 and HDAC7 co-expressed genes relate to apoptosis. It can be noted that SHB also plays a role for apoptosis in a context-dependent manner, both in vitro and in vivo.5,10 Taken together, the data suggest a network consisting of at least five genes, SHB, PAX5, HDAC7, BCORL1, and TET1, which has an impact on at least three phenotypic features (immune cell characteristics, vascular function, and apoptosis) in the AML cohort.
In conclusion, SHB gene expression plays an important role for phenotypic characteristics of AML or its subgroup APL. The most apparent feature is shorter survival time, and this finding is reinforced by the observation of reduced proliferation of a leukemic cell line upon SHB knockdown in vitro. In addition, “immune cell” and “vascular feature” characteristics of the leukemic cells were also noted. The consequences of the latter features in AML remain largely unexplored. It is our belief that assessment of SHB gene expression may become useful as a prognostic marker for this subgroup of AML.
Footnotes
Acknowledgements
The authors acknowledge the support from Bioinformatics Infrastructure for Life Sciences (BILS) at Uppsala University for helping in data presentation and Katja Gabrysch at Uppsala Clinical Research Center for statistical analysis. The authors are grateful to Richard Rosenquist at Uppsala University, Michael Kharas at the Memorial Sloan Kettering Cancer Center, and Andrius Kazlauskas at Roche Pharma Research for suggestions. Maria Jamalpour and Xiujuan Li performed the experiments; Michael Welsh analyzed the online databases; Martin Höglund provided the clinical information on the patient samples; and Lucia Cavelier provided the clinical information on the tumor RNA samples. All authors have provided suggestions on the experimental design and interpretation of the results. Michael Welsh wrote the paper with the help of Maria Jamalpour, Lucia Cavelier, Karin Gustafsson, and Martin Höglund All authors have read and commented the text.
Availability of data and materials
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by the Swedish Cancer foundation, the Swedish Research Council, the Family Ernfors fund, and EXODIAB.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
