Abstract
Lung cancer is by far the leading cause of cancer death in the world. Despite the improvements in diagnostic methods, the status of early detection was not achieved. So, a new diagnostic method is needed. The aim of this study is to obtain the highly specific nucleic acid aptamers with strong affinity to tumor markers in the serum of the lung cancer patients for targeting the serum. Aptamers specifically binding to tumor markers in the serum of the lung cancer patients were screened from the random single-stranded DNA library with agarose beads as supports and the serum as a target by target-substituting subtractive SELEX technique and real-time quantitative polymerase chain reaction technique. Subsequently, the secondary single-stranded DNA library obtained by 10 rounds of screening was amplified to double-stranded DNA, followed by high-throughput genome sequence analysis to screen aptamers with specific affinity to tumor markers in the serum of the lung cancer patients. Finally, six aptamers obtained by 10 rounds of screening were identified with high specific affinity to tumor markers in the serum of the lung cancer patients. Compared with other five aptamers, the aptamer 43 was identified both with the highest specificity to bind target molecule and without any obvious affinity to non-specific proteins. The screened aptamers have relatively high specificity to combine tumor markers in the serum of the lung cancer patients, which provides breakthrough points for early diagnosis and treatment of lung cancer.
Keywords
Introduction
Lung cancer is by far the leading cause of cancer death in the world, and its incidence increased greatly over the past 50 years. 1 In some large industrial cities located in the Western developed countries, the incidence of lung cancer has been ranked first for men and second for women, which means that lung cancer has become a major disease endangering lives and health of people.2,3 However, many patients found to have lung cancer at an advanced stage have lost the best chance of treatment. 4 In addition, the 5-year survival rate of lung cancer patients is very low with only about 5%–10%.5,6 Therefore, early diagnosis and treatment are crucial to improve the survival rate of the patients with lung cancer. 7
Up to now, lung cancer was clinically diagnosed through joint detection of tumor markers in the serum of the lung cancer patients. 8 The most common tumor markers include mainly neuron-specific enolase (NSE), carcinoembryonic antigen (CEA), cytokeratin 19 fragment (CYFRA21-1), gastrin-releasing peptide (GRP), and squamous cell carcinoma antigen (SCC-Ag).9–11 However, the sensitivity of traditional lung cancer diagnosis techniques is too low to detect early lung cancer patients. 12
Aptamer is one kind of single-stranded DNA (ssDNA) or RNA with highly specific affinity to target molecules. 13 Compared with the detection of the antibody with the sensitivity at the level of picomole/liter, the detection of nucleic acid aptamers has the highest sensitivity at the level of nanomole/liter and other advantages including specific affinity to more target molecules, no immunogenicity, higher stability, and easier modification. 14 Therefore, nucleic acid aptamers can be used for clinical detection and treatment in place of antibody to achieve early diagnosis.15,16
Herein, we report for the first time that, with the serum of the lung cancer patients as a target, six DNA aptamers with specific affinity to tumor markers in the serum of the lung cancer patients were obtained through a novel, improved SELEX technique that can screen specific aptamers binding to the tumor markers of lung cancer more rapidly than the traditional SELEX technique, which provides a new efficient and accurate method for early diagnosis of lung cancer.17,18
Materials and methods
Materials
Random ssDNA library (88 n, 1.2 nmol; 5′-CTATAGCAATGGTACGGTACTTCC (40N) CAAAAGTGCACGCTACTTTGCTAA-3′; N = A, G, T, and C), P7 primer (5′-CTATAGCAATGGTACGGTACTTCC-3′), P11 primer (5′-TTAGCAAAGTAGCGTGCACTTTTG-3′), and P11 biotin-labeled primer (5′biotin-TTAGCAAAGTAGCGTGCACTTTTG-3′) are purchased from Shanghai Brilliance power grid Biotechnology Co., Ltd. (Table 1). The sera of lung cancer patients and healthy persons are supplied by Qinhuangdao First Hospital, Hebei Province in China. All related polymerase chain reaction (PCR) reagents are gained from Shanghai Biological Technology Co., Ltd. today Mai.
Double-stranded DNA reaction system.

PCR: polymerase chain reaction.
Blood treatment
The whole blood samples of 50 lung cancer patients were collected and then centrifuged at 1500 r/min for 5 min at room temperature and successively at 12,000 r/min for 5 min at 4°C to collect serum samples after removal of red blood cells. After removal of high-abundance proteins by successive 50,000, 30,000, and 10,000 ultrafiltration interception, followed by the centrifugation at 15,000 r/min for 15 min at 20°C, the supernatant was stored at −80°C for further use.
Screening mixed serum of healthy persons and lung cancer patients by subtractive SELEX technique
Each round of screening consists of two steps: negative screening and positive screening. Carboxyl agar beads (0.3 mL) and mixed serum of lung cancer patients (200 µL) were mixed and incubated at 37°C for 2 h in a positive sieve tube, and carboxyl agar beads (0.3 mL) and mixed serum of healthy persons (200 µL) were mixed and incubated at 37°C for 2 h in a negative sieve tube. The magnetic beads in the two tubes were recovered, followed by the addition of transfer RNA (tRNA; 10 µmol/L). The mixture in both tubes was incubated in a rotating incubator for 30 min at 37°C. tRNA was used as non-specific competitive agent for screening to increase the specificity of screening. Subsequently, the recovered magnetic beads in these two cubes were rinsed twice with screening buffer twice (each time 1 min) and binding buffer once for 1 min. The primary ssDNA libraries were added into the negative sieve tube containing the recovered magnetic beads, and the mixture was incubated at 37°C for 1 h. Subsequently, the supernatant containing the ssDNA libraries uncombined to the magnetic beads in the negative sieve tube was added into the positive sieve tube containing the recovered magnetic beads, and the mixture was incubated at 37°C for 1 h. The recovered magnetic beads were rinsed three times with screening phosphate-buffered saline (PBS) containing 0.1% Tween-20 three times (each time 1 min) to remove the ssDNA libraries with weak affinity to the proteins in the sera of the lung cancer patients. The recovered magnetic beads in these two tubes were mixed with deionized water (200 µL). After the above mixtures in these two tubes were treated at 95°C for 5 min and 0°C for 10 min, the supernatants were collected. Some recovered supernatants in the negative sieve tube and the positive sieve tube were used as a negative group and a positive group, respectively. The positive and negative groups were simultaneously measured to determine the number of cycles through a real-time quantitative PCR technique and to monitor the enriched content of mixed serum of lung cancer patients. The residual supernatant in the negative sieve tube was amplified through a real-time quantitative PCR technique and subsequent streptavidin–biotin magnetic beads–screening method to be used as a secondary library for the next round of screening. Each subsequent round of screening consists of two steps: negative screening and positive screening. The conditions for screening are shown in Table 2.
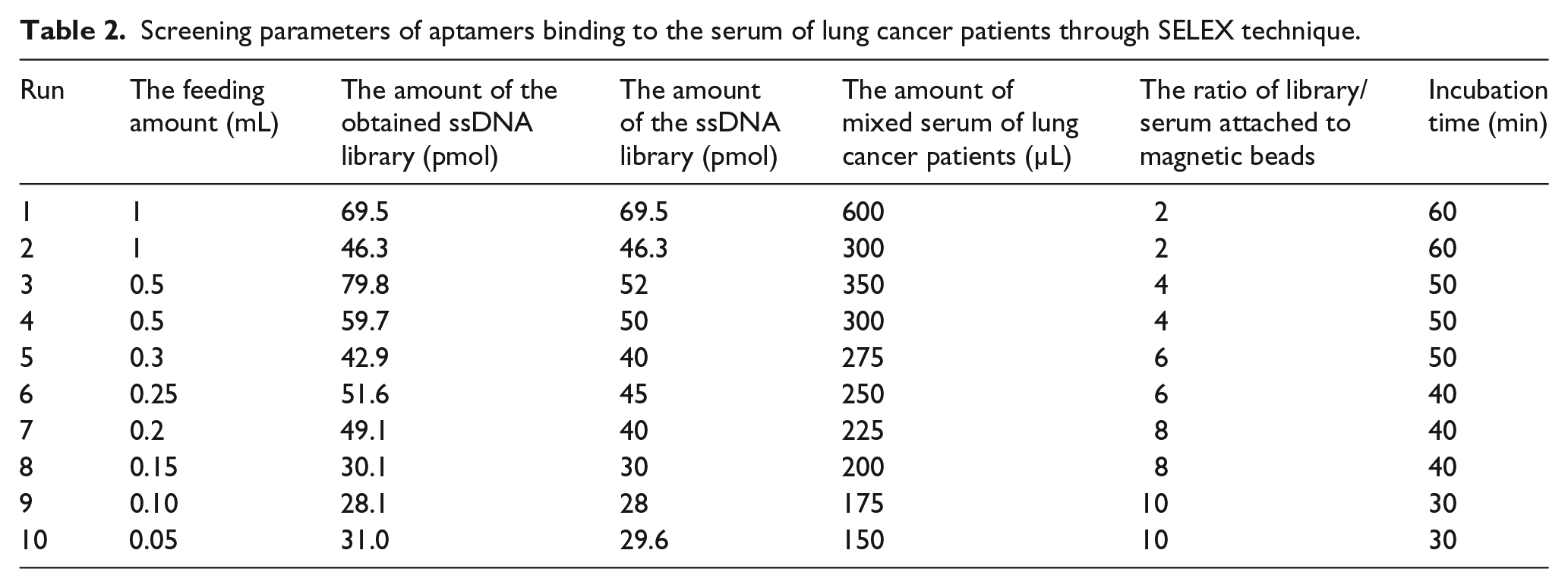
Screening parameters of aptamers binding to the serum of lung cancer patients through SELEX technique.
Preparation of ssDNA using magnetic bead method
A volume of 0.1 mL of streptavidin–biotin magnetic beads and 500 µL of streptavidin–biotin was added into a 1.5 mL Eppendorf (EP) tube and incubated in a rotating incubator for 1 h at 37°C. Subsequently, the recovered magnetic beads were washed three times with the PBS solution (each for 1 min), and then, double-stranded DNA (dsDNA) obtained through PCR amplication was added. The mixture mentioned above was incubated in a rotating incubator for 30 min at 37°C and the supernatant was discarded. In the tube containing the recovered magnetic beads, 1 mL of 1 wt% Tris phosphate-buffered saline (TPBS) solution was added, and the mixture was incubated for 10 min at 40°C. Subsequently, the magnetic beads were separated and mixed with 200 µL of LDEPC solution. The obtained mixture was incubated for 5 min at 95°C and 10 min at 0°C. Finally, the supernatant was collected.
Preparation of dsDNA through amplification of real-time polymerase chain reaction
A volume of 160 µL of template was added into the PCR reaction system, and the homogeneous PCR reaction solution was divided into eight equal portions and amplified through the real-time polymerase chain reaction (RT-PCR) technique. dsDNA was obtained when the S-type curve reached the peak.
High-throughput genome sequence analysis
The supernatant obtained during the 10 round of screening was taken as the template, and 240 µL of PCR solution including 48 µL of the template was packed equally in eight tubes to accomplish PCR amplification. The PCR products in a frozen environment were delivered to Shanghai Bioengineering Company for high-throughput sequencing analysis. The 10,000 sequences of aptamers were obtained and the aptamer with the highest specificity was screened through software analysis.
Identification of the specificity of aptamers
A volume of 200 µL of the serum of lung cancer patients and healthy persons was mixed with 0.1 mL of carboxyl agar beads. The corresponding mixture was then incubated at 37°C for 1 h. The recovered magnetic beads were washed with screening buffer solution three times for 1 min each time and then mixed with 100 µL of the obtained nucleic acid aptamers that have specific affinity to tumor markers in the serum of lung cancer patients. Subsequently, these two mixtures were incubated at 37°C for 30 min. These recovered magnetic beads were washed with a 1% mixed Tween and PBS solution three times and mixed with 100 µL of deionized water. After the mixture was incubated at 95°C for 10 min, these two supernatants were collected to test through RT-PCR technique.
Results
Screening results of mixed serum of lung cancer patients used as combined target through SELEX technique
A total of 10 rounds of aptamer screening were carried out, and each round of screening conditions was gradually enhanced. As shown in Table 2, 10 rounds of screening were performed. Accompanied by an increase in the rounds of screening, the screening condition becomes rigid, the ratio of library/serum attached to magnetic beads goes up, and incubation time reduces; thus, the aim of screening was achieved (Figure 1).

Flowchart of aptamer screening.
Sequencing results
We sent samples to Shanghai Brilliance power grid Biotechnology for high-throughput sequencing, and the samples were analyzed by sequence analysis software DNAMAN. The DNA sequence from 5′- to 3′-end on two fixed extremities is given in Table 3. Six DNA aptamers with specific affinity to tumor markers in the serum of the lung cancer patients were screened from 1000 aptamer sequences obtained through high-throughput sequencing analysis. The aptamer 43 has the largest Kd value of 64.0 ± 3.6, the aptamer 49 has a Kd value of 61.0 ± 5.9, and the aptamer 5 has the smallest Kd value of 25.0 ± 4.9.
Sequences of selected aptamers.

Secondary structure prediction of four nucleic acid aptamers that have specific affinity to tumor markers of the serum of lung cancer patients
We used NUPACK software to analyze the secondary structure of the nucleic acid aptamer sequence of lung cancer we screened. Figure 2 shows secondary structure prediction spectra of six nucleic acid aptamers to tumor markers of the serum of lung cancer patients, the secondary structures of the six aptamers are predicted by NUPACK, demonstrating that all the six obtained aptamers form the stem loop structure which features multiple stem-loop structures as well.

Secondary structure prediction spectra of six nucleic acid aptamers that have specific affinity to tumor markers of the serum of lung cancer patients: (a) Apt-3, (b) Apt-5, (c) Apt-15, (d) Apt-25, (e) Apt-43, and (f) Apt-49.
Identification of the specificity of four nucleic acid aptamers that have specific affinity to tumor markers of the serum of lung cancer patients
We used real-time quantitative PCR to test the specificity of the six nucleic acid aptamers. As shown in Figure 3, the number of cycles in the positive and negative groups was about five cycles. The six obtained aptamers have specific affinity to the serum of lung cancer patients and do not have any specific affinity to the serum of healthy persons, demonstrating that the obtained aptamers have high specific affinity to the serum of lung cancer patients.

Identification of the specificity of six nucleic acid aptamers that have specific affinity to tumor markers of the serum of lung cancer patients: (1 and 2) the serum of lung cancer patients and (3 and 4) the serum of healthy persons.
Lung cancer serum test results
We used 20 cases of both lung cancer patients and non-lung cancer patients to detect the specificity of the six nucleic acid aptamers. Figure 4 plotted by real-time quantitative PCR test results shows that the difference in Ct values of cycles between the six lung cancer serum and non-lung cancer serum is 4–6. Indicating that the subtractive SELEX technique used for screening the six nucleic acid aptamers can identify the lung cancer serum specifically.

In all, 20 patients with lung cancer were tested for serum and non-lung cancer serum. The results were obtained by real-time quantitative analysis. The average number of cycles was obtained. It can be seen that the Ct values of Apt-43 (5.6) and Apt-49 (5.2) were the best; Apt-15 has a Ct value of 4.8, Apt-3 has a Ct value of 4.3, Apt-25 has a Ct value of 4, and Apt-5 has a Ct value of 3.8.
Discussion
The aim of this study is to screen specific nucleic acid aptamers with strong affinity to the sera of lung cancer patients through a subtractive SELEX technique. The carboxylated agar beads were combined with serum proteins of lung cancer patients under different screening parameters to finally obtain six specific nucleic acid aptamers. In order to remove targeted molecules, for example, nucleic acid aptamers, of non-target species, negative screening was first performed. Subsequently, the recovered supernatant and magnetic bead combined with the sera of lung cancer patients were incubated together to conduct positive screening, followed by the treatment at 95°C to obtain specific ssDNA libraries for the sera of lung cancer patients. The obtained ssDNA libraries were amplified though a real-time quantitative PCR technique and subsequent combination of streptavidin–avidin beads to prepare the secondary ssDNA libraries for the next screening of the aptamers. Finally, the second-generation gene high-throughput sequencing analysis was used to determine the enriched aptamer sequences and the enriched content of the aptamer sequences specifically binding targeted molecules.19The successful screening is greatly dependent on both the used ssDNA libraries and the feeding amounts of ssDNA library and the serum of lung cancer patients. In order to obtain the highly specific nucleic acid aptamers with strong affinity to the sera of lung cancer patients, the amounts of ssDNA libraries and the sera of lung cancer patients should gradually decrease with the increase in the screening round, but the feeding ratio of ssDNA library and the sera of lung cancer patients should gradually go up.
The SELEX technique is an efficient route to obtain the aptamers with high specificity and strong affinity through several rounds of screening of exponentially enriched oligonucleotides specifically binding targeted molecules from large-capacity random oligonucleotide library combined with an in vitro PCR amplification technique.20,21 The ssDNAs in random oligonucleotide library could form many spatial structures including a hairpin, false knot, convex ring, and G- four split, and these structures could stably bind targeted molecules. 22,23The nucleic acid aptamer sequences with high specificity and strong affinity were finally obtained though screening and enrichment. Compared with the antibodies such as a single-strand DNA and RNA fragment, the nucleic acid aptamers show not only similar functionality but also higher stability, easier tunability, and higher specificity to targeted molecules.24
Up to now, the serum is still the best specimen for in vitro diagnosis, and the detection of tumor markers is the only non-invasive diagnostic method in clinical treatment.25 Lung cancer, one of the malignant tumors threatening seriously human health and life, has attracted much attention. Early diagnosis and treatment are crucial to improve the survival rate of lung cancer patients. The aptamers with diverse structures could show high specificity and strong affinity to many targeted proteins. Accordingly, the diagnosis method using the aptamers as the probes is superior to the traditional antigen–antibody method in the detection of proteins, holding a great promise in practice. Our six screened nucleic acid aptamers were used to combine with the sera of lung cancer patients and healthy persons from Qinhuangdao First Hospital to diagnose lung cancer early. First, the aptamers were combined with serum proteins adhered onto magnetic beads, followed by elution and thermal denaturation. Subsequently, the aptamers attached on the magnetic beads combined with the sera of healthy persons and lung cancer patients were attained, respectively. The enrichment content of the aptamers was monitored though a real-time quantitative PCR technique. The difference between the Ct value of ssDNA combined with the sera of lung cancer patients and that with the sera of healthy persons is defined as the delta Ct value. The real-time quantitative PCR test results reveal that all the delta Ct values of six aptamers range from 4 to 6 rounds. In particular, the delta Ct value of the aptamer, Apt-43, with the highest specificity, is 5.6 rounds. These results mentioned above demonstrate that all our six screened aptamers have strong affinity to serum proteins of lung cancer patients.
In summary, six aptamers with high specificity and strong affinity to the sera of lung cancer patients have been successfully screened though a subtractive SELEX technique. These screened aptamers could be used as the simple, efficient, and stable DNA molecular probes for early diagnosis of lung cancer. Moreover, the tumor markers in the sera of lung cancer patients could be extracted through our aptamers, followed by protein sequencing analysis to establish spectral library of the corresponding tumor markers and investigate the interaction mechanism between the aptamers and the proteins, which provides a new breakthrough for the diagnosis and treatment of lung cancer.
Footnotes
Compliance with ethical standards
The study protocol was approved by the Ethics Committee of the First Hospital of Qinhuangdao, with informed consent signed by all participants.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by “Qinhuangdao Science and Technology Research and Development Plan,” China (nos 201501B034, 201401A105, 201501B051, and 201502A193); Special Research Fund for the Doctoral Program of Higher Education (no. 20121333120017); and Hebei Province Key Research and Development Projects (no. 17272402).
