Abstract
Virus-directed enzyme prodrug therapy is one of the major strategy of increasing cytotoxicity of bioreductive agents. This research intended to examine new selected benzimidazole derivatives as a substrate for nitroreductase, the enzyme involved in nitroreduction which is responsible to the production of cytotoxic metabolites. In this way, the selectivity and strength of cytotoxicity can be raised. The effect of benzimidazoles on virus transfected cells and non-virus transfected cells A549 cell line was established by Annexin V + propidium iodide test, western blot, and polymerase chain reaction analysis of specific pro- and anti-apoptotic proteins in the corresponding gene expression and additionally nitroreductase gene expression. Our results proved the pro-apoptotic properties of all tested compounds in normoxia and hypoxia, especially according to virused A549 cells where the time of exposition was reduced from 48 to 4 h. In this shorten period of time, the strongest activity was shown by N-oxide compounds with nitro-groups. The apoptosis was confirmed by generation of BAX gene and protein and reduction of BCL2 gene and protein.
Keywords
Introduction
Taking into consideration increasing number of cancer diagnosis all over the world, novel cancer treatments are strongly investigated. One of such approaches is gene therapy, which has been successfully performed for the first time in 1990.1,2 Until 2016, 2300 clinical trials were conducted (majority in phase I). 3 Basic principle of gene therapy is to deliver nucleic acid polymers (DNA or RNA) into targeted cells as a drug which should treat specific disease. This is a key point and the greatest difficulty in gene therapy—achieving the best possible treatment selectivity.4–6
It can be done through insertion of correct copy of a gene (in case of lack of the gene or insufficient level of expression), insertion of correct DNA fragment to replace fragments where single-point mutations occur, silencing incorrect copy of a gene (by non-coding DNA/RNA sequences), and insertion of a DNA fragment coding pro-apoptotic factors which will cause natural process of cell death. One of the methods of introduction of nucleic acid into cells is using plasmid or viral vectors. Most commonly used are adenovirus, thymidine kinase and cytosine deaminase, retroviruses, lentiviruses, adeno-associated virus (AAVs), herpes simplex virus, Sindbis virus, and vaccinia virus.7–12 Currently, the ways of delivering an enzyme/prodrug therapy are being investigated the most: introducing genes that encode prodrug-activating enzymes into tumor cells (GDEPT), the use of viral vectors for transgene introduction (virus-directed enzyme prodrug therapy (VDEPT)), and an antibody designed against tumor cells linked to an enzyme and injected to the blood (ADEPT).4,8,9,13
VDEPT is one of the therapies allowing to enter therapeutic enzyme to the cancer cells and target it with a prodrug in order to obtain activated and cytotoxic for cancer cells drug.4,14,15
Enzyme prodrugs are promising when it comes to improving therapy selectivity. 16 This therapy is a two-step process. First step is to target and express drug-activating enzyme in tumor cells. Second step is to apply a non-toxic prodrug which is a substrate of targeted enzyme. Prodrug is converted into active form, and this way a high concentration of an anticancer drug is obtained in cancer cells. In this approach, enzyme should be human protein (which is absent or expressed at low concentrations in normal tissue and has a high catalytic activity and adequate level of expression in the cancer cells) or it should be a non-human origin. The prodrug should meet the following requirements:
It should not be activated in non-tumor cells but should be a good substrate for targeted enzyme in tumor cells.
It should cross cell membranes of tumor cells.
Prodrug should be as non-toxic as possible, but an active form should have a high cytotoxicity level.
An active drug should be diffusible and show a “bystander effect” (killing neighboring cancer cells).
An active drug should have a half-life time sufficient to start “bystander effect” but short enough to prevent drug to reach systemic circulation.17,18
VDEPT strategy has a high therapeutic potential in cancer treatment and is worth investigating. Described method was used in this study in order to trigger cytotoxic activity of new bioreductive compounds—benzimidazole derivatives. These compounds are introduced into an organism as an inactive prodrug and then are reduced to active form by an enzyme—nitroreductase. In our research, the carrier of a nitroreductase gene was viral vector—adenovirus. Moreover, tested compounds were compared to reference compounds—tirapazamine and 2-nitroimidazole (NI). The evaluation of cytotoxic activity has been done by assessing the stage of cell death as early- and late-apoptotic or necrotic cells.15,18,19
Results and discussion
Tested compounds
The effectiveness of gene-directed enzyme prodrug therapy was tested using four representatives of the benzimidazole derivative class, identified by numbers from 1 to 4, respectively (Table 1). Additionally, tirapazamine and NI were used as reference compounds due to their high selectivity to hypoxic conditions. NI also shows increased affinity to the nitroreductase enzyme. Concentrations of listed compounds were chosen in accordance to its IC50 values designed previously. Benzimidazole derivatives have been prepared using procedures described earlier. 20
Tested benzimidazole derivatives (1–4) and reference compounds (T and NI).

Benzimidazoles and their induction of the apoptosis in normal and virused A549 cells
To determine the cytotoxic activity of the new benzimidazole derivatives, the Annexin V + propidium iodide (PI) assay was carried out (Figures 1 and 2). This analysis allowed indicating the pathway of tumor cell death which occurs after addition of tested compounds. The participation of alive cells, cells in early and late apoptosis, or necrotic cells in the whole population of A549 cell line was also specified. The main problem referred to the evaluation and comparison of the phase of induced apoptosis and necrosis. Conducted experiment showed the dominance of early-apoptotic cells over late-apoptotic and necrotic cells, both in the culture performed in normoxia and hypoxia for normal A549, in time up to 48 h (Figure 1). Pro-apoptotic effects of tested compounds were also indicated after incubation of virused cells in normoxic and hypoxic conditions for 4 h for virused A549 (Figure 2). Results show that treatment using tested and reference compounds, compared to control, revealed increase in number of apoptotic cells of about 60% under hypoxic conditions, while in normoxia it was increased by 50%. Hypoxic conditions induced early apoptosis with much higher efficiency than normoxia for all tested substances, especially for compound 3, which showed the biggest difference between normoxic (58%) and hypoxic (74%) conditions. Taking into consideration the influence of tested compounds on apoptosis pathway induction, the strongest impact was obtained in samples with compound 4 (N-oxide) in normoxia and hypoxia, followed by compound 3—analogue of compound 1. Compounds having the nitrophenyl substituent (compounds 1 and 2) demonstrate higher impact on apoptosis pathway induction that compounds with chlorophenyl substituent (compounds 3 and 4). Also, N-oxide compounds increase the number of apoptotic cells with higher selectivity.

Results of Annexin V + PI test for normoxia (N) and hypoxia (H) on normal A549 cells. The A549 cells were exposed to IC50 of the tested compounds 1–4 for 48 h.

Results of Annexin V + PI test for normoxia (N) and hypoxia (H) on virused A549 cells. The A549 cells were exposed to IC50 of the tested compounds 1–4 for 4 h.
Much more evident results were obtained using virused A549 cell line. The amount of early apoptotic cells in samples treated by tested compounds was of about 2 times higher in normoxia and of about 3 times higher in hypoxia, comparing to the control samples. However, control sample in hypoxic conditions revealed about 2 times more cells in the phase of early apoptosis than control sample in normoxia, thus overall results indicate much higher efficiency of compound treatment approach in hypoxia. Similarly, to test with normal cells, virused cells showed higher susceptibility on induction of apoptosis pathway when treated by N-oxide compounds. However, in that case, the chlorophenyl substituent played greater role in treatment, as both compounds 1 and 4 had the highest impact but compound 2 had the lowest.
In both normal and virused cell lines, despite the conditions, the highest efficiency of treatment by tested compounds was indicated in samples with compounds 1 and 3, and these results were comparable to samples treated by two reference compounds—tirapazamine and nitroimidazole. Also, in all tested assays, it can be observed that hypoxic conditions had higher impact on induction of cell apoptosis than normoxia. Moreover, combination of tested compounds’ treatment and nitroreductase gene vector transfection showed stronger influence on apoptosis induction than in the case of normal cells, especially if we notice that virused cells were treated only for 4 h, comparing to 48-h incubation of non-virused cells. The increase in number of necrotic cells shown in Figures 1 and 2 should be considered as a result of a natural cell death through apoptosis pathway, not through a strong cytotoxic action of tested compounds.
Polymerase chain reaction analysis for nfsB gene expression
The analysis was made using A549 cell line transfected by adenovirus transferring nfsB gene. After transfection, cells were cultured, and after 24 h, the cells were treated with tested compounds. Samples are given in Table 2.
Samples prepared for agarose gel electrophoresis for nfsB gene.

K: control sample for transfection (non-transfected cells); I and II: samples of cells without tested compounds; 1–4: tested compounds; NI: 2-nitroimidazole (reference compound); A and B: samples of non-transfected cells.
Qualitative analysis
To confirm the efficiency of polymerase chain reaction (PCR), reaction with β-actin gene (ACTB) was carried out. ACTB gene is a basic metabolism gene, which is present in every cell on stable level. Obtained results of agarose gel electrophoresis showed a single band (about 500 bp) for all samples (excluding sample N, which is a negative control of PCR (without template)). These results confirmed correct amplification of DNA in all tested samples (results not shown). Later, for all investigated cases, qualitative analysis was performed in order to show expression of nfsB gene. Positive results of nfsB gene amplification during PCR were observed in samples I, II, 1–4, and NI. Negative results were observed in sample N (negative control of PCR (without template)) and sample K (control of transfection process (non-transfected cells)). Results of PCR amplification are presented in Figure 3.

Results of PCR amplification of nfsB gene.
To confirm the specificity of the PCR analysis and the absence of the nfsB gene in A549 cell line, PCR amplification was repeated for samples A and B, which were collected before cells’ transfection with adenovirus. Figure 4 shows no visible bands, which confirms that in samples A and B, expression of nfsB gene was not observed.

Results of PCR amplification of nfsB gene for non-transfected cells.
Quantitative analysis
Evaluation of the expression level of nfsB gene in transfected A459 cell line treated with tested compounds was made in triplicate using real-time PCR (CORBETT Research—ROTOR GENE 300). As a reference, β-actin gene (ACTB) was used. After amplification, melting curves were obtained, which confirm product specificity (Figure 5). The results were calculated using the double-delta method (comparative analysis). Ct values (cycle threshold) for investigated nfsB gene and reference gene ACTB are presented in Table 3.

Melting curve graph for investigated gene nfsB.
Ct values after real-time PCR reaction with use of SYBR Green I for tested nfsB gene samples and reference ACTB gene samples.

1–4: samples with tested compounds; NI: sample with 2-nitroimidazole as reference compound; N: negative control of PCR (without template).
Data are expressed as mean (n = 3).
The influence of tested benzimidazole derivatives on the nfsB expression was analyzed based on the comparative Ct method. It involves comparing the Ct values of the sample genes of interest with a calibrator (non-treated sample). The Ct values of both, the calibrator and the investigated samples, are normalized to an ACTB endogenous. It was previously proved that the amplification efficiency of investigated and reference genes is on the same level of expression and every cycle causes doubling of output matrix amount. It allowed to use comparative Ct method of analysis. Results presented in Figure 4 and R values for each sample show that relative expression of nfsB gene was the highest in sample 3 and the lowest for sample 2 (Table 4).
Obtained R values for tested nfsB gene samples and reference ACTB gene samples.

1–4: samples with tested compounds; NI: sample with 2-nitroimidazole as reference compound; N: negative control of PCR (without template).
ΔCt (sample) = Ct (investigated gene) − Ct (reference gene); ΔΔCt = ΔCt (sample) − ΔCt (calibrator); R = 2−ΔΔCt.
The main peak values of Tm for investigated nfsB gene are observed above temperature of 80°C. This result confirmed that PCR reaction was specific. Also, for reference ACTB gene, obtained results were similar and confirmed specificity of reaction (figure not shown).
Fluorescence tested during the PCR reaction was linear in the logarithmic phase of PCR reaction (Figure 6) which allowed to determine Ct (cycle threshold) value. For every sample, product amplification was exponential which means that PCR reaction was successful.

Amplification plots for nfsB and reference ACTB genes.
Real-time PCR for BAX and BCL2 analysis
Expression of BAX and BCL2 gene was analyzed in PCR analysis to show the impact of tested compounds on apoptosis. The virused A549 cells were treated with tested compounds in normoxia and hypoxia, and the expression of pro-apoptotic and anti-apoptotic genes was analyzed. The results proved that all tested substances reduce the expression of BCL2. Minimum reduction was observed in normoxia for compounds 3 and 4. The most interesting outcomes are seen in hypoxia where the strong stimulation of BAX (pro-apoptotic gene) was observed for compounds 1 and 2. It proved that the N-oxide bond and nitro-group in the structure of benzimidazole ring correlate with increase in the cell apoptosis, especially in hypoxia. Thus, it shows potential cytotoxic selectivity of N-oxide nitro-benzimidazoles. Tirapazamine and NI were used as positive reference control (Figure 7).
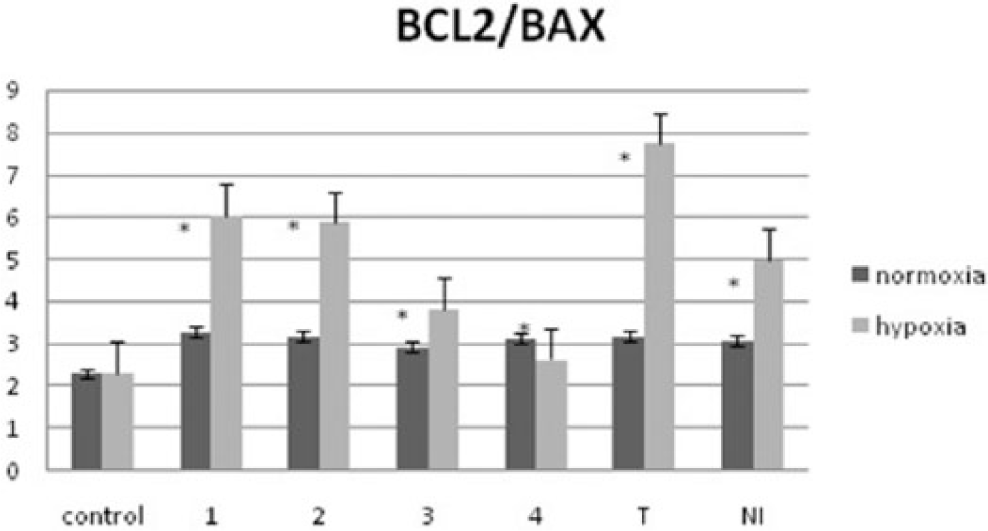
Level of BAX and BCL2 gene expression under normoxic and hypoxic conditions.
Western blot analysis for BAX and BCL2
To prove the pro-apoptotic properties of tested benzimidazoles, the levels of BAX and BCL2 proteins were checked. Virused A549 cells were treated with tested compounds. The highest level of pro-apoptotic protein BAX was indicated for compounds 1 and 2, especially in hypoxia. It might be possible due to the presence of N-oxide bond and nitro-group in their structure. Tirapazamine and NI were used as positive reference control (Figure 8).

Level of BAX and BCL2 protein biosynthesis under normoxic and hypoxic conditions.
Experimental section
Cell culture
Human lung adenocarcinoma A549 cell line was purchased from the Health Protection Agency Culture Collections (ECACC, Salisbury, UK) and cultured in F12K medium (HyClone, Loughborough, UK) supplemented with 10% heat-inactivated (sterilized) fetal bovine serum (FBS; Lonza, Basel, Switzerland), penicillin (10,000 U/mL), streptomycin (10,000 µg/mL), and amphotericin B (250 µg/mL). In case of normoxia, cells were incubated in conditions of 37°C and 5% CO2. Hypoxic cells were obtained by incubation of A549 cells in a hypoxic incubator in 1% O2 and 5% CO2, also in 37°C. In both conditions, cells were cultured for 24 h before treatment.
Morphological changes in cultured cells, launched by the influence of conditions mentioned above, were evaluated after 48 h using a phase-contrast microscope at 100-fold magnification (OPTA-TECH, Warsaw, Poland).
The statistical data are expressed as mean ± standard deviation (SD) using Student’s t-test. For each experiment, n = 3; *p < 0.01 is considered as statistically significant.
Assessment of apoptosis and flow cytometry analysis for normal cells
In T25 flasks, 5 × 105 of A549 cells were seeded for 24 h and then treated with benzimidazoles/reference compounds (100 µL) for 48 h. Samples were divided into normoxic and hypoxic based on their incubation conditions. After incubation, cells were washed using phosphate-buffered saline (PBS) in order to remove dead ones and then detached from flask surface by addition of the accutase enzyme. Detached cells were diluted in buffer (100 µL/1 × 105 cells) containing 4 µL of allophycocyanin (APC)-conjugated Annexin V + PI. Next, cells were incubated for 20 min at dark area and at room temperature. After that time, the flow cytometry analysis was carried out (FACS Canto II; Becton Dickinson, San Jose, CA, USA). The apoptosis and necrosis populations were assessed by PI versus Annexin V correlation. The A549 cells stained only with Annexin V indicated early apoptosis, while the Annexin V and PI double-stained cells indicated late apoptosis or necrosis.
Transfection of the nitroreductase gene using adenoviral vector
Determination of the multiplicity of infection coefficient
Multiplicity of infection (MOI) stands for the average amount of virus molecules able to infect every single cell. Too high values of this parameter can cause cytopathic effect (morphological changes connected to virus overexpression), too low values lead to decreased transfection efficiency.
In our study, the MOI parameter was calculated to estimate the number of virus molecules needed to infect 1 × 106 A549 human adenocarcinoma cells. In order to determine the optimal MOI value, transfected cells were seeded on 96-well plate (5000 cells per well). MOI values of 0, 100, 500, and 1000 were used. After 24-h incubation, medium was changed, and after next 24 h, cell growth was observed under the microscope. MOI value which gives optimal nfsB gene expression was determined as MOI 100.
PI test
PI is a specific stain used for necrotic cells or late-apoptotic cell detection, as it does not penetrate cell membrane of living cells. The stain binds to DNA molecules and excites in blue light. Dead cells are visible under the fluorescent microscope as red and orange dots. The test was performed after 72 h of incubation. Cells were centrifuged and density of 105 cells were suspended in 100 μL of PBS.To each sample, 0.4 µL of PI was added and cells were incubated for 20 min at room temperature. After that, samples were analyzed by cytometric analysis.
Assessment of apoptosis and flow cytometry analysis for virused cells
In T25 flasks, 5 × 105 of A549 previously virused cells were seeded for 24 h and then treated with benzimidazoles/reference compounds (100 µL) for 4 h. Samples were divided into normoxic and hypoxic based on their incubation conditions. After incubation, virused cells were washed using PBS in order to remove dead ones and then detached from flask surface by addition of the accutase enzyme. Detached cells were diluted in buffer (100 µL/1 × 105 cells) containing 4 µL of APC-conjugated Annexin V + PI. Next, cells were incubated for 20 min at dark and room temperature. After that, the flow cytometry analysis was carried out (FACS Canto II). The apoptosis and necrosis populations were assessed by PI versus Annexin V correlation. The A549 virused cells stained only with Annexin V indicated early apoptosis, while the Annexin V and PI double-stained cells indicated late apoptosis or necrosis.
Measurement of DNA content by flow cytometry analysis for both normal and virused A549 cells
Tested compounds (100 µL) were introduced to 5 × 105 of A549 normal cells and incubated for 24 h. Also, control sample (without addition of compounds) was prepared. Later, cells were detached by accutase, collected, washed with PBS, and fixed in 70% ethanol. Incubation with RNase (100 µg/mL; Sigma-Aldrich, St. Louis, MO, USA) and PI (50 µg/mL; Sigma-Aldrich, Munich, Germany) was held for 30 min (dark space at room temperature). Each sample containing cells was scanned by FACS Scan Canto II (Becton Dickinson) and the amount of DNA was determined.
Virused A549 cells (5 × 105 cells) were incubated with tested compounds (100 µL) under normoxic and hypoxic conditions for 4 h. Control sample (without addition of compounds) was prepared. Cells were detached by accutase enzyme, washed with PBS, and fixed in 70% ethanol. Before analysis, a buffer (100 µL/1 × 105 cells) was added containing Annexin and PI (4 µL/1 × 105 cells). After that, the cytometric analysis was carried out (FACS Canto II) for determination of DNA content.
RNA isolation and reverse transcription PCR analysis for nitroreductase gene expression
RNA was isolated from A549 cell line (1 × 106 cells) by Total RNA Prep Plus Minicolumn Kit (A&A Biotechnology, Gdynia, Poland) based on RNA isolation method developed earlier. 21 The isolated RNA has an A260/280 ratio of 1.6–1.8. Ultraviolet (UV) absorbance was used to determine the amount of RNA added to a complementary DNA (cDNA) reverse transcription (RT) reaction. PCRs are then set up using cDNA derived from the same amount of input RNA obtained during RT reaction. RT reaction was done using Enhanced Avian HS RT-PCR Kit (Sigma, Aldrich, Poznan, Poland) according to the manufacturer’s protocol. The cDNA was used immediately or stored at −20°C. PCR reaction mixture for PCR amplification consisted of a cDNA template, 0.5 µM of each primer, 10× AccuTaq Buffer, 0.5 U of AccuTaq LA DNA Polymerase Mix, 0.2 mM each 2′-deoxynucleoside 5′-triphosphate (dNTP), and water to a final volume of 20 µL. Negative control was included in each experiment (sample without a cDNA template). The primer sequences for the genes nfsB and reference ACTB were planned using software Primer 3. The primer sequences are presented in Table 5.
Primers sequences.

Amplification conditions for qualitative PCR: 35 cycles, initial denaturation at 94°C for 2 min, denaturation at 94°C for 15 s, primer annealing at 59°C for 45 s, extension at 72°C for 45 s, and final extension for 3 min at 72°C. The PCR products were analyzed with a 100 bp DNA ladder on 1.2% agarose gel (BioRad, Warszawa, Poland; for 1 h at a constant temperature of 90 C°) in Tris/Borate/EDTA (TBE) buffer. The gel was stained with ethidium bromide, visualized under UV light, and photographed using DNR Bio-Imaging System MiniBIS Pro (Israel).
Quantitative analysis by real-time PCR
Real-time PCR was performed on the corresponding cDNA synthesized from each sample using an MX3005P™ System. The nfsB gene and a reference ACTB were amplified in parallel for each sample in separate wells, during the same PCR run. ACTB was utilized as an internal positive control and as a normalizer for the correction of gene expression. For each PCR run, a reaction mixture was prepared consisting of 7.5 µL SYBR® Green JumpStart™ Taq ReadyMix™, 0.7 µL forward primer (final concentration 0.2 µM), 0.7 µL reverse primer, 4.1 µL nuclease-free water, and 2.0 µL template cDNA. The thermal cycling conditions comprised an initial denaturation step at 95°C for 2 min, 35 cycles at 94°C for 30 s, 59°C for 30 s, and 72°C for 30 s and a final extension step at 72°C for 3 min. After reaction, a melting curve was performed to confirm reaction specificity. Experiments for all samples were performed in triplicate.
Total RNA extraction and cDNA generation for BAX and BCL2 analysis
Total RNA was extracted from cells using RNeasy mini kits (QIAGEN) according to the manufacturer’s instructions. RNA content and purity were measured using PicoDrop spectrophotometer (PicoDrop Ltd.). The quality of RNA samples was analyzed by measuring the ratio of absorptions at 260/280 nm. The purified total RNA was immediately used for cDNA synthesis or stored at −80°C.
Generation of cDNA was performed with Maxima First-Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Warszawa, Poland) according to the protocol of the manufacturer; 500 ng of total RNA was used as starting material, and RT was performed in conditions optimized for use with this kit (25°C for 10 min, 50°C for 30 min, and 85°C for 5 min). The cDNA samples were kept frozen at −20°C.
Real-time PCR for BAX and BCL2 analysis
TaqMan gene expression experiments were performed in 10 µL reactions including 20 ng cDNA, 5 µL KAPA PROBE FAST qPCR Kit Master Mix ABI Prism (Kapa Biosystems; Sigma-Aldrich, Poznan, Poland), and 0.5 µL appropriate TaqMan Gene Expression Assay (20×). Specific Pre-made TaqMan assays (Thermo Scientific™) B-cell CLL/lymphoma 2 (BCL2, Hs00608023_m1), BCL2-associated X protein (BAX, Hs00180269_m1), and β-actin (ACTB, Hs01060665_g1) as the endogenous control were used in this study. TaqMan PCR assays were performed on a 7900HT Fast Real-Time PCR System (Applied Biosystems, Thermo Fisher Scientific, Warszawa, Poland) in FastGene Fast 96-well PCR plates (Nippon Genetics Europe GmbH, Dueren, Germany). The following thermal cycling specifications were performed: 20 s at 95°C and 40 cycles each for 3 s at 95°C and 30 s at 60°C. Expression values were calculated using Sequence Detection System 2.3 Software. Fold induction values (RQ) were calculated according to the equation 2−ΔΔCt, where ΔCt represents the differences in cycle threshold numbers between the target gene and β-actin, and ΔΔCt represents the relative change in these differences between examined and control cells (calibrator).
Western blot analysis of the expression of BAX and BCL2 proteins
The cells were lysed with M-PER™ Mammalian Protein Extraction Reagent and Health™ Protease Inhibitor Cocktail (Thermo Scientific™) on ice. The protein content of lysates was determined by the bicinchoninic acid (BCA) method. Cell lysate aliquots containing appropriate equal total protein were boiled with 5× concentrated sample buffer, separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), and then electrophoretically transferred to polyvinylidene difluoride (PVDF) membrane (BioRad). The membranes were blocked in 5% non-fat milk (BioRad) for 2 h at RT, washed with Tris-buffered saline (TBS)/0.1% Tween 20, and incubated overnight at 4°C with the following primary antibodies: anti-BAX or anti-BCL2 (Cell Signaling Technology) polyclonal antibodies. After washing and incubation with goat anti-rabbit horseradish peroxidase (HRP)-secondary antibody (Santa Cruz Biotechnology, Lodz, Poland), immunodetection was performed using an enhanced chemiluminescence kit (BioRad). Membranes were scanned on a Chemidoc (BioRad) imaging and gel documentation system.
Conclusion
The most common globally diagnosed cancers are the lung cancer in men and breast cancer in women. 3 Cancer treatment generally includes surgery, non-selective chemotherapy, radiotherapy, and hormonal therapy. While the last listed method is strictly based on the environmental change in tumor tissue, the cytostatic chemotherapy agents or cytotoxic drugs generally do not possess the selective antitumor activity. Most of them affect cell proliferation and can damage normal tissue as well. Systemic toxicity is one of the major shortcomings of therapies listed above.5,22 Effective anticancer therapy should consider knowledge of the effect of the compounds on the cell cycle. One of the new strategies of anticancer therapy includes application of bioreductive drugs with tirapazamine as major representative. The bioreductive therapy uses hypoxia as the target for main differences between normal and cancer cells. The distinguishing characteristic feature of these compounds from the group of cytotoxic substances is their selectivity for hypoxic cells which in turn provides an opportunity to reduce the systemic toxic effects.23,24
This study provided the evaluation of the cytotoxicity of new benzimidazole derivatives using virus-directed enzyme prodrug therapy (VDEPT). Our goal was to determine the influence of tested compounds on the cell death pathway differentiation, to assess the level of nfsB gene expression after introducing to human adenocarcinoma A549 cell line through adenoviral vector, and to provide the analysis of the level of BAX/BCL2 proteins synthesis. The highest expression level of nfsB gene among all tested samples was observed for N-oxide compound 3, while the lowest for compound 1. It is necessary to conduct further studies to determine the effect of nfsB gene on IC50 value reduction in investigated benzimidazole derivatives samples.
Flow cytometry analysis with Annexin V + PI assay was carried out in order to evaluate different pathways of cell death. Moreover, the division into early- or late-apoptotic cells and necrotic cells was specified as a function of the pro-apoptotic properties of tested benzimidazole derivatives.
All tests were done using virus transfected cells and non-virus transfected cells A549 cells treated with tested compounds under normoxic and hypoxic conditions. Tirapazamine and NI were used as reference compounds which are proved to be highly selective for nitroreductase enzyme, especially in hypoxia.
The performed tests showed pro-apoptotic properties of the tested compounds related to their ability to induce early apoptosis in both normoxia and hypoxia conditions and elimination of undesirable necrosis. Especially, their strong pro-apoptotic activity against virused A549 cells was indicated. In our research, viral vector played a role of a nitroreductase gene carrier. This vector was introduced into the cell line in order to increase the level of reduction of tested compounds into their bioactive forms. This led to increased pro-apoptotic properties in virused cells. Conducted research proved the stability of these properties and clearly indicated the structure of the N-oxide as the factor ensuring the selectivity in hypoxia. Regarding the reference compound, the level of activity of all tested compounds is high, which demonstrates the accuracy of the selection of the studied structure with the characteristic substituents of the benzimidazole ring.
The study of benzimidazole biological activity should be continued especially as an aspect of broadening cell line types. This would allow evaluating their features against different types of tumors.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by the Medical University of Lodz, Poland (503/3-021-01/503-31-004).
