Abstract
Acid-sensing ion channels, a proton-gated cation channel, can be activated by low extracellular pH and involved in pathogenesis of some tumors such as glioma and breast cancer. However, the role of acid-sensing ion channels in the growth of lung cancer cell is unclear. In this study, we investigated the expression of acid-sensing ion channels in human lung cancer cell line A549 and their possible role in proliferation and migration of A549 cells. The results show that acid-sensing ion channel 1, acid-sensing ion channel 2, and acid-sensing ion channel 3 are expressed in A549 cells at the messenger RNA and protein levels, and acid-sensing ion channel–like currents were elicited by extracellular acid stimuli. Moreover, we found that acidic extracellular medium or overexpressing acid-sensing ion channel 1a promotes proliferation and migration of A549 cells. In addition psalmotoxin 1, a specific acid-sensing ion channel 1a inhibitor, or acid-sensing ion channel 1a knockdown can abolish the effect of acid stimuli on A549 cells. In addition, acid-sensing ion channels mediate increase of [Ca2+]i induced by low extracellular pH in A549 cells. All these results indicate that acid-sensing ion channel–calcium signal mediate lung cancer cell proliferation and migration induced by extracellular acidosis, and acid-sensing ion channels may serve as a prognostic marker and a therapeutic target for lung cancer.
Introduction
Lung cancer, a common malignant solid tumor, is the leading cause of cancer-related death worldwide, which accounts for 18% of the total number of deaths.1,2 Moreover, there is a rapid increase in morbidity and mortality of lung cancer in recent years. 3 Non–small cell lung cancer (NSCLC), the most common subtype of lung cancer, accounts for about 85% of lung cancer cases. There are various treatment methods for NSCLC such as surgery, chemotherapy, radiotherapy, and genetic therapy, but the 5-year survival rate of NSCLC patients is less than 5% because of complicated pathogenesis of lung cancer. 4
The tumor microenvironment (TME), characterized by chronic inflammation, immune suppression, hypoxia, and metabolic and biochemical alterations, is a milieu that influences the survival, growth, and evolution of tumor cells.5–7 Increasing evidences have shown that TME plays a key role in tumor progression and metastasis.8–11 In addition, metabolic acidosis is considered as a common phenomenon of tumor microenvironment and affects the phenotype of tumor cells. 12 Previous studies have shown that acidosis can degrade extracellular matrix and promote tumor metastasis.13–16 Acidic microenvironment in tumor biology is becoming more and more important for clinical treatment.
Acid-sensing ion channels (ASICs) are cation channels which belong to the degenerin/epithelial Na+ channel (DEG/ENaC) superfamily and can be activated by extracellular protons. 17 To date, at least eight ASIC subunit proteins, encoded by five genes, have been identified including ASIC1a, ASIC1b1, ASIC1b2, ASIC2a, ASIC2b, ASIC3, ASIC4, and ASIC5.18,19 Except for ASIC5, they are widely expressed in central and peripheral nervous system and involved in various physiological and pathological processes, such as brain ischemia, learning and memory, mechanosensation, nociception, and pain.19–21 In addition, ASICs are also present in non-neuronal tissues such as testis, bone, dendritic cells, vascular smooth muscles, and endothelial cells.22–24 Many reports have shown that ASICs play an important role in acidosis-associated physiological and pathological functions.23,25–29 Recent studies have shown that ASICs are expressed in glioma cells and mammary carcinoma and are involved in tumor pathogenesis.30,31 However, it remains unclear whether ASICs are present in lung cancer cells and affect cell growth.
In this study, we detected the expression of ASICs in human lung adenocarcinoma A549 cells, a NSCLC cell line, and investigated the role of ASICs in A549 cell proliferation and migration induced by extracellular acidosis. Our results indicate that ASICs are expressed in A549 cells and are involved in acidosis-induced cell proliferation and migration.
Materials and methods
Reagents and antibodies
Dulbecco’s Modified Eagle’s Medium (DMEM) and fetal bovine serum were obtained from Gibco (Invitrogen, Carlsbad, CA, USA). Hoechst 33258, trypsin, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), and amiloride were purchased from Sigma-Aldrich (St. Louis, MO, USA). Psalmotoxin 1(PcTX1) and primary antibodies of ASIC1, ASIC2, and ASIC3 were obtained from Alomone Labs (Jerusalem, Israel). β-actin antibody and horseradish peroxidase–conjugated secondary antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). pCMV6-ASIC1a plasmid was purchased from Bioyong Technologies Inc. (Beijing, China). ASIC1 small interfering RNA (siRNA; ID No. 113602) was obtained from Ambion (Austin, TX, USA). Other general agents were commercially available.
Cell culture and transfection
Lung cancer A549 cells were obtained from Chinese Type Culture Collection and were maintained in DMEM medium containing 10% fetal bovine serum, 100 U/mL penicillin, and 100 mg/mL streptomycin in a humidified atmosphere of 95% air and 5% CO2 at 37°C. Then, the cells were seeded onto 96- or 6-well plates at 5 × 104 cells per well and used during the logarithmic phase of growth. The medium was adjusted to acidic pH (pH 7.0, pH 6.5, and pH 6.0) with HCl. To suspended medium at pH 7.0, 6.5, or 6.0, cell cultures were maintained at 37°C in a humidified atmosphere with 6%, 7%, and 8% CO2, respectively. Overexpressing ASIC1a was constructed as described previously; 32 briefly, the A549 cells were transfected with plasmid pCMV6-ASIC1a using the liposome transfection reagent (Lipofectamine 2000, Invitrogen, USA), which contains rat ASIC1a complementary DNA (cDNA). siRNA molecules were transfected according to the manufacturer’s instructions. Following 4 h incubation, cultures were supplemented with growth media for 72 h before study. Control siRNA was used as a negative control.
Reverse transcription polymerase chain reaction
Total RNA was isolated from cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. cDNA synthesis was performed using a PrimeScript 1st Strand cDNA Synthesis Kit (TaKaRa, Tokyo, Japan). Polymerase chain reaction (PCR) amplifications of cDNA were performed by standard methods. The following specific primers were used: ASIC1—forward: 5′-TGCTCTCCTGCCACTTCC-3′ and reverse: 5′-GCT TTGCTGGGGATCTTG-3′&0x44; ASIC2—forward: 5′-TGCAAGTTCAAAGGGCAG-3′ and reverse: 5′-TGGCTGATGTCTTGCTGG-3′, ASIC3—forward: 5′-AGTGGCCA CCTTCC TCTA-3′ and reverse: 5′-CAGTCCAGCAGCATGTCATC-3′, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH)—forward: 5′-GGTGAAGGTCGGTGTGAACG-3′ and reverse: 5′-TTGGCTCCA CCCTTC AAGTG-3′. PCR products were analyzed by 2% agarose gel electrophoresis.
Western blotting
Cells were lysed on ice in extraction buffer containing 50 mM Tris-base (pH 7.4), 100 mM NaCl, 1% NP-40, 10 mM ethylenediaminetetraacetic acid (EDTA), 20 mM NaF, 1 mM phenylmethylsulfonyl fluoride (PMSF), 3 mM Na3VO4, and protease inhibitors. Then, the cells were centrifuged at 12,000g at 4°C for 15 min. Supernatant was separated and protein concentration was determined using the BCA Protein Assay Kit (Pierce Biotechnology, Rockford, IL, USA). Protein samples (30 µg) were separated by 10% sodium dodecyl sulfate (SDS)-polyacrylamide gel and then transferred onto a polyvinylidene fluoride membrane (Millipore, Billerica, MA, USA). After blocking with 5% non-fat milk in Tris-buffered saline containing 0.1% Tween-20 (TBST) for 1 h at room temperature, membranes were incubated overnight at 4°C with different primary antibodies (anti-ASIC1, anti-ASIC2, and anti-ASIC3; 1:100 dilution), followed by incubation with horseradish peroxidase–conjugated secondary antibodies (1:10 000) in TBST with 1% non-fat milk for 1 h at room temperature. Then, the membranes were reacted with enhanced chemiluminescence reagents (Amersham Pharmacia Biotech, Inc., Piscataway, NJ, USA) for 5 min and visualized using chemiluminescence detection system (Bioshine, Shanghai, China).
Immunofluorescence
A549 cells were fixed with 4% paraformaldehyde for 30 min. After being rinsed with phosphate-buffered saline (PBS), cells were permeabilized with 0.3% Triton X-100 for 30 min and blocked with 3% bovine serum albumin (BSA)–PBS for 30 min. Then, the cells were incubated with 1:50 anti-ASIC antibodies overnight at 4°C, followed by incubation with 1:100 fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit IgG for 1 h at room temperature. After being washed with PBS, cells were incubated with Hoechst 33258 for 15 min. Finally, the cells were mounted with coverslips and visualized with confocal microscopy.
Whole-cell patch-clamp recording
The whole-cell patch-clamp recording was performed as described in our previous reports.23,32 The current signals were acquired at a sampling rate of 10 kHz using an EPC-10 amplifier (HEKA, Lambrecht, Germany) and Pulse/PulseFit software (HEKA, Southboro, Germany). All the recordings were made at room temperature (20–22°C).
Calcium imaging
Digital calcium imaging was performed as described previously. 32 The cells were loaded with 1 mM Fura-2/AM in artificial cerebrospinal fluid (ACSF) for 30 min at 37°C. Fluorescence was excited at wavelengths of 340 nm for 150 ms and 380 nm for 50 ms at 1 s interval by calcium imaging system (PTI, Birmingham, NJ, USA). F340/F380 fluorescence ratio was recorded and analyzed by MetaFluor version 6.3 software (Molecular Devices, San Francisco, CA, USA).
Cell viability assay
After various treatments, cells were incubated with MTT solution (5 mg/mL) for 4 h at 37°C. Then, the medium was discarded and dimethyl sulphoxide (DMSO) was added to plate, followed by shaking for 15 min. Absorbance at 492 nm was measured with a microplate reader (ELx800, BioTek, Winooski, VT, USA). Cell viability was expressed as a percentage of the value in control group which was defined as 100%.
Wound-healing assay
A549 cells were seeded on six-well plates. After formation of a confluent monolayer, the cells were scratched to form a 1-mm-wide wound using a sterile pipette tip, followed by washing the cells with PBS to remove debris. Then, the cells were treated with or without acidic extracellular medium for 24 h. Microscopic photographs were taken at 0 and 24 h.
Statistical analysis
Data from experiments were analyzed with the statistical program SPSS (SPSS, Chicago, IL, USA). Comparison between two groups was evaluated by a two-sided and unpaired Student’s t test. Data are expressed as mean ± standard error of the mean (SEM). p < 0.05 was considered to be statistically significant.
Results
ASICs are expressed in A549 cells
To determine whether ASICs are expressed in A549 cells, we first investigated the expression of ASICs at messenger RNA (mRNA) level. Reverse transcription PCR (RT-PCR) results have shown that mRNAs of ASIC1, ASIC2, and ASIC3 were detected in A549 cells, and their PCR products were of 649, 642, and 335 bp, respectively (Figure 1(a)). Then, we investigated the expression of ASICs at protein level. Western blotting results have also shown that ASIC1, ASIC2, and ASIC3 proteins were detected in A549 cells. The ASIC proteins cannot be detected after pretreatment with the corresponding antigen (Ag) peptide (Figure 1(b)). Furthermore, we observed cellular distributions of ASICs in A549 cells using immunofluorescence. As shown in Figure 2, ASIC1 and ASIC3 proteins were present in the nucleus, cytoplasm, and plasma membrane. In addition, ASIC2 is predominantly expressed in the nucleus. Similarly, ASIC proteins cannot be detected after pretreatment with the corresponding Ag peptide. All these results suggest that ASICs are expressed in A549 cells.

Expression of ASICs in A549 cells. (a) ASIC 1, ASIC2, and ASIC3 transcripts were detected in A549 cells by RT-PCR. (b) Representative immunoreactive bands showing ASIC1, ASIC2, and ASIC3 proteins are expressed in A549 cells. ASIC proteins cannot be detected after pretreatment with the corresponding Ag peptide. Cortex was used as positive control.

Localization of ASICs in A549 cells. Detection of the localization of ASICs in A549 cells by double-staining immunofluorescence (original magnification: 400×). Nuclei were counterstained with Hoechst33258 (blue). ASIC1, ASIC2, and ASIC3 were labeled with FITC (green). ASIC1 and ASIC3 were found in the nucleus, cytoplasm, and plasma membrane, and ASIC2 was located in the cell nucleus. ASIC proteins cannot be detected after pretreatment with the corresponding Ag peptide.
Electrophysiological characteristics of ASIC-like currents in A549 cells
To further determine whether ASIC proteins have a function in A549 cells, the currents were observed by whole-cell patch-clamp recording. Cells were held at −80 mV and then extracellular pH was rapidly reduced from 7.4 to 6.0. In 83% (n = 24/29) of the recorded cells, low extracellular pH elicited inward ASIC-like currents. In addition, amiloride (100 µM), a blocker of ASICs, significantly inhibited the ASIC-like currents in A549 cells. After washing, ASIC currents can return to the control level (n = 6; Figure 3(a) and (b)). Similarly, the ASIC-like currents in A549 cells were also reversibly inhibited by PcTX1 (50 nM), a specific ASIC1a inhibitor (n = 6; Figure 3(c) and (d)), indicating that ASIC-like currents in A549 cells are mainly ASIC1 currents. All these data indicate that ASICs are functionally expressed in A549 cells.

Electrophysiological characteristics of ASIC-like currents in A549 cells. (a) Representative traces and (b) statistical results showing pH 6.0 elicited ASIC-like currents in A549 cells and the currents were reversibly inhibited by amiloride (100 µM). (c) Representative traces and (d) statistical results showing ASIC-like currents elicited by acid in A549 cells were also reversibly inhibited by PcTX1 (50 nM). Data are expressed as mean ± SEM (n = 6; ##p < 0.01 vs control and **p < 0.01 vs amiloride or PcTX1).
Extracellular acidosis promotes proliferation and migration of A549 cells
Previous studies have shown that acidosis promotes proliferation or migration in some tumor cells such as glioma cells, mammary carcinoma, and AT-1 cells.12,30,31 To determine the effect of extracellular acidosis on the proliferation of A549 cells, cells were pretreated with extracellular medium at different pH (7.0, 6.5, and 6.0) for 24 h, and then, cell viability was measured by MTT assay. As shown in Figure 4(a), pH 7.0 has no influence on cell viability. However, pH 6.5 and pH 6.0 significantly increases cell viability, indicating acidic extracellular medium promotes the proliferation of A549 cells in a pH dependent manner. Then, we further investigated the effect of extracellular acidosis on the migration of A549 cells by wound-healing assay. Similarly, acidic extracellular medium also significantly stimulates A549 cell migration in a pH dependent manner (Figure 4(b)). These data suggested that extracellular acidosis promotes the proliferation and migration of A549 cells.
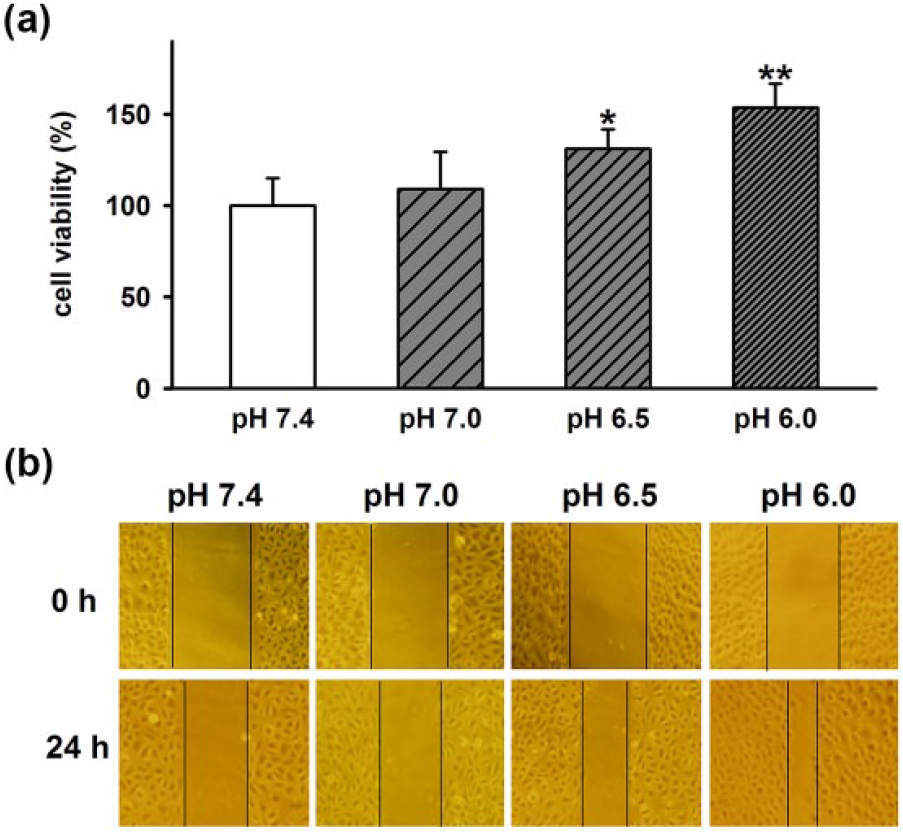
Effect of extracellular acidosis on A549 cell proliferation and migration. (a) Statistical results showing acidic extracellular medium (pH 6.0) promoted A549 cell proliferation in a pH dependent manner. (b) Representative micrographs (original magnification: 100×) showing acidic extracellular medium (pH 6.0) stimulated A549 cell migration in a pH dependent manner. Data are expressed as mean ± SEM (n = 6; *p < 0.05 and **p < 0.01 vs pH 7.4).
ASICs are involved in A549 cell proliferation and migration induced by extracellular acidosis
The role of ASICs in A549 cell proliferation and migration was determined. As shown in Figure 5(a) and (b), overexpressing ASIC1a does not influence A549 cell proliferation and migration in neutral pH, but it significantly promotes A549 cell proliferation and migration under acidic stimuli (pH 6.0). Moreover, specific ASIC1a inhibitor PcTX1 and control siRNA have no influence on A549 cell proliferation and migration at neutral pH. However, PcTX1 (50 nM) and ASIC1a siRNA significantly prevented A549 cell proliferation and migration induced by acidic extracellular medium (pH 6.0; Figure 5(c)–(f)), indicating that ASICs mediated A549 cell proliferation and migration induced by extracellular acidosis.

ASICs involved in A549 cell proliferation and migration induced by acidosis. (a, c, and e) Statistical results showing the effects of overexpressing ASIC1a, PcTX1 (50 nM), and ASIC1a siRNA on A549 cell proliferation under acidic environment. (b, d, and f) Representative micrographs (original magnification: 100×) showing the effects of overexpressing ASIC1a, PcTX1 (50 nM), and ASIC1a siRNA on A549 cell migration induced by low extracellular pH. Data are expressed as mean ± SEM (n = 6; #p < 0.05 vs control or pH 7.4 and *p < 0.05 vs pH 6.0).
ASICs mediate increase of [Ca2+]i induced by extracellular acidosis in A549 cells
Calcium is considered as a crucial mediator of cancer cell migration and invasion. To further determine molecular mechanism of acidosis promoting A549 cell proliferation and migration. We investigated the effect of extracellular acidosis on [Ca2+]i in A549 cells. As shown in Figure 6, low extracellular pH significantly increases [Ca2+]i in A549 cells. This effect was blocked by specific ASIC1a inhibitor PcTX1, suggesting that ASICs mediated the increase of [Ca2+]i induced by extracellular acidosis. These indicates that ASIC–calcium signal contribute to the effect of extracellular acidosis on A549 cell proliferation and migration.

Extracellular acidosis increases [Ca2+]i by ASICs in A549 cells (a) Representative traces and (b) statistical results showing PcTX1 (50 nM), specific ASIC1a inhibitor, inhibited increase of [Ca2+]i induced by extracellular acidosis in A549 cells. Data are expressed as mean ± SEM (n = 6, ##p < 0.01 vs pH 7.4 and *p < 0.05 vs pH 6.0).
Discussion
In this study, we demonstrated that ASICs are functionally expressed in A549 cells and contribute to A549 cell proliferation and migration induced by extracellular acidosis.
Different from normal tissue environment, the TME is normally acidic, especially in solid tumors.5,31 Because solid tumors grow rapidly, followed by poor supply of oxygen and nutrients; to maintain sufficient supply of oxygen and nutrients, tumor cells need to change metabolic pathway, for instance, switching energy metabolism toward glycolysis.33,34 This causes substantial lactic acid and carbon dioxide accumulation and lead to pronounced acidic extracellular pH (pHe) of the tumor, which is approximately between 5.8 and 7.4. 35 In addition, tumor-derived protons, crucial transporters, such as the Na+/H+ exchanger 1 (NHE-1), vacuolar–adenosine triphosphatase (ATPase) and carbonic anhydrases, also contribute to the formation of the acidic TME.12,36 The effect of acidity on cell growth and proliferation is markedly different between normal cell and tumor cell, which is usually harmful to normal cell and beneficial to tumor cell. 31 It has been shown that extracellular acidosis affects proliferation or migration of different tumor cells such as mammary carcinoma (MCF-7 and LM-4142 cells), glioma cells, and AT-1cells.12,30,31 More evidences have also demonstrated that the acidic pHe offers an advantage for tumor progression and metastasis.37,38 In this study, our results showed that extracellular acidosis promotes lung cancer A549 cell proliferation in a pH dependent manner. Furthermore, acidic extracellular medium also stimulates A549 cell migration (Figure 4), indicating the acidic microenvironment may be a critical factor for growth of lung cancer cells.
Acidic microenvironment is important for tumor progression and metastasis, but the underlying mechanism is poorly understood. Ion channels were considered to play an important role in tumor pathogenesis. 12 Previous studies have shown that several potassium channels, for instance, KV1.3, KV11.1, and TRPM8, are involved in tumor cell migration by activating intracellular signaling cascades. 39 In addition, calcium can induce reactive oxygen species (ROS) production and is known to be crucial for cancer cell migration and invasion. 31 Besides K+ and Ca2+ channels, Na+ channels have also been demonstrated to be involved in tumor cell migration. 40 ASICs are H+-gated cation channels and activated by extracellular acid. ASIC1 is Na+ and Ca2+ permeable, whereas other types of ASICs are only permeable to Na+. 21 Recent studies have shown that ASICs are functionally expressed in mammary carcinoma. Acidic tumor microenvironment activates ASICs and mediates ROS generation, which contributes to the pathogenesis of breast cancer. 31 Similar to breast cancer cells, lung cancer cells are also epithelia-derived carcinoma. Moreover, acidic extracellular medium, which can activate ASICs, promote proliferation and migration of A549 cells. So, we proposed that ASICs may be implicated in A549 cell proliferation and migration induced by extracellular acidosis.
To test this hypothesis, we investigated the expression of ASICs in A549 cells. As expected, ASIC1, ASIC2, and ASIC3 are expressed at the mRNA and protein levels in A549 cells (Figure 1) and human NSCLC tissue (Supplementary Figure 1). Then, we examined ASICs localization in A549 cells by immunofluorescence. The results showed ASIC1 and ASIC3 are located in the nucleus, cytoplasm, and plasma membrane, whereas ASIC2 is found mainly in the nucleus (Figure 2). Furthermore, acidic pH does not influence immunolocalization of ASICs (Supplementary Figure 2). We further examined the function of ASICs in A549 cells. We found that low extracellular pH elicits inward ASIC-like currents in A549 cells, which were reversibly inhibited by ASICs blocker amiloride and specific ASIC1a inhibitor PcTX1 (Figure 3), suggesting that A549 cells expressed functional ASICs and ASIC-like currents are mainly ASIC1 currents. To further determine the role of ASICs on A549 cell proliferation and migration under extracellular acid stimuli, we upregulated the ASICs by overexpressing ASIC1a and blocked ASICs by PcTX1 and ASIC1a knockdown. The results showed that overexpressing ASIC1a significantly promoted A549 cell proliferation and migration induced by extracellular acidosis, while acidosis-induced A549 cell proliferation and migration were prevented by PcTX1 and ASIC1a knockdown (Figure 5). We also found that acidosis induced increase of [Ca2+]i was inhibited by PcTX1 (Figure 6). These indicate that ASIC–calcium signal contribute to the effect of acidic microenvironment on lung cancer cell proliferation and migration
Conclusion
This study demonstrated that ASICs are functionally expressed in human lung cancer A549 cells. Extracellular acidosis can activate ASICs and increase [Ca2+]i, which contribute to the effect of acidic microenvironment on A549 cell proliferation and migration. Our study indicates that ASICs may serve as a prognostic marker and a new therapeutic target for lung cancer. However, further efforts will be made to clarify the precise mechanism that ASICs involve in lung cancer pathogenesis in response to acidic microenvironment.
Footnotes
Acknowledgements
Y.W., B.G., and Q.-J.X. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (NSFC; No. 81671327 and 81201020), the Doctor Foundation of Anhui Medical University (No. XJ201103) to W.-N.W., and the NSFC (No. 81300963) to Q.-J. X.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
