Abstract
Neoadjuvant chemotherapy aims to improve the outcome of breast cancer patients, but only few would benefit from this treatment. Pathological complete response has been proposed as a surrogate marker for the prediction of long-term clinical benefits; however, 50%–85% patients have an unfavorable pathological complete response to chemotherapy. MicroRNAs are known biomarkers of breast cancer progression; nevertheless, their potential to identify patients with pathological complete response remains poorly understood. Here, we investigated whether a microRNA profile could be associated with pathological complete response in triple-negative breast cancer patients receiving 5-fluorouracil, adriamycin, cyclophosphamide–cisplatin/paclitaxel as a novel neoadjuvant chemotherapy. In the discovery cohort, the expression of 754 microRNAs was examined in tumors from 10 triple-negative breast cancer patients who achieved pathological complete response and 8 without pathological complete response using TaqMan Low-Density Arrays. Unsupervised hierarchical cluster analysis identified 11 microRNAs with significant differences between responder and no-responder patients (fold change ≥ 1.5; p < 0.05). The differential expression of miR-30a, miR-9-3p, miR-770, and miR-143-5p was validated in an independent group of 17 patients with or without pathological complete response. Moreover, Kaplan–Meier analysis showed that expression of these four microRNAs was associated with an increased disease-free survival. Gene ontology classification of predicted microRNA targets indicated that numerous genes are involved in pathways related to chemoresistance, such as vascular endothelial growth factor, focal adhesion kinase, WNT, ERbB, phosphoinositide 3-kinase, and AKT signaling. In summary, we identified a novel microRNA expression signature associated with pathological complete response in breast cancer. We propose that the four validated microRNAs could be used as molecular biomarkers of clinical response in triple-negative breast cancer patients with pathological complete response to neoadjuvant therapy.
Keywords
Introduction
Breast cancer is the leading cause of cancer deaths among females worldwide. 1 Triple-negative breast cancer (TNBC) is a heterogeneous disease that occurs in 12%–15% of cases and is associated with poor prognosis. Neoadjuvant chemotherapy has been increasingly used as a first-line treatment for operable breast cancer being useful strategy for patient’s management. 2 Recent studies have demonstrated a marked sensitivity of TNBC to neoadjuvant chemotherapy.3,4 Pathological complete response (pCR) which is defined by the absence of residual disease in breast and lymphatic nodes after cytotoxic treatment is the criteria used to discriminate between patients with favorable and unfavorable outcomes to neoadjuvant chemotherapy. A number of studies, mainly in locally advanced TNBC, indicated that patients who benefit better from therapy are those who achieve a pCR which is associated with improved disease-free survival (DFS) and overall survival (OS) compared to those with pathological evidence of residual cancer.5–8 Remarkably, pCR rates of 10%–50% have been reported in TNBC patients having a low risk of local or distant recurrence and an improved survival in the first 5 years.4,9,10 Patients who fail to achieve pCR have adverse prognosis in terms of DFS and OS. 11 Remarkably, a large study (CTNeoBC) in 12,000 women with primary breast cancer established a definitive association between pCR and both the DFS and OS events. 12 However, no specific tumor biomarkers that can predict the clinical response and outcome of patients with TNBC have been clearly defined yet.
MicroRNAs (miRNAs) are small non-coding RNAs of 25 nucleotides length that function as negative regulators of gene expression by either inhibiting translation or inducing deadenylation-dependent degradation of target messenger RNAs (mRNAs). 13 Deregulated expression of miRNAs has been involved in the development and progression of breast cancer, and they have also been recently investigated as potential predictors of clinical response to therapy.14,15 Several studies have investigated the role of miRNAs in tumor chemoresistance. For instance, overexpression of miR-200b-3p and miR-190a was associated with pCR and breast conservative surgery in TNBC patients. 11 Moreover, high level of miR-621 predicted a better response to paclitaxel/carboplatin neoadjuvant therapy in breast cancer patients who achieved pCR. 16 In addition, recent evidences suggested that changes in miRNA expression can affect the sensitivity of breast cancer cell lines to chemotherapeutic agents.17,18 However, the relevance of miRNAs as predictors of clinical response of patients diagnosed with breast cancer has not been completely addressed. Here, we report a novel set of deregulated miRNAs that can be used as potential markers to predict clinical outcomes in TNBC patients receiving 5-fluorouracil, adriamycin, cyclophosphamide–cisplatin/paclitaxel (FAC–CDDP/PTX) neoadjuvant chemotherapy. Importantly, we describe for the first time the deregulation of miR-30a, miR-9-3p, miR-770, and miR-143-5p in response to neoadjuvant therapy in breast cancer and their impact on biological pathways related to chemoresistance, such as vascular endothelial growth factor (VEGF), focal adhesion kinase (FAK), WNT, ERbB, phosphoinositide 3-kinase (PI3K), and AKT signaling pathways.
Methods
Tissues samples
Tumors from TNBC patients with or without pCR were collected at the National Institute of Cancerology, Mexico. Biopsies were obtained after selection following the regulations approved by the Ethics Committee including patients’ informed consent. All recruited patients received neoadjuvant chemotherapy based on FAC–CDDP/PTX regimen. After tumor resection, specimens were formalin-fixed and paraffin-embedded (FFPE). Pathologists confirmed the triple-negative status of tumors by immunohistochemistry using specific antibodies against estrogen, progesterone, and HER2 receptors in FFPE clinical specimens. Only tissues containing at least 80% of tumor cells were used for downstream analysis.
RNA total isolation
Total RNA was isolated using the RNeasy FFPE Kit (QIAGEN, Valencia, CA) following the manufacturer’s protocol. Briefly, 5–10 sections of 10 µm were incubated twice in xylene for 1 h at 63°C for deparaffinization. Total RNA was extracted using TRIzol protocol. RNA concentration and purity were evaluated by spectrophotometry (NanoDrop Technologies, Wilmington, DE), and RNA integrity was analyzed through 1% agarose gel electrophoresis.
MiRNA expression analysis
The Megaplex TaqMan Low-Density Array (TLDA) v 3.0 (Applied Biosystems, Foster City, CA) platform was used to measure the expression of 754 human specific miRNAs in parallel. Briefly, 600 ng of total RNA was retrotranscribed using stem-loop primers to obtain complementary DNA (cDNA) templates. Subsequently, a pre-amplification step of 12 cycles was included to increase the concentration of low-level miRNAs. The pre-amplified products were loaded into the TLDA and reactions were started using the 7900 FAST real-time thermal cycler (ABI). RNU44 and RNU48 expression was used as internal control.
Statistical analysis of miRNA expression
MiRNA levels were measured by quantitative reverse transcription polymerase chain reaction (qRT-PCR) in TLDA using the comparative Ct (2ΔΔCt) method. All analyses were done using R (HTqPCR and gplots-bioconductor). The Ct raw data were determined using an automatic baseline and a threshold of 0.2. A fold change (FC) (log2 RQ) value >1.5 was used to define the differentially expressed miRNAs. An adjusted t test was used to evaluate the significant differences in Ct values between groups. To identify subgroups defined by miRNA expression profiles, an unsupervised clustering analysis using Spearman correlation and average linkage was used.
Reverse transcription and real-time PCR
qRT-PCR analysis of individual miRNA was performed using MicroRNA Assays (Thermo Fisher Scientific, Waltham, Massachusetts, USA). Briefly, 10 ng of total RNA was reverse transcribed using a specific looped RT primer, 0.15 µL of deoxynucleotide triphosphates (dNTPs; 100 mM), 1.0 µL of Reverse Transcriptase MultiScribe (50 U/µL), 1.5 µL of 10× buffer, 0.19 µL of RNase inhibitor (20 U/µL), and 4.16 µL of RNase-free water. Then, retrotranscription reaction (1:15 dilution) was mixed with 10 µL of master mix TaqMan Universal PCR Master Mix, No AmpErase UNG 2×, 7.67 µL of RNase-free water, and 1.0 µL of PCR probe. PCR reaction was performed in a GeneAmp System 9700 (Applied Biosystems) as follows: 95°C for 10 min and 40 cycles at 95°C for 15 s and 60°C for 1 min. Tests were normalized using RNU44 as an internal control.
Bioinformatics analysis
MiRNA targets were identified using TargetScan 7.0 (http://www.targetscan.org/vert_71/), miRWalk 2.0 (http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/), and PicTar (http://www.pictar.org/) softwares. Only target genes that were predicted by the three algorithms were selected for further analysis. Gene ontology and enrichment cellular pathway analyses were performed using David and miRPath v.3 tools, respectively.
Statistical analysis
Experiments were performed three times by triplicate and results were represented as mean ± standard deviation (SD). One-way analysis of variance (ANOVA) followed by Tukey’s test were used to compare the differences between the mean values. A p < 0.05 was considered as statistically significant.
Results
Clinicopathological features of patients
Tumor biopsies were collected from a small cohort of 18 patients diagnosed with TNBC. Patients enrolled in this study have all signed informed consent. Tissue samples were histologically analyzed to confirm at least 80% of tumor cells and then processed for RNA isolation. An overview of the clinical and pathological features of tumors and patients included in the study is given in Table 1. Based on the clinical data, patients of this discovery cohort were grouped as pCR (n = 10), and no-pCR (n = 8). Patients were aged between 28 and 65 years old; mean age was 46.4 years in the pCR group and 53.3 in the no-pCR group. The majority of patients were diagnosed at stages IIIA (50%), IIIB (22.2%), IIIC (11.1%), IV (5.5%), IIA (5.5%), and IIB (5.5%). Histopathologically, breast tumors were classified as ductal (83.3%), ductal/metaplastic (5.5%), metaplastic (5.5%), and lobular (5.5%) subtypes. After FAC–CDDP/PTX neoadjuvant chemotherapy, all patients of the no-pCR group had residual breast tumor or recurrence into lymph nodes, 5 brain (n = 1), bone (n = 1), and skin (n = 1). The expression of the proliferation marker Ki67 was greater in pCR (64%) in comparison to no-responder patients (51%), suggesting that highly proliferating tumor cells are more sensitive to the cytotoxic treatment.
Clinical and pathological characteristics of patients.

pCR: pathological complete response.
Profiling of miRNAs in TNBC patients with pCR
In order to identify miRNAs that are differentially expressed between primary breast tumors from patients who achieved pCR in comparison to no-responders, we profiled 754 mature miRNAs using stem-loop qRT-PCR in TLDA. Breast tumors were collected from a small cohort of patients diagnosed with TNBC who achieved pCR (n = 10) and without pCR (n = 8).
Immunohistochemical analysis using specific monoclonal antibodies confirmed that all breast tissue samples were negative for estrogen, progesterone, and HER2/neu receptors expression (Figure 1). To identify a molecular profile of miRNAs that could discriminate between the responder and no-responder groups, an unsupervised classification approach based on expression values for each miRNA t test was employed. After comparative 2−ΔΔCt analysis, a set of 11 differentially expressed miRNAs were identified in pCR versus no-PCR groups (adjusted p value ≤ 0.05; FC ≥ 1.5; Table 2). Of these, three miRNAs were significantly downregulated (miR-143-5p, miR-770-5p, and miR-584) and eight were upregulated (miR-135b, miR-380-5p, miR-941, miR-30a-3p, miR-652, miR-9-3p, miR-181c*, and miR-135*) in responder relative to no-responder patients. Moreover, this panel of deregulated miRNAs was able to separate the pCR from no-pCR group of tumors in the two-way unsupervised hierarchical cluster as shown in Figure 2(a). To confirm the differential expression of miRNAs identified in the TLDA screening, four deregulated miRNAs were analyzed by individual qRT-PCR. Results showed that the expression levels of miR-30a, miR-9-3p, miR-770, and miR-143-5p were similar in both the assays (Figure 2(b)).
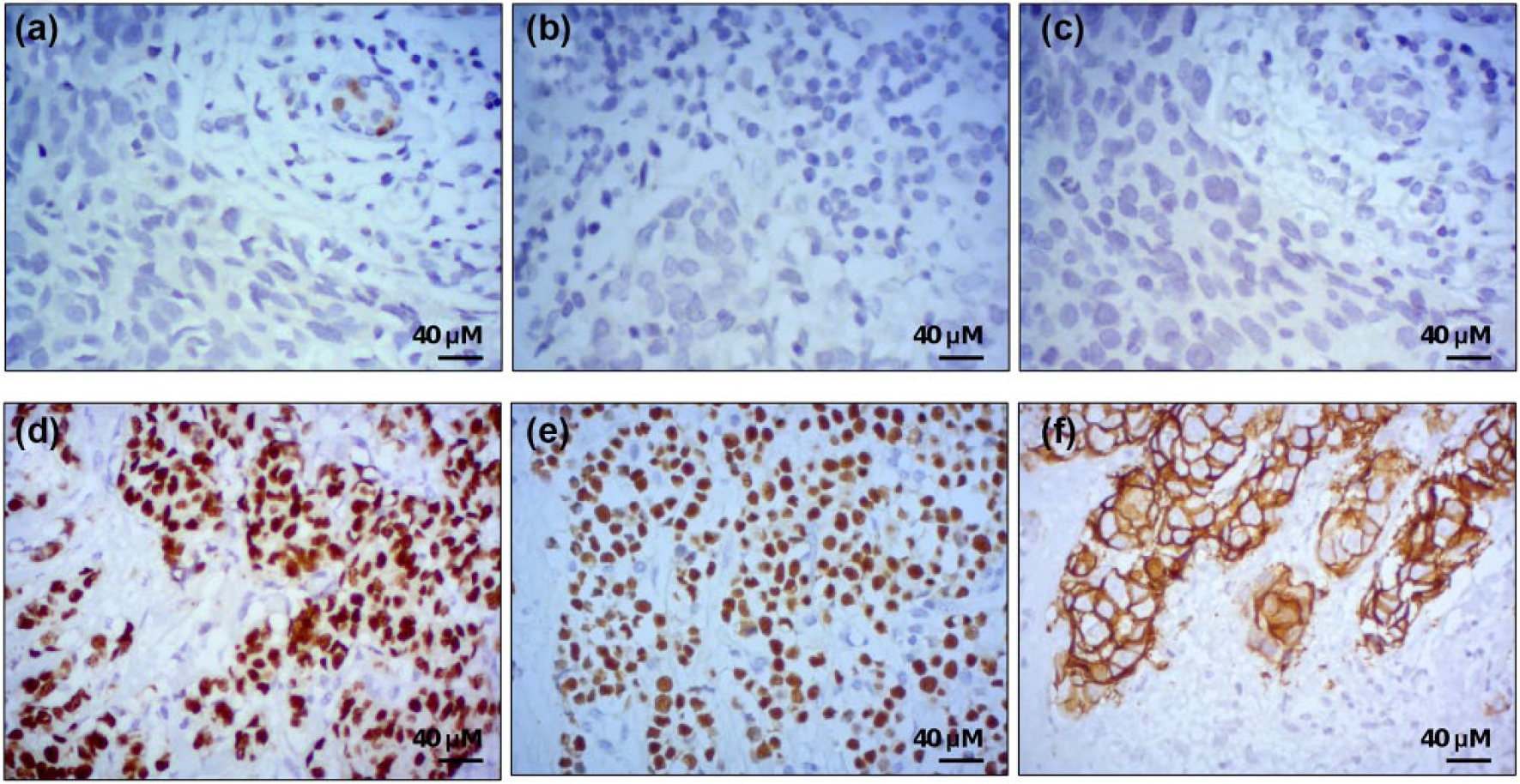
Representative images of the immunohistochemical reactions in triple-negative breast cancer tissues. (a) Estrogen receptor: (d) positive control. (b) Progesterone receptor: (e) positive control. (c) HER2/neu: (f) positive control.
MicroRNAs differentially expressed in pCR group.

pCR: pathological complete response.

Supervised hierarchical clustering of miRNAs differentially expressed in patients with pathological complete response. (a) Clustering of 18 breast cancer samples using data from 11 differentially expressed miRNAs. Heatmap function and Euclidean correlation were used; a p < 0.05 was considered to identify significantly differentially expressed miRNAs in patients with no response to experimental neoadjuvant therapy (FAC–CDDP/PTX). Columns display the clustering of tumor samples; rows indicate the clustering of miRNA names. Patients’ clinical outcome is indicated by colored squares: blue for no-pCR and red for pCR. (b) Validation of four miRNA levels using Taqman qRT-PCR assay (gray) in comparison with data obtained from TLDA (white). Data were expressed as mean ± SD.
Overview of deregulated miRNAs in patients with pCR
Of the 11 miRNAs differentially expressed, miR-584, miR-181c*, miR-380-5p, miR-135b-3p, miR-30a-3p, and miR-9-3p have not been previously associated with clinical response to chemotherapy in breast cancer. However, the functions of other miRNAs have been described in cancer. For instance, transforming growth factor beta (TGF-β) suppressed the miR-584 expression resulting in enhanced protein phosphatase and actin regulator 1 (PHACTR1) expression, actin cytoskeleton rearrangements, and migration of breast cancer cells. 19 TGF-β and other interleukins in the microenvironment have been reported to stimulate the adaptive immune response against TNBC.20,21 However, miR-380-5p targets B7-H3, an immunoregulatory protein of T cells. Overexpression of B7-H3 induces metastasis and is associated with poor prognosis in breast cancer. 22 On the contrary, overexpression of miR-941 was associated with ionizing irradiation (2 Gy) in human lymphoblast cell line TK6, but no data were described in patients. 23 Besides, miR-135b expression has been widely associated with chemotherapy response and prognosis in myeloma bone disease, non–small cell lung cancer, as well as colorectal and esophageal cancers.24–27
Validation of deregulated miRNAs by qRT-PCR
To validate the miRNA signature in terms of clinical response, we assessed the expression of a subset of four miRNAs (miR-30a, miR-9-3p, miR-770, and miR-143-5p) in an independent cohort of 17 tumor specimens from TNBC patients with pCR and no-pCR using qRT-PCR. Data confirmed that the differential expression of these set of miRNAs was able to distinguish between pCR and no-pCR groups (Figure 3).
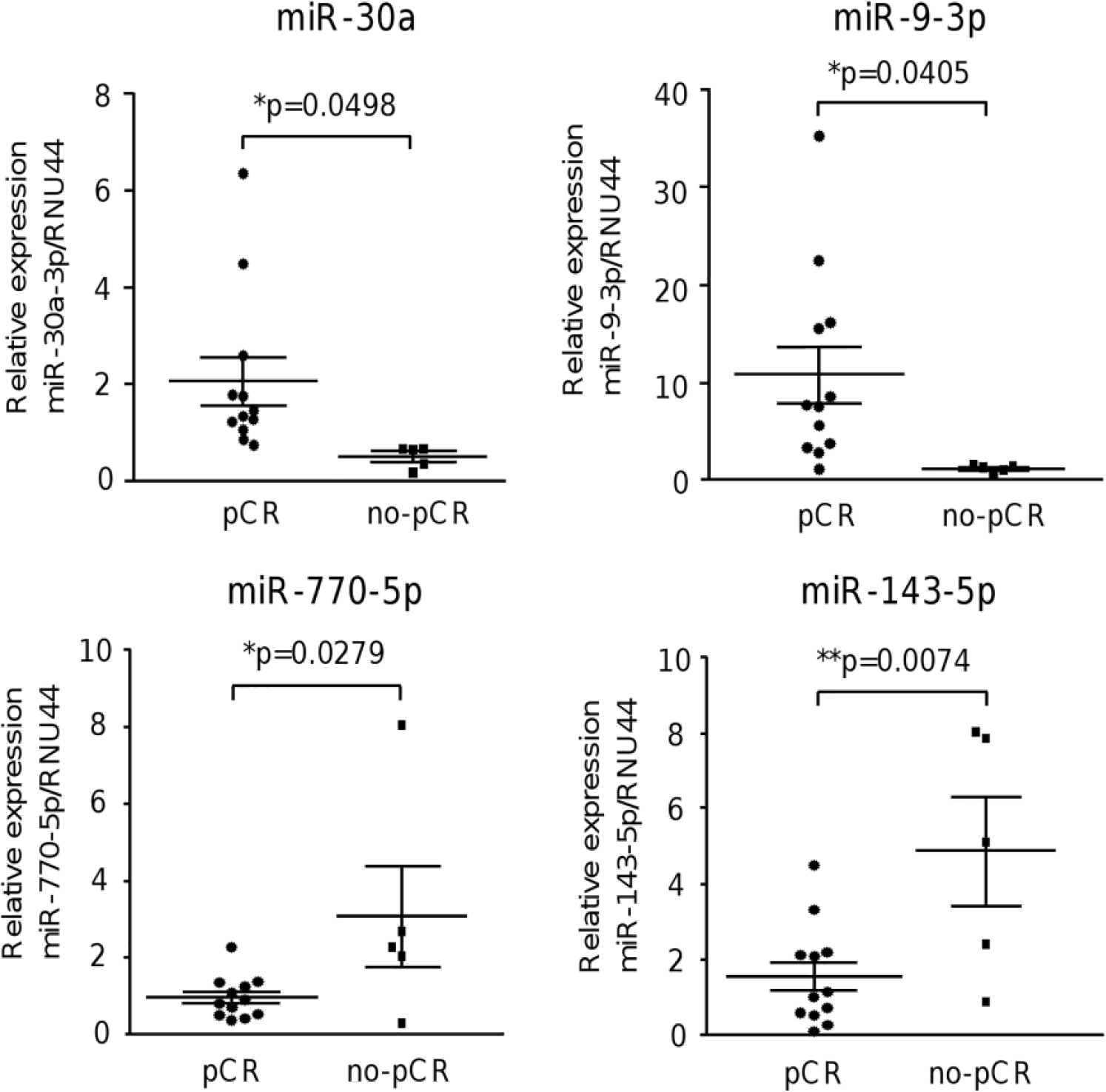
Validation of miRNA expression by qRT-PCR. Expression levels of miR-30a, miR-9-3p, miR-770-5p, and miR-143-5p were validated in an independent cohort of 17 patients using TaqMan MicroRNA Assays. Data were normalized with RNU44 expression and represented as mean ± SD.
Signaling pathways modulated by deregulated miRNAs
Classification of predicted targets of deregulated miRNAs based on gene ontology categories indicated that several genes were involved in cellular processes and pathways frequently deregulated in human cancers. A large number of predicted target genes belong to key cellular pathways that might modulate the hallmarks of cancer including focal adhesion (15 genes), mammalian target of rapamycin (mTOR) signaling (19 genes), p53 signaling (7 genes), TGF-β signaling (21 genes), and PI3K signaling among others (Figure 4). To better understand the putative targets and signaling pathways associated with treatment resistance in TNBC patients, we used a bioinformatics strategy that involved the identification of integrative miRNA–mRNA regulatory pathways. For this analysis, we used the list of 11 significantly deregulated miRNAs between both groups of patients. We selected those miRNA–mRNA pairs that had three matches in different databases. One of the databases was miRWalk, which includes information from experimentally validated data; therefore, these identified mRNAs were considered as bona fide miRNA target genes. The selected gene targets were associated with diverse oncogenic signaling pathways. To obtain a complete visualization of the signaling pathways based on mRNAs and their respective miRNA regulators, mRNA targets and their associated miRNAs were visualized using Kyoto Encyclopedia of Genes and Genomes (KEGG)-mirPath v.3. With this approach, we identified 100 unique mRNA targets. The miRNA–mRNA interactions were represented in KEGG annotated pathways that are associated with TNBC patients who had no response to treatment. Integrating the global miRNA–mRNA target information, we found that the most significant biological pathways have been previously involved in the maintenance of the tumor phenotype including VEGF, focal adhesion, WNT, ErbB, cyclic adenosine monophosphate (cAMP), mTOR, mitogen-activated protein kinase (MAPK), VEGF, and Janus kinase–signal transducer and activator of transcription (JAK/STAT) signaling pathways (data not shown). All together these pathways are responsible for oncogenic cellular processes such as apoptosis, cell cycle, differentiation, angiogenesis, metabolism, and resistance to chemotherapy. In particular, we found that PI3K/AKT pathways were affected by deregulated miRNAs (Figure 5). These pathways illustrate some of the most significant pathways that may also maintain chemotherapy-resistant phenotypes in TNBC.

Supervised hierarchical clustering of signaling pathways affected by deregulated microRNAs. MicroT-CDS function and Euclidean correlation were used; a p < 0.05 was considered to identify significantly differentially expressed miRNAs in patients with no response to experimental neoadjuvant therapy (FAC–CDDP/PTX). Columns display the clustering of cellular pathways with the number of genes involved. Rows indicate the clustering of miRNA names, and pathways are denoted at bottom.

MiRNA–mRNA interactions. Pathways modulated by deregulated miRNAs are depicted. Specific miRNA and mRNA interactions are highlighted in red for upregulated miRNAs and in blue for the downregulated one. Predicted gene targets are depicted in yellow and orange boxes.
Disease free survival
Kaplan–Meier survival analysis using the four validated miRNA signature was performed to estimate the DFS in TNBC. Patients were dichotomized into two groups with low- and high-expression levels using the median value for each miRNA as the cutoff. Expression of miRNA signature in comparison with the median level was considered as a significant predictor of no response to FAC–CDDP/PTX treatment (p < 0.001). Data showed that both groups were separated within the first month. In addition, the no-pCR group had a mean DFS of 38 months, while DFS was not reached in the pCR group (Figure 6).
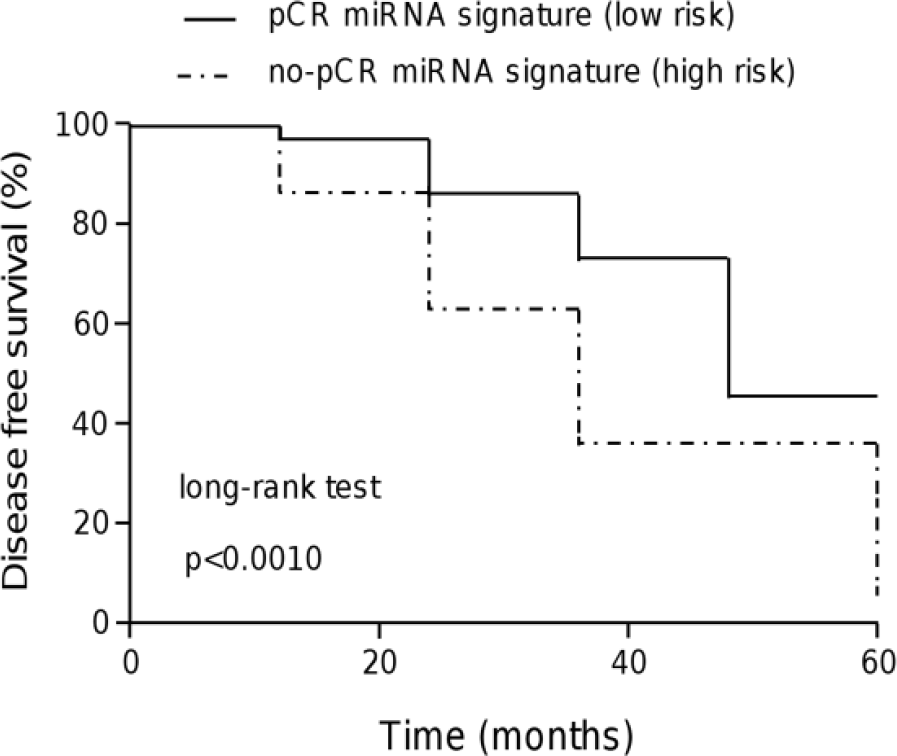
Kaplan–Meier survival analysis based on four validated miRNAs. The behavior of the four miRNAs for patients with no response to treatment is shown by a dotted line. Continuous line denotes the behavior for patients with a complete response. The log-rank (Mantel–Cox) test identified significant differences (p < 0.001) between the groups of patients (95% CI: 0.014–0.052).
Discussion
Neoadjuvant chemotherapy has been shown to improve the outcome in patients with breast cancer and is considered as a standard care. The neoadjuvant chemotherapy is an optimal model for research on TNBC because it provides opportunities to test novel or conventional therapeutic regimens that may lead to better clinical response and outcome in TNBC. pCR has been proposed as a surrogate marker for prediction of long-term clinical benefits after neoadjuvant therapy; however, most breast cancer patients have an unfavorable pCR to therapy. As only a minority of breast cancer patients would benefit from conventional or novel regimens, there is a need to identify novel biomarkers that may predict clinical response to neoadjuvant therapy. These data may be used to improve treatment strategy and to reorient the oncologic management of patients. Several retrospective analyses in diverse types of cancer, especially colorectal cancer, suggest that miRNAs associated with pathological stage of disease after neoadjuvant treatment has a significant prognostic impact on DFS and OS. In particular, the subgroup of patients who achieved a pCR has a very low risk of local or distant recurrence.28–31 These data highlight the prominent role of miRNAs as potential predictors of pCR in cancer.
In order to contribute to the discovery of novel potential markers for clinical response to neoadjuvant treatment, we studied a small cohort of TNBC patients treated with a novel therapeutic FAC–CDDP/PTX–based experimental treatment and achieved pCR. Our data revealed a set of 11 miRNAs that were significantly deregulated in pCR patients. Interestingly, several miRNAs identified in this study were previously associated with resistance to chemotherapy, which validates our results. For instance, the most downregulated miRNA identified here, miR-770-5p, inhibits cisplatin chemoresistance in human ovarian cancer (OVC) cells by targeting DNA repair gene ERCC2. 32 In addition, miR-770-5p expression was proposed as a prognostic biomarker and a predictor of response to cisplatin treatment and survival among OVC patients. In our study, we found that miR-770-5p was drastically suppressed in patients who exhibit a good response to chemotherapy, thus it could represent a feasible marker for clinical response to cisplatin-based therapy. Other miRNA deregulated in the pCR patients was miR-143, which has been recently studied in highly glycolytic TNBC tumors. MiR-143-based therapeutics targeting hexokinase 2 in breast tumors have been reported; however, the therapeutic role of miR-143 in patients was not evaluated. 33 However, a miRNA profile, which includes miR-652, was reported as a survival predictor for colon adenocarcinoma. 34 In addition, overexpression of miR-941 was associated with ionizing irradiation (2 Gy) in human lymphoblast cell line TK6. 23 Moreover, miR-135b expression has been widely associated with chemotherapy response and prognosis in myeloma bone disease, non–small cell lung cancer, colorectal, and esophageal cancers.24–27 These findings suggest a common role for this set of miRNAs in mechanisms leading to chemotherapy resistance in diverse human tumors. We suggested that alterations of miRNA levels found in TNBC cohorts analyzed here could be involved in the positive response to therapy and pCR, probably through the stimulation of diverse signaling pathways. Our data also suggested that these miRNAs could be associated with the development of chemotherapy resistance through the regulation of key genes involved in VEGF, focal adhesion, WNT, ErbB, cAMP, mTOR, MAPK, VEGF, and JAK/STAT signaling pathways (Figures 3 and 4). In contrast, several miRNAs identified here (miR-584, miR-181c*, miR-380-5p, miR-135b-3p, miR-30a-3p, and miR-9-3p) have not been previously associated with clinical response of patients to chemotherapy, thus they represent novel and potential therapeutic targets and deserve further clinical and functional investigation.
In conclusion, here we identified a novel signature of miRNAs related to pCR in a small cohort of TNBC, which deserves further validation on larger groups of patients and in other subtypes including HER2 and luminal breast tumors. Our results were sufficiently robust to demonstrate that the four validated miRNAs could be associated with response to therapy and DFS; therefore, we suggested that they could be useful as biomarkers of pCR after neoadjuvant therapy in TNBC patients.
Footnotes
Acknowledgements
The authors acknowledge Universidad Autónoma de la Ciudad de México and CONACYT (Grants: 222335 and 233370) for support. The authors also thank Julieta Santamaría for patients’ database management.
Data accessibility
All relevant data are available with this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Consejo Nacional de Ciencia y Tecnología (CONACYT; México), fondo de Investigación Científica Básica (Grant No. 222335), and fondo SSA/IMSS/ISSSTE (Grant No. 233370). R.G.-V. received CONACYT fellowship (Grant No. 441111).
Informed consent
Informed consent was obtained from all individual participants included in the study.
