Abstract
Gastric cancer is a common malignancy with limited treatment options and poor prognosis. Introduction of novel pathways of gastric cancer will provide candidates for target therapy. Hepatocyte growth factor activator inhibitor type 1 is an integral-membrane proteinase inhibitor. Hepatocyte growth factor activator inhibitor type 1 abnormality is found in various cancers and correlates with tumor progression and metastasis. However, the mechanisms underlying the dysregulation of hepatocyte growth factor activator inhibitor type 1 expression in gastric cancer remain unclear. Although microRNAs have been reported to be involved in the development of cancer, the roles of miR-221 and miR-222 in gastric cancer have not been reported yet. In this study, we showed that hepatocyte growth factor activator inhibitor type 1 protein was downregulated, while miR-221 and miR-222 were significantly increased in gastric cancer tissues. Bioinformatic predictions and luciferase assay verified that the 3′-untranslated region of the
Keywords
Background
Gastric cancer (GC) is the fifth most common malignant disease and the third leading cause of cancer-related mortality worldwide. 1 Surgical resection and systemic chemotherapy have improved the treatment outcomes during the past decade, but prognosis of GC remains poor with a 5-year–overall survival is only 29%.2,3 Targeting major pathways involved in tumorigenesis, like proliferation or metastasis, is a promising strategy for the treatment of GC. 4 However, effective targeted drugs for GC treatment are still limited. 2 Studies on new pathways of GC carcinogenesis will provide candidates for target therapy.
Hepatocyte growth factor activator inhibitor 1 (HAI-1) is a type I transmembrane serine protease inhibitor encoded by the serine protease inhibitor Kunitz type I (
MicroRNAs (miRNAs) are a novel class of endogenous, single-stranded noncoding RNAs (~22 bp) that are believed to regulate gene expression by directly binding to the 3′-untranslated region (3′-UTR) of target messenger RNAs (mRNAs), which results in either degradation of the transcript or impairing their translation. 9 Researchers indicate that over 30% of human genes are regulated by miRNAs.10,11 Altered miRNA expression profiles are reported in various types of tumors, including gastric, colorectal, breast, and lung cancer. 12 Acting as tumor suppressors or oncogenes, miRNAs refer to several aspects of the cancer pathogenesis, including cell proliferation, invasion, and metastasis. 13 Recently, some have studies shown that miRNAs can be secreted into the extracellular environment through microvesicles (MVs) and function as secretory signaling molecules that influence the recipient cell phenotypes.14,15
miR-221 and miR-222 (miR-221/222) are encoded on the X chromosome in tandem and are highly conserved in vertebrates. The expression of miR-221/222 has been found to be dysregulated in several cancers. 16 Overexpression of miR-221/222 in prostate carcinoma, breast cancer, and hepatocelluar carcinoma was found to promote cell proliferation and suppress apoptosis. 16 It has been reported that miR-221/222 are also significantly upregulated in GC.17,18 However, the molecular mechanisms by which miR-221/222 regulate tumorigenesis in GC remain largely unknown.
In this study, we have shown that miR-221/222 were upregulated, while the expression of HAI-1 protein was significantly suppressed in GC tissues. The subsequent luciferase assay showed that the 3′-UTR of HAI-1 is a direct target site of miR-221/222. Overexpression of miR-221/222 in GC cells leads to the inhibition of HAI-1 protein expression, thus promoting cell proliferation and migration. Moreover, we found that the expression level of HGFA protein was increased when HAI-1 was downregulated in GC cells. Thus, GC is probably an autocrine tumor, and the antitumor mechanism of HAI-1 in vitro might be mediated by regulating the expression of HGFA protein. Therefore, our data illustrated a novel pathway comprising miR-221/222 and HAI-1 in GC, which is a potential target for future clinical use.
Methods
Tissue specimens
Human GC tissues and paired adjacent noncancerous tissues were obtained from patients undergoing a surgical procedure at the Tianjin Medical University Cancer Institute and Hospital (Tianjin, China). Tissue fragments were promptly frozen in liquid nitrogen at the time of surgery and stored at −80°C. Both the tumor tissues and noncancerous tissues were histologically confirmed. The pathological type of each cancer was determined to be adenocarcinoma. This study was approved by the Ethics Committee of Tianjin Medical University Cancer Institute and Hospital. Written consents were obtained from each patient.
Cell line and cell culture
The human GC cell line MGC-803 was purchased from the Shanghai Institute of Cell Biology, Chinese Academy of Sciences (Shanghai, China). The human embryo kidney epithelial cell line HEK293T was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). MGC-803 and HEK293T were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, Rockville, MD, USA) supplemented with 10% fetal bovine serum (FBS, Gibco, Rockville, MD, USA), with routine addition of ampicillin and streptomycin.
Immunohistochemical staining
Formalin-fixed, paraffin-embedded specimens of tissue specimens were sectioned and stained with an anti-HAI-1 antibody (ab189511, Abcam, Cambridge, MA, USA) at a 1:50 dilution. Positive reaction was observed with 3,3′-diaminobenzidine (DAB Substrate Kit, Zhongshan Jinqiao, Beijing, China) as chromogen. Sections were counterstained with hematoxylin. At least five random fields were selected for each specimen.
RNA isolation and quantitative reverse transcription polymerase chain reaction
Total RNA was extracted from the tissues and cultured cells using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Assays to quantify mature miRNAs, HAI-1, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA were conducted using methods as described previously. 19 All the reactions were run in triplicate. After the reactions were completed, the cycle threshold (CT) data were determined using fixed threshold settings, and the mean CT was determined from triplicate polymerase chain reactions (PCRs). A comparative CT method was used to compare each condition to the control reactions. U6 small nuclear RNA (snRNA) was used as an internal control of miRNAs, and the mRNA levels of HAI-1 was normalized to GAPDH. The relative amount of gene normalized to control was calculated with the equation 2−ΔCT, in which ΔCT = CTgene − CTcontrol. Primers of HAI and GAPDH were as follows: GAPDH—5′-AGAAGGCTGGGGCTCATTTG-3′ (sense) and 5′-AGGGGCCATCCACAGTCTTC-3′ (antisense); HAI-1—5′-CAGCAGTGCCTCGAGTCTTGTC-3′ (sense) and 5′-GATGGCTACCACCACCACAATG-3′ (antisense).
The miRNA target prediction
The miRNA target prediction was analyzed with the bioinformatics tools of TargetScan (http://www.targetscan.org/), miRanda (http://www.microrna.org/), and PicTar (http://pictar.mdc-berlin.de/).
Cell transfection
MGC-803 cells were seeded on six-well or other plates and transfected with Lipofectamine 2000 (Invitrogen) on the following day according to the manufacturer’s instructions. miRNA overexpression was achieved by transfecting cells with miRNA mimics (RiboBio, Guangzhou, China), whereas knockdown was achieved by transfecting the cells with miRNA locked nucleic acid (LNA) inhibitors (Ribobio). For the purpose of HAI-1 upregulation, HAI-1 over expressing lentiviral particles (sc-404011-LAC, Santa Cruz Biotechnology, CA, USA) were put into use and an empty lentiviral was used as a negative control. A small interfering RNA (siRNA) targeting human HAI-1 (sc-39554, Santa Cruz Biotechnology) was used for HAI-1 downregulation and a scrambled siRNA was used as a negative control.
Luciferase reporter assay
The 3′-UTR of the wild type and mutated human HAI-1, containing the predicted miR-221/222 targeting regions, was synthesized and inserted into a p-MIR-reporter plasmid (GenScript, Nanjing, China). For luciferase reporter assays, HEK293T cells were seeded on six-well plates and cotransfected with 2 mg of β-galactosidase expression vector (Ambion, Austin, Texas, USA), 2 mg of firely luciferase reporter plasmid, and equal amounts (200 pmol) of miR-221/222 mimics, inhibitors, or scrambled negative control RNA using Lipofectamine 2000 (Invitrogen). And the β-galactosidase vector was used as a transfection control. After 24 h of transfection, the cells were harvested and analyzed for luciferase activity using a luciferase assay kit (Promega, Wisconsin, USA).
Western blotting
The HAI-1, tyrosine-protein kinase Met (c-MET), hepatocyte growth factor (HGF), and HGFA protein expressions were assessed by western blotting and the protein levels were normalized to GAPDH or β-actin. Total proteins were extracted from tissues and cultured in radioimmunoprecipitation assay (RIPA) buffer with freshly added protease inhibitor. Total cell lysates were separated on sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes (Roche, Basel, Switzerland). After blocking with 2% bovine serum albumin (BSA) at room temperature for 1 h, the membranes were incubated overnight at 4°C with anti-HAI-1 (1:2000, Abcam), anti-c-Met (1:2000, ImmunoWay, Newark, New Jersey, USA), anti-HGF (1:2000, Abcam), anti-HGFA-L (1:5000, Santa Cruz Biotechnology), anti-GAPDH (1:5000, ImmunoWay), and anti-β-actin (1:5000, Santa Cruz Biotechnology) antibodies, respectively. Then, the membranes were incubated with the secondary antibodies and visualized with an enhanced chemiluminescence system kit (Millipore, Bedford, MA USA) according to the manufacturer’s protocol.
Cell proliferation assay
Cells were seeded on 24-well plates and transfected with miR-221/222 mimics, inhibitors, HAI-1 siRNA, and the relevant negative controls. At 24 h post-transfection, a Cell-Light EdU DNA cell kit (Ribobio) was used to measure the cell proliferation ability according to the manufacturer’s instructions. Briefly, 5-ethynyl-2′-deoxyuridine (EdU) at a concentration of 50 µM/mL was added into the medium and cultured for 5 h. After being fixed with 4% paraformaldehyde for 30 min and treated with 0.5% Triton X-100 for 15 min, the cells were then incubated in dark with Apollo for 30 min and in Hoechst 33342 for another 30 min. Eventually, EdU-labeled cells were counted manually in five fields of view randomly selected from each well. All the staining procedures were performed in triplicate.
Wound scratch assay
MGC-803 cells were seeded on 12-well plates and transfected with miR-221/222 mimics, inhibitors, HAI-1 siRNA, and the relevant negative controls as described in “cell transfection.” When densities of the transfected cells were more than 90%, each well was scratched to create two empty linear regions with a 10 µL pipette tip. Then, the cells were cultured in DMEM medium without FBS in the humidified incubator. The wound healings were observed and pictured at 0, 12, 24, and 36 h after scraping. The distance of the wound zone was measured at least in three randomly selected fields.
Transwell cell migration assay
The capability of cell migration was also assessed using a 24-well Boyden chambers with 8.0-µm pore polycarbonate membrane insert (Corning, New York, NY, USA), according to the manufacturer’s instructions. Briefly, at 24 h post-transfection, approximately 105 cells were seeded on the upper chamber with 200 µL serum-free medium. Simultaneously, 600 µL of medium with 10% FBS was added into the lower chamber for chemotaxis. Another 24 h later, the membranes were fixed with methanol and stained with a three-step stain set (Thermo Fisher Scientific, Runcorn, UK). All assays were performed in triplicate. To minimize the bias, at least three randomly selected fields were counted with 200× magnification under a microscope, and the average number was taken.
Statistical analysis
SPSS 20.0 was used to analyze the data. Statistical analysis was performed using an unpaired Student’s
Results
HAI-1 is downregulated in human GC tissues
The expression pattern of HAI-1 in GC is still not elucidated. In this study, immunohistochemistry (IHC) was first applied to analyze the expression characteristics of HAI-1 protein in the GC. Six pairs of GC tissues and corresponding adjacent noncancerous tissues derived from patients undergoing radical gastrectomy were collected. Results showed that HAI-1 protein had low expression in the GC tissues, and four patients had high expression of HAI-1 protein in the paracarcinoma tissues (Figure 1(a)). These four patients were further analyzed for HAI-1 protein expression using western blotting. In accordance with the results of the IHC, HAI-1 protein is lowly expressed in the cancer tissues compared to the corresponding noncancerous tissues (Figure 1(b) and (c)). Thus, the expression of HAI-1 protein is downregulated in human GC tissues.

The expression patterns of HAI-1 and miR-221/222 in GC tissues. (a) Immunohistochemical staining of the human GC tissues and paired noncancerous tissues (400×). (b) Western blot analysis of HAI-1 expression in the GC cancer tissues and paired noncancerous tissues. (c) Quantitative analysis of (b) (
Identification of miR-221/222 as potential upstream regulator of HAI-1
The mechanism of downregulation of HAI-1 protein expression in GC is largely unknown. miRNA can regulate gene expression by directly binding to the 3′-UTR of target mRNAs. To investigate whether a miRNA regulation mechanism is involved in the post-transcriptional regulation of HAI-1 protein expression, bioinformatics tools were put out to predict the miRNAs that can potentially bind with HAI-1 3′-UTR. Results showed that miR-221/222 can directly bind with a region in the 3′-UTR of HAI-1 mRNA (Figure 1(d)). Then, the four patients in which HAI-1 proteins were highly expressed in the paracarcinoma tissues were further analyzed for the levels of miR-221/222. As expected, miR-221/222 are lowly expressed in the paracarcinoma tissues and obviously increased in the paired tumor tissues (Figure 1(e)). Therefore, miR-221/222 are most likely to be important regulator of HAI-1 in GC.
Validation of HAI-1 as a direct target of miR-221/222
The levels of miR-221/222 and HAI-1 protein in GC showed inverse correlation. In addition, bioinformatics predictions showed that the 3′-UTR of human HAI-1 mRNA contains a potential miR-221/222 binding site with 6-mer seeds (Figure 2(a)). However, the direct evidence of the interaction between miR-221/222 and HAI-1 mRNA given by luciferase assay is still needed. As shown in Figure 2(b) and (c), the relative luciferase activity of the reporter gene was significantly inhibited when HEK293T cells were cotransfected with the luciferase reporters containing the predicted target regions of HAI-1 mRNA and miR-221/222 mimics, while the inhibition was lost when the binding sites in 3′-UTR were mutated. Moreover, the cotransfection of miR-221/222 inhibitors and the plasmid with the wild HAI-1 3′-UTR resulted in a relative increase in the luciferase signal. These data verified that HAI-1 is a direct target of miR-221/222.

Direct recognition of the HAI-1 3′-UTR by miR-221/222. (a) Schematic description of the base-pairing interaction between miR-221/222 and HAI-1 mRNA. (b and c) Luciferase reporter assay. HEK293T cells were cotransfected with firefly luciferase reporters containing either wild type (WT) or mutant (Mut) HAI-1 3′-UTR with miR-221/222 mimics and inhibitors (
miR-221/222 regulates HAI-1 expression in MGC-803 cells
To further clarify the relationship between miR-221/222 and HAI-1 in GC, miR-221/222 mimics or inhibitors were transfected into MGC-803 cells, and relative levels of miR-221/222 and the expression of HAI-1 protein and mRNA were detected using quantitative reverse transcription PCR (qRT-PCR) or western blotting, respectively. As shown in Figure 3(a), miR-221/222 mimics significantly increased the levels of miR-221/222 in the MGC-803 cells, while inhibitors downregulated the levels of miR-221/222 dramatically. Overexpression of miR-221/222 led to the reduction of HAI-1 protein expression, whereas downregulation of miR-221/222 led to an increased expression of the HAI-1 protein (Figure 3(b)). Furthermore, miR-221/222 did not affect the mRNA levels of HAI-1 in the MGC-803 cells (Figure 3(c)). These results suggested that miR-221/222 are vital post-transcriptional regulators of HAI-1, and miR-221/222 can regulate HAI-1 protein expression in the GC cells.

Effect of miR-221/222 on HAI-1 expression in MGC-803 cells. (a) Quantitative RT-PCR analysis of the relative miR-221/222 levels in MGC-803 cells transfected with mimics or inhibitors (
miR-221/222 regulate proliferation and migration of MGC-803 cells
We next analyzed the biological roles of miR-221/222 on the GC cells in vitro. Wound healing method was first performed to evaluate the migration capacity of MGC-803 cells transfected with miR-221/222 mimics or inhibitors. Results showed that upregulation of miR-221/222 promoted cell migration, whereas downregulation of miR-370 suppressed cell migration (Figure 4(a)). We also conducted a transwell assay to evaluate the migration ability of the transfected cells. Accordance with the results of the wound healing, MGC-803 cells transfected with miR-211/222 mimics showed a higher ratio in migration, while inhibition of miR-221/222 caused a decrease in the cell migration (Figure 4(b)). Cell proliferation rate was evaluated by the Cell-Light EdU DNA cell kit (RiboBio, Guangzhou, China). As shown in Figure 4(c), high levels of miR-221/222 resulted in an increase in the cell growth rate; in contrast, inhibition of miR-221/222 had the opposite effect on cell proliferation. These results demonstrated that miR-221/222 synergistically increase in proliferation and migration in GC cells.
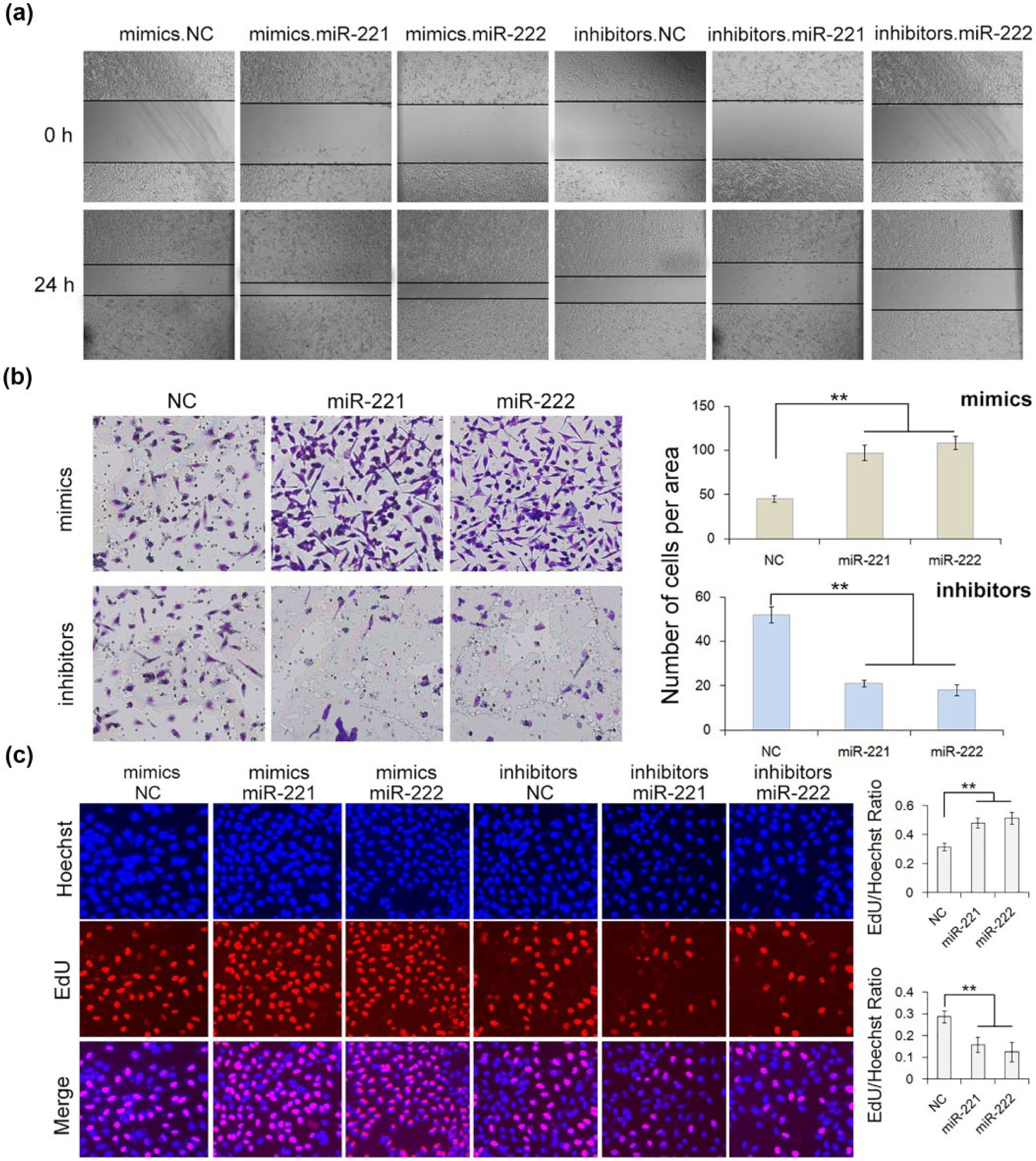
Effect of miR-221/222 on proliferation and migration of MGC-803 cells. (a) Validation of cell migration promotion mediated by miR-221/222 using wound healing method (
Effect of HAI-1 silencing on MGC-803 cells
We next investigated the effects of HAI-1 silencing on cell proliferation and migration. The siRNA sequence targeting human HAI-1 was used to knockdown the expression of HAI-1 protein. The silencing efficiency was assessed by qRT-PCR for RNA and by western blotting for protein. As shown in Figure 5, cells transfected with HAI-1 siRNA led to an obvious decrease in the expression levels of protein and mRNA. MGC-803 cells in which HAI-1 was knocked down exhibited a significant increase in migration and higher rate of proliferation (Figure 6(a)–(c)). These results demonstrated that HAI-1 acts as a cancer suppresser in the GC, the dysregulation of which may promote the proliferation and migration of cancer cells.

Effects of HAI-1 silencing on HAI-1 expression in MGC-803 cells. (a) Quantitative RT-PCR analysis of the relative levels of HAI-1 mRNA in MGC-803 cells transfected with siRNA (

Effects of HAI-1 silencing on cell proliferation and migration of MGC-803 cells. (a) Validation of HAI-1-mediated cell migration using wound healing method (
Effect of HAI-1 overexpression on MGC-803 cells transfected with miR-221/222 mimics
To provide more evidence that miR-221/222 promote tumorigenesis in GC through the silencing of HAI-1, we used miR-221/222 mimics and lentivirus particles to overexpress miR-221/222 and HAI-1 simultaneously in MGC-803 cells. As shown in Figure 7(a), cotransfection of miR-221/222 mimics and HAI-1 lentivirus particles partly rescued HAI-1 protein expression compared with the single transfection of miR-221/222 mimics. The overexpression of HAI-1 in MGC-803 cells partly decreased miR-221/222-induced cell proliferation and migration (Figure 7(b)–(e)).

Effects of miR-221/222 on proliferation and migration of MGC-803 cells overexpressed with HAI-1. (a) Western blot assay of HAI-1 protein expression in MGC-803 cells transfected either with miR-221/222 mimics and HAI-1 lentivirus or alone (
HAI-1 regulates the expression level of HGFA protein in MGC-803 cells
HGF is produced as an inactive precursor called pro-HGF, which requires proteolytic conversion by HGFA to evoke a biological response. c-MET is specific receptor for HGF. To give a better understanding of the antitumor mechanisms of HAI-1 in vitro, we checked the HGF, HGFA, and c-MET protein expression in the MGC-803 cells. As shown in Figure 8, HGF, HGFA, and c-MET were also expressed in the MGC-803 cells. Furthermore, the expression level of HGFA protein was increased when HAI-1 was knocked down using siRNA (Figure 8(a)–(d)) or was downregulated by miR-221/222 mimics (Figure 8(e)–(h)). Thus, GC is probably an autocrine tumor, and the antitumor mechanism of HAI-1 in vitro seemed to be mediated, at least partly, by regulating the expression of HGFA.

Effects of HAI-1 downregulation on the expression of HGFA protein in MGC-803 cells. (a) Western blot assay of the HGFA, HGF, and c-MET protein expression in MGC-803 cells transfected with HAI-1 siRNA. (b–d) Quantitative analysis of (a) (
Discussion
In this study, we obtained a number of results regarding HAI-1 and miR-221/222 in GC. HAI-1 is a target gene of the miR-221/222 cluster. miR-221/222 act as onco-miRNAs in the development of GC. The important roles of HAI-1 in GC are mediated, at least partly, by regulating the expression of HGFA protein. Our results supported these suggestions. Computer-aided algorithm predicted that miR-221/222 could directly bind with the 3′-UTR of HAI-1 mRNA. Luciferase assay validated that miR-221/222 were regulators of HAI-1. Overexpression of miR-221/222 promoted the proliferation and migration of the GC cells, while downregulation of miR-221/222 inhibited the proliferation and migration. In addition, GC was probably an autocrine tumor, and downregulated HAI-1 protein expression was accompanied by significantly increased expression level of HGFA protein.
Low expression of HAI-1 protein has been observed in a number of advanced malignancies such as cervical, endometrial, and colorectal carcinomas, as compared with their paired paracarcinoma tissues.20–22 In addition, reduced HAI-1 mRNA levels have been reported in breast, colorectal, and renal cell carcinoma tissues.22–24 Further detailed analysis indicated that reduced HAI-1 expression levels were associated with worse clinicopathological parameters and poor prognosis. 25 However, the regulatory mechanisms that result in the aberrant HAI-1 expression levels under different circumstances remain unclear. In our study, we found that HAI-1 protein expression in GC was controlled by a post-transcriptional regulation. Thus, our data illustrated a novel miR-221/222-HAI-1 pathway in the tumorigenesis of GC, which could act as a potential target in the future treatment of GC. To our knowledge, this is the first attempt to investigate the miRNA-HAI-1 pathway in the GC.
MiR-221/222 could be served as potential therapeutic targets for GC. Several reports indicated that miR-221/222 are dysregulated in several cancers and could be used as a therapeutic tool. 16 Pineau et al. 26 reported that antago-miRNAs can reduce the growth of liver cancer cells in which miR-221/222 were overexpressed. Galardi et al. 27 showed that miR-221/222 knockdown through antisense LNA oligonuceleotides strongly reduced the clonogenicity of human prostate cancer cells in vitro. Recently, one study from Chun-Zhi et al. 28 demonstrated that miR-221/222 can regulate radiosensitivity via direct modulation of phosphatase and tensin homolog (PTEN) expression and this suggested that inhibition of miR-221/222 might form a novel therapeutic strategy for GC. In addition, our group and others have reported that miRNA and other small RNAs trafficked by MVs or plasmids can be used for the treatment of diseases, including cancer, in mouse models.29–31 Therefore, miR-221/222 have a good prospect in clinical application.
To date, several studies have suggested a pathogenic role of HAI-1 in the invasion and migration of carcinoma cells in vitro. Forced expression of HAI-1 protein significantly inhibited the invasion and migration of cervical, endometrial, and uterine cancers. 25 Furthermore, breast, prostate, pancreatic, and oral carcinoma cells also exhibited enhanced invasiveness in vitro in response to HAI-1 knockdown. 25 However, the precise mechanisms regulating cellular invasiveness and motility by HAI-1 in cancer cells remain poorly understood. Studies have shown that downregulation of HAI-1 protein expression was accompanied by significant increase in the expression levels of HGFA or matriptase, and HAI-1 knockdown-induced enhanced migration is partially reversed by silencing of serine protease expression.7,25,32,33 Our study also showed that the expression level of HGFA protein was upregulated after HAI-1 was downregulated in MGC-803 cells. Thus, GC is probably an autocrine tumor, and the antitumor mechanism of HAI-1 in vitro seemed to be mediated, at least partly, by regulating the expression of HGFA protein. However, this needs more evidence in further studies.
In conclusion, we demonstrate that the expression of HAI-1 protein in GC is regulated by miR-221/222, and the miR-221/222-HAI-1 pathway is involved in the process of cell growth and migration. This novel pathway may provide potential therapeutic targets for GC in the future.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by grants from the National Natural Science Foundation of China (Y.B.: 81372394, H.Z.: 81602158, D.H.:81572321 and T.D.: 81602156) and Tianjin Health and Family Planning Commission Foundation of Science and Technology (Y.B.: 15KG142). This work was also funded by Tianjin Science Foundation (D.H.: No.15JCYBJC28200) and Doctoral Foundation of Tianjin Medical University Cancer Institute and Hospital (H.Z.: B1502).
Informed consent
Informed consent was obtained from all individual participants included in the study.
