Abstract
RNA editing is a widespread post-transcriptional mechanism that confers specific and reproducible nucleotide changes in selected RNA transcripts and plays a critical role in many human cancers. However, little is known about how RNA editing operates in non-small-cell lung cancers. Here, we measured the sequence and expression level of genes of antizyme inhibitor 1 and adenosine deaminase acting on RNA family in 30 non-small-cell lung cancer patient samples and 13 cell lines and revealed RNA editing S367G in antizyme inhibitor 1 is a high-frequent molecular events. We determined overexpression of antizyme inhibitor 1 with RNA editing, implying the oncogenic function of this alteration. We also detected the association of adenosine deaminase acting on RNA overexpression with RNA editing occurred in antizyme inhibitor 1. Furthermore, the RNA editing could cause a cytoplasmic-to-nuclear translocation of antizyme inhibitor 1 protein and conferred the malignant phenotype of non-small-cell lung cancer cells. The in vivo experiment confirmed that this RNA editing confers higher capacity of tumor migration as well. In conclusion, antizyme inhibitor 1 RNA editing and its involvement in tumorigenesis of non-small-cell lung cancer pave a new way for potential clinical management of non-small-cell lung cancer.
Introduction
Lung cancer is the most frequently diagnosed malignancy and the leading cause of cancer death worldwide. An estimated 1.8 million new lung cancer cases were diagnosed in 2012, accounting for 13% of total human cancers. 1 Lung cancer is also the most prevalent cancer in China, 2 while non-small-cell lung cancers (NSCLC) being the most common histological subtype. 3 Tobacco exposure, occupational carcinogens, and environmental pollution are known etiological factors in NSCLC tumorigenesis;4,5 however, understanding of biological factors in NSCLC is still required for more effective diagnosis and clinical management.
Similar to other human cancers, NSCLC is essentially promoted by the accumulation of somatic genetic or epigenetic alterations. Beyond genomic DNA level, RNA editing is a post-expression regulation system defined as the nucleotide sequence change of RNA transcripts following transcription of an encoding DNA sequence (gene). RNA editing enhances the RNA diversity and may lead to changes in amino acid sequence and alternative splicing, thereby increasing the complexity of gene expression.6,7 The RNA editing has been connected to cancer development and progression.8,9 Recently, several studies systematically characterized the landscape and clinical relevance of RNA editings in a variety of cancers.10,11 RNA editing is globally controlled by tumor interferon and adenosine deaminase acting on RNA (ADAR) copy number, and both factors are highly prevalent among human cancers.11,12 The high frequency of a site-specific RNA editing event, which is the serine-to-glycine substitution at residue 367 (S367G) in antizyme inhibitor 1 (AZIN1), was uncovered in hepatocellular and esophageal carcinoma.13,14 Here, we illustrated how AZIN1 RNA editing occurred in NSCLC patients’ specimens and cell line models as well as demonstrated the role of ADAR-mediated AZIN1 RNA editing in NSCLC development.
Methods
Tumor sample collection and cell line preparation
Paired tumor and normal adjacent tissues from 30 NSCLC patients were retrospectively collected, and 13 human cell lines were used in this research. All tumors were surgically resected and pathologically proven as primary NSCLC at the First Affiliated Hospital of Guangzhou Medical University, with no pre-operative chemotherapy or radiotherapy. Fresh tumors were macro-dissected within 30 min after surgical resection and stored at −80°C. All samples were obtained with informed consent and agreement. The study was approved by the Medical Ethics Committee of the First Affiliated Hospital of Guangzhou Medical University. In this study, we employed 12 lung cancer cell lines, including SPC-A1, H1299, A549, A0907, PC9, H460, H1395, Calu3, H520, H1650, H358, and HCC827, as well as a cell line derived from normal lung tissue, human bronchial epithelial (HBE). In principle, they were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS; Gibco), and penicillin–streptomycin (Invitrogen) was used routinely to prevent infection. All cells were incubated at 37°C in a humidified incubator with 5% CO2.
Complementary DNA synthesis, real-time polymerase chain reaction analysis, and RNA editing test
Total RNA was extracted using TRIzol reagent (Invitrogen) from NSCLC cell lines and clinical samples. To quantify gene expression in cells or clinical samples, complementary DNA (cDNA) was synthesized from equal amounts of RNA, using the PrimeScript™ RT reagent Kit (TaKaRa), and used for quantitative polymerase chain reaction (qPCR) analysis. We performed qPCR using the KAPA SYBR FAST qPCR master mix (Kapa Biosystems) and the primers of ADAR, ADARBP1, ADARBR2, and AZIN1, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as an internal control. The genotype of candidate RNA editing site was tested using direct Sanger sequencing, performing on PCR products with the primers AZIN1mut. The editing degrees were calculated by software ImageJ (http://rsb.info.nih.gov/ij/). All primers are listed in Table S1.
Immunohistochemical staining
We sectioned the paraffin-embedded tissue blocks for immunohistochemical (IHC) staining. Briefly, sections were deparaffinized and rehydrated. The endogenous peroxidase activity was blocked with 3% hydrogen peroxide (H2O2) for 10 min. For antigen retrieval, the slides were immersed in 10 mM citrate buffer (pH 6.0) and boiled for 15 min in a microwave oven. Non-specific binding was blocked with 5% normal goat serum for 10 min. The slides were incubated in a 1:100 dilution of AZIN1-specific antibody (ab57169; Abcam) at 4°C overnight in a humidified chamber. The slides were then sequentially incubated with biotinylated goat anti-mouse/rabbit IgG (MXB Bio-technologies) for 30 min at room temperature. Finally, the 3,3′-diaminobenzidine (DAB) Substrate Kit (Dako) was used for color development followed by Mayer’s hematoxylin counterstaining.
Lentiviral vector construction and infection
Vectors for expression of the wild-type and mutant AZIN1 fused to green fluorescent protein (GFP) gene were constructed by PCR, and fragments were amplified using the primers AZIN1-EcoRI-F and AZIN1-BamHI-R. The fragment was inserted between the EcoRI and BamHI sites of pLVX-mCMV-ZsGreen-PGK-puro. The AZIN1 mutant was created using a two-round PCR procedure. Briefly, two separate DNA fragments were generated in a first PCR: fragment-1 was amplified by primers AZIN1-EcoRI-F and AZIN1-mutantion-R, while fragment-2 was amplified by primers AZIN1-mutantion-F and AZIN1-BamHI-R. Two fragments were purified, mixed, and re-amplified by AZIN1-EcoRI-F and AZIN1-BamHI-R to produce a final PCR product, which was then cloned into pLVX-mCMV-ZsGreen-PGK-puro vector. Virus particles were harvested 48 h after the lentiviral vector (or the control lentivirus vector) co-transfection with the packaging vectors into human embryonic kidney (HEK) 293T cells using Lipofectamine 2000 (Life Technologies Corporation). Virus-containing supernatants were collected for subsequent transduction into NSCLC cells. Puromycin 5 µg/mL (Life Technologies Corporation) was used to select for stably transduced cells.
Western blot analysis and antibodies
Protein lysates were quantified and separated on a sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel. After the separation, proteins were transferred onto a polyvinylidene difluoride (PVDF) membrane (Bio-Rad Laboratories, Inc.), and immunoblotted with a primary antibody, followed by an incubation with a secondary antibody. The luminescence was visualized on the Tanon-5200 Chemiluminescent Imaging System (Tanon Science & Technology Co., Ltd.). The following antibodies were used: AZIN1 (1:1000, ab57169; Abcam), β-tubulin (1:5000, AP0064; Bioworld), goat anti-mouse IgG (1:5000, HS201; TransGen Biotech), and goat anti-rabbit IgG (1:5000, HS101; TransGen Biotech).
Adhesion assay
An amount of 45,000 cells were seeded onto a 96-well plate coated with Matrigel basement membrane in 200 µL normal medium and incubated for 40 min. Adherent cells were then fixed in 4% formaldehyde (v/v) in balanced salt solution (BSS) before being stained in 0.5% crystal violet solution (w/v). Absorbance was obtained using a plate reader (MultiskanTM FC; Thermo Fisher Scientific) after an extraction of the dye with 10% acetic acid. This was modified from the method previously reported. 15
Cell proliferation assay
Cells were seeded in 96-well plate at a density of 3 × 103 cells per well. The cells were fixed in 4% formaldehyde for 20 min and stained with 1% crystal violet. The crystal was dissolved in 10% glacial acetic acid. The absorbance was measured using the plate reader (Thermo Scientific) at a wavelength of 490 nm. Three independent experiments were performed in triplicate. This was modified from the method previously reported. 15
Wound-healing assay
Cells were seeded into six-well plate and grown to confluence. A wound was created by scraping the cell monolayer with a 200-µL pipette tip. Culture medium was changed to remove detached cells and cell debris, and then, wound closure was monitored microscopically at different time points. The cells were allowed to migrate for 24 h. Different time points of scratched gap breadth were measured using the light microscope (Nikon). This was modified from the method previously reported. 15
Matrigel invasion assay
Cell culture inserts were seeded with 5 × 103 cells in 100 µL of medium without FBS, and 24-well Transwell chambers were used according to the manufacturer’s instructions. The insert chamber was precoated with Matrigel (50 µL, 1 mg/mL; BD Biosciences). Medium with 10% FBS (800 µL) was added to the lower chamber and served as a chemotactic agent. Noninvasive cells were wiped from the upper side of the membrane, meanwhile cells in the lower side were fixed in formaldehyde and air dried. Cells were stained with 0.1% crystal violet (dissolved in methanol) and counted using the inverted microscope. Each individual experiment had triplicate inserts, and six microscopic fields were counted per insert.
Xenograft assay
Animal experiments were performed in the Laboratory Animal Center of Guangzhou Institutes of Biomedicine and Health (GIBH), and all animal procedures were approved by the Animal Welfare Committee of GIBH. NOD-scid-IL2Rg−/− (NSI) mice were derived at GIBH; 1 × 106 A549 control cells and A549 AZIN1edited cells were injected intravenously via the retro-orbital route into 8- to 10-week-old NSI mice, respectively. After 4 weeks, lung tumors and adjacent lung tissues were collected and fixed with 10% neutral buffered formalin, embedded in paraffin, and sectioned using a microtome (Leica).
Fluorescent antibody technique (immunofluorescence colony)
The cells were fixed in 100% methanol (10 min), permeated with 0.5% Triton X-100/PBS for 10 min, and blocked with 10% BSA/0.2% PBS-Tween for 30 min at 37°C. The cells were then incubated with the AZIN1 antibody (ab57169; Abcam) used at a 1/100 dilution overnight at 4°C. The secondary antibody (pseudo-colored red) was Alexa Fluor® 594 goat anti-mouse IgG (Life Technologies Corporation), which was used at a 1/1000 dilution for 45 min at room temperature. ProLong® Gold Antifade Reagent with DAPI (Life Technologies, USA) was used to stain the cell nuclei (pseudo-colored blue).
Result
The extensive spread of RNA editing of AZIN1 in NSCLC
Genomic DNA and RNA derived from 30 tumors and adjacent normal tissues of identical NSCLC patients, as well as 12 NSCLC cell lines, were applied to determine A-to-I RNA editing incidence in AZIN1. AZIN1 S367G RNA editing occurred in 20% clinical samples (6/30) and 17% NSCLC cell lines (2/12; Figure 1(a), using P0105 and H358 as examples). The range of editing degrees (or, namely, expressed mutant allele frequency) was 11%–49% in tumor tissues and found to be a little higher in cell lines (44% and 73%). The prevalence of AZIN1 RNA editing in our result showed a little higher than previous genome-wide studies,10,11 which may be due to sensitivity distinction in different experiments. Moreover, unlike the variant’s distribution in other cancer types, the AZIN1 RNA editing was exclusively occurred in tumors compared to adjacent normal tissues. Meanwhile, for cell line samples, the editing is absent in the HBE cells as well, which is originated from normal lung tissue. It is noticed that AZIN1 rarely carries somatic DNA mutation and copy number variation among NSCLC patients, according to a pan-lung cancer study 16 and a Chinese cohort study. 17 Combining with our data, RNA editing is the major form of molecular alteration that may confer AZIN1 function in NSCLC progression. Finally, combining with clinical information, this post-transcriptional modification tended to occur in advanced NSCLC, although the p value is not significant due to limited sample size (Table 1).
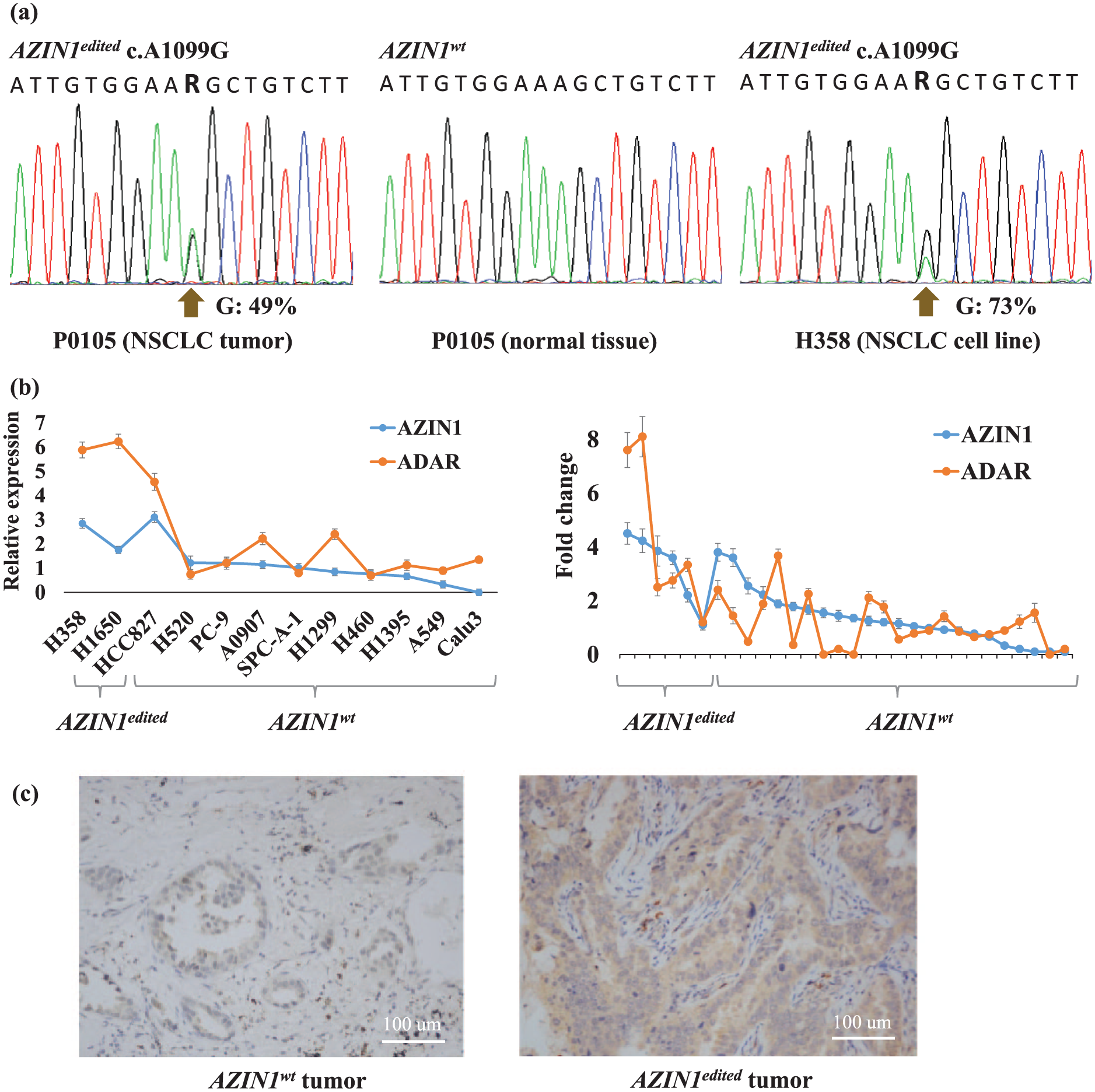
The RNA editing of AZIN1, and expression level of AZIN1 and ADAR in NSCLC. (a) The representative sequence chromatograms show editing of AZIN1 in a lung tumor (left), adjacent normal tissue (middle), and NSCLC cell line (right). (b) The mRNA expression level of AZIN1 in NSCLC cell lines (left) and patient’s specimens (right), and the RNA edited cells are noted, and standard error of mean (SEM) is depicted. For cell line samples, y axis shows expression value of the given genes in NSCLC cell lines relative to HBE. For clinical samples, y axis shows fold change value comparing tumors with normal tissue of identical patients. (c) IHC staining of AZIN1 protein in tumor with (left) AZIN1wt and (right) AZIN1edited. The protein expression was increased in AZIN1edited NSCLC cells (scale bar = 100 µm).
Summary of the clinical features of 30 NSCLC tested for AZIN1 RNA editing.

NA: not applicable; NSCLC: non-small-cell lung cancer.
Chi-squared test.
Wilcoxon rank-sum test.
AZIN1edited expression is upregulated in NSCLCs
For dissecting the role of AZIN1 RNA editing in the development of NSCLC, we explored AZIN1 mRNA expression in the above panel of NSCLC cell lines. AZIN1 expression was detected in 11/12 tested cell lines, including H358 and H1650 which carry the RNA editing event, and AZIN1 expression in HBE was also observed. Furthermore, the expression levels of AZIN1 in H358 and H1650 were higher than HBE and most of AZIN1wt NSCLC cells (Figure 1(b)). Then, we examined the AZIN1 expression data in TCGA NSCLC data using the cBioPortal resource and found that 18% (95/515) adenocarcinoma and 12% (62/501) squamous cell carcinoma showed overexpression in cancer tissue. 18 Next, we quantified the AZIN1 mRNA expression in clinical samples by real-time PCR, and 83% (5/6) of AZIN1edited tumors and 17% (4/24) of AZIN1wt tumors showed overexpression comparing to their normal adjacent counterparts (fold change > 2), respectively (Figure 1(b)). Moreover, AZIN1 mRNA abundance showed increase in AZIN1edited samples. We also determined the AZIN1 protein expression by immunohistochemistry (IHC) staining in six tumors, three derived from AZIN1wt patients and three derived from AZIN1edited patients, and confirmed that AZIN1 protein expressed increasingly in the AZIN1edited tumors (Figure 1(c)). In an overall view, this specific site of nucleotide change in transcript and overexpression status of AZIN1 suggest that the edited AZIN1 may function as an oncogene in a portion of NSCLCs. 19
The overexpression of ADAR in NSCLC and its association with AZIN1 RNA editing
The A-to-I editing, which accounts for most RNA editing events including the one found in AZIN1, is an enzymatic process mediated by ADAR. 7 The ADAR family has three members, including ADAR, ADARBP1, and ADARBP2 (also known as ADAR1, ADAR2, and ADAR3, respectively). The previous study revealed that ADAR and ADABP1 are expressed in most human tissues, while ADABP2 is exclusively detected in the central nervous system. 8 Here, we examined the expression of those three ADAR genes in above-mentioned NSCLC cell lines. All of them expressed ADAR, 50% (6/12) cell lines expressed ADARBP1 in a relatively low level, and ADARBP2 expression was barely detected. Comparing to HBE, most of NSCLC cell lines showed overexpression of ADAR (Figure 1(b)). Consequently, we focus on ADAR for further research in the case of AZIN1 RNA editing in NSCLC. The ADAR mRNA overexpression is detected in 14% (73/515) lung adenocarcinoma and 15% (73/501) lung squamous cell carcinoma, with the merit of cBioPortal data resource. 18 Here, in tested clinical biopsies, 83% (5/6) of AZIN1edited tumors and 25% (6/24) AZIN1wt tumors showed overexpression of ADAR comparing to their normal adjacent counterparts (fold change > 2), respectively (Figure 1(b)). Notably, the AZIN1edited samples showed upregulated expression of ADAR in accordance with sample showed AZIN1 overexpression, implication of their potential regulatory relationship.
AZIN1 RNA editing induced the malignant phenotype of NSCLC models
To investigate the functional impact of RNA editing of AZIN1 in NSCLC development, we introduced GFP-labeled wild-type AZIN1 (AZIN1wt) and edited AZIN1 (AZINedited) expression constructs into two NSCLC cell lines (A549 and H1299) without AZIN1 RNA editing. Cells were transduced and overexpressed the edited AZIN1 (A549 AZIN1edited and H1299 AZIN1edited) and wild-type AZIN1 (A549 AZINwt and H1299 AZINwt), which were verified by quantitative PCR (Figure 2(a)) and western blot (Figure 2(b)) in RNA and protein levels, respectively, and the AZIN1 editing site was validated by Sanger sequencing as well (Figure 2(c)).

The validation of manipulating AZIN1 RNA editing in NSCLC cell lines. (a) Quantitative real-time PCR verifies wild-type AZIN1 (AZIN1wt) and edited AZIN1 (AZIN1edited) transfected and expressed in the indicated cells. (b) Western blot verifies AZIN1 wt and AZIN1 edited protein expression in the indicated cells using β-tublin as control. (c) The sequence chromatograms of the AZIN1wt and AZIN1edited transcript by RT-PCR and Sanger sequencing, and A549 was used as an example.
First, we found that the AZIN1edited significantly accelerated NSCLC cell proliferation than cells transduced with the AZIN1wt lentivirus (A549 AZIN1edited vs control, p = 0.0179; H1299 AZIN1edited vs control, p = 0.0206; Figure 3(a)). Next, the effect of cell-matrix adhesion was assessed using an in vitro adhesion assay. The absorbance of adhered AZIN1edited cells was significantly increased, compared to control cells (A549 AZIN1edited vs control, p = 0.0362; H1299 AZIN1edited vs control, p = 0.0203; Figure 3(b)); while the absorbance of AZINwt cells was also higher than it in control cells, which was also significant (A549 AZINwt vs control, p = 0.0142; H1299 AZIN1wt vs control, p = 0.0026). Then, the invasion potential of cells transfected with AZIN1wt, AZIN1edited, and the scramble RNA was examined by a Transwell system. The results indicated a significant increase in the number of cells migrating through Matrigel when the primary NSCLC cells were transfected with AZIN1edited, whereas those with AZIN1wt showed no apparent difference, compared to the control (A549 AZIN1edited vs control, p = 0.0107; H1299 AZIN1edited vs control, p = 0.0135; Figure 3(c)). In line with the clinical features of patients carrying the RNA editing, the AZIN1edited cells had increased capacity of migration than AZIN1wt cells (Figure 3(d)). The healing and migration speed of AZIN1edited cells after wounding was obviously higher compared to the control cells. More than three points at the front edge were measured under microscope and converted to the migrated distance based on Time 0. The AZIN1edited cells migrated 300–350 µm at the final time point (12 h), while the highest migration distance of control cells was ~240 µm.
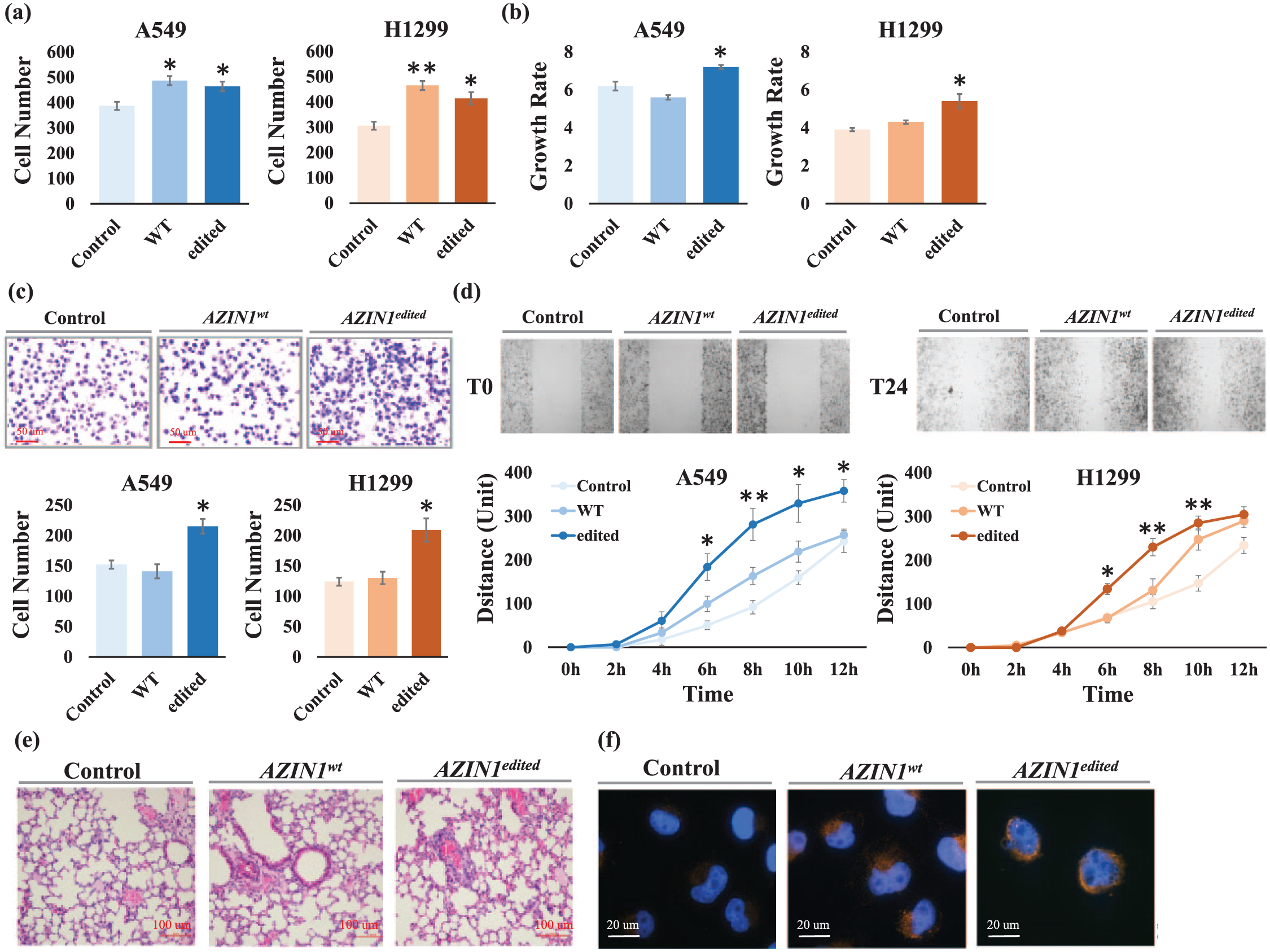
The edited AZIN1 confers tumorigenic phenotypes in NSCLC models. (a) The adhesive capacity was increased in AZIN1 overexpressed A549 and H1299 cells, compared to control cells. (b) The growth rate was markedly increased after the AZIN1 editing in NSCLC cells, compared to empty control cells on Day 3. (c) The invade capacity of NSCLC cells was significantly enhanced after AZIN1 editing. (d) The migration speed of NSCLC cells was markedly increased after AZIN1 editing. For (a)–(d), three independent experiments were performed (each with n = 3); *p < 0.05, **p < 0.01, unpaired Student t test, and standard error of mean (SEM) were depicted in bar and line charts. (e) The HE staining of mice models shows the tumor cells with AZIN1edited protein forming more lung nodules compared with control. Scale bar = 100 µm. (f) As GFP-tagged wild-type (wt) and edited (edt) AZIN1 was used to be expressed in A549 and H1299 cells, the overlay of the Alexa Fluor® 594-labelled AZIN1 and the nuclear DAPI (blue) staining of the same field was shown in the images (scale bar = 10 µm).
Moreover, xenograft studies in mice demonstrated that 4 weeks after intravenous inoculation, tumors derived from AZIN1edited A549 cells formed more lung nodules, which illustrated that the capacity of migration was higher than those derived from AZIN1wt A549 and non-transfected cells in vivo (Figure 3(e)). All these data indicate that the AZIN1 editing acts as a “gain-of-function” way and promotes the tumorigenicity during NSCLC progression.
To further investigate the underlying mechanism, we examined the AZIN1 protein expression in NSCLC cells before/after editing. According to the immunofluorescence colony (IFC) imaging, the wild-type AZIN1 protein was dominantly located in cytoplasm, whereas edited AZIN1 protein was abundant in the nuclei than cytoplasm, indicating that the AZIN1 editing results in a cytoplasmic-to-nuclear translocation (Figure 3(f)), which is consistent with the findings reported in liver cancer. 13 The authors proved that the edited AZIN1 promotes cell proliferation through the neutralization of antizyme-mediated degradation of ornithine decarboxylase (ODC) and cyclin D1 (CCND1), and lung cancer cells may also take advantage of this molecular machinery.
Discussion
RNA editing is considered as a common and essential RNA modification in mammals, with millions of editing sites catalogued so far in human samples. 20 It is reasonable that dysregulation of a particular RNA editing may give rise to tumorigenesis by either inactivating a tumor suppressor or activating genes that promote tumor progression. In fact, many genes in which RNA editing occurs have been connected with multiple malignant phenotypes, and increased editing activity is associated with poor prognosis. 11 For example, RNA editing of BLCAP and FLNB functions within hepatocellular carcinoma,21,22 RHOQ in colorectal cancer, 23 CDC14B in glioblastoma, 24 and under-editing of GluR-B in glioma. 25 Especially, AZIN1 is a well-characterized RNA editing target in liver and esophageal cancer.13,14 In our study, we revealed that the specific A-to-I RNA editing in AZIN1 also was a significant molecular aberration in NSCLC. Although A-to-I RNA editing is not frequently occurred in patients with NSCLC, 10 comparing with various human cancers, our findings have highlighted that the RNA editing is one of the molecular machineries occurring in NSCLC tumorigenesis and cannot be neglected in lung cancer research. The authors demonstrated that lung adenocarcinoma is an over-editing cancer type, and more RNA editing events occurred in lung adenocarcinoma than in squamous cell carcinoma; 10 however, for the particular RNA editing sites in AZIN1, we did not observe preference between those two histological types. Similar to many other cancers, we found ADAR1 overexpression among NSCLC tumors, in accordance with Paz-Yaacov et al.’s 11 report. It is associated with RNA editing of AZIN1, which has also been observed in liver and esophageal cancer.13,14 However, we did not see a correlation between level of edited AZIN1 and mRNA expression level of ADAR. In addition to genomic variations, RNA editing is another source for transcriptome diversification in cancers, and it remains unknown whether other RNA editing events play a causative role in NSCLC progression. Future works, such as modulating the expression of RNA editing enzyme ADARs by overexpressing or silencing ADARs and reinstating specific hyper-edited or under-edited transcript, are required to explore this question.
The protein encoded by AZIN1 gene belongs to the antizyme inhibitor family, which plays a role in maintaining polyamine homeostasis. The polyamines are important for a variety of cellular functions, including cell growth. 26 Antizyme inhibitors are homologs of ODC (ODC, the key enzyme in polyamine biosynthesis) that have lost the ability to decarboxylase ornithine; however, they retain the ability to bind to antizymes. 27 Former study found that decreased ODC is associated with prostate cancer. 28 The gene encodes AZIN1, the first member of AzI family that is ubiquitously expressed. It was reported previously that cells overexpressing AzI exhibited improved capacity of proliferation and gave rise to tumors when injected into nude mice, 29 whereas knockdown of AzI using small interfering RNA (siRNA) or short hairpin RNA (shRNA) decreased cell proliferation both in vitro and in vivo.30,31 Interestingly, although the researchers were not aware of the sequence changes in AzI, they found that its expression was substantially elevated in cancers of the prostate, brain, breast, and liver. 32 Until recently, Chen et al. 13 reported that RNA editing was a novel unusual mechanism for antizyme-AZIN1 inter-regulation. Similarly, we found that AZIN1 mRNA expression is upregulated in lung cancer and in association with the RNA sequence change. Within NSCLC cell lines, the in vitro and in vivo data demonstrated that AZIN1 RNA editing significantly induced the malignant phenotypes of cancer cells, including proliferation, invasion, and migration. In addition, we found AZIN1 protein translocated from cytoplasm to nucleus through RNA editing modification. So, as is illustrated in liver cancer, A-to-I recoding of AZIN1 may confer a gain of function that drives NSCLC tumorigenesis through blocking antizyme-mediated degradation of ODC and CCND1. In summary, our study revealed AZIN1 RNA editing and its involvement in tumorigenesis of NSCLC, which could be a valuable prognosis predictor and a potential therapy target.
Footnotes
Author contributions
JL and XH conceived and designed the experiments; JC, XS, FF, YC, and YC performed the experiments; KWL analyzed the data; XH wrote the paper; and all the authors have read and approved the final manuscript to be publish.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All applicable international guidelines for the care and use of animals were followed. All procedures performed in studies involving human cohorts were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is supported by the National Natural Science Foundation of China (81672270, 81402300) and Guangdong Province Natural Science Foundation (2015A030313474, 2015A030310528).
Informed consent
Informed consent was obtained from all individual participants included in the study.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
