Abstract
Cervical carcinoma is a frequent malignancy in developing countries despite being a preventable disease. For the first time, four screening tests were used simultaneously for identifying women with a risk of developing cervical cancer, to help clinicians and policy makers to implement the best strategy for reducing the burden of this disease. Women visiting a hospital in India were enrolled after institutional ethics clearance and informed consent. Visual inspection using acetic acid and Pap smear tests were performed on 2683 women, and 104 had abnormal cytology: atypical squamous cells of undetermined significance (n = 29), low-grade squamous intraepithelial lesion (n = 41), high-grade squamous intraepithelial lesion (n = 17), and squamous cell carcinoma (n = 17). These and 96 samples, with normal cytology, were subjected to high-risk human papilloma virus testing and fluorescent in situ hybridization evaluation. Women with abnormal cytology were followed for 5 years and evaluated with colposcopy-guided biopsy. Three accepted methods of screening and one novel fluorescent in situ hybridization assay were carried out in 200 cases. Cutoffs for fluorescent in situ hybridization were established. The screening methods had 88%–96% negative predictive value, while positive predictive value was low (20%) for visual inspection using acetic acid, 47% for fluorescent in situ hybridization, 56% for high-risk human papilloma virus, and 73% for combined high-risk human papilloma virus and fluorescent in situ hybridization. Combined high-risk human papilloma virus and fluorescent in situ hybridization had 94% sensitivity, specificity, and negative predictive value, suggesting that simultaneous screening with these two tests is appropriate for identifying women progressing to cervical cancer and not visual inspection using acetic acid, which has low positive predictive value and Pap cytology which requires to be repeated. Policy makers and clinicians can assess feasibility of incorporating this screening strategy to prevent cervical cancer.
Keywords
Introduction
Carcinoma of the uterine cervix is the most frequent gynecological malignancy diagnosed in women from developing countries despite being a potentially preventable disease. Globally, it is the second most frequent cancer affecting women with a steady rise in incidence among younger women. The implementation of a regular cytology-based screening program, or annual repeated Pap smear analysis between the ages of 21 and 65 years, has greatly reduced the incidence and mortality of cervical cancer in developed countries. 1 The absence of such a large-scale program in a low-resource country like India is responsible for its high incidence and mortality. It is a fact that only 10% of the Pap smear–screened population exhibits a cytological abnormality and only 15% of these cytological dysplastic lesions progresses to cancer.2,3 Apart from this, cytological examination needs to be repeated annually and the test result is available after 3–5 days making this a cumbersome method for countries with a large population. Hence, it has been debated that visual inspection using acetic acid (VIA), which is an inexpensive method of identifying an abnormal cervix, may be recommended as a screening method in developing countries like Africa, China, and India. 4
It is well established that human papilloma virus (HPV) plays a role in the etiology of cervical cancer, and detection of this in cells obtained from the endocervix was considered relevant for assessing cases which progress to cancer. 5 But, persistent infection with only 13 of the known 130 HPV subtypes is likely to progress to cervical cancer;6,7 therefore, not just HPV detection, but its subtyping to identify high-risk HPV (hrHPV) types is important to identify women at risk of getting this cancer. Literature indicates that less than 10% of women with hrHPV strains like 16, 18, and 31 will progress to develop the malignancy. 5 Hence, the new guidelines for cervical cancer screening by American College of Gynecologists (ACOG) recommend a lengthened 5-year interval screening for women aged between 30 and 65 years with a combination of Pap smear and HPV typing. 8
Chromosomal abnormalities such as amplifications, deletions, and rearrangements of whole or parts of chromosomes are a hallmark of cancer and have been described in many tumor types.9–11 The common chromosomal abnormalities identified in cervical cancer are gain of chromosomal regions 3q26 followed by 5p15, 20q13, and cen 7.12,13 A new fluorescent in situ hybridization (FISH)-based test to identify these specific alterations has been developed. 9 However, field data of this test are very limited and its potential as a screening tool is not evaluated yet.
The aim of this study is to use all the above-mentioned four tests for identifying women with a high risk of developing cervical cancer and thereby to help clinicians and policy makers to implement the best strategy for reducing the burden of this disease.
Study design
Patient study group and specimen collection
During the study collection period from May 2010 to April 2011, 2683 women underwent gynecologic examination either for a routine healthcare check-up or seeking care for a specific health condition. Institutional Ethics Committee approved this study. Clinical details, personal history, and sociodemographic details were recorded in a well-designed case proforma. Gynecologic examination included visual inspection using 3%–5% acetic acid (VIA) which was carried out by a senior gynecologist with documentation of acetowhite areas. 14 A cervical cytology specimen was obtained, using a spatula, from each woman, and 15 min after VIA, three smears were made on glass slides, two for cytology review and one reserved for FISH, pending the cytology result. After routine Pap staining, all cytology slides were reviewed by experienced pathologists and classified based on the Bethesda system. 15 Remaining endocervical material on the collection device was repeatedly washed off with phosphate-buffered saline into a tube for pelleting for HPV detection.
HPV detection and subtyping
DNA was isolated from pelleted endocervical material, using the salting out technique after standardizing the protease treatment as discussed earlier by our group. 16 Polymerase chain reaction (PCR) was performed on the DNA using the primers PGMY09 and PGMY11 from the consensus L1 region of the HPV genome, 17 followed by electrophoresis. 16 Cases showing a band in the expected size range were considered positive for HPV and subjected to subtyping for high-risk subtypes using PCR-based screen assay (Sacace Biotechnologies, Italy) following the manufacturer’s instructions.
FISH
The cervical smears on slides reserved for FISH were placed in Carnoy’s fixative (methanol:acetic acid, 3:1) for 10 min and air-dried. Slides were aged at 37°C overnight prior to carrying out FISH using a four-color FISH-based HPV Associated Cervical Cancer Test (FHACTTM) DNA-FISH probe (3q26 (red), 5p15 (green), 20q13 (gold), and cen7 (aqua; Cancer Genetics, Inc., USA) by a new method standardized by us. Briefly, the slides were dehydrated (70%, 85%, and 100% alcohol) for 5 min each at room temperature and air-dried. Following denaturation of the slides, they were hybridized with 10 µL of FISH probe (FHACT) at 80°C using 35% formamide solution in 2× saline sodium citrate (2×SSC) and hybridized for 18 h at 37°C. Slides were washed briefly in 2×SSC (pH 7.0), three times in 2×SSC/0.1% Tween-20 at 45°C for 5 min each, and finally rinsed in distilled water. After air-drying, DAPI/Antifade (1:10, VECTASHIELD; Vector Laboratories, USA) was applied to the hybridized area. Interphase FISH signal viewing and enumeration were performed using the Olympus BX41 system (Olympus America Inc., USA) and ASI software (Applied Spectral Imaging, USA) equipped with tetramethylrhodamine isothiocyanate (TRITC), fluorescein isothiocyanate (FITC), AQUA, and GOLD filters (Chroma Technologies, USA). For each specimen, the signal patterns in up to 2000 nuclei were enumerated and recorded, without bias of nuclei scored with respect to size or nuclear membrane appearance. For each locus, nuclei with three or more signals were considered positive for gain and expressed as a percentage of total nuclei scored. The cutoff values for each locus are 3q26, >0.62%; 5p15, >0.32%; 20q13, >0.22%; and cen7, >0.08%.
Results
Study group and demographics
For the 11-month duration of the study period, 2683 women were included whose reproductive details are provided in Table 1. As part of the study, the women underwent VIA and three cytology smears were collected, after which excess cells were washed off and stored for DNA extraction. Of the total women screened, 104 showed an abnormal cytology comprising atypical squamous cells of undetermined significance (ASCUS, n = 29), low-grade squamous intraepithelial lesion (LSIL, n = 41), high-grade squamous intraepithelial lesion (HSIL, n = 17), and squamous cell carcinoma (SCC, 17). The ASCUS and LSIL were categorized together in a group as <HSIL for all analyses. These cases and an additional 96 randomly selected samples with normal cytology were then subjected to HPV testing and FISH evaluation. The details of these 200 cases are also given in Table 1 for comparative purposes and to ensure that the case dataset is representative of the entire study group. No significant differences were found for all demographic features between the selected 96 cases with normal cytology and the entire 2683 dataset. However, a noticeable increase in current age was evident in women with increasing cytologic severity, which was similarly reflected by years of sexual activity and parity (Table 1).
Reproductive details of all women assessed during the study period, including the total 2683 women screened by VIA and Pap smear, as well as 200 samples selected for performing all four screening tests based on cytology result.

NILM: negative for intraepithelial lesion or malignancy; HSIL: high-grade squamous intraepithelial lesion; SCC: squamous cell carcinoma.
HPV subtypes and chromosome gain in abnormal and normal cytologies
All cases with an abnormal cytology (n = 104) and the 96 selected representative negative for intraepithelial lesion or malignancy (NILM) cases were subjected to HPV testing, first for the presence of HPV by PCR using primers across the L1 consensus region. By this method, 90 cases of the 200 were positive for the presence of the HPV genome (45%). HPV genotyping for 12 hrHPV types (16, 18, 31, 33, 35, 39, 45, 52, 56, 58, 59, and 66) was then performed on these samples, 35 (17.5%) of which tested positive for hrHPV. Table 2 shows the distribution of the hrHPV subtypes found across the cytology groups. As expected, 16 of the cervix cancer samples were positive for hrHPV (94%), but only 10 were positive for HPV16 and 18 (62.5%), while the remaining 337.5% had HPV 59, 31, and 33 sub types. Of the HSIL cases, 24% were positive, as were 17% of the abnormal cases with <HSIL; 3 of the 96 NILM cases were positive for hrHPV (3%), thus as expected with increasing cytologic severity, there was an increase in the proportion of cases positive for hrHPV.
Distribution of hrHPV genotypes and chromosome gain in normal and abnormal cytology specimens.
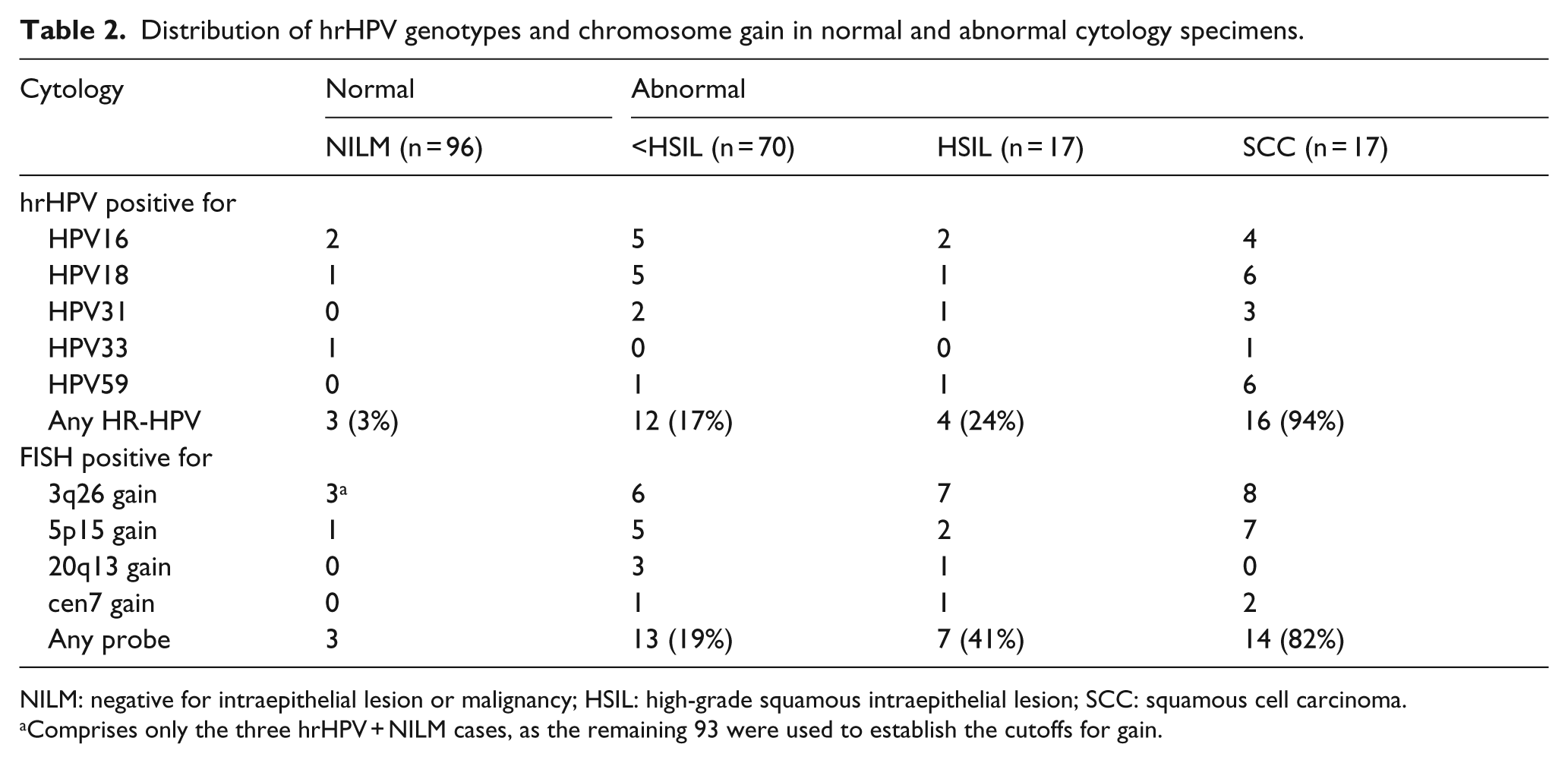

NILM: negative for intraepithelial lesion or malignancy; HSIL: high-grade squamous intraepithelial lesion; SCC: squamous cell carcinoma.
Comprises only the three hrHPV + NILM cases, as the remaining 93 were used to establish the cutoffs for gain.
All 200 cases were also subjected to FISH on the smear collected to assess clonal gain of four chromosomal loci commonly detected in HPV-associated cancers. 9 For each locus, nuclei with more than two signals were considered positive for gain. Although there were nuclei showing more than three signals for 3q26, they were all classified as >3 signals indicating the copy number variability. Figure 1 shows representative images for FISH signal pattern. A median of 2000 nuclei were enumerated per case (range = 1300–2000). The 93 NILM cases that were negative for hrHPV served to establish cutoffs for each locus: 3q26, >0.62%; 5p15, >0.32%; 20q13, >0.22%; and cen7, >0.08%. Using these cutoffs (mean plus 3 standard deviation (SD)), the remaining 107 cases were called positive or negative for clonal gain of each locus. Table 2 lists the results where it was evident that 3q26 was the most frequently gained genomic locus, followed by 5p15, 20q13, and cen7. Across all loci, chromosome gain was observed to increase with cytologic severity, as consistently reported in other studies.9,10
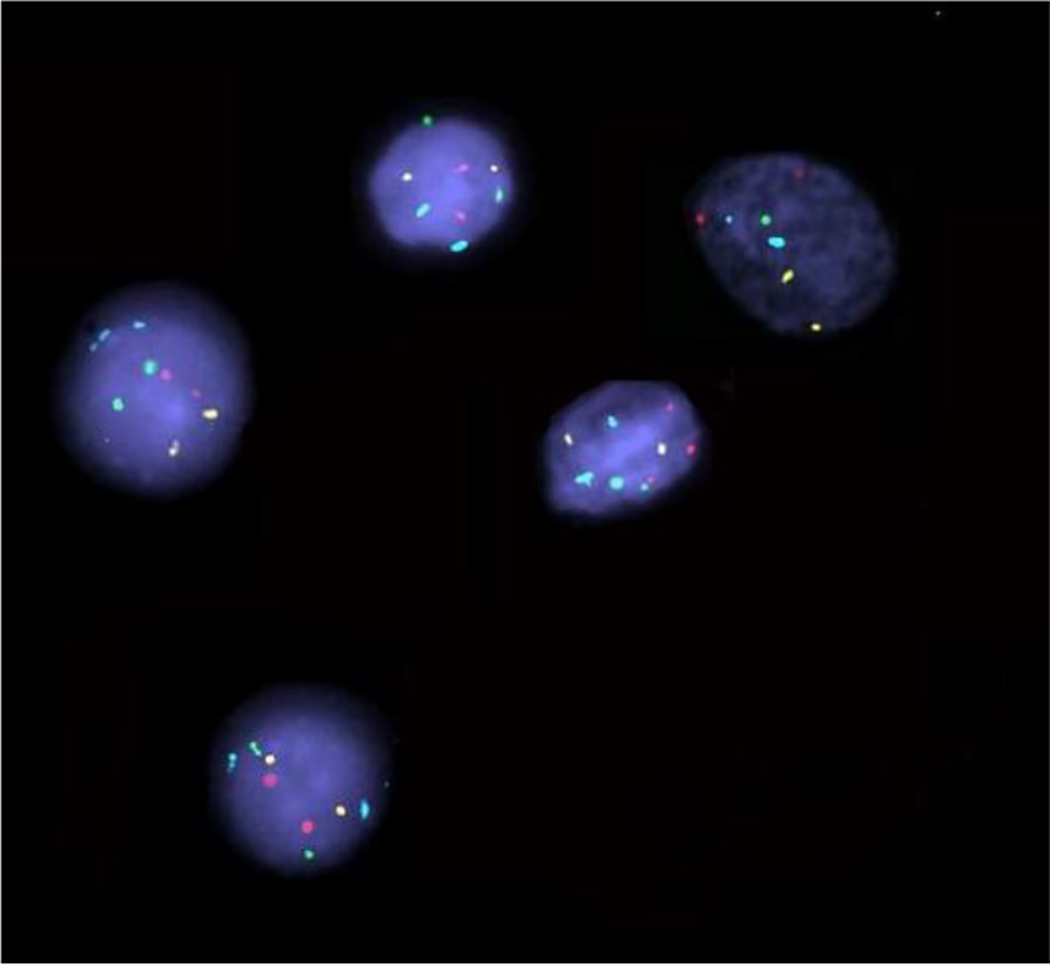
Representative FISH images showing nuclei with normal signal pattern with two signals for each region (2 red—3q26, 2 green—5p15, 2 gold—20q13, and 2 aqua—cen7) and a nucleus with gain of red signal—3q26 (three signals) in the center.
For 180 of the 200 cases, the results of all four screening tests were available. VIA was not performed for the women with SCC identified during the initial inspection, and three women with NILM were not cooperative. Figure 2 shows the percent of cases according to cytology group that were positive for VIA, hrHPV, and FISH (any locus). As described above, since the FISH cutoffs were established on the 93 NILM (hrHPV−), no results are shown for NILM other than VIA and hrHPV. Similarly, results for VIA are not available for SCC as applying acetic acid to a cervix bleeding on touch is not an acceptable clinical practice. The percent of women positive by VIA increased with cytologic severity but 43% of those women with NILM were VIA positive. In cases with <HSIL, 18 cases were positive for hrHPV or FISH, and of these, 7 (10%) were positive by both assays (Figure 2). For abnormal cytology like HSILs, all four cases that were hrHPV+ were also positive for chromosome gain by FISH.

Combined chart for VIA, hrHPV, and FISH positivity in all cytological categories.
Correlation with colposcopy-directed biopsy histology
The 17 SCC cases were referred to Oncology Unit for further management. As part of routine care in a 5-year follow-up period, 65 of the 87 women with abnormal cytology (12 HSIL and 53 <HSIL) had colposcopy-guided biopsy that comprised 54 <cervical intraepithelial neoplasia grade 2 (CIN2) and 11 CIN2+. Table 3 lists the calculated performance measures for sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) for the detection of high-grade cervical disease (CIN2+). Cytology was excluded due to the inclusion in this analysis only of cases with an abnormal cytology (ASCUS and LSIL—combined as <HSIL and HSIL). It is evident from these data (Table 3) that the NPV for all three screening methods is high (88%–96%), but the PPV is very low for VIA (20%) and slightly better for FISH (47%) and hrHPV (56%), but combined hrHPV and FISH screening gives the best PPV (73%). Even sensitivity and specificity reached 94% when both hrHPV and FISH tests are carried out together.
Performance measures of VIA, hrHPV testing, and FISH to detect high-grade cervical disease (≥CIN2) in 65 cases with abnormal cytology and follow-up colposcopy-directed biopsy within 5 years of detecting abnormal cytology.

PPV: positive predictive value; NPV: negative predictive value; VIA: visual inspection using acetic acid; hrHPV: high-risk human papilloma virus; FISH: fluorescent in situ hybridization.
Discussion
Cervical cancer prevention program is extremely important for developing countries especially India where the incidence and mortality of this disease are very high. The burden of cervical cancer patients in India is alarming when compared to developed countries where there are established screening programs which have effectively reduced the incidence of cervical cancer with repeated annual Pap smear evaluation from the age of 30 years. 18 Such a cytology-based screening is not feasible in many developing countries owing to limited resources, lack of trained personnel, and a large uneducated population which fails to report for annual screening. 1 Apart from this, ASCUS and LSIL cytologies (<HSIL) can revert to normal and cannot be used as ideal indicator of progression to cervical cancer. Hence, it is important to identify other screening modalities and develop a feasible program for screening women with a high risk of progression to cervical cancer. In this study, for the first time, four screening tests were used simultaneously to identify women at risk of developing cervical cancer.
VIA has been recommended as a suitable screening test for low-resource countries.4,19 Our study showed that despite being a sensitive test, its PPV is only 20% which will lead to unnecessary referrals and follow-up, thereby defeating the purpose of identifying at-risk individuals. Similar results of Tota et al. 18 and Basu et al. 19 also suggest that VIA is a suboptimal screening method. Therefore, using it as an isolated screening tool may not be sufficient.
In this study, hrHPV was positive in 94.11% women with squamous cell cancer of cervix and 24% with HSIL and 17% with <HSIL. Women with normal Pap smears showed 3% HPV positive cases. Since the PPV of the presence of hrHPV is 56% and its specificity 87%, identifying the presence of high-risk subtypes may be a valuable screening method (Table 3), and some developed countries have included it in their screening programs, thereby reducing the frequency of screening women from annually to once in 3 years.
HPV subtyping results also showed that in our population, high-risk subtypes other than HPV 16 and 18 are also prevalent in SCC, these include HPV 31, 33, and 59 in this study (Table 2), with HPV 59 having the same prevalence frequency as HPV 18. Another study in a North Indian population reported HPV high-risk subtypes like 62, 71, and 63 along with HPV 16 and 18. 20 These results indicate that there is a need to do more regional studies to identify hrHPV subtypes prevalent in different parts of India and develop a hrHPV panel suitable for country-/region-specific screening. This will in future help in the HPV vaccination program, but that is a discussion suitable for another publication.
Genomic alterations specific to cervical cancer established by array comparative genomic hybridization 9 were evaluated in samples by FISH analysis. Other studies have indicated that 3q26, 5p15, 20q13, and Cen 7 have relevant genes located, which are involved in the progression of cervical cancer. 21 This is the first study of its kind to evaluate the four genomic regions responsible for cervical cancer in the population, using FISH technique on a direct Pap smear slide obtained during first visit as a screening tool.
Majority of FISH positive cases exhibited 3q26 gain followed by 5p15 and 20q13. Evaluating the samples using all four regions simultaneously increases the sensitivity of FHACT probe for cervical cancer screening as also indicated by Luhn et al. 9 Of the 87 women with abnormal cytology who were followed up for a period of 5 years, 65 showed a progression to CIN1+ and CIN2+ in those cases which were FISH positive. Interestingly, it was observed that combined hrHPV and genomic amplification in any of the four chromosomal regions by FISH evaluation has a 94% sensitivity and specificity. Thus, these two tests are good indicators of cervical cancer progression.
The limitation of our work is that the women with normal cytology who showed the presence of hrHPV and/or FISH positivity were not followed up. Such a study would help to establish non-equivocally that simultaneous screening with hrHPV and FISH would be the appropriate modality for screening, instead of repeated VIA or pap cytology. However, based on our present data, it may be recommended that hrHPV testing combined with FISH positivity has a high PPV when compared to the current available screening tests. This will help the policy makers and clinicians in assessing the feasible incorporation of these tests in the current screening strategy to prevent cervical cancer.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Pavani Upendram received fellowship support from Kamineni Hospitals. Vasavi Mohan received SR/FT/LS-106/2009 DST Grant for HPV work. Cancer Genetics Inc. partially supported the FISH work.
