Abstract
Chronic inflammation increases the risk of development of various cancers, including colorectal cancer. Interleukin-6 has been described as a key regulator of colorectal cancer development and is important in the process of colorectal tumorigenesis largely through the regulation of tumor-promoting inflammation. Several studies have reported the association of various polymorphisms in human interleukin-6 gene including IL-6 −174G/C single nucleotide polymorphism with various cancers, including colorectal cancer, but the results are mixed and inconclusive.
The aim of this study was to analyze the association of IL-6 −174G/C promoter single nucleotide polymorphism with colorectal cancer risk and also to evaluate the modifying effects of possible IL-6 −174G/C single nucleotide polymorphism genotypes on different risk factors of colorectal cancer or the reciprocal effect in ethnic Kashmiri population through a case control setup. The genotype frequencies of IL-6 −174G/C promoter single nucleotide polymorphism were compared between 142 colorectal cancer patients and 184 individually matched healthy controls by using polymerase chain reaction–restriction fragment length polymorphism method. The association between the IL-6 −174G/C single nucleotide polymorphism and colorectal cancer risk was examined through conditional logistic regression models adjusted for multiple possible confounding (third) variables. The possible effect measure modification of the association between the relevant single nucleotide polymorphism genotypes and colorectal cancer risk by various colorectal cancer risk factors including age, gender, and smoking status was also evaluated. Furthermore, the associations between these single nucleotide polymorphisms and various clinicopathological parameters, demographic variables, and environmental factors within the case group subjects with regard to colorectal cancer risk were also analyzed.
The overall association between the IL-6 −174G/C single nucleotide polymorphism and the modulation of colorectal cancer risk was found to be highly significant (p = 0.001). The variant genotype (CC) was significantly associated with a decreased risk of colorectal cancer (odds ratio, 0.15 (95% confidence interval, 0.04–0.54); p = 0.004). Furthermore, the less common IL-6-174C allele was associated with a decreased risk of colorectal cancer (odds ratio, 0.49 (95% confidence interval, 0.33–0.73); p = 0.0006). The combined variant genotype (GC + CC) was also significantly associated with a decreased risk of colorectal cancer (odds ratio, 0.54 (95% confidence interval, 0.33–0.89); p = 0.015).
This study demonstrates that there is a strong and highly significant association between the IL-6 −174G/C promoter single nucleotide polymorphism and a decreased risk of colorectal cancer in ethnic Kashmiri population. However, in order to substantiate our findings, this study needs to be replicated with larger sample size and with other ethnically defined populations with comparable colorectal cancer incidence.
Keywords
Introduction
Colorectal cancer (CRC) includes the cancers of the colon, rectum, and appendix, and is one of the most common cancers and leading cause of cancer-related mortality and morbidity worldwide. Globally, it is the third most frequent cancer among the men and second among the women, and is the third most frequent cause of cancer-related mortality with nearly 1.2 million new cases diagnosed each year and approximately 600,000 deaths reported yearly.1,2 The Kashmir Valley, part of Jammu and Kashmir State, located in Northern India has a unique ethnically defined population with highly distinctive food habits compared with the rest of the world. In Kashmir Valley, CRC represents the third most common gastrointestinal (GI) cancer after esophageal cancer and gastric cancer 3 and is the fourth common cancer among the males and the third among the females.3,4
The role of chronic inflammation in the pathogenesis of various cancers is well established. The cancer-related chronic inflammation is now regarded as an important mediator of cancer and is known to influence and predominantly promote tumor growth, survival, progression, angiogenesis, invasion, and metastasis.5,6 The role of chronic inflammation as an essential tumor initiating and/or promoting event has been established in about 25% of all tumors. 7 The CRC is prominent among the cancers in which inflammation is present as an important pathological factor at the earliest stages of tumor progression and is capable of transforming the pre-neoplastic lesions characteristic of colorectal tumorigenesis into aggressively invasive tumors.8,9 Cytokines, both pro-inflammatory and anti-inflammatory, play a vital role in the mediation and regulation of immune response, including inflammation, and they are equally important in the process of tumorigenesis largely through the regulation of tumor-promoting inflammation. 10
Interleukin-6 (IL-6) is a pleiotropic cytokine involved in T-helper 2 (Th2)-type of humoral immune response and exhibits both pro-inflammatory and anti-inflammatory activities. 11 It plays an essential role in various cellular processes, including cell proliferation, differentiation, apoptosis, and angiogenesis, and is involved in various physiological functions, including bone metabolism, hematopoiesis, endocrine activity, and the regulation of immune response, including acute-phase response and balance between pro-inflammatory and anti-inflammatory pathways. 12 IL-6 plays an essential role in tumor development, progression, and differentiation, and modulation of tumor microenvironment which also promotes metastasis.11,13,14 These precancerous activities involve increased angiogenesis through enhanced expression of growth factors and adhesion receptors on endothelial cells, and inhibition of tumor cell apoptosis by promoting the expression of or delaying the degradation of anti-apoptotic factors. 15 Higher expression of IL-6 has been reported in almost every tumor type and has been associated with the entire process of tumorigenesis, including initiation, promotion, invasion, and metastasis.16,17 The elevated serum or circulating levels of IL-6 have been reported to correlate positively with advanced tumor stages, poor prognosis, and worse clinical outcome, and reduced overall survival in various cancers.11,16,18,19
IL-6 has been described as a key regulator of CRC development. 20 The elevated expression accompanied by increased serum and tumor tissue levels of IL-6 have been reported in CRC.21,22 Furthermore, the increased expression and serum and tumor tissue levels of IL-6 in CRC have been associated with increased tumor cell proliferation, advanced tumor stage, increased tumor size, increased angiogenesis and invasiveness, metastasis, poor prognosis, worse clinical outcome, and decreased overall survival.21,23,24 IL-6 has been reported to act as an autocrine growth-promoting factor for CRC cells both in vivo 21 and in vitro. 25 Furthermore, IL-6, mainly through the regulation of immune response, has also been reported to play an important role in the pathogenesis of inflammatory bowel diseases (IBDs), including Crohn’s disease (CD) and ulcerative colitis (UC). 26 IBD has an established relationship with an increased risk of CRC development.27,28 Increased IL-6 expression and elevated serum and colon tissue levels have also been reported in IBD. 29 It is clear from the above discussion that IL-6 plays an essential role in almost all aspects of colorectal tumorigenesis including the pathological conditions that increase predisposition to CRC, and therefore it is reasonable to argue that IL-6 is a potential risk modulatory or even causative factor for CRC that acts mainly through its modulation of immune response. The pro-tumorigenic immune functions mediated by IL-6 also establish it as an important link between chronic inflammation and tumor development.
The IL-6 gene is located on chromosome 7 at position p21–24 and its expression like that of other cytokines is regulated tightly and mainly at the transcriptional level. The promoter region of IL-6 contains a variety of DNA responsive or regulatory elements, including binding sites for nuclear factor interleukin-6 (NF-IL-6), nuclear factor-kappaB (NF-κB), activator protein 1 (AP-1), Fos/Jun, CAAT/enhancer-binding protein beta (C/EBP-beta), and glucocorticoid receptor. 30 The expression of IL-6 gene is tightly regulated and effectively modulated through the binding of various transcriptional factors to the specific responsive elements or sequences in the promoter region. The 5′ and 3′ flanking regions of IL-6 gene are highly polymorphic, and several pathogenic mutations (https://portal.biobase-international.com/cgi-bin/portal/login.cgi) and single nucleotide polymorphisms (SNPs) have been identified in the promoter region of this gene.31,32 The SNPs in the IL-6 gene promoter region may modify specific DNA responsive sequences and alter the binding of various transcription factors to these cis-acting elements which may result in differential gene expression and secretion of IL-6 and contribute to the individual-specific differences in relevant biological functions and subsequently the differences in susceptibility among different individuals to various diseases, including various cancers such as breast cancer, hepatocellular carcinoma, and CRC.33,34 Among various SNPs in IL-6 gene promoter, the functional IL-6 −174G/C (rs1800795) SNP involving G to C transition at −174 nucleotide (nt) position has been reported to be biologically and clinically important.
Several studies have shown the association of IL-6 −174G/C SNP with various cancers, including neuroblastoma, pancreatic cancer, prostate cancer, gastric cancer, hepatocellular carcinoma, multiple myeloma, ovarian cancer, gallbladder cancer, squamous cell esophageal cancer,35–43 and CRC.44–50 Some studies have reported no association of this SNP with various cancers.17,33,51,52
In this study, we systematically analyzed the possible association between IL-6 −174G/C SNP and susceptibility to CRC in Kashmiri population through a case control study. We also evaluated the possible effect modification of CRC risk by age, gender, and smoking status. Furthermore, we analyzed the possible relationship of this IL-6 SNP with various clinicopathological parameters, demographic variables, and environmental factors, including smoking habit, and evaluated their role in modulating the risk of CRC in the population under study.
Materials and methods
Study subjects
This study involved two subject groups: case and control. The case group included 142 patients recruited consecutively irrespective of their age and gender with primary CRC who underwent surgical resection for primary CRC tumors at the Department of General Surgery, Sher-I-Kashmir Institute of Medical Sciences (SKIMS), Srinagar, Kashmir. The diagnosis of CRC was confirmed histopathologically. The tumor stage and the tumor grade were classified according to the 8th edition of tumor–node–metastasis (TNM) classification of Union International Control of Cancer (UICC). Only those cases who had not received any neoadjuvant chemo- or radiotherapy were chosen for this study. All the cases were more than 18 years old and had no prior history of any malignancy. Blood and tissue samples were obtained from these CRC patients. The control group included 184 healthy individuals with no history or prior diagnosis of any malignant disorder or any other serious disease who were recruited during the same time period and from the same geographic area and from whom blood was collected and used as controls for this study. The control group included both general-population-based subjects and hospital-based subjects. The control group subjects were matched to the case group subjects individually for age (±5 years), sex, place of residence (rural/urban), smoking habit, and ethnicity in order to minimize the confounding effect of these various relevant factors. Both the case and the control subjects chosen for this study were ethnic Kashmiris.
Data collection
The data relevant to the study concerning all the CRC patients including various clinicopathological parameters, demographic variables, and the environmental factors were obtained and evaluated from the patient medical records (files), pathology reports, and also from the personal interviews with the patients and/or their guardians (for those who were illiterate or unable to communicate). The interviews were conducted in local language for easy and direct communication, which also helped to gather maximum possible relevant information. The data collected included tumor location, Dukes’ stage, lymph node status, age, sex, place of residence, ethnicity, smoking habit, and the family history of cancer, among several other potential confounding parameters. The relevant data were also obtained for each of the recruited controls mostly through personal interviews and included parameters like age, sex, place of residence, ethnicity, and smoking habit. The data collection was carried out by research professionals only in order to fulfill requisite quality standards during the course of this study. All the patients and/or their guardians were informed about the study and their willingness to participate in this study was documented using a predesigned questionnaire, and same procedure was followed for the controls. All the procedures concerning the study subjects including sample procurement and the data collection were carried out in accordance with the ethical standards laid down by the Institutional Ethics Committee (IEC), SKIMS, World Health Organization (WHO), and the Code of Ethics of the World Medical Association (WMA) 53 (Declaration of Helsinki, 1964, and its Seventh Amendment, 2013) for experiments in humans.
Sample preparation and DNA extraction
The tumor tissue samples collected after surgical resection were immediately snap frozen in liquid nitrogen and then stored at −80°C until further use for DNA extraction and other experimental purposes. Prior to DNA extraction, tumor tissue samples were washed two to three times in phosphate-buffered saline (PBS), and adipose and connective tissue portions if any were dissected away. Peripheral blood sample (3–5 mL) from each case and control group individual was collected by venipuncture into ethylenediaminetetraacetic acid (EDTA)-coated blood vacutainer collection tubes (purple capped tubes; ADS Hitech Polymers, India) and stored at −80°C until further use. The genomic DNA was extracted from both the tumor tissue and blood specimens using DNeasy™ Blood and Tissue Kit (catalog number 69504; QIAGEN, Germany) and Quick-gDNA™ MiniPrep kit (catalog number D3024; Zymo Research, USA) according to the manufacturers’ instructions. The extracted DNA was stored at −20°C until further use. The qualitative and the quantitative assessments of the extracted genomic DNA samples were carried out by absorbance measurements at 260 and 280 nm using ultraviolet (UV)-visible spectrophotometric analysis and also by agarose gel electrophoresis. The DNA extracted from blood samples of case and control group subjects was used for this study.
SNP analysis or genotyping
The IL-6 −174G/C (rs1800795) SNP was genotyped using polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) assay.
IL-6 −174G/C PCR
The PCR for IL-6 −174G/C SNP was carried out in a total volume of 25 µL containing 100 ng–1 µg of genomic DNA, 0.7–1 U Taq DNA polymerase with l× Standard Taq reaction buffer (New England Biolabs, UK), 2.1 mM MgCl2, 0.28 mM deoxynucleotide triphosphate mix (New England Biolabs), 0.56 µM forward and reverse oligonucleotide primers (Integrated DNA Technologies, India), and nuclease-protease free water (QIAGEN). In addition, Phusion DNA Polymerase with Phusion HF Buffer (New England Biolabs) were used alternatively and randomly instead of Taq DNA polymerase with standard Taq reaction buffer to check for any Taq polymerase–induced amplification errors.
The PCR conditions used for the amplification of IL-6 gene region containing −174G/C SNP were as follows: initial denaturation at 95°C for 6 min followed by 35 cycles of denaturation at 95°C for 45 s, annealing at 58°C for 60 s, and extension at 72°C for 60 s followed by a single final extension step at 72°C for 10 min. The oligonucleotide primers used for this amplification were 5′-ATGACTTCAGCTTTACTCTT-3′ (forward) and 5′-ATAAATCTTTGTTGGAGGGT-3′ (reverse). The desired PCR product obtained for IL-6 −174G/C SNP was 244 bp in size (see Figure 1 for PCR gel picture).

Electrophoresis of IL-6 −174G/C SNP PCR products on a 2.5% agarose gel.
Genotyping
The IL-6 −174G/C SNP was genotyped using the restriction enzyme Hin1II (NlaIII; Thermo Fisher Scientific, USA). The digestion was carried out, according to the manufacturer’s instructions, in a 30-µL reaction volume containing 10 µL of PCR product and 10 U of Hin1II enzyme and incubated at 37°C overnight. Wild genotype (GG) is cleaved by Hin1II enzyme yielding two fragments of size (216 and 28 bp), whereas the variant genotype (CC) yields three fragments in size (122, 94, and 28 bp). Heterozygous genotype (GC) yields four fragments in size (216, 122, 94, and 28). The digestion products of IL-6 −174G/C SNP were separated on a 4% agarose gel stained with ethidium bromide (HiMedia, India) to a final concentration of 0.5 µg/mL (see Figure 2 for RFLP digestion gel picture).

Electrophoresis of IL-6 −174G/C SNP genotyping by PCR–restriction fragment length polymorphism on a 4% agarose gel.
Quality control
The quality control included the assessment of genotyping errors including the false estimates of a particular allele frequency and the evaluation of the reproducibility of the genotyping done. For these assessments, approximately 10% of the patient and control samples selected randomly were re-genotyped. In addition, in each PCR-RFLP setup, previously amplified and genotyped samples representing different genotypic scenarios were included as a reference control. The genotyping reproducibility was very high for the IL-6 SNP studied with a weighted kappa coefficient of 0.98, which corresponds to a high concordance rate of 98%.54,55
Statistical analyses
The frequencies of genotypes and alleles for the SNPs under study were obtained through direct counting. The numbers and percentages were calculated and presented for each of the categorical variables along with means, standard deviations (SDs), median, and inter-quartile range for continuous variables. Conditional logistic regression analysis was carried out to calculate unadjusted and adjusted odds ratios (ORs) and corresponding 95% confidence intervals (CIs) to assess the possible association of the relevant SNP genotypes with CRC risk and to assess the possible gene–environment interactions, if applicable. The more common homozygous genotype and/or more common haplotype was used as the baseline or reference group in the conditional logistic regression model, and the genotypes in order of more common/wild homozygous, heterozygous, and less common/variant homozygous were, respectively, represented by the ordinal values 1, 2, and 3 during the statistical analyses. In order to eliminate the possible confounding (third) variables, the conditional logistic regression models were adjusted for the known CRC risk factors like gender, age, and smoking habit, and also with the place of residence. The comparison of the genotype and allele distributions of the SNP under study between the CRC case group and the control group using conditional logistic regression involved two-sided chi-square test. The possible effect measure modification or effect modification of the association between various genotypes relevant to the SNP under study and CRC risk by various risk factors including age, gender, and smoking status was also included in the conditional logistic regression models and analyzed. The correlation between the genotypes and the clinicopathological parameters, demographic variables, and environmental factors including smoking habit within the case group was analyzed by Fisher’s exact test. The fitness of the genotype distributions to Hardy–Weinberg equilibrium (HWE) for the allele and the genotype frequencies in the population under study was tested using the chi-square test. A two-sided probability value of or less than 5% (p ≤ 0.05) was considered statistically significant for all types of analyses. All statistical analyses were performed using IBM SPSS Statistics v21 software.
The effective sample size and the statistical power were computed using the “Genetic Power Calculator” developed by Purcell et al. (http://zzz.bwh.harvard.edu/gpc/). The statistical power of 80% is widely used in genetic association studies to avoid type II errors and to determine a cost-effective sample size under the assumption of 10%–25% variant allele frequency, 1:1 case-to-control ratio, and 5% type I error rate (α). We obtained a healthy power score of about 85% for the SNP under study in our case control study design with 142 case subjects and 184 control subjects.
Results
This study included a total of 142 CRC case subjects and 184 control subjects with prior consent of each participant. The rate of participation was 100% for both the study groups because only those study subjects were considered who fully agreed to participate in this study. This was done to ensure exhaustive data collection relevant to the study concerning all the CRC patients and control subjects. The frequencies of various clinicopathological parameters, demographic variables, and environmental factors in CRC case subjects and relevant parameters in control subjects are given in Table 1. The mean age of the case group subjects was 52.68 years; for males it was 54.70 years and for females it was 50.66 years, and the age range was 21–82 years. Furthermore, 53.52% (76/142) of case group subjects were >50 years old and 46.48% (66/142) of subjects were ≤50 years of age. This group consisted of 59.86% (85/142) males and 40.14% (57/142) females (male-to-female ratio = 1.49). The mean age of the control group subjects was 52.22 years; for males it was 53.64 years and for females it was 50.80 years, and the age range was 21–80 years. Furthermore, 50.54% (93/184) of subjects were >50 years old and 49.46% (91/184) of subjects were ≤50 years of age. The control group consisted of 55.43% (102/184) males and 44.57% (82/184) females (male-to-female ratio = 1.24). The difference in the distribution of gender and age among the case group and control group subjects was not statistically significant (p > 0.05; Table 1). The case group subjects consisted of 61.27% (87/142) rural residents and 38.73% (55/142) urban residents, and the control group consisted of 54.89% (101/184) rural residents and 45.11% (83/184) urban residents. Furthermore, the case group consisted of 56.34% (80/142) smokers and 43.66% (62/142) non-smokers, while the control group consisted of 51.09% (94/184) smokers and 48.91% (90/184) non-smokers. No statistically significant dwelling and smoking status–related differences were observed between the case and control group subjects (p > 0.05; Table 1).
General characteristics of study subjects.

SD: standard deviation; SEM: standard error of mean.
Various clinicopathological parameters, demographic variables, and environmental factors in colorectal cancer case subjects and relevant parameters in control subjects from Kashmir. Pearson’s chi-square test (χ2) was used to calculate the p values for categorical variables.
N denotes the number of subjects or individuals.
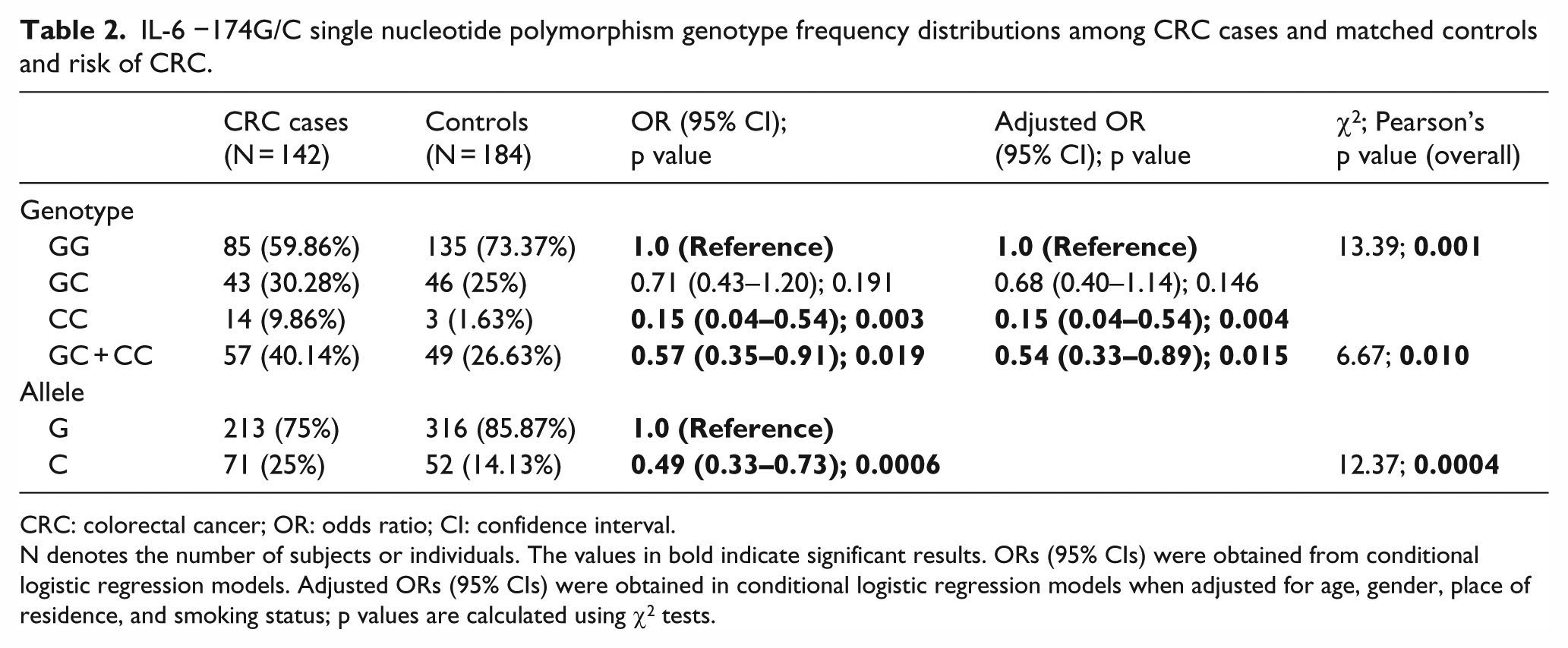
The frequencies of the genotypes of IL-6 −174G/C SNP for both the case and the control groups are listed in Table 2. The more common GG genotype of IL-6 −174G/C SNP was less frequent among the case group (59.86% (85/142)) in comparison to the control group (73.37% (135/184)). The frequency of the heterozygous genotype (GC) in the case group (30.28% (43/142)) was higher than that of the control group (25% (46/184)). The variant genotype (CC) also showed a higher frequency in the case group (9.86% (14/142)) in comparison to the control group (1.63% (3/184)). Furthermore, the frequency of the more common IL-6-174G allele was found to be 75% (213/284) among the case group subjects and 85.87% (316/368) among the control group subjects. The frequency of the less common IL-6-174C allele was found to be 25% (71/284) among the case group subjects and 14.13% (52/368) among the control group subjects. The frequency of the combined variant genotype (GC + CC) in the case group was found to be 40.14% (57/142) and among the control group subjects it was found to be 26.63% (49/184; Table 2). The overall association between the IL-6 −174G/C SNP and the modulation of CRC risk was found to be significant (p = 0.001; Table 2). The variant genotype (CC) was significantly associated with a decreased risk of CRC (OR, 0.15 (95% CI, 0.04–0.54); p = 0.004). Furthermore, the less common IL-6-174C allele showed an overall significant association with CRC (p = 0.0004). The IL-6-174C allele was associated with a decreased risk of CRC (OR, 0.49 (95%CI, 0.33–0.73); p = 0.0006). The combined variant genotype (GC + CC) also showed an overall significant association with CRC (p = 0.010). The combined variant genotype (GC + CC) was significantly associated with a decreased risk of CRC (OR, 0.54 (95% CI, 0.33–0.89); p = 0.015; Table 2). Furthermore, the genotype frequencies for IL-6 −174G/C SNP among the control group subjects were found to be in agreement with HWE (χ2 = 0.168; p = 0. 682) but not among the case group subjects (χ2 = 5.261; p = 0.022).
IL-6 −174G/C single nucleotide polymorphism genotype frequency distributions among CRC cases and matched controls and risk of CRC.
CRC: colorectal cancer; OR: odds ratio; CI: confidence interval.
N denotes the number of subjects or individuals. The values in bold indicate significant results. ORs (95% CIs) were obtained from conditional logistic regression models. Adjusted ORs (95% CIs) were obtained in conditional logistic regression models when adjusted for age, gender, place of residence, and smoking status; p values are calculated using χ2 tests.
The possible effect measure modification or effect modification of association between IL-6 −174G/C SNP genotypes and CRC risk by various CRC risk factors including age, gender, and smoking status is summarized in Table 3. This involved studying the effect exerted on the association between the more common or wild genotype (GG) and the combined variant genotype (GC + CC) and the CRC risk by these various risk factors. On analyzing the effect modification of IL-6 −174G/C SNP genotypes by age, gender, and smoking status, it was found that the effect of the combined variant genotype (GC + CC) on CRC risk was significantly dependent on gender (p = 0.022). A decreased CRC risk was observed in males (OR, 0.46 (95% CI, 0.24–0.89); p = 0.021). It was also observed that the effect of the combined variant genotype (GC + CC) on CRC risk was also significantly influenced by age (p = 0.037). However, the nature of this association could not be deciphered further in a statistically significant manner.
Effect modification of IL-6 −174G/C SNP genotypes in the presence of various risk factors of CRC in ethnic Kashmiri population.

SNP: single nucleotide polymorphism; CRC: colorectal cancer; OR: odds ratio; CI: confidence interval.
N denotes the number of subjects or individuals. The p values in bold indicate significant results. ORs (95% CIs) were obtained from conditional logistic regression models. Adjusted ORs (95% CIs) were obtained from conditional logistic regression models when adjusted for age, gender, place of residence, and smoking status. The variable under consideration was excluded in the time of analysis; p values are calculated using χ2 tests.
Wild refers to GG genotype and variant refers to GC + CC genotype.
The numbers and the frequencies of the subsets of various characteristics of the case group subjects under study, that is, age, gender, dwelling, smoking status, tumor location, tumor grade, and lymph node status for IL-6 −174G/C SNP, are listed in Table 4. We analyzed the correlation of the IL-6 −174G/C promoter SNP with the subsets of these various characteristics of the case group subjects. The IL-6 −174G/C SNP was significantly associated with age (p = 0.025). The risk of developing CRC in the subjects who carried the variant genotype (CC) increased with age (OR, 6.75 (95%CI, 1.42–32.02); p = 0.0087). The IL-6 −174G/C SNP was also significantly associated with lymph node status (p = 0.021). The non-involvement of lymph node was significantly greater in the subjects who carried the variant genotype (CC) of IL-6 −174G/C SNP (p = 0.0191; Table 4). The ORs are not shown in Table 4 for the sake of simplification of data presentation.
Association of IL-6 −174G/C polymorphism with various clinicopathological parameters, demographic variables, and environmental factors in CRC cases.
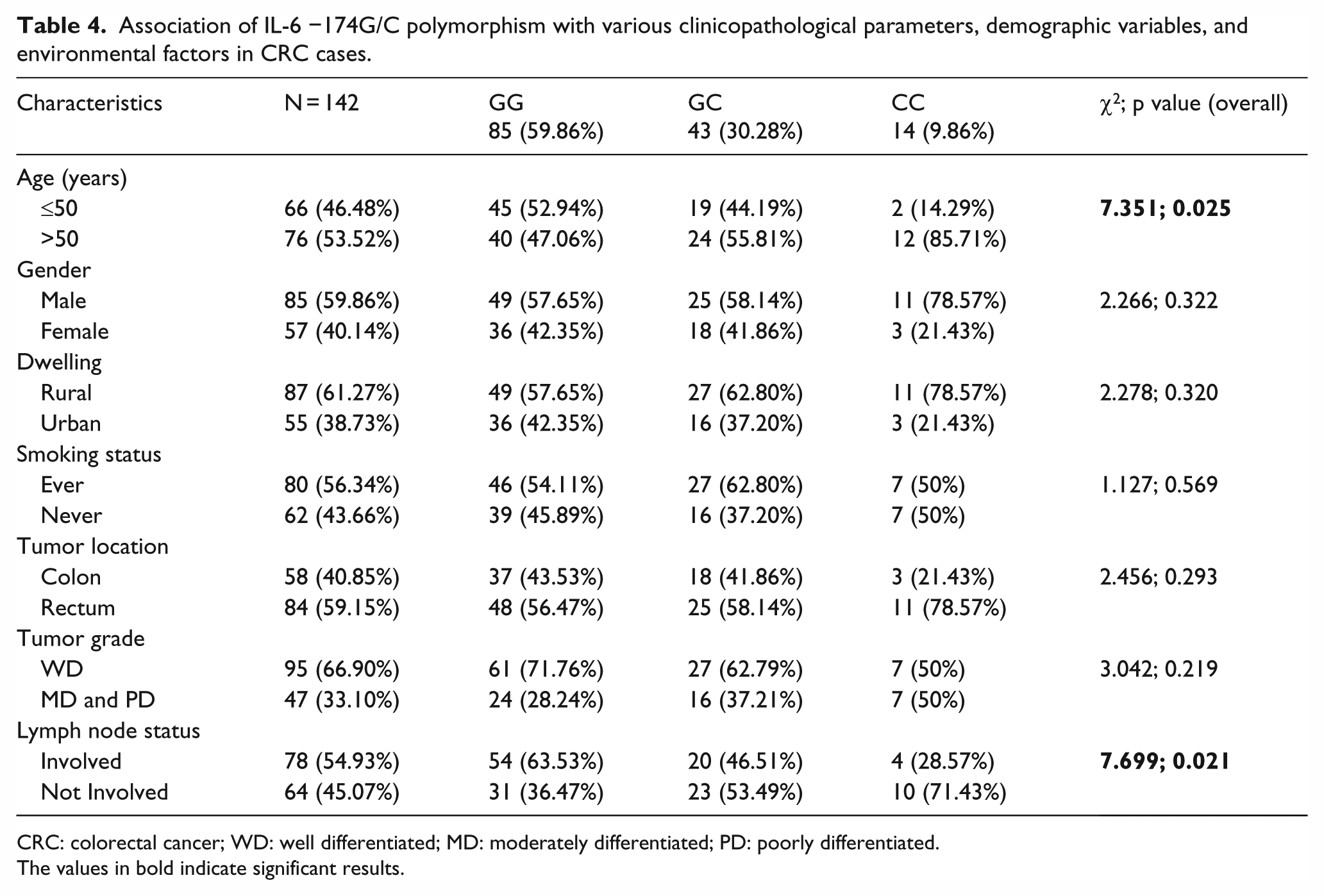
CRC: colorectal cancer; WD: well differentiated; MD: moderately differentiated; PD: poorly differentiated.
The values in bold indicate significant results.
Discussion
In this study, we investigated the role of functional IL-6 −174G/C SNP in the promoter region of IL-6 gene as a potential CRC risk factor in a case control study design with 142 case subjects and 184 control subjects.
The IL-6 −174G/C SNP is located near C/EBP-binding site and has been reported to influence the binding of the glucocorticoid receptor and transcription factor GATA1. This polymorphism results in reduced transactivation due to decreased ability of the promoter to bind glucocorticoid receptor and provides promoter access to the transcriptional repressor GATA1. The net effect is that the CC genotype and the C allele of IL-6 −174G/C SNP are significantly associated with reduced expression and lower serum or circulating levels and lower inducible responses of IL-6 in comparison to GC and more prominently GG genotypes and G allele of this SNP. 56
In our study, we found a strong association between the IL-6 −174G/C SNP and the modulation of CRC risk. We found a strong association of this SNP with decreased CRC risk across different allelic parameters. The variant genotype (CC), less common IL-6-174C allele, and the combined variant genotype (GC + CC) all were significantly associated with a decreased risk of CRC. These findings are in agreement with several studies mentioned earlier that have reported the association of CC genotype of IL-6 −174G/C SNP with a decreased risk for the development and progression of CRC.44,45,47,48 Furthermore, this effect is possibly due to the reduced IL-6 expression (& activity) associated with the CC genotype which results in decrease or inhibition of the pro-tumorigenic activity of this cytokine. As mentioned already, IL-6 plays an essential role in tumor development, progression, and differentiation, and modulation of tumor microenvironment which also promotes metastasis,11,14 and it is reasonable to argue that the decreased IL-6 expression and serum levels associated with the CC genotype may result in decrease or inhibition of these tumorigenic activities, which may reflect through inter-individual differences in susceptibility to CRC associated with this SNP. However, our finding is not in agreement with one study that has reported the association of IL-6 −174G/C SNP with an increased risk of CRC 46 and few others that have reported no association of this SNP with CRC risk.49,57 A plausible explanation for this discrepancy is the possible population-specific differences, which may result in differential susceptibility patterns shown by the same specific SNP genotype across different populations. Furthermore, varied and complex gene–environment interactions across different populations or geographical regions may modify the expression and activity of IL-6 resulting in different functional and susceptibility outcomes for the same SNP genotype.
In this study, we also evaluated the possible effect modification of association between IL-6 −174G/C SNP genotypes and CRC risk by age, gender, and smoking status. We found a significant effect modification of association between the combined variant genotype (GC + CC) of IL-6 −174G/C SNP and CRC risk by gender and age.
The male gender decreased the CRC risk. The males carrying the variant allele, IL-6-174C, in heterozygous form (GC) or variant homozygous form (CC) were at a decreased risk of developing CRC in comparison to females. A plausible explanation of this finding is that the immune response in mammals including humans has been naturally programmed to exhibit a gender-based dimorphism,58,59 and the immune response is regulated in a gender-specific manner in which sex hormones, the androgens and the estrogens, play an essential role but affect the immune system in opposite ways.60,61 The androgens have been reported to favor the T-helper 1 (Th1)-type response, 62 whereas estrogens promote Th2-type immune response. 60 It is now known through various studies that the cytokines, interferon gamma (IFN-γ) in males, and IL-6 in females are the main players and regulators of immune system in a gender-specific manner.63–65 This indicates that possibly the expression of IL-6 may be more tightly regulated in females in comparison to males and similarly the expression of IFN-γ may be more stringently regulated in males in comparison to females. A possible support for this hypothesis comes from various studies, which have reported that estrogens repress or inhibit IL-6 gene transactivation by decreasing its promoter activity through direct interactions with IL-6 trans-activators, including NF-IL6, NF-κB, and C/EBP-beta.66,67 Therefore, with respect to our findings, it is reasonable to argue that the affect of IL-6-174C allele on the expression, serum levels, and activity of IL-6 will be more profound in males in comparison to females considering the relative importance of IL-6 in immune function and its tight regulation in females. This may explain our finding that the males carrying IL-6-174C were at a decreased risk of developing CRC in comparison to females However; this hypothesis needs to be mechanistically validated in order to arrive at a final conclusion. Furthermore, it is important to emphasize here that the immune response may be regulated differently in a gender-specific manner, but the final outcome, that is, the preservation of immune homeostasis, is similar in both males and females. Furthermore, the functional alterations in IFN-γ in males and IL-6 pathways in females have been reported to be associated prognostically with the development of an adenoma and its progression into full blown CRC. 65 This further underlines the importance of cytokines such as IL-6 in the pathogenesis of CRC.
We also found a significant effect modification by age. However, the nature of this association could not be deciphered further in a statistically significant manner. Further studies involving a large cohort of CRC patients and healthy controls may help to decipher this association in a statistically significant manner and may explain its influence on CRC risk in a more conclusive manner.
In this study, we also evaluated the association of IL-6 −174G/C SNP with the numbers and the frequencies of the subsets of various characteristics of the case group subjects under study, that is, age, gender, dwelling, smoking status, tumor location, tumor grade, and lymph node status. We found that the risk of developing CRC in subjects who carried the variant genotype (CC) increased with age. The age is an established CRC risk factor. 68 The incidence of CRC increases with age, and there is a sharp increase after 50 years of age. 69 About 90% incidence of CRC is reported in people aged 50 years or above. 69 Furthermore, it has been reported that the incidence rate of CRC in individuals aged more than 65 years is more than 12 times higher in comparison to those with age less than 45 years. 69 Therefore, our finding that the CRC risk in the carriers of otherwise low-risk IL-6-174CC genotype increases with age is in agreement with these various studies.
We also found that the carriers of the variant genotype (CC) of IL-6 −174G/C SNP were at a significantly reduced risk of lymph node infiltration. As discussed already, the variant genotype (CC) exhibits relatively lower promoter activity and decreased protein expression and circulating levels of IL-6 in comparison to the more common homozygous GG genotype and the heterozygous GC genotype. Furthermore, the increased IL-6 expression and increased IL-6 serum and tumor tissue levels in CRC have been reported to be associated with an increased tumor cell proliferation, advanced tumor stage, increased tumor size, increased angiogenesis, and increased invasiveness and metastasis.21,23,24,44,70 Therefore, with reference to our finding that the carriers of the variant genotype (CC) were at a reduced risk of lymph node infiltration, it is reasonable to argue that the decreased IL-6 expression and eventually the decreased IL-6 activity associated with the CC genotype may be responsible for the decreased colorectal tumor invasion and metastasis which is partly reflected through decreased lymph node infiltration. However, further studies involving comprehensive mechanistic evaluation of the expression and the circulating levels of IL-6 protein in various phases of colorectal tumorigenesis are warranted to arrive at a conclusive explanation.
In our study, we found a significant deviation from HWE among the case group subjects. The main factors responsible for the deviations from HWE include population stratification, inbreeding and consanguinity, small population size, migration, mutations, and genotyping errors.71–73 Some of these factors are possibly responsible for the deviations from HWE observed in our study. The study population is inhabited in Kashmir region of India and possesses several characteristics that characterize a pure ethnic population. 74 The population is a Muslim majority—one which commonly prefers consanguineous intra-familial marriages, 75 and consanguineous marriages are traditional among several subpopulations within this population. Furthermore, the study subjects represent a population that has largely remained isolated genetically from the rest of the world, and most of the marriages occur within the population, which has led to a considerable degree of population stratification. Therefore, a high degree of both the consanguinity and population stratification present in the study population may lead to the deviations from HWE observed in our study. Also the population is relatively small in size, 74 and an important characteristic of the Hardy–Weinberg law is infinite population size, and therefore small size of the study population may be another contributory factor toward the HWE deviations observed in our study. The genotyping errors, due to either technological or human reasons or both, may also contribute toward deviations from HWE in association studies. However, it is unlikely that the deviations from HWE observed in our study are due to genotyping errors. This is because we maintained standard quality control procedures recommended for genotyping and other experimental procedures in our study. This is obvious from a high degree of the genotyping reproducibility of the samples for the SNP under study and this is reflected by a weighted kappa coefficient of 0.98 obtained in our study, which meant a high concordance rate of 98%. Furthermore, there may be an additional reason behind the deviations from HWE observed among the case group subjects in our study. These deviations may actually represent or may be due to the true associations of the SNP under study with the disease and may be indicative of the concerned SNP as a true disease marker. However, further studies involving comprehensive mechanistic evaluation of the role of the gene carrying this SNP in the disease under study are warranted to substantiate or prove this hypothesis.
This study is the first of its kind with regard to exploring the association of IL-6 −174G/C SNP with CRC risk in our population. The major strengths of this study are the use of clinically diagnosed and histopathologically confirmed CRC samples, and the involvement of population-based controls in addition to the hospital-based controls who were recruited from the same geographical area during the same time period and were matched to the case group subjects individually for age, sex, place of residence (rural/urban), smoking habit, and ethnicity in order to minimize the confounding effect of these relevant factors. Further in our study, the results were adjusted for multiple potential confounding (third) variables. The major limitation of this study is the modest sample size to detect comprehensively the gene–gene and gene–environment interactions, which usually require much larger sample size. Further studies incorporating a larger sample size and/or another ethnic population in our study are required to substantiate our findings or confirm in depth the role of this SNP in relation to CRC susceptibility. However, these limitations are unlikely to affect the final outcome of this study.
Conclusion
We have demonstrated through this study that the IL-6 −174G/C promoter SNP is strongly and significantly associated with a decreased risk of CRC in ethnic Kashmiri population. We have also shown that there is a significant effect modification of the association between IL-6 −174G/C SNP genotypes and CRC risk by gender and age. Furthermore, we have also demonstrated that there is a significant association between IL-6 −174G/C SNP and some characteristics of the case group subjects, including age and lymph node status.
Footnotes
Acknowledgements
M.Z.B. acknowledges the Department of Biotechnology, University of Kashmir, for providing all the research facilities and University Grants Commission (UGC), Government of India (GOI) for UGC–Basic Scientific Research (BSR) fellowship. M.Z.B. would like to thank his colleagues Taseen Gul, Insha Zahoor, and Zeenat Hamid from Department of Biotechnology, University of Kashmir, and Idrees Ayoub Shah from the Department of Biochemistry, University of Kashmir, Hazratbal, Srinagar. The authors wish to thank each and every colorectal cancer (CRC) patient and control subject who took part in this study and cooperated during the interview and sample collection. They also thank the head and technical staff of the operating theater in the Department of General Surgery, Sher-I-Kashmir Institute of Medical Sciences (SKIMS), Soura, Kashmir, who helped us with tissue procurement, and the anonymous pathologists at the Department of Pathology, SKIMS, Soura, Kashmir, for the histopathological assessment of the tumor tissues.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
