Abstract
Background:
Recently experimental validation of the networks revealed bach1, a basic leucine zipper transcription factor, as the common regulator of several functional invasive genes. The expression of bach1 and its target genes was linked to the higher risk of breast cancer recurrence in patients. The aim of this study was to investigate the effect of specific bach1 small interfering RNAs, on the invasive and expression level of miR-203, miR-145, matrix metalloproteinase-9, and CXCR4 receptor which play a role in cancer metastasis, in MDA-MB-468 cell lines.
Methods:
Small interfering RNA transfection was performed with transfection regent. The survival effects of small interfering RNA were determined using trypan blue assay cells. The expression level of messenger RNA and matrix metalloproteinase-9 to assess cell invasion and the expression level of miR-203, miR-145, and CXCR4 receptor were measured by quantitative real-time polymerase chain reaction analysis on the MDA-MB-468 cell lines.
Results:
Transfection with small interfering RNA significantly suppressed the expression of bach1 gene in dose-dependent manner after 48 h (
Conclusions:
Our results suggest that the bach1-specific small interfering RNA effectively decrease CXCR4 receptor, matrix metalloproteinase-9 expression and breast adenocarcinoma cells invasive, also increased the expression of tumor-suppressive microRNA-203 and miR-145. Thus, these microRNAs may play a role in invasive/metastasis of carcinogenic breast cancer cells. Therefore, bach1 knockdown can be considered as a potent adjuvant in breast cancer therapy.
Introduction
Cancer comprises a group of diseases characterized by unregulated cell growth, invasion, and metastasis of cells from the primary site of infection to other parts of the body. After skin cancer, breast cancer is the second most common cancer diagnosed. International studies show an increase in the number of patients with this cancer. 1 This disease can be treated surgically by removing the tumor and by radiotherapy, immunotherapy, chemotherapy, hormone therapy, gene therapy, and tyrosine kinase inhibitors. Modern medicine has moved toward a more specialized treatment to enhance the therapeutic properties and to reduce the side effects. Gene therapy is one of the novel therapeutic methods for fighting the disease with genetic origin, and it is used for elevating the sensitivity of tumor cells to therapeutic methods. 2 The advantages of gene therapy for cancer-related gene silencing are as follows: its high power and performance, induced silencing capabilities in advanced stages, the ability to transfer it to the next generation, and cost-effective compared to the other methods. 3 The transfection of RNA has become extremely important with the discovery of RNA interference (RNAi)-mediated gene silencing using small interfering RNA (siRNA). RNAi is a process in which a double-stranded RNA molecule prevents the expression of certain genes. This gene, essentially with regard to sequence, is homologous double-stranded RNA. Inhibition of gene expression through messenger RNA (mRNA) degradation which is why it is called post-translational silencing. 4 One of the most important strategies in gene therapy experiments is suppressing the expression of genes involved in diseases, using synthetic siRNA molecules at the laboratories. Now siRNA molecules are going through clinical trial phases 1–3 testing their therapeutically effect on infectious diseases and colon cancer, breast cancer, pancreas cancer, and leukemia.5,6 SiRNA is small double-stranded RNA molecules, and despite microRNA (miRNA), they are synthetic with foreign origins, such as viral RNA. The sequences of the synthetic siRNA are complementary to the sequences of the target miRNA, which is why they act with high specificity. With the completion of the Human Genome Project in 2003 and several studies, many genes that are involved in tumor progression and drug resistance were identified. 7 Recently, several genes involved in breast cancer invasive/metastasis were identified, bach1 protein (BRCA1-associated C-terminal helicase) is a transcription factor of the leucine zipper-binding protein family, 8 encoded by chromosome 21, consists of 730 amino acids. That has critical role in the development of the invasive phenotypes of cancer metastasis and inhibits apoptosis in tumor cells.9,10 CXCR4 receptor is one of the molecules playing a role in breast cancer metastasis. In 2001, the role of CXCR4 in metastatic breast cancer was proven for the first time.11,12 Thereafter, there were several studies on CXCR4 in which the proliferative role of this receptor in breast cancer metastasis has been highlighted. When CXCL12 binds to CXCR4, it stimulates cell migration in breast cancer. 13 Matrix metalloproteinase (MMP) plays a vital role in the invasion and cancer progression. 14 A recent study showed that high expression of matrix metalloproteinase-9 (MMP-9) is associated with the invasion and metastasis of breast cancer. So they introduced this protein as a diagnostic marker of metastatic breast cancer. 15 Increased plasma concentrations of MMP-9 in a variety of tumors, including breast, colon, lung, head, and neck; hepatocellular carcinoma; and gastric cancer, have been observed.16,17 MiRNA expression can be used to classify cancers dominant in groups with different characteristics, such as cell type and etiology. MiRNA structure, and how it functions, shows that many miRNAs are abnormally expressed in cancer samples. Furthermore, the functional differences between tumor types and stages of cancer associated with the expression of miRNA. Changes in miRNA expression by reducing the expression of genes involved in proliferation or survival of cells are necessary, leading to tumor formation. This does not mean that miRNAs are directly involved in tumor genesis and cancer progression. Although still not fully understood is that changes in the expression level of microRNA is a cause of cancer or the cancer is the direct cause of this change of expression. However, many changes occur in cancer cells that can directly or indirectly influence the expression of miRNA. 18 Increased or decreased expression changes of some miRNAs that lead to cancer affect cell growth through interference with cell cycle regulators. The aim of this study was to turn off oncogene bach1 by RNAi in metastatic breast cancer cell line and evaluate its effect on miR-203, miR-145, MMP-9, and CXCR4 receptor.
Materials and methods
This was an experimental study performed at the laboratory (in vitro), and since it had no clinical sample, there was no need for laboratory animals.
Materials and reagents
The following reagents were used: RPMI-1640 medium containing
Cell culture
RPMI-1640 culture medium was used in this study in powder form containing glutamate and no sodium bicarbonate. The study was conducted on MDA-MB-468 cell lines. This study was conducted on MDA-MB-468 cancer cell lines related to human adenocarcinoma origin. To cultivate this cell line, RPMI-1640 base medium containing 10% FBS was used, and it was incubated under the following conditions: temperature: 37°C, 5% carbon dioxide, and adequate moisture.
Bach1-specific siRNA transfection applied to determine the effective time and dose of siRNA
First, three 6-well plates were chosen for different periods of time: 24, 48, and 72 h. Three wells for bach1 gene and three wells for control were chosen for each of the plates. In this step, the selected dose for all wells was 60 µg/µL. The experiments were performed in triplicate. To perform the experiment, 20 × 105 cells were distributed in each of the wells, and MDA-MB-468 cells were stored in an incubator at 37°C, 95% humidity, and 5% carbon dioxide for 24 h. Transfection was carried out after 24 h of incubation of the cells; after being analyzed by microscope, the confluence was between 50 and 60, and spindle-shaped cells were observed. The wells containing the cells were washed in a fresh medium, free of antibiotics and FBS, and then 2 mL of medium, free of antibiotics and without FBS, was added to each of the wells. Then, the plate was placed in the incubator with the following conditions for an hour: temperature set at 37°C, 95% humidity, and 5% carbon dioxide. For each well, 6 µL of siRNA with 94 µL of transfection medium and 6 µL of special transfection reagent with 94 µL of transfection medium were added, and the solution obtained was incubated at room temperature for 30 min. Finally, 200 µL of the mixture solution mentioned above was added to each of the wells, and the plate was incubated for 6 h at 37°C, 95% humidity, and 5% carbon dioxide. After 6 h of incubation, 1 µL of RPMI-1640 medium, containing 20% FBS and antibiotics, was added to each well.
RNA extraction
After opening the cell sediment, 1 mL of RNX™-PLUS was added to the cell sediment, and after being vortexed, it was incubated at room temperature for 5 min. After the addition of 200 µL of chloroform to the sample tube, it was gently shaken for 15 s. The tube containing the sample was incubated on ice for 5 min. Then, it was centrifuged, revolving at 12,000 r/min and 4°C for 15 min. The aqueous phase was removed, and the same volume of isopropanol was added. Then, it was incubated at 4°C for 15 min. After centrifugation for 15 min at 12,000 r/min and temperature of 4°C, 1 mL of cold ethanol (75%) was added to the sediment, which was then centrifuged at 7500 r/min and 4°C for 10 min. After centrifugation the ethanol was removed and the pellet were allowed to dry to the air and finally diluted in 20 µL of DEPC-treated water. Thereafter, it was directly transferred to −80 to be stored.
Determination of the viability of the cells using trypan blue dye
In order to stabilize the cells and let them attach on the plate, 50,000–70,000 cells were distributed in each well of 24-well plates and then the plate was incubated under the following conditions: with CO2, at 37°C and for 24 h. Next step was the transfection of cells at different concentrations of 40, 60, and 80 µg/µL for 24, 48 and 72 h using siRNA. After the incubation period, the wells were emptied, and cells were washed with 200 µL of phosphate-buffered saline (PBS) and trypsinized. Then, 50 µL of cell suspension were mixed with 50 µL of trypan blue, which was incubated for 1 min and vortexed for 15 s. Finally, 20 µL of the mixture were poured on a hemocytometer, so that the number of live and dead cells on the slide was counted.
Real-time PCR for the analysis of the expression of MMP-9, CXCR4 receptor, and bach1 genes
Total RNA was isolated from cells using RNX-Plus reagent according to the manufacturer’s protocol. Total RNA of 1 µL (5 ng) was converted to complementary DNA (cDNA) using 1 µL random hexamer primer, 1 µL MMLV, 4 µL 5× reaction buffer, and 2 µL of 10 mM dNTP. cDNA (500 ng) was amplified by real-time (RT) PCR using SYBR Green-1 dye universal master mix and Rotor-Gene TIM 6000 (Corbett Life Science, Mortlake, NSW, Australia) and LightCycler® 96 System (Roche, Roche Life Science) Sequence Detection System. To confirm the PCR specificity, PCR products were subjected to a melting-curve analysis. The PCR conditions were 95°C for 10 min followed by 45 cycles at 95°C for 10 s, 59°C for 30 s, and 72°C for 20 s. The ratio of gene expression was calculated using 2−(ΔΔCT) method and β-actin was used as an internal control. For the record, the primers used in this study are shown in Table 1. A tube containing all the substances used in the reaction except for cDNA was set as blank and DEPC water was added to this blank tube. All procedures were performed on ice and under the laminar-flow hood to prevent contamination.
Primer sequence.

FW: Forward; RV: Reverse.
MiRNA extraction and its cDNA synthesis
In order to synthesize, Total RNA must be extracted from the cell culture using miRCURY RNA Isolation kit—cell and plant (Exiqon) with the catalog numbers of 205914 and 204483. In Total RNA extraction, a suitable method should be used in a way that species of small RNAs remain. For this purpose, the special buffers for cell lysis, washing, and extraction (based on the protocol provided by the kit manufacturer) were used. For quality control and determination of the purity of the extracted Total RNA, UniSp RNA provided in the kit was used. After Total RNA extraction, cDNA synthesis was started. cDNA synthesis kit was used for this purpose. First, the Total RNA was diluted and then the reaction solution containing the enzyme reaction buffer and nuclease-free water was mixed according to the protocol. The diluted Total RNA was added to the reaction solution, the mixture obtained was incubated at 45°C for 60 min and at 95°C for 5 min, and then it was cooled at 4°C quickly. Quantitative real-time polymerase chain reaction (qRT-PCR) was performed as follows: 95°C for 10 min, 40 cycles of 95°C for 10 s and 60°C for 10 s, and ramp rate 100% under standard condition. U6 snRNA was used as references. Target sequence of miR-203 was 5′-GUGAAAUGUUUAGGACUAG-3′.
Statistical analysis
In this study, all tests were performed in triplicate. Statistical analysis was performed using GraphPad Prism 6, and the central indexes consisting of average value and standard deviation were calculated by descriptive statistics. A
Results
Before starting the experiments, it was essential to determine the efficient period of time and effective dose of siRNA on MDA-MB-468 cells; thus, for this purpose, a medium dose (60 pmol) was selected. First, the selected dose of siRNA (60 pmol) was tested on treated cell and control cell for different periods of time of 24, 48, and 72 h. After the efficient time was determined by analyzing the expression of the genes, in order to determine the efficient dose, three doses of 40, 60, and 80 pmol were chosen to be tested for an effective period of time chosen.
Specific siRNA reduces the expression of bach1 gene of breast cancer cells
The effect of siRNA on the expression of mRNA in MDA-MB-468 cancer cell line was tested. Cells were transfected with bach1-specific siRNA, and the relative expression of each gene was analyzed using qRT-PCR. Each of the samples was compared to beta-actin internal control gene. For this purpose, the siRNA-treated cells were evaluated with untreated cells (Figures 1 and 2). Finally, a significant decrease in bach1 gene was observed. The effective period of time of 48 h and an effective dose of 60 pmol were obtained for bach1 gene (Figure 1). In this study, the

The amplification and melting curves for bach1 to obtain the effective time. (a) The amplification curve of beta-actin and bach1 gene in cells treated with bach1-specific small interfering RNA (siRNA) and control ones at duration of 24, 48, and 72 h. (b) The melting curve for beta-actin and bach1 gene in cells treated with bach1-specific siRNA and control ones at duration of 24, 48, and 72 h in Rotor-Gene 6000 system. (c) Chart for effective time for bach1 gene suppression using siRNA in MDA-MB-468 cell lines. Data are shown as mean ± SD. ****
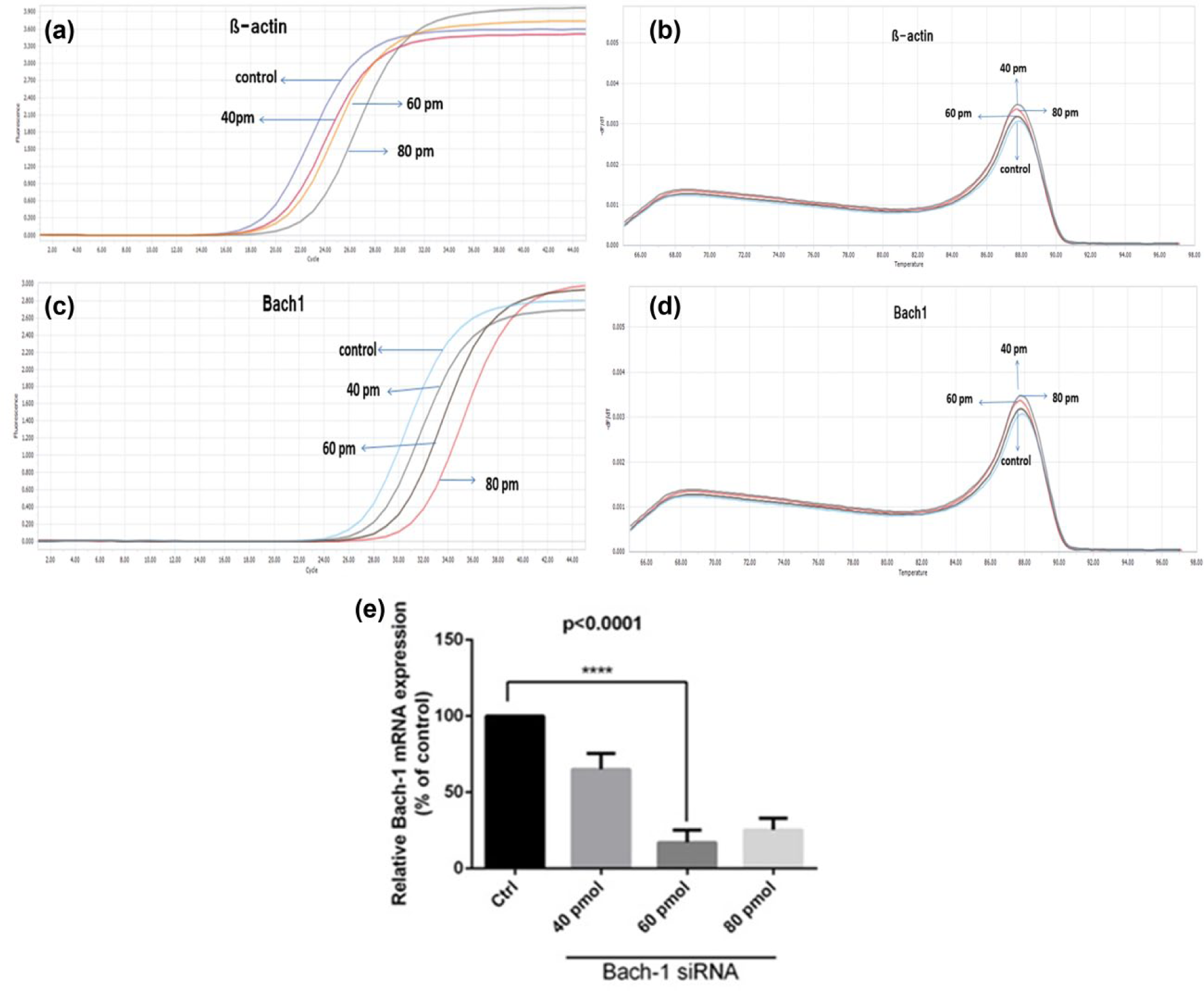
The amplification and melting curves for beta-actin gene and bach1 to obtain the effective dose. (a) The amplification curve of beta-actin gene in cells treated with bach1-specific small interfering RNA (siRNA) and control ones at concentrations of 40, 60 and 80 picomol. (b) The melting curve for beta-actin gene in cells treated with bach1-specific siRNA and control ones at concentrations of 40, 60, and 80 picomol. (c) The amplification curve of bach1 gene in cells treated with bach1-specific siRNA and control ones at concentrations of 40, 60, and 80 picomol. (d) The melting curve for bach1 gene in cells treated with bach1-specific siRNA and control ones at concentrations of 40, 60 and 80 picomol in LightCycler® 96 SW 1.0 system. (e) Chart for effective dose for bach1 gene suppression using siRNA in MDA-MB-468 cell lines. Data are shown as mean ± SD. ****
Bach1-specific siRNA decreases the expression of MMP-9 gene in metastatic MDA-MB-468 cells
As shown in Figure 3, a significant decrease in MMP-9 gene expression was observed after the bach1 knockdown process. The MMP-9 gene expression in untreated cells was considered as 100 so that we could calculate the percentage of the treated sample. The relative mRNA expression of MMP-9 gene after bach1 knockdown was 23%. (Figure 3(c)). Since it is proven that MMP-9 has a role in breast cancer metastasis, so by knockdown of MMP9 gene, the metastasis process would be inhibited significantly.

The effect of bach1-specific siRNA on matrix metalloproteinase-9 (MMP-9) in MDA-MB-468 breast cancer cell line. (a) The amplification curve of MMP-9 gene in cells treated with bach1-specific siRNA and control ones. (b) The melting curve for MMP-9 gene in cells treated with bach1-specific siRNA and control ones. (c) Percentage of treated cells with bach1-specific 60 picomol siRNA compared to untreated cells. Data are shown as mean ± SD. ****
Bach1-specific siRNA decreases the expression of CXCR4 receptor gene
A significant decrease in CXCR4 receptor gene expression was observed after the bach1 knockdown process (

The effect of bach1-specific siRNA on CXCR4 receptor in MDA-MB-468 cell lines. (a) The amplification curve of CXCR4 receptor gene in cells treated with bach1-specific small interfering RNA (siRNA) and control. (b) The melting curve for CXCR4 receptor gene in cells treated with bach1-specific siRNA and control. (c) Chart for effective dose for bach1 gene suppression using siRNA in MDA-MB-468 cell lines. Data are shown as mean ± SD. ****
The effect of bach1 gene–specific siRNA on the morphology of MDA-MB-468 cells
Using the effective dose and period of time obtained, metastatic MDA-MB-468 cells were analyzed on the morphological basis, before and after being transfected with the specific genes mentioned. Apoptosis and cell growth inhibition were shown in cells after transfection with the specific gene (Figure 5). So we can come to the conclusion that bach1 knockdown induces apoptosis.

The effect of bach1-specific siRNA on the morphology of MDA-MB-468 breast cancer cell line. (a) The microscopic image of untreated cells with siRNA using a 40× lens. (b) The microscope image of cells treated with bach1 gene–specific siRNA using a 40× lens.
Bach1 knockdown increased expression of miR-203 and miR-145 genes in MDA-MB-468 cell lines
Using the effective dose and period of time obtained, it was proven that when we set bach1 gene as the specific siRNA target, expression of miR-203 and miR-145 results in metastatic MDA-MB-468 breast cancer cell line, compared to untreated ones with siRNA (Figures 6 and 7). These data indicate that bach1 knockdown led to significant increase in the expression of miR-203 and miR-145.

Study of amplification and melting for miR-203 in MDA-MB-468 cell line. (a) The amplification curve of miR-203 in cells treated with bach1-specific siRNA and control cells. (b) The melting curve for miR-203 in cells treated with bach1-specific siRNA and control cells in LightCycler 96 SW 1.0 system. (c) Percentage of treated cells with bach1-specific 60 picomol siRNA compared to untreated cells. Data are shown as mean ± SD. ****

The amplification and melting curves for miR-145. (a) The amplification curve of miR-145 in cells treated with bach1-specific siRNA. (b) The melting curve for miR-145 in cells treated with bach1-specific siRNA in LightCycler 96 SW 1.0 system. (c) Expression percentage of miR-145 in treated cells with bach1-specific 60 picomol siRNA compared to untreated cells. Data are shown as mean ± SD. ****
Discussion
Recently, several genes involved in breast cancer metastasis were identified. According to the study on breast cancer metastasis and the genes involved, by Liang et al., 19 it was reported that bach1 had a role in the expansion of breast cancer metastasis to bones. Bach1 inhibition in a mouse model of metastasis reduced metastasis dramatically, and uncontrolled overexpression of bach1 leads to increased malignant breast cancer. According to the studies on humans, bach1 causes severe metastasis in breast cancer. 19 Our results were similar to Liang’s results; moreover, we proved that bach1 knockdown in metastatic breast cancer cell lines leads to less invasion and growth of breast cancer and leads to increased expression of miR-203 and miR-145; also the expression of MMP-9 and CXCR4 receptor decreased significantly through the bach1 knockdown. So far, it has been proven that miRNA has a role in carcinogenesis of some of the cancers. MiRNA, for its great sensitivity, can be the potential biomarker for the purpose of detection, diagnosis, classification, and treatment of cancer. Furthermore, it can show the stage of the tumor, receptor status, and patient’s survival chance.20,21 One of the miRNAs involved in carcinogenesis can be miR-203, on which only a few studies have been carried out. In recent studies, it has been shown that this miRNA has a tumor-suppressive effect on prostate, lung, colorectal, and larynx cancer.22–26 However, until recently the role of miR-203 in the pathogenesis of breast cancer has not been studied. Zhao et al. 25 carried out a study on miR-203 in a variety of breast cancer cell lines, including healthy, non-metastatic, and metastatic cell lines. Their results showed that the expression of miR-203 in healthy breast cancer cell lines is less than in metastatic breast cancer cell lines. Considering the overexpression of the miR-203 in the non-metastatic breast cancer cell lines, we conclude that this miRNA has a significant role in carcinogenesis and breast cancer metastasis. Moreover, they showed that overexpression of miR-203 prevents the breast cancer cell growth and invasion, and when inhibited by the antisense strand of this miRNA, there can be seen a significant increase in the growth and invasion of breast cancer cells. Overexpression of some oncogenes causes tumor progression in many human cancers.25,26 According to the study by Zhao et al. in 2012, overexpression of miR-203 in breast cancer leads to inhibition of growth and invasiveness of breast cancer cells. 25 We observed that by bach1 gene knockdown, the expression of miR-203 increased significantly so that we can conclude that by this bach1 gene knockdown, the growth and invasiveness of the cancer cells diminish and miR-203 expression is increased. Also miR-145 has been known as a tumor suppressor in a variety of cancers, including ovarian, 27 prostate, 28 bladder, 29 lung, 30 and colon. 31 In several studies, it has been indicated that expression of miR-145 decreases in different cancers, while the function of miR-145 and its mechanism have not been elucidated in breast cancer. 32 In 2015, Zheng et al. showed that the expression of miR-145 is reduced in metastatic breast cancer. From 2005 to 2011, researchers had been studying miR-145 in breast cancer and shown that this miRNA expression levels are reduced in cancer.33,34 In our study, we have set the invasion and growth of breast cancer cells as the target, and by stopping the growth and invasion process, miR-145 levels were increased significantly.
In invasive cancer cases, the cancer cells penetrate the basement membrane of epithelial tissue and eliminate the extracellular matrix. At this stage, protease enzyme—that digests the extracellular matrix and its compounds—plays a key role by facilitating the invasion and metastasis of cancer cells to other tissues. 35 The complete role of the proteases, particularly MMP in metastasis and invasion of tumor cells, has been demonstrated, which facilitates the process of breaking the bonds of the connective tissue and digesting the extracellular matrix components.36,37 Increased plasma concentrations of MMP-9 have been observed in a variety of malignant tumors such as breast cancer, colon cancer, lung cancer, head and neck cancer, hepatocellular carcinoma, and gastric cancer. 38 Merdad et al. 15 showed that high expression of MMP-9 is associated with the invasion and metastasis of breast cancer; so they suggested this protein as a diagnostic marker for metastatic breast cancer. Our results showed that the expression of MMP-9 decreased significantly through the bach1 knockdown in MDA-MB-468 cells. CXCR4 receptor is one of the molecules playing a role in breast cancer metastasis. In 2001, the role of CXCR4 in metastatic breast cancer was proven for the first time. 11 Thereafter, there were several studies on CXCR4 in which the proliferative role of this receptor in breast cancer metastasis has been highlighted. Yagi et al. 39 showed that CXCR4 is required for breast cancer cell migration. When CXCL12 binds to CXCR4, it stimulates cell migration in breast cancer. 13 In this study, we showed that by bach1 gene knockdown in metastatic cancer cells, a significant reduction in CXCR4 receptor expression and a significant increase in the expression of miR-145 can be seen. So, hopefully, the suppression of this gene can reduce the breast cancer metastasis. The common therapeutic methods such as chemotherapy and surgery are not responsive for the treatment of metastatic breast cancer, and all human beings aim is to inhibit this cancer using gene therapy and target therapy. However, early diagnosis of this cancer is another issue that concerns scientists trying to find markers by the level of which early diagnosis of this cancer can be made. In this experiment, bach1 and miR-145 were both markers that were studied.
When MDA-MB-468 cells were transfected with bach1 specific-siRNA, increased cells apoptosis and decreased cells growth have been observed that was evaluated by comparing the morphology of the cells before and after treatment.
Conclusion
According to the results obtained from this study, we can conclude that bach1 can have a role in invasion and breast cancer metastasis and by knockdown this gene using specific siRNA, we can dramatically decrease the intensity of the invasion and metastasis in breast cancer cells, also by knockdown of bach1, the miR-203 and miR-145 expression was increased and the expression of MMP-9 and CXCR4 receptor was decreased. Therefore, bach1 and miR-203, miR-145, MMP-9 and CXCR4 can be introduced as malignant breast cancer markers and bach1 knockdown can be possible therapeutic targets for breast cancer. Therefore, bach1 and miR-203, miR-145, MMP-9, and CXCR4 can be introduced as malignant breast cancer markers. Also, bach1 knockdown can be possible therapeutic targets for breast cancer.
Footnotes
Acknowledgements
This work was sponsored by Maragheh University. We thank Mohammadreza Alivand, PhD, Molecular Genetic in Tabriz University of Medical Sciences.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was sponsored by Maragheh University. We thank investigators, Mansoor Altaha, PhD, Biotechnology, Pathology Department, University of Shiraz, and Mohammadreza Alivand, PhD, Molecular Genetic in Tabriz University of Medical Sciences.
