Abstract
Glioma is an aggressive malignancy with limited effective treatment and poor prognosis. Cytoplasmic polyadenylation element binding protein 4 is a regulator of gene transcription and has been reported to be associated with biological malignancy in cancers. However, the mechanisms that cytoplasmic polyadenylation element binding protein 4 contributes to tumor migration and invasion remain unknown. Here, cytoplasmic polyadenylation element binding protein 4 expression was assessed using immunohistochemistry, and the results were compared with clinicopathological parameters, including survival. Using glioma cell lines (SKMG-4 and T98G), we measured cytoplasmic polyadenylation element binding protein 4 messenger RNA and protein expression and studied the effects of cytoplasmic polyadenylation element binding protein 4 expression on cell migration and invasion. Cytoplasmic polyadenylation element binding protein 4 expression was significantly higher in tumor tissues than that in normal brain tissues. Clinicopathological analysis showed that cytoplasmic polyadenylation element binding protein 4 expression was significantly correlated with advanced World Health Organization grade (p < 0.001) and lower Karnofsky Performance Status (KPS) score (p = 0.001). Cytoplasmic polyadenylation element binding protein 4 positive as opposed to the cytoplasmic polyadenylation element binding protein 4 negative patients had lower overall survival (p < 0.001). Multivariate analysis suggested that cytoplasmic polyadenylation element binding protein 4 expression might be an independent prognostic indicator (hazard ratio = 2.091, 95% confidence interval: 1.093–3.999, p = 0.026) for glioma patients. Moreover, upregulated cytoplasmic polyadenylation element binding protein 4 expression could promote T98G cell migration and invasion, and downregulated cytoplasmic polyadenylation element binding protein 4 expression could inhibit SKMG-4 cell migration and invasion. Furthermore, downregulated cytoplasmic polyadenylation element binding protein 4 could reduce the protein expression of matrix metalloproteinase-2 and matrix metalloproteinase-9. In conclusion, our studies indicated that positive cytoplasmic polyadenylation element binding protein 4 expression predicted a worse prognosis in glioma patients, and cytoplasmic polyadenylation element binding protein 4 could represent a useful biomarker or therapeutic target for glioma.
Introduction
Glioma is the most common type of malignant intracranial tumor. Glioma accounts for 50% of all intracranial tumors. 1 Glioma has high capacity of proliferation and invasion into healthy brain tissue. 2 The current standard of care for gliomas is surgery, radiotherapy, and alkylating agent chemotherapy, which result in a 2-year survival rate of 15%–26% and extremely frequent tumor recurrence. 3 Therefore, identifying novel prognostic markers, carcinogenic pathways, and therapeutic targets is crucial for the treatment of glioma.
Cytoplasmic polyadenylation element binding protein 4 (CPEB4) is a sequence specific RNA-binding protein that participated in translational control and belongs to the CPEB family, which includes CPEB1, CPEB2, CPEB3, and CPEB4. The members of CPEB family shared structure and sequence identity in the C-terminal RNA-binding domain (RBD). Recent studies suggested that CPEB proteins control a variety of biological processes, including cell proliferation, apoptosis, migration, invasion, cell cycle, senescence, and so on.4–8 CPEBs have vital function in cell fate, the loss- or gain- of CPEBs could give rise to some genetic diseases, including carcinogenesis. For instance, as tumor suppressor gene, CPEB1 could inhibit tumor metastasis through mediating epithelial-to mesenchymal transitions in breast cancer. 5 In glioblastoma, Kochanek et al. 9 reported that CPEB4 could bind to and regulate translation of AEG-1 in the rat glioblastoma cell lines and finally inhibited cell migration. But Xiaoping et al. 10 demonstrated that CPEB1 is an oncogene and overexpressed in glioma cells and tissues; and the knockdown of CPEB1 reduced cell senescence by regulating the expression or distribution of P53. In triple negative breast cancer, knockdown of CPEB2 promoted the resistance of AnR TNBC cells to anoikis and increased metastatic potential. 11 Acting as a tumor suppressor, downregulation of CPEB3 could promote tumor invasion and migration. 12 Similarly, in colorectal cancer, Wang et al. 13 reported that CPEB3 is downregulated in colorectal cancer tissues. Thus, a number of studies showed that CPEB proteins may be attractive targets for therapeutic intervention in certain cancers.
CPEB4 is an 80.2-kDa protein of the CPEB family, which could modulate translation by controlling the polyadenylation of target genes.8,14 In recent years, some studies demonstrated that CPEB4 not only expressed in a variety of somatic tissues but also has essential functions in regulating the balance between senescence and proliferation and tumor development.15,16 Although the relationship between CPEB4 and cancers has been investigated in several studies, the molecular mechanism of CPEB4 modulates tumor invasion, and migration has not been detailedly demonstrated. Therefore, we investigated the significance of CPEB4 expression with respect to clinical features and prognosis in glioma patient, and the potential molecular pathway of CPEB4 on tumor invasion and migration.
Materials and methods
Patients
Ethical approval for human subjects and the use of tissue from these subjects was obtained from the Institutional Review Board of the Huai He hospital of He Nan University, and written consent was obtained from each patient. Samples from 89 glioma patients, who underwent surgery at Huai He hospital of He Nan University between 2008 and 2011, were collected, confirmed to be glioma, and then made available for this study. Follow-ups were performed until December 2015.
Fresh tumor samples were obtained from the resection specimens of primary glioma patients treated by surgery at Huai He hospital of He Nan University in 2015 without prior radiotherapy or chemotherapy (N = 30). All excised tissues were frozen immediately in liquid nitrogen and then stored at −80°C until use.
Cell culture
The two glioma cell lines (SKMG-4 and T98G) were obtained from the cell bank of He Nan University. The two cell lines were grown in Dulbecco’s modified Eagle’s medium (DMEM; GIBCO Laboratories, Grand Island, NY, USA). The medium contained 10% fetal bovine serum, 100 U/mL penicillin G, and 100 µg/mL streptomycin (GIBCO Laboratories). All cells were cultured at 37°C in a humidified incubator containing 5% CO2.
Immunohistochemical staining
Immunohistochemistry (IHC) was conducted as described. 17 Paraffin-embedded glioma tissues were obtained from the Department of Pathology. For immunohistochemistry, sections were taken, deparaffinized, and pretreated with 10 mM sodium citrate buffer for antigen unmasking (pH 6.0, boiling temperature, 30 min). Then they were blocked in normal serum (Vectastain ABC kit; Vector Laboratories, Inc., Burlingame, CA, USA), incubated with primary antibodies at 4°C overnight, rinsed, and incubated with the secondary antibody (Vectastain ABC kit). Signals were amplified using the Vectastain ABC kit as per manufacturer’s instructions. The target protein was visualized using diaminobenzidine as the substrate.
The results were interpreted by two independent pathologists who were blinded to the specific diagnosis and prognosis for each case and were scored using a semi-quantitative method where the staining of more than 10% of the tumor cells was considered positive. The staining intensity was scored as “no” (immunohistochemical score 0), “weak” (immunohistochemical score 1+), “moderate” (immunohistochemical score 2+), and “strong” (immunohistochemical score 3+) staining. According to CPEB4 expression, glioma patients were divided into the CPEB4-negative and CPEB4-positive groups.
Western blot
Western blot assay was conducted as described. 17 Total cell or tissue extracts were extracted using cell lysis buffer followed by immunoblotting with anti-CPEB4 (1:500; Abcam, Cambridge, MA, USA), anti–matrix metalloproteinase-2 (MMP2) (1:1000; Cell Signaling Technology, Danvers, MA, USA), anti-MMP9 (1:1000; Cell Signaling Technology), and anti-GAPDH (1:1000; Jietewei Biotechnology, Guangzhou, China).
Quantitative reverse-transcriptase polymerase chain reaction
Total RNA was isolated using the RNA plus reagent (Takara, Dongjido, Japan). Complementary DNA was prepared using oligo-dT primers according to the protocol supplied with the PrimerScript™ RT Reagent (Takara, Dongjido, Japan). Expression levels of CPEB4 messenger RNA (mRNA) were determined by quantitative reverse-transcriptase polymerase chain reaction (qRT-PCR) using Power SYBR green PCR master mix (Applied Biosystems, USA). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) levels were used for normalization. The PCR primers used were as follows: CPEB4 (sense: TGGGGATCAGCCTCTTCATA, antisense: CAATCCGCCTACAAACACCT) and GAPDH (sense: GAGTCAACGGATTTGGTCGT, antisense: GACAAGCTTCCCGTTCTCAG).
RNA interference
The T98G cell was seeded in six-well plates and cultured for 24 h before use. CPEB4-specific small interfering RNAs (siRNAs) and scrambled sequence were followed in Huang et al.’s 18 study and were synthesized in Ribobio (Guangzhou, China). Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) was used to transfect siRNA into cells following the manufacturer’s instructions. After transfection of 48 h, the cells were collected and used for the further experiments.
Plasmid transfection
The SKMG-4 cell was seeded in six-well plates and cultured for 24 h before use. The CPEB4 overexpression plasmids were purchased from Heyuan (Shanghai, China). Lipofectamine 2000 (Invitrogen) was used to transfect plasmid into cells following the manufacturer’s instructions. After transfection of 48 h, the cells were collected and used for the further experiments.
Migration and invasion assay
To examine the migratory ability of cells in vitro, the transwell chamber assay was performed. Cells were placed in the upper chamber of a 24-well transwell unit (7 × 104 cells/well) with 8-µm polycarbonate nucleopore filters (Corning Costar, Cambridge, MA, USA). The upper compartment contained serum-free medium, while the lower compartment contained medium with 10% fetal bovine serum; the cells were incubated for 24 h in a humidified atmosphere of 5% CO2 at 37°C. The cells adhering to the lower surface of the filter were fixed and counted. The cells from at least five representative fields were analyzed. For the invasion assay, the membrane of the transwell unit was coated with 40 µL Matrigel (BD Biosciences, San Jose, CA, USA) at 37°C for 4 h to form a reconstructed basement membrane. The cells were treated in the same way as the migration assay.
Statistical analysis
SPSS 18.0 was used for the statistical analysis. All values are expressed as the mean ± standard error of the mean (SEM), and all experiments were repeated at least three times. Student’s t-tests were used to determine the statistical significance of the differences between groups. Paired t-test was used for the clinical sample analysis. Differences with p < 0.05 were considered significant. GraphPad Prism 5.0 was used to draw survival curve and graph.
Results
CPEB4 level in glioma tumor tissues
CPEB4 protein and mRNA expression in glioma and non-tumor tissues were analyzed by western blot (Figure 1(a)) and qRT-PCR (Figure 1(b)) in a cohort of 30 patients. We found that compared with glioma tissues, CPEB4 protein and mRNA levels were upregulated in 73.33% (22/30) paired glioma samples and were downregulated in 26.67% (8/30) paired tissues. In this part, we demonstrated that CPEB4 expression was upregulated in glioma tumor tissues (p = 0.019).

CPEB4 protein and mRNA levels are increased in glioma tissues: (a) expression of the CPEB4 protein was analyzed by western blot in tumor tissues (T) and adjacent normal brain tissues (N) in two typical samples, and (b) expression of CPEB4 mRNA was analyzed by qRT-PCR in tumor and adjacent normal brain tissues (N = 30).
Association between CPEB4 protein expression and clinicopathological factors in patients with glioma
CPEB4 potentially contributed to tumor progression, and increased CPEB4 expression could be associated with more advanced stages of malignant disease. IHC was performed to evaluate CPEB4 protein expression in 89 glioma samples. CPEB4 protein in glioma tissues mainly accumulated in the cytoplasm and partially in the nucleus. Positive staining was observed in 57.3% (51/89) of patients, and negative staining (Score: 0, Figure 2(a)) was observed in 42.70% (38/89) of patients. Weak staining (Score: 1+, Figure 2(b)) was observed in 13.48% (12/89) of patients, moderate staining (Score: 2+, Figure 2(c)) was observed in 20.22% (18/89) of patients, and strong staining (Score: 3+, Figure 2(d)) was observed in 23.60% (21/89) of patients. Furthermore, as shown in Table 1, chi-square test suggested that positive CPEB4 expression (Scores: 1+, 2+, 3+) in glioma tissue samples significantly correlated with advanced World Health Organization grade (WHO grade; p < 0.001) and lower KPS score (p < 0.001). However, there were no statistically significant relationships between CPEB4 expression and other clinicopathological variables, such as age (p = 0.52), gender (p = 0.501), and tumor size (p = 0.887).

Immunohistochemical staining of CPEB4 protein in glioma tissues: (a) no, (b) weak, (c) moderate, and (d) strong CPEB4 staining in glioma tissues. Original magnifications of ×400 fold are shown.
Relationship between CPEB4 expression and clinicopathological characteristics in glioma patients (N = 89).

CPEB4: cytoplasmic polyadenylation element binding protein 4; WHO grade: World Health Organization grade; KPS: Karnofsky Performance Status.
CPEB4-positive expression correlated with poor glioma patient survival
For all patients, the follow-up period ranged from 4 to 120 months, with the mean survival time of 42.475 ± 3.211 months. The mean survival time of negative CPEB4 expression group was 62.411 ± 4.743 months, and the positive CPEB4 expression group was 27.037 ± 2.741 months. The Kaplan–Meier analysis was used to compare survival between patient groups with negative CPEB4 expression (N = 38) and positive CPEB4 expression (N = 51). The result indicated that patients with positive CPEB4 expression predicted a worse prognosis (p < 0.001; Figure 3).
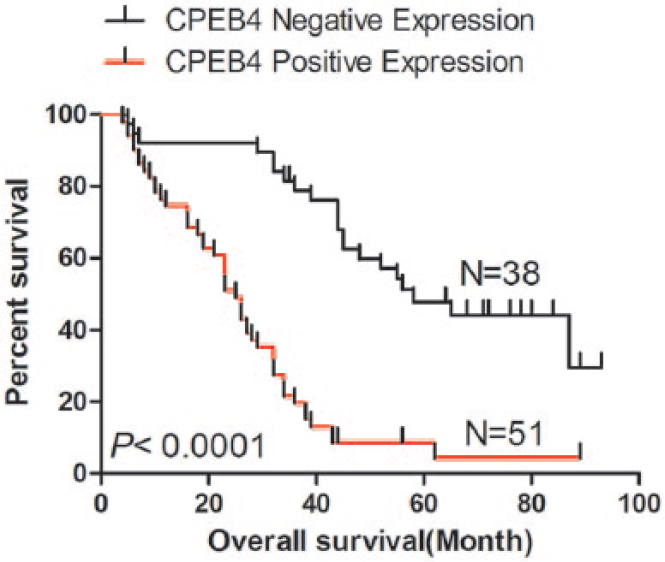
Kaplan–Meier survival curves for glioma patients according to CPEB4 expression levels.
As shown in Table 2, a univariate analysis showed that clinicopathological characteristics including gender (hazard ratio (HR) = 3.109, 95% confidence interval (CI): 1.734–5.574, p < 0.001), KPS score (HR = 0.221, 95% CI: 0.118–0.417, p < 0.001), WHO grade (HR = 2.919, 95% CI: 2.058–4.137, p < 0.001), and CPEB4 expression (HR = 2.059, 95% CI: 1.066–3.975, p = 0.034) were significantly correlated with overall survival. Furthermore, multivariate Cox regression analysis was also performed to evaluate the potential of CPEB4 expression as an independent predictor of overall survival in glioma patients. Besides gender (HR = 3.045, 95% CI: 1.721–5.389, p < 0.001), KPS score (HR = 0.222, 95% CI: 0.118–0.416, p < 0.001), and WHO grade (HR = 2.902, 95% CI: 2.072–4.066, p < 0.001), CPEB4-positive expression (HR = 2.091, 95% CI: 1.093–3.999, p = 0.026) was also an independent prognostic predictor for glioma patients.
Univariate and multivariate analysis showing the overall survival rate for glioma patients.
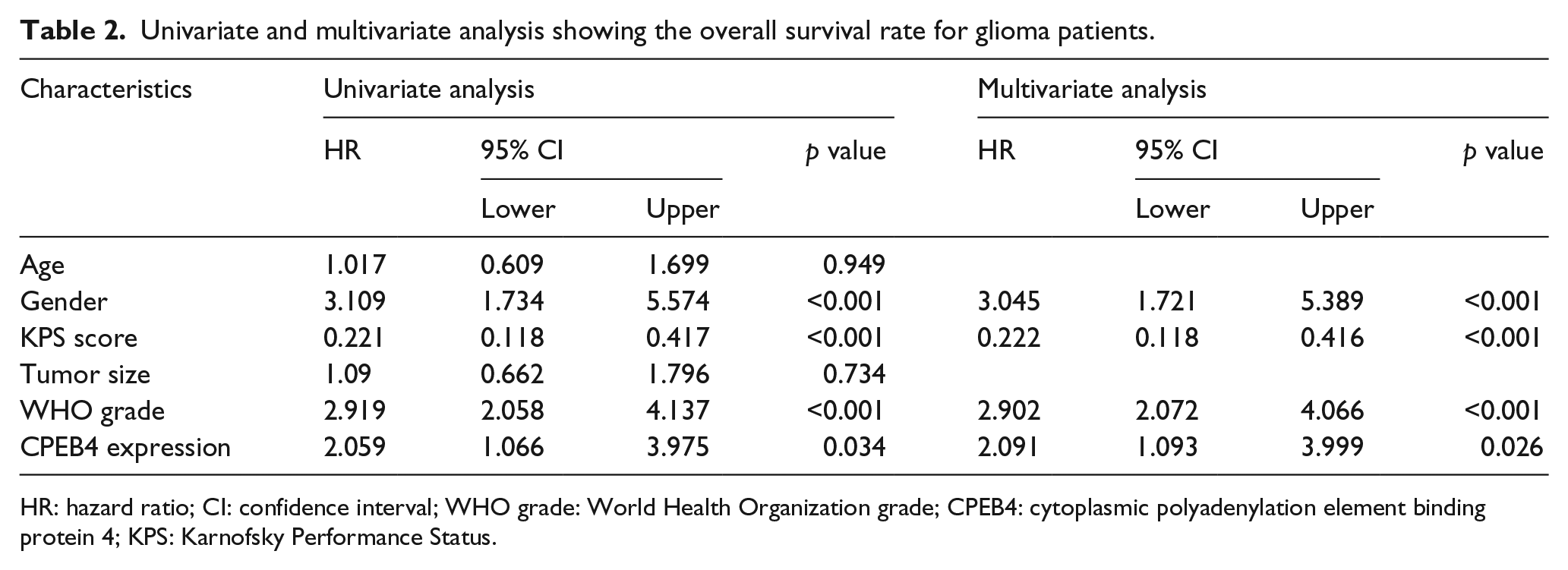
HR: hazard ratio; CI: confidence interval; WHO grade: World Health Organization grade; CPEB4: cytoplasmic polyadenylation element binding protein 4; KPS: Karnofsky Performance Status.
CPEB4 expression in glioma cells
In order to explore the affection of CPEB4 on cell migration and invasion, we first detected CPEB4 expression in glioma cells SKMG-4 and T98G by western blot and qRT-PCR. As shown in Figure 4(a) and (b), we found that CPEB4 level was higher in SKMG-4 cell line than that in T98G cell line. For the following study, we would up- or downregulate CPEB4 expression by CPEB4 overexpression plasmid or siCPEB4 in T98G and SKMG-4 cell lines.

CPEB4 protein and mRNA levels in glioma SKMG-4 and T98G cell lines: (a) expression of the CPEB4 protein was analyzed by western blot in glioma SKMG-4 and T98G cell lines, and (b) expression of CPEB4 mRNA was analyzed by qRT-PCR in glioma SKMG-4 and T98G cell lines.
Effect of CPEB4 downregulation on SKMG-4 cell invasion and migration
To address the relevance of CPEB4 downregulation in cell invasion and migration, we employed siRNA approach to decrease CPEB4 endogenous expression in SKMG-4 cell. The decreased expression of CPEB4 was demonstrated using western blot (Figure 5(a)) and qRT-PCR (Figure 5(b)) in cells transfected with siCPEB4#1, 2, 3. The transfection efficiency showed that siCPEB4#1, 2, 3 could downregulate CPEB4 expression. For the following study, we chose siCPEP4#2 to downregulate CPEB4 expression in glioma SKMG-4 cell. As shown in Figure 5(c), downregulated CPEB4 expression significantly inhibited cell invasion (Figure 5(c), top panel) and migration (Figure 5(c), bottom panel) in SKMG-4. And as shown in Figure 5(d), western blot analysis demonstrated that downregulation of CPEB4 expression could decrease MMP2 and MMP9 protein levels compared with control cells.

Knockdown of CPEB4 inhibited cell migration and invasion in SKMG-4 cell: (a) western blot was applied to detect the protein expression level of CPEB4 in SKMG-4 cells transfected with siCPEB4; (b) qRT-PCR was applied to detect the protein expression level of CPEB4 in SKMG-4 cells transfected with siCPEB4; (c) the transwell assays with (top panel) or without (bottom panel) Matrigel were performed in SKMG-4 cells transfected with siControl and siCPEB4#2; and (d) expression of MMP2 and MMP9 was analyzed by western blot in glioma SKMG-4 cells transfected with siControl and siCPEB4#2.
Effect of CPEB4 upregulation on T98G cell invasion and migration
In SKMG-4 glioma cells, we found that downregulation of CPEB4 could promote cell invasion and migration, and decrease MMP2 and MMP9 expression. In this part, we would detect the effect of CPEB4 expression on cell invasion and migration. The overexpression of CPEB4 was demonstrated using western blot (Figure 6(a)) and qRT-PCR (Figure 6(b)) in cells transfected with pcDNA-CPEB4 plasmid. The pcDNA-CPEB4 plasmid could effectively upregulate CPEB4 protein and mRNA level in T98G cell lines. In the transwell migration assay, upregulation of CPEB4 significantly increased the migration cells of T98G (p < 0.05; Figure 6(c), top panel). Matrigel invasion assay showed that the average cell counts crossing Matrigel-coated membrane in the control group were less than that in the overexpression group of T98G cells, which indicated that upregulation of CPEB4 significantly increased the invasion of glioma (p < 0.05; Figure 6(c), bottom panel). These results show a critical role of CPEB4 in the promotion on glioma migration and invasion. And as shown in Figure 6(d), western blot analysis demonstrated that upregulation of CPEB4 could increase MMP2 and MMP9 protein levels compared with control cells.

Upregulation of CPEB4 promoted cell migration and invasion in T98G cell: (a) western blot was applied to detect the protein expression level of CPEB4 in T98G cells transfected with pcDNA-CPEB4; (b) qRT-PCR was applied to detect the protein expression level of CPEB4 in T98G cells transfected with pcDNA-CPEB4; (c) the transwell assays with (top panel) or without (bottom panel) Matrigel were performed in T98G cells transfected with pcDNA-CPEB4; and (d) expression of MMP2 and MMP9 was analyzed by western blot in glioma T98G cells transfected with pcDNA-CPEB4.
Discussion
CPEB4 has been described as a critical transcription factor that regulated cell biological functions in certain tumors. As a candidate oncogene, CPEB4 plays a crucial role in carcinogenesis. CPEB4 has emerged as a potential biomarker in several cancers, such as colorectal cancer, breast cancer, astrocytic tumor, non-small cell lung cancer, pancreatic cancer, and glioma.18–24
In this study, we found that compared to adjacent normal brain tissues, CPEB4 protein and mRNA levels were significantly increased in glioma tissues. In addition, CPEB4 expression was positively correlated with advanced WHO grade (p < 0.001) and lower KPS score (p < 0.001). Moreover, positive CPEB4 expression was correlated with worse prognosis in patients undergoing surgery for glioma. In invasive ductal carcinoma (IDC), Sun et al. 20 reported that high CPEB4 expression was possibly related to increased histological grading (p = 0.037) and N stage (p < 0.001); patients with high expression of CPEB4 showed shorter overall survival (p = 0.001). High CPEB4 expression was an independent prognostic factor for overall survival. Chen et al. 21 demonstrated that high expression of CPEB4 was significantly correlated with WHO grade, KPS score, tumor size, intracranial hypertension symptom, and epilepsy symptom; and high expression of CPEB4 indicated a poor prognosis for patients with astrocytic tumors. With the WHO grade increasing, the high expression of CPEB4 was further increased in high-grade gliomas, and CPEB4 expression level correlated with advanced WHO classification. 24 Our results were similar with other studies.
In pancreatic cancer, Ortiz-Zapater et al. 25 first explored CPEB4 expression and cancer progression. They found that CPEB4 is overexpressed in human pancreatic cancer; downregulation of CPEB4 in vitro could strongly reduce cell migration and colony formation, while it cannot affect cell proliferation. However, in the functional studies on glioma cell lines, we found that invasion and migration were significantly decreased after CPEB4 knocking down, suggesting that CPEB4 may promote glioma tumor progression by increasing invasiveness; CPEB4 high expression could promote pancreatic cancer invasion and metastasis through transforming growth factor-beta (TGF-β) signaling pathway. 23 In pancreatic cancer, CPEB4 is upregulated in cancer tissues, and could promote cell proliferation, migration, invasion and tumor vascularization; while, CPEB2 is also a biomarker for chemotherapy resistance in pancreatic cancer.23,26,27 CPEB4 also could target BCL2, SMAD3, MMP7, and Vimentin, which indicate that CPEB4 might impact tumor progression in several ways.21,28–31 Although CPEB4 could modulate other genes, it could be modulated by other genes. In hepatocellular carcinoma, gain-of-function and loss-of-function assays revealed that miR-550a markedly promoted hepatocellular carcinoma cell (HCC) migration and invasion through binding and increasing CPEB4. 32 In non–small cell lung cancer and colorectal cancer, miR-1246 and miR-203 may promote tumor metastasis and progression by targeting CPEB4.18,22
MMPs play an important role in tumor metastasis. MMP2 and MMP9 are considered to be overexpressed and degrade type-IV collagen in highly metastatic cancers.33–35 Yang et al. 36 reported that thymoquinone (TQ) could inhibit cell migration and invasion by decreasing the expression of MMP2 and MMP9. He et al. 37 demonstrated that elevated STMN1 could promote tumor migration and invasion by increasing MMP2 and MMP9. In this study, we firstly reported that knockdown of CPEB4 in glioma cells could decrease the expression of MMP2 and MMP9; and upregulation of CPEB4 could increase the expression of MMP2 and MMP9. The results indicated that MMP2 and MMP9 may be regulated by CPEB4-mediated translational control in glioma.
In conclusion, this study suggests that CPEB4 is an independent prognostic factor and could be used to identify glioma patients at high risk of a poor clinical outcome. Furthermore, the potential for developing CPEB4 as a candidate therapeutic target for glioma should be further investigated. The discovery of the interaction between CPEB4 and MMP2 and MMP9 might provide us a new insight for gene-targeted tumor-specific therapy in glioma.
Footnotes
Acknowledgements
L.Z. and W.D. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Funding
This study was supported by the Science and Technology Research Project of He Nan Province (Grant No. 162102310162).
