Abstract
One of the mechanisms for epigenetic silencing of tumor suppressor genes is hypermethylation of cytosine residue at CpG islands at their promoter region that contributes to malignant progression of tumor. Therefore, activation of tumor suppressor genes that have been silenced by promoter methylation is considered to be very attractive molecular target for cancer therapy. Epigenetic silencing of glutathione S-transferase pi 1, a tumor suppressor gene, is involved in various types of cancers including breast cancer. Epigenetic silencing of tumor suppressor genes can be reversed by several molecules including natural compounds such as polyphenols that can act as a hypomethylating agent. Curcumin has been found to specifically target various tumor suppressor genes and alter their expression. To check the effect of curcumin on the methylation pattern of glutathione S-transferase pi 1 gene in MCF-7 breast cancer cell line in dose-dependent manner. To check the reversal of methylation pattern of hypermethylated glutathione S-transferase pi 1, MCF-7 breast cancer cell line was treated with different concentrations of curcumin for different time periods. DNA and proteins of treated and untreated cell lines were isolated, and methylation status of the promoter region of glutathione S-transferase pi 1 was analyzed using methylation-specific polymerase chain reaction assay, and expression of this gene was analyzed by immunoblotting using specific antibodies against glutathione S-transferase pi 1. A very low and a nontoxic concentration (10 µM) of curcumin treatment was able to reverse the hypermethylation and led to reactivation of glutathione S-transferase pi 1 protein expression in MCF-7 cells after 72 h of treatment, although the IC50 value of curcumin was found to be at 20 µM. However, curcumin less than 3 µM of curcumin could not alter the promoter methylation pattern of glutathione S-transferase pi 1. Treatment of breast cancer MCF-7 cells with curcumin causes complete reversal of glutathione S-transferase pi 1 promoter hypermethylation and leads to re-expression of glutathione S-transferase pi 1, suggesting it to be an excellent nontoxic hypomethylating agent.
Introduction
Breast cancer remains the most common malignancy in women worldwide and is the leading cause of cancer-related mortality in females in developed and developing regions. 1 Epigenetic silencing of tumor suppressor genes (TSGs) is a well-established carcinogenic process. Therefore, reactivation of TSGs that have been silenced by promoter methylation is a very striking molecular target for cancer therapy. 2 Epigenetic silencing of TSGs can be reversed by several natural compounds such as curcumin, a yellow spice and the active component of the perennial herb Curcuma longa, which can act as a hypomethylating agent. Curcumin covalently blocks the catalytic thiolate of DNA methyltransferase 1 (DNMT1) to exert its inhibitory effect on DNA methylation. Curcumin exists predominantly in solution as the enol form, which serves as an acceptor to covalently block the catalytic thiol group in DNMT1 through the C3 keto-enol moiety of the curcumin compounds.
Glutathione-S-transferases (GSTs) are a supergene family of isoenzymes implicated in the detoxification of a wide range of xenobiotics and chemotherapeutic agents. GSTs catalyze the conjugation of glutathione with electrophilic compounds including carcinogens and exogenous drugs, resulting in less toxic and more readily excreted metabolites. There are four distinct classes (α, µ, θ, and π) of isozymes in the GST superfamily, each encoded by a different gene at different loci and with peculiar structural and functional characteristics. The pi-class glutathione-S-transferase (GST-π) is of particular interest in the study of cancer biology. GST-π is expressed in normal tissues at varying levels in different cell types, and abnormal GST-π activity and expression have been reported in a wide range of tumors including those of the breast and kidney.3,4 GST-π is encoded by the glutathione-S-transferase pi 1 (GSTP1) gene located in chromosome 11. The 5′ region of GST-π contains a CpG island, and in cancer cells, the hypermethylation of the CG-rich area in the promoter region of TSGs correlates with its loss of transcription, as demonstrated for many TSGs.
Hypermethylation of regulatory sequences at GST-π associated with the loss of GST-π expression has been found in the vast majority of human prostate carcinomas with poor prognosis. 5 GSTP1 gene is also hypermethylated in 31% of primary tumor tissues and 55% in breast cancer cell lines. 6 Undoubtedly, it is the best DNA methylation marker for cancer detection and one of the most likely genes to be succeeded as an epigenetic biomarker. However, little is known about epigenetic silencing of GST-π gene by promoter hypermethylation in the precursors of breast cancer and other tumor types. To understand the mechanisms of regulation of the human π class, GSTP1 gene in breast cancer cells is of particular importance to study breast carcinogenesis which opens new avenues for cancer chemoprevention based on the inhibition or reversal of epigenetic alterations before the onset of cancer using DNA methylation as cancer biomarkers for better patient prognosis.
There are several synthetic demethylating agents currently being evaluated in preclinical and clinical studies. 5-azacytidine and 5-aza-2-deoxycytidine are the most studied and were developed over 30 years ago as classical cytotoxic agents but were subsequently discovered to be effective DNA methylation inhibitors. Some other drugs such as procainamide and hydralazine are also in different stages of trial. 2 As most of the synthetic compounds may have cytotoxic effects, the focus is on natural products for the epigenetic reversal of phytochemicals derived from fruits and vegetables, referred to as chemopreventive agents, including genistein, diallyl sulfide, S-allyl cysteine, allicin, lycopene, curcumin, 6-gingerol, ursolic acid, silymarin, anethol, catechins, and engenol. 7 Curcumin is a natural phytochemical and is presently under a great deal of inspection from cancer investigators because of its chemopreventive properties against human malignancies. Curcumin has great potential as an epigenetic agent. Previous studies have shown that curcumin, an herbal antioxidant, can reverse the hypermethylation of TSGs like retinoic acid receptor beta (RAR-β) gene in cervical cancer. 8
Unlike genetic alterations, epigenetic changes can be modified by the environment, diet, or pharmacological intervention. This characteristic has increased enthusiasm for developing therapeutic strategies by targeting the various epigenetic factors, such as histone deacetylases (HDAC), histone acetyltransferases (HAT), DNA methyltransferases (DNMTs), and micro RNAs (miRNAs) by dietary polyphenols such as curcumin.
Considering the potential role of promoter hypermethylation in silencing of TSG in cancer and the role of GSTP1, this study has been designed to study the hypermethylation status of GSTP1 and to study the reversal of hypermethylation of GSTP1 using a nontoxic herbal compound curcumin.
Materials and methods
Materials
The breast cancer cell line, MCF-7, was procured from National Centre for Cell Sciences (Pune, India). MCF-7 cell line was well maintained in culture growth media DMEM (PAN-Biotech GmbH, Aidenbach, Germany), supplemented with 10% fetal bovine serum (FBS) (Sigma-Aldrich, St. Louis, MO, USA), 1% penicillin/streptomycin (Sigma-Aldrich) and incubated at 37°C, 5% CO2, and 95% humidity. Curcumin and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) were obtained from Sigma-Aldrich.
Treatment of MCF-7 breast cancer cell line with curcumin
The MCF-7 cells were grown until 60%–70% of confluence was reached. The cells were treated with curcumin at various concentrations, that is, 0, 1, 3, 5, 10, 20, 30, 50, and 100 µM for different time periods, that is, 24, 48, and 72 h after that. The treated cells were used for further experiments. The cytotoxic effects of curcumin against MCF-7 were determined by MTT dye uptake method. The cells were incubated in triplicate in a 96-well plate in the presence or absence of curcumin in a final volume of 0.1 mL for 24, 48, and 72 h at 37°C in a CO2 incubator.
MTT assay
MTT assay was performed as described previously. 9 Briefly, 5000 exponentially growing cells per well were seeded in 96-well plates. After curcumin treatment and 4 h prior to completion of incubation period, 10 µL of MTT (Sigma-Aldrich) reagent was added to each well. After 4 h, MTT solution was removed, and the blue crystalline precipitate in each well was dissolved in dimethyl sulfoxide. The optical density at a wavelength 570 nm was measured using a 96-well multiscanner autoreader (Biotek, Winooski, VT, USA) with the lysis buffer serving as blank. Cell viability was estimated using the following formula
Morphological changes
Morphological changes in curcumin-treated MCF-7 cells were observed through a phase-contrast microscope (Nikon Eclipse E400, Nikon Corporation, Tokyo, Japan) after 24, 48, and 72 h of treatment with curcumin at IC50 values along with proper controls.
Genomic DNA extraction from curcumin-treated MCF-7 cells
The genomic DNA was extracted following treatment of MCF-7 cells with 10 µM/mL curcumin for 24 h using phenol and chloroform method. 10 In brief, pelleted cells were first washed with 1× phosphate-buffered saline (PBS) and then 400 µL of 1× Tris–ethylenediaminetetraacetic acid (EDTA) (TE) was further added. After mixing well, 200 µL of tissue lysis buffer (3% sodium dodecyl sulfate (SDS) in 2× Tris–EDTA) was added followed by addition of 6 µL proteinase K (Sigma-Aldrich). After overnight incubation at 37°C, equal amount (600 µL) of Tris–EDTA equilibrated phenol was added and subjected to overhead shaker for 15 min at room temperature. It was then centrifuged (Sigma 1-14K, Osterode am Harz, Germany) at 10,000 r/min at 4°C for 10 min, and supernatant was carefully aspirated with the help of micropipette. To the supernatant, equal volume of phenol and chlorofom-isoamyl alcohol (CIA) (Sigma-Aldrich) in the ratio 25:24:1 was added. After overhead shaking for 15 min at room temperature, it was centrifuged at 10,000 r/min at 4°C for 10 min. The supernatant was again collected carefully, and to this, an equal amount of CIA was added and shaked overhead for 15 min at room temperature. After centrifugation at 10,000 r/min at 4°C for 10 min, the supernatant was aspirated, and to this, around 1/10th volume of chilled sodium acetate (~50 µL) and equal volume of isopropanol (Sigma-Aldrich) was added. After keeping at −70°C for 2 h or at −20°C for overnight, it was centrifuged for 15 min at 10,000 r/min at 4°C. The pellet was washed with 70% ethanol by centrifuging at 8000 r/min at 4°C for 5 min. The pellet was air dried at room temperature overnight and then dissolved in 200 µL of 1× Tris–EDTA buffer (Sigma-Aldrich).
DNA fragmentation assay
During DNA fragmentation assay, 1 × 106 cells were treated with curcumin at the IC50 value for 48 h. Cellular genomic DNA of treated cells was extracted from the cells using phenol-CIA method. 10 Briefly, treated and untreated cells were trypsinized with 0.25% trypsin (Sigma-Aldrich) and collected using centrifugation (200g, 10 min), washed twice in cold PBS (10 mM), and resuspended in hypotonic lysis buffer (5 mM Tris, 20 mM EDTA, pH 7.4) containing 0.5% Triton X-100 (Sigma-Aldrich) for 30 min at 4°C. The lysates were centrifuged at 13,000g for 15 min at 4°C. Genomic DNA was extracted from the supernatant with equal volume of phenol-CIA, precipitated by addition of two volumes of absolute ethanol and 0.1 volume of 3 mM sodium acetate and treated with RNase A (500 U/mL) 37°C for 3 h. The pattern of fragmentation was analyzed on 2% agarose gel.
Sodium bisulfite modification of DNA
Bisulfite modification was done using EZ DNA Methylation-Gold™ Kit (Zymo Research, Irvine, CA, USA). To 130 µL of the CT Conversion Reagent (20 µL) of DNA, sample was added in Eppendorf tube, sample tube was incubated at 98°C for 10 min, 64°C for 2.5 h. M-binding buffer (600 µL) to a Zymo-Spin™ IC Column (Zymo Research, CA, USA) was added and column was placed into the provided collection tube, sample was loaded in column containing the M-binding buffer and was centrifuged at >10,000g for 30 s, and flow-through was discarded then 100 µL of M-wash buffer was added to the column and centrifuged. Then, 200 µL of M-desulfonation buffer was added to column and incubated for 15–20 min, 200 µL of M-wash buffer was added to the column and centrifuged, and column was placed into a 1.5 mL microcentrifuge tube, 10 µL of M-elution buffer was added directly to the column matrix followed by centrifuged at >10,000g for 30 s to elute the DNA.
Methylation-specific polymerase chain reaction
Methylation-specific polymerase chain reaction (PCR) was carried out on the bisulfite-modified DNA samples. The PCR mixture contained 10× PCR buffer (16.6 mM ammonium sulfate/67 Mm Tris, pH 8.8/6.7 mM MgCl2/10 mM 2-mercaptoethanol), deoxynucleotide triphosphates (dNTPs) (each at 1.25 mM; Sigma-Aldrich), primers (300 ng each per reaction), and bisulfite-modified DNA (50–100 ng) or unmodified DNA (50–100 ng) in a final volume of 50 µL. Then, 1.25 units of Taq polymerase (Bangalore Genei, Bangalore, India) was used for the final volume. Amplification was carried out in a BioRad thermal cycler for 39 cycles (30 s at 95°C, 30 s at the annealing temperature listed in Table 1, and 30 s at 72°C), followed by a final 7-min extension at 72°C. Controls without DNA were performed for each set of PCRs. Each PCR product along with loading dye (10 µL + l µL) was directly loaded onto 2% agarose gels, stained with ethidium bromide, and directly visualized under ultraviolet (UV) illumination.
PCR primers used for GSTP1 MSP.
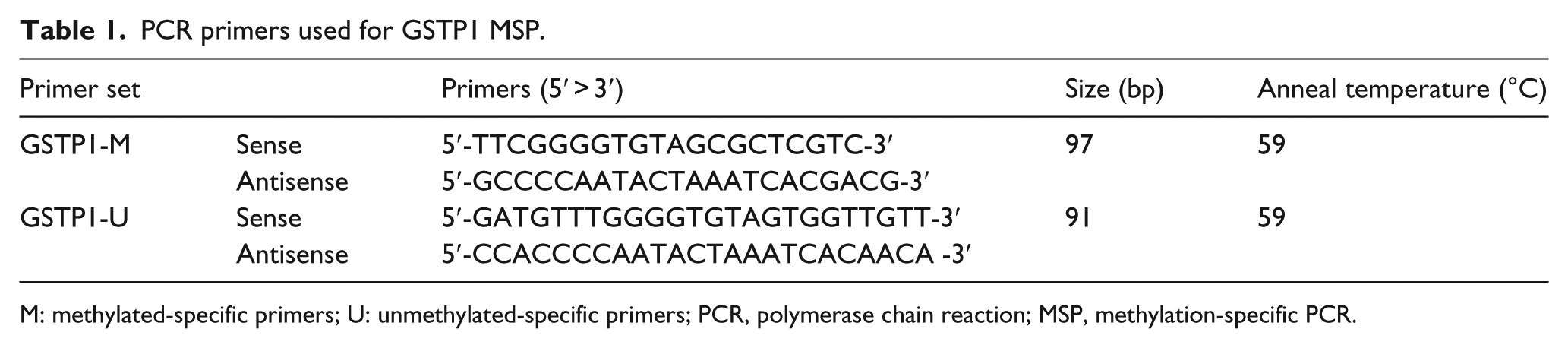
M: methylated-specific primers; U: unmethylated-specific primers; PCR, polymerase chain reaction; MSP, methylation-specific PCR.
Protein extraction and immunoblotting
Whole-cell lysate was extracted following treatment of MCF-7 cells with 10 µM/mL curcumin for 24 h, resolved by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), electrotransferred to Immobilon-P membranes (Millipore Corporation, Bedford, MA, USA) by standard method described by Dignam. 11 Briefly, PBS-washed cells were extracted in buffer C, freshly supplemented with dithiotheriotol (DTT) and polymethyl sulfonic acid (PMSF) and were kept in −70°C or liquid nitrogen. Then, 15–20 µg of cellular protein extracts were separated in 12% SDS-polyacrylamide gels and electrotransferred to Immobilon-P Millipore membrane and probed with monoclonal mouse antibody to the corresponding protein (GSTP1, sc-66000; Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA). The desired protein bands were detected by anti-mouse IgG antibody conjugated with horseradish peroxidase (sc-2302; Santa Cruz Biotechnology, Inc.), using the Amersham ECL detection system (GE Healthcare Life Sciences, Buckinghamshire, UK). The expression was semi-quantitated with respect to expression of β-actin (sc-47778, mouse monoclonal antibody; Santa Cruz Biotechnology, Inc.) which was used as internal control.
Statistical analysis
Data were presented as mean ± standard deviation (SD). Statistical analysis was performed with SPSS program (version 11.5; SPSS Inc., Chicago, IL, USA). Comparisons of mean values among different groups were performed using analysis of variance (ANOVA). A p value of <0.05 was considered significant.
Results
Effect of curcumin treatment on cell viability
Within the concentration range of 0–100 µM, curcumin reduced the viability of the MCF-7 cells in a dose-dependent manner. There was a significant decreased in cell viability after 3–100 µM of curcumin treatment (p = 0.001; Figure 1). The IC50 value of the curcumin was detected to be 20 µM in MCF-7 breast cancer cell line. This substantiates the anti-proliferative effect of the curcumin in vitro on the tested MCF-7 breast cancer cell line.

(a) 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay-dose kinetic curve and (b) viability percentage inhibition in MCF-7 after 24 h of treatment with various concentration of curcumin, that is, 0, 1, 3, 10, 20, 30, 50, and 100 µM.
Effect of curcumin treatment on morphology of cells
The MCF-7 cells were treated with different concentrations (0, 1, 3, 5, 10, 20, and 30 µM) of the curcumin for 24 h and viewed under phase-contrast microscope to visualize any morphological changes. The distinct morphological changes in the cells were observed which showed not much visible change in the cells up to a concentration of 10 µM of curcumin. But at higher concentration of 20 and 30 µM, there was more growth inhibition with increased amount of cell deaths (Figure 2).

Morphological changes observed in MCF-7 after treatment with various concentrations of curcumin, that is, 1, 3, 10, 20, 30 µM. UT: untreated (magnification, ×20).
Curcumin-induced DNA damage in MCF-7 cells
DNA laddering assay is often considered as a common marker for apoptosis.12,13 Accordingly, we set out to examine whether DNA laddering was happening in the curcumin-treated MCF-7 cells. As evident from the results presented in Figure 3, smearing of the genomic DNA was observed with a DNA ladder/fragments at the bottom of the gel in the curcumin-treated cell line at concentration of 100 µM cells.

Curcumin-induced DNA fragmentation indicative of apoptosis in MCF-7 cells. Multiple arrows show the fragmented DNA.
Methylation-specific PCR for GSTP1 gene
Methylation-specific PCR (MSP) of GSTP1 gene for analysis of promoter hypermethylation was performed using methylation-specific primers in breast cancer cell line MCF-7 treated with various concentrations of curcumin (1, 3, 5, 10, 20, and 30 µM) for time intervals of 48 and 72 h. Untreated MCF-7 was used as a positive control because it is hypermethylated for GSTP1, and MDA-MB-231 was used as a negative control because it is unmethylated for GSTP1 promoter region.
GSTP1 promoter was confirmed to be hypermethylated in DNA isolated from untreated MCF-7 cell line with the amplification of specific band of 97 bp by MSP (Figure 4). The GSTP1 promoter remained methylated at 1 and 3 µM concentration of curcumin, as shown in Figure 4(a) and (b). However, at 10-µM curcumin treatment, partial reversal of hypermethylation was observed after evident from the partial and complete absence of MSP band after 42 and 72 h of treatment. Intriguingly, the methylation-specific band reappeared at 20 and 30 µM of curcumin concentration both at 48 and 72 h of treatment (Figure 4(a) and (b)).

(a) and (b) Methylation-specific PCR products showing a band of GSTP1 gene in MCF-7 cell line after (a) 48 h and (b) 72 h of treatment with various concentrations of curcumin, respectively. M represents ΦX174 DNA-Hae III digested marker, lane P is positive control (untreated MCF-7 DNA), and lane N is negative control (MDA-MB-231 cell line DNA).
Reversal of GSTP1 promoter methylation in treated as well as untreated MCF-7 cells was performed using unmethylation-specific primers (Figure 5). DNA isolated from GSTP1 promoter region of untreated MCF-7 served as a negative control because it is methylated for GSTP1 promoter region.

(a) and (b) Unmethylation-specific PCR band of GSTP1 gene in MCF-7 cell line after (a) 48 h and (b) 72 h of treatment with various concentrations of curcumin, respectively. M represents ΦX174 DNA-Hae III digested marker, Lane P is positive control (MDA-MB-231 cell line DNA), and lane N is negative control (untreated MCF-7 DNA).
As shown in Figure 5(a) and (b), GSTP1 promoter was found to be methylated in DNA isolated from untreated MCF-7 cell line because unmethylation-specific band of 91 bp was not detected. Corroborating the results of PCR of methylation-specific bands, it was found that at 1 and 3 µM concentration of curcumin treatment, unmethylation-specific band was not detected. Similarly, at 10 µM curcumin concentration, unmethylation-specific band of 91 bp was detected showing reversal of hypermethylation, whereas at 20 and 30 µM, intensity of unmethylation-specific band decreased both at 48 and 72 h.
GSTP1 protein expression in curcumin-treated MCF-7 cell line
The western blot analysis of breast cancer cell line MCF-7 was performed to check protein expression of GSTP1 before and after treatment with various concentrations of curcumin (1, 3, 10, 20, and 30 µM) for various time periods (48 and 72 h). Untreated MCF-7 was used as a negative control because GSTP1 gene is not expressed in MCF-7 cell line, and MDA-MB-231 was used as a positive control because GSTP1 gene is expressed in this cell line. GSTP1 protein expression was not observed in the untreated MCF-7 cell line (Figure 6(a) and (b)).

(a) and (b) Protein expression of GSTP1 gene in MCF-7 cell line after 48 and 72 h of treatment with different concentrations of curcumin, respectively. (c) Expression of β-actin as a control.
Similarly, at 1 and 3 µM concentrations of curcumin protein, expression was not observed. However, at 10 µM curcumin concentration, GSTP1 gene expression was maximum that gradually decreased at 20 and 30 µM. There was a time-dependent increase in expression of GSTP1 gene where treatment for 72 h showed more GSTP1 protein expression as compared to 48 h of treatment (Figure 6(a) and (b)).
Discussion
Epigenetic silencing of TSGs is emerging as a well-established oncogenic process. 2 Epigenetic alteration is a reversible process, and this phenomenon establishes the potential use of DNMT as a smart concept of cancer therapy. 14 Much of contemporary research focused on the study of epigenetic changes such as promoter hypermethylation of GSTs and BRCA1 (breast cancer type1 susceptibility protein), resulting in breast carcinogenesis and their possible reversal using natural compounds such as curcumin and tea polyphenol EGCG.
GSTP1 is involved in the detoxification of electrophilic compounds (such as carcinogens and cytotoxic drugs) by glutathione conjugation and believed to play a role in the protection of DNA oxidative damage. Abnormal GSTP1 activity and expression have been reported in a wide range of tumors including those of the breast.3,4 GSTP1 gene promoter has been found to be hypermethylated in more than 31% of primary breast cancer tumor tissue and 55% in breast cancer cell lines. 6 Hypermethylation of TSGs leading to silencing of their expression is a common feature in majority of cancer cases, and therefore, attempts are made to use hypomethylating agents to make these genes to express normally in order to reverse tumorigenicity.
Curcumin, a phytochemical present in C. longa, has been shown to inhibit the DNMT activity thereby causing promoter demethylation. Molecular docking studies on the interaction of curcumin and DNA methyltransferases suggest that curcumin covalently block the catalytic thiolate of C1226 of DNMT1 to exert its inhibitory effects. 15 Tea polyphenol (catechin epicatechin and (−)EGCG) and bioflavonoids (quercetin myricetin) inhibit DNMT and DNMT1-mediated DNA methylation in concentration-dependent manner. 16 EGCG has been shown to cause demethylation of CpG island in promoter and reactivation of methylation silence genes such as p16, retinoic acid receptor β2 (RAR-β2), O6-alkylguanine DNA alkyltransferase (O 6 -MGMT), and GSTP1 in human esophageal, colon, prostate, and mammary cancer cell lines. 17
This study was designed to analyze reversal of GSTP1 promoter hypermethylation by curcumin in breast cancer cell line MCF-7 in dose-dependent manner. MSP was carried out to study the ability of curcumin to cause reversal of hypermethylation and led to reactivation of GSTP1 protein expression in MCF-7 cells. The IC50 of curcumin was found to be at 20 µM, whereas the effective concentration of curcumin for DNA demethylation and reactivation of GSTP1 was found to be 10 µM. At concentration less than 10 µM, that is, at 1 and 3 µM, curcumin treatment did not alter the promoter methylation pattern. It indicates that at these concentrations, reversal of hypermethylation did not occur because these concentrations could not be enough to inhibit DNMTs.
The intensity of unmethylation-specific band as well as protein expression of GSTP1 was detected to maximum at 10 µM concentration of curcumin and that interestingly expression decreased in progressive manner at 20 and 30 µM curcumin. Such biphasic response of curcumin has also been observed on the proteosomal activity in human fibroblast and telomerase-immortalized mesenchymal bone marrow stem cells and human keratinocyte. 18 This biphasic dose response is known as hormesis where low doses of compound show beneficial effects, whereas higher doses lead to deleterious effects. 19
The reversal of hypermethylation and re-expression of the GSTP1 gene by the curcumin treatment were found almost similar to that produced by unmethylated GSTP1 gene in MDA-MB 231. Similar to this study, reversal of hypermethylation by curcumin in RAR-β2 has been reported in SiHa, a cervical cell line. 8 In a recent study by Khor et al., 20 demethylation of Nrf2 (nuclear factor erythroid 2; (NF-E2)-related factor 2), master regulator of cellular antioxidant defense system, by curcumin was found to be associated with the re-expression of Nrf2 and its downregulation target gene, NQO-1; NAD(P)H dehydrogenase (quinone 1) both at the messenger RNA (mRNA) and protein levels. Similarly, curcumin treatment on prostate cancer cells LNCaP also led to demethylation of the first 14 CpG sites of the CpG island of the Neurog1 (neurogenin 1) gene and restores its expression. 21 Curcumin has also been shown to re-induce the expression of Wnt inhibitory factor-1 gene, via demethylation, that is known to be hypermethylated at promoter region in lung cancer cell lines and tissues. 22
Since curcumin is an important component of our diet and does not have any cytotoxic effect on normal cells unlike other demethylating chemicals, it may be used as part of adjunct therapy to reverse promoter hypermethylation of cancer-associated genes. Therefore, it may serve as a lead compound in combinatorial cancer therapy.
Conclusion
Treatment of breast cancer MCF-7 cells with curcumin for 72 h at 10 µM causes complete reversal of GSTP1 promoter hypermethylation and leads to re-expression of GSTP1 protein suggesting curcumin to be an excellent nontoxic hypomethylating agent. Curcumin at lower concentration causes reversal of hypermethylation of GSTP1, and at higher concentration (20 and 30 µM) re-expression of GSTP1 decreases due to hormesis.
It is intriguing that although curcumin acted as a biphasic molecule, at the concentration higher than the optimum concentration of 10 µM, methylation of GSTP1 promoter gradually reappeared.
Footnotes
Acknowledgements
The authors would like to acknowledge Indian Council of Medical Research, Govt. of India, New Delhi, for the award of Senior Research Fellowship (3/2/2/202/2009.NCDIII) to Umesh Kumar.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.

