Abstract
Microvesicles are released by different cell types and shuttle mRNAs and microRNAs which have the possibility to transfer genetic information to a target cell and alter its function. Acute myeloid leukemia is a malignant disorder, and leukemic cells occupy all the bone marrow microenvironment. In this study, we investigate the effect of leukemia microvesicles on healthy umbilical cord blood hematopoietic stem cells to find evidence of cell information transferring. Leukemia microvesicles were isolated from acute myeloid leukemia patients and were co-incubated with healthy hematopoietic stem cells. After 7 days, cell count, hematopoietic stem cell–specific cluster of differentiation (CD) markers, colony-forming unit assay, and some microRNA gene expressions were assessed. Data showed a higher number of hematopoietic stem cells after being treated with leukemia microvesicles compared with control (treated with no microvesicles) and normal (treated with normal microvesicles) groups. Also, increased levels of microRNA-21 and microRNA-29a genes were observed in this group, while colony-forming ability was still maintained and high ranges of CD34+, CD34+CD38−, CD90+, and CD117+ phenotypes were observed as stemness signs. Our results suggest that leukemia microvesicles are able to induce some effects on healthy hematopoietic stem cells such as promoting cell survival and some microRNAs deregulation, while stemness is maintained.
Introduction
Microvesicles are created by different cells in physiological or pathological stages via direct budding of the plasma membrane into the extracellular space with a maximum size of 1 µm in diameter.1–3 Their contents include mRNA4,5 and microRNA 6 as well as protein. 7 Their interactions with target cells directly impact phenotypic and functional characteristics of these cells. 8 Reprogramming of hematopoietic stem cells (HSCs)/progenitor cells by embryonic stem cell–derived microvesicles was one of the first signs of cellular phenotype transfer by microvesicles reported by professor Ratajczak and his team. 9 Pulmonary epithelial cell–specific mRNA expression in marrow cells after co-culturing with rat lung fragments separated by 0.04-µm pore-size membrane is further evidence of the power of microvesicles to induce gene expression and transform phenotype in a target cell. 4
Acute myeloid leukemia (AML) is an aggressive clonal myeloid neoplasm, initiated by a leukemic stem cell (LSC) and characterized by an arrest in differentiation and an uncontrolled proliferation of myeloid precursors in the bone marrow. While leukemic cells occupy all spaces in bone marrow microenvironment, healthy HSCs exist in this place and proliferate and differentiate to healthy blood progenitors.10,11 Although LSCs and healthy HSCs are similar in self-renewing ability and differentiate to progenitor cells, different microRNA signatures exist between them which indicates the role of microRNAs in transforming healthy HSCs to LSCs in the early stage of leukemogenesis. 12 MicroRNAs, as one of the most important post-transcription regulatory elements, are transferred among the cells by microvesicles and affect them based on the information of the cell origin.8,13
In the present study, for the first time, we investigated the probable effect of leukemic bone marrow cell–derived microvesicles on healthy HSCs from umbilical cord blood to find evidence for transferring genetic information between leukemic and healthy cells.
Materials and methods
Patient and normal samples
Bone marrow samples from patients and normal cases were obtained in heparin tubes after written informed consent forms were signed. Patients were suspected to have AML and referred to the hospital from April to November 2014 for diagnosis. Normal samples were obtained from healthy volunteers for bone marrow transplant, and use of all these samples was approved by the Medical Research Ethics Committee of Tarbiat Modares University.
Cell culture
Bone marrow samples mixed with heparin were subjected to ammonium chloride for red blood cell lysis. The remaining cells including both adherent and suspended cells were cultured in Roswell Park Memorial Institute (RPMI) medium supplemented with 0.5% bovine serum albumin (BSA) at 37°C, 5% CO2, and at least 90% humidity overnight. The next day, the cell supernatants were collected and frozen at −20°C until microvesicle isolation.
Microvesicle isolation, characterization, and measurement
Isolation
Cell-free supernatant was obtained by ultra-centrifugation at 500g for 10 min. Cell debris was excluded by centrifugation at 7000g for 20 min at 4°C. Microvesicle pellets were observed after 1 h of centrifugation at 20,000g at 4°C; this step was repeated to wash microvesicles with phosphate-buffered saline (PBS). Suspended microvesicles in PBS were used fresh for both size characterization and protein measurement.
Dynamic light scattering technique
Suspended microvesicles in PBS solvent were used for size analysis by Zetasizer Nano Range Instrument (Malvern, UK) which measures the size of each particle based on viscosity and refractory indices of the solvent at a wavelength of 630 nm. Viscosity and refractory indices of PBS were 1.08 and 1.33, respectively.
Transmission electron microscopy
Ten microliters of suspended microvesicles was placed on formvar–carbon-coated grid and then subjected to 2% uranyl acetate for 30 s as negative staining to show the quality of isolated microvesicles by electron microscope.
Protein measurement
Microvesicle protein was estimated by Bradford assay, using BSA as a standard protein. Each sample’s optical density (OD) was read by a spectrophotometer instrument, and protein concentration was calculated based on the normal curve.
Umbilical cord blood HSC sorting
Cord blood samples were collected in citrate phosphate dextrose adenine (CPDA1) reagent. Mononuclear cells (MNCs) were isolated by Ficoll-Hypaque gradient centrifugation at 500g at 21°C for 30 min. HSCs were then sorted by magnetic associated cell sort (MACS) technique using CD34 magnetic beads (Miltenyi Biotec, Germany). Briefly, 100 µL of FcR blocking reagent and 100 µL of CD34 MicroBeads were added for up to 108 MNCs according to the manufacturer’s instructions. Cells were incubated at 4°C for 30 min and then washed by MACS buffer at 500g for 10 min in 4°C. Suspended cells in MACS buffer were passed through magnetic column, and only HSCs with CD34 antigen were captured and then released by exiting the column from magnetic area.
Co-incubation of HSCs and microvesicles
Three groups were designed in this study, namely, a control group containing HSCs without any microvesicles, a normal group containing HSCs incubated with normal bone marrow cell–derived microvesicles, and a leukemia group containing HSCs incubated with leukemic bone marrow cell–derived microvesicles. A total of 55,000 sorted HSCs were seeded in each well of a 24-well plate in 500-µL Stemline medium (Sigma–Aldrich, USA) containing 50 ng/mL Fms-like tyrosine kinase 3 (FLT3; ORF Genetics, Iceland) and thrombopoietin (TPO; PeproTech, UK) recombinant growth factors. In normal and leukemia groups, 30 µg/mL isolated microvesicles were also added to each well. Plates were incubated at 37°C, 5% CO2, and at least 90% humidity for 7 days.
Cell count
After 7 days, cells were centrifuged at 500g to completely separate them from probable remained microvesicles in supernatant, and then, they were washed by PBS. Viable cells were counted in a hemocytometer chamber using Trypan Blue staining.
Flow cytometry analysis for HSC-specific CD markers
Cells were stained with CD34 (PE-eBioscience, USA), CD38 (PE-CY5-eBioscience, USA), CD117 (PE-Becton Dickinson), and CD90 (APC-eBioscience, USA) antibodies in flow cytometry staining buffer to be analyzed by BD FACSAria cell sorter (BD Biosciences, San Jose, CA), and data were analyzed by FlowJo 2.7.4 software program.
CFU assay
Three thousand cells in 100-µL RPMI supplemented with 2% fetal bovine serum (FBS) were added to 1-mL MethoCult (Stemcell Technologies, USA) in a 6-well plate and incubated at 37°C, 5% CO2, and complete humidity for 14 days. Colony-forming unit-granulocyte macrophage (CFU-GM), colony-forming unit-erythrocyte (CFU-E), and colony-forming unit-granulocyte erythrocyte monocyte megakaryocyte (CFU-GEMM) were separately counted for each sample.
Quantitative polymerase chain reaction
RNA extraction and cDNA synthesis
Total RNA was extracted from washed cells in PBS using RNX Plus (CinnaGen, Iran) according to the manufacturer’s instructions, and complementary DNA (cDNA) was specifically synthesized (Fermentas) for microRNA-21 and microRNA-29a and Snord 47 using stem loop primers 14 as shown in Table 1.
Primer sequences.

PCR: polymerase chain reaction; F: forward; R: reverse; RT: reverse transcription.
Real-time polymerase chain reaction
Gene expression fold change of selected microRNAs was assessed by relative quantitative real-time polymerase chain reaction (PCR) using Applied Biosystems StepOne real-time system (Applied Biosystems, Foster City, CA, USA) and SYBR Green PCR master mix (TaKaRa, Japan). Primer sequences are provided in Table 1 and ΔΔCt method was used to evaluate the gene expression fold change, normalized with Snord 47.
Statistical analysis
SPSS 22 (Microsoft, Chicago, IL, USA) was used to statistically analyze the results. Three patients, three control, and three normal samples participated in this study in three different experiments. For comparing the means between groups, the one-way analysis of variance (ANOVA) was used for normal distributions and the Kruskal–Wallis test was used for abnormal distributions. p < 0.05 was considered statistically significant.
Results
Characteristics of patients and normal cases
Three bone marrow aspiration samples from patients with AML at the diagnosis phase without any medication or treatment were obtained, and diagnosis was then confirmed by different laboratory and microscopy tests. Also, three bone marrow aspiration samples were obtained from healthy volunteers. The characteristics of all patients and normal cases are summarized in Table 2.
Characteristics of patients.

Hb: hemoglobin; Plt: platelet.
Size characterization of isolated microvesicles
Both dynamic light scattering (DLS) and transmission electron microscopy (TEM) techniques, quantitatively and qualitatively, showed the size of isolated microvesicles to be between 90 and 1000 nm as shown in Figure 1, which proved the isolation protocol. Also, electron microscope image showed whole microvesicles without any damage in cell membrane which allowed us to use them for further experiments.

Size characterization of microvesicles (a) DLS histogram of one sample shows two peaks with total average diameter of 274.4 nm and (b) transmission electron microscopy image of isolated microvesicles treated with uranyl acetate. The biggest microvesicle is 1000 nm in diameter, with undamaged cell membrane (6000×).
Cell count
During the first 4 days of HSCs co-culture with microvesicles, a majority of HSCs in normal and control groups died rapidly, and after that, the rate of death slowed. But, no event was observed in the leukemia group. After 7 days, viable cells, which were significantly higher in the leukemia group compared with normal and control groups, were counted as shown in Figure 2 (p = 0.02, p = 0.03). However, no significant difference was observed between normal and control groups’ cell count (p > 0.05).

Cell count. Significant higher HSCs number in the leukemia group after 7 days co-incubation with leukemia microvesicles than in normal and control groups. No significant difference was observed between cell number of the leukemia group and HSCs at day 0. Significant decrease in cell number of normal and control groups was present compared with HSCs number at day 0 (Control: HSCs co-incubation with no microvesicle, Normal: HSCs co-incubated with normal bone marrow microvesicles, and Leukemia: HSCs co-incubated with leukemia bone marrow microvesicles).
HSCs stemness
HSC-specific CD markers
After 7 days, flow cytometry analysis of CD34+, CD34+CD38−, CD90+, and CD117+ phenotypes showed no significant difference among studied groups (Figure 3(a)–(d)). In fact, different situations of cell culture had no effect on the most important HSC CD markers.

HSC-specific CD markers in different studied groups at day 0 and after 7 days incubation: (a) CD34+ population, (b) CD34+CD38− population (Q3), (c) CD117+ population, and (d) CD90+ population.
Colony-forming ability
After 7 days, cells were tested to show the ability of differentiation to various blood cell types as a stemness proof which lasts 14 days. Colonies containing more than 50 cells were counted as shown in Figure 4(a). No significant change of colony count, totally and separately, was observed among the studied groups (Figure 4(b) and (c)).
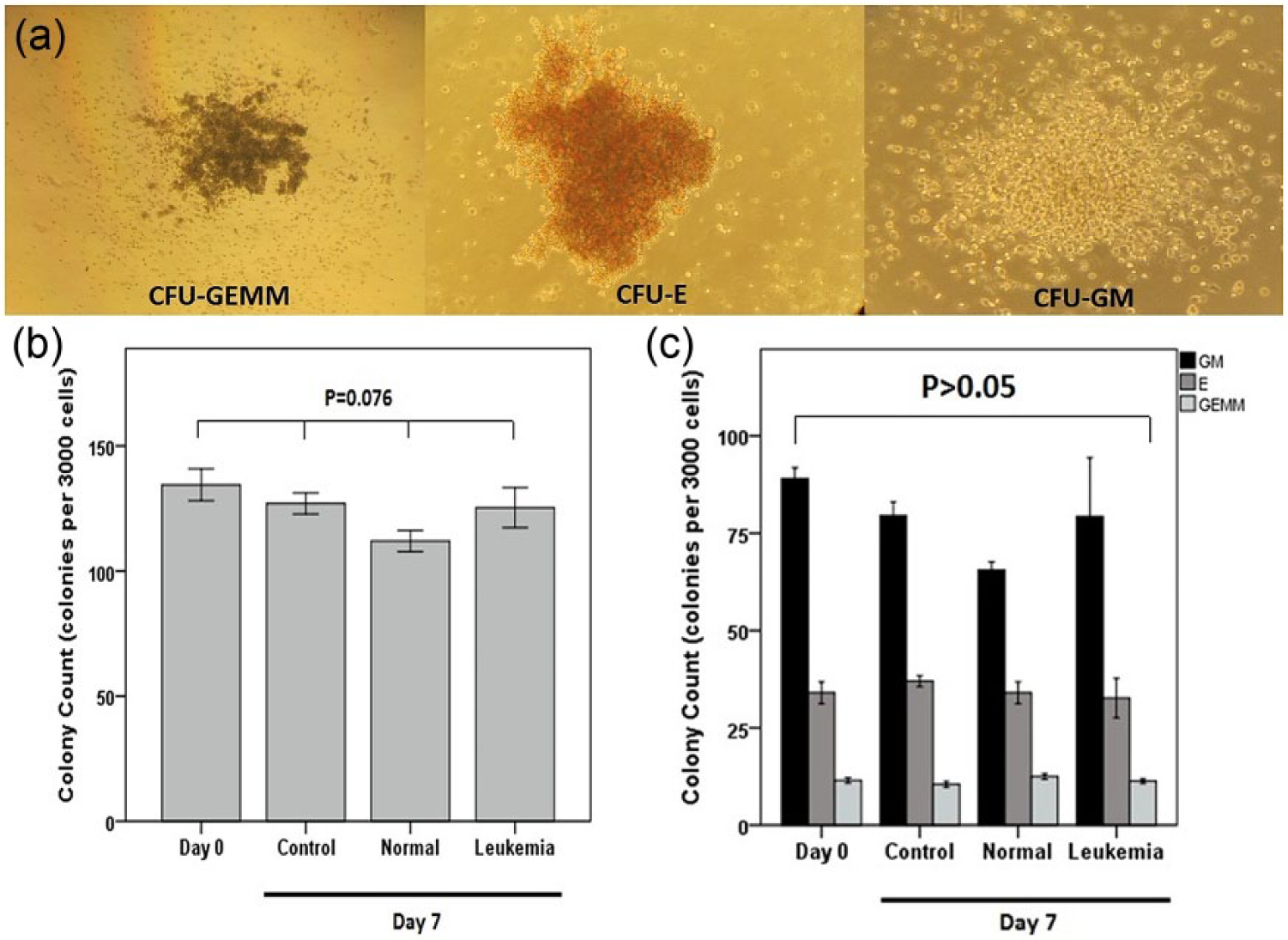
Colony assay of cells at day 7: (a) different shapes of colonies which contain more than 50 cells (CFU-GM: colony-forming unit-granulocyte macrophage; CFU-E: colony-forming unit-erythrocyte; CFU-GEMM: colony-forming unit-granulocyte erythrocyte monocyte megakaryocyte), (b) total colony count (CFU-GM+ CFU-E+ CFU-GEMM) in different studied groups at days 0 and 7, and (c) separate colony count in different studied groups at days 0 and 7.
MicroRNA-21 and microRNA-29a genes expression
Quantitative real-time PCR analysis in the leukemia group showed a 2.7-fold increase of microRNA-21 gene expression in comparison to control and normal groups as shown in Figure 5 (p = 0.03). In addition, microRNA-29a gene expression considerably increased in the leukemia group compared to the control (4.1-fold) and to the normal groups (3.9-fold; p = 0.03 and p = 0.04).

MicroRNA-21 and microRNA-29a genes expression. HSCs in the leukemia group show significant over-expression of these two microRNAs after 7 days of co-incubation with leukemia microvesicles compared with normal and control groups.
Discussion
In the bone marrow niche, where one HSC converts to an LSC, all normal and malignant cells are close to each other. There is a hypothesis that malignant cell–derived microvesicles are capable of penetrating into the adjacent normal cells, such as healthy HSCs, transferring some information and transforming them. This was the first factor in investigation of the probable effect of these microvesicles on healthy HSCs. We used bone marrow aspiration samples, including all cells in complete niche, to consider the whole leukemic microenvironment–derived microvesicles. Thus, a primary model was designed for the first time to seek genotype or phenotype changes related to leukemia disease, in healthy HSCs using these microvesicles.
Leukemia microvesicles promote survival in healthy HSCs
Our results showed a higher cell count in the leukemia group, indicating a role for leukemic bone marrow cell–derived microvesicles to provide a better situation for cells compared with the normal group containing normal bone marrow cell microvesicles and the control group containing no microvesicles. These data became more interesting when no change in HSC especial CD markers including CD34+, CD34+CD38−, CD90+, and CD117+ phenotypes was observed. Also, the high ability of cells to make colonies and differentiate to various blood cells compared to day 0 cells proved that survived cells were still stem cells and different groups of microvesicles did not affect this property. Therefore, leukemia microvesicles increased the stem cells survival while normal microvesicles did not. Professor Ratajczak and his colleagues also showed that embryonic stem cell–derived microvesicles enhanced survival in murine SKL (Sca-1+/Kit+/Lin−) progenitors. 9 Moreover, our team previously showed this important effect by Jurkat (acute T lymphoblastic leukemia) cell line–derived microvesicles on healthy HSCs, 15 and again, in this study, it was confirmed by leukemia microvesicles from primary cells. Resistance to death and long survival are the first signs of cell transformation from normal to malignant stage. Escaping from programmed death induced by chromosomal abnormalities and genome instability provides the required time for accumulating mutations and emergence of the cancer. So, leukemia microvesicles are probably able to transfer this important property from leukemia cells to healthy HSCs.
Leukemia microvesicles dysregulate the famous oncomir: microRNA-21
The most important finding in our study was dysregulation of microRNA-21 gene expression which is up-regulated in almost all epithelial cell–derived solid tumors including breast, lung, gastric, prostate, and colon cancers. Its over-expression has also been reported in hematological malignancies such as leukemia, lymphoma, and multiple myeloma.16–19 Moreover, higher microRNA-21 gene expression was reported in LSCs compared with healthy HSCs, which proves its role as one of the first microRNAs in leukemogenesis. 20 A recent study by Leighton Grimes and colleagues showed that inhibition of microRNA-21 inhibited in vitro leukemic colony-forming activity and, in vivo, depleted leukemia-initiating cell activity, leading to leukemia-free survival in the murine AML model. 21 Also, its expression is completely correlated with long survival. 18 In this study, microRNA-21 was about threefold over-expressed in the leukemia group, a group with better survival, reflecting the ability of leukemic microvesicles in inducing aberrant expression of the most known oncomir, microRNA-21, in healthy HSCs.
Leukemia microvesicles increase microRNA-29a gene expression: leukemia-initiating microRNA
Han and colleagues 22 showed that ectopic expression of microRNA-29a in non-self-renewing myeloid progenitor cells transforms them to self-renewing cells which precedes myeloproliferative disorder progression to AML, suggesting that aberrant acquisition of self-renewing in myeloid progenitors may be an early event during myeloid leukemogenesis. They also showed that myeloid progenitors with over-expressed microRNA-29a are functional LSCs, suggesting a role in human myeloid leukemogenesis. In our study, microRNA-29a expression was about fourfold higher in the leukemia group, which shows that while cells are still stem cells with maintained colony-forming ability, some leukemic-like changes have occurred by leukemic microvesicles.
Long survival and dysregulation of microRNA-21 and microRNA-29a gene expression in healthy HSCs are not enough to acclaim “leukemic transformation.” But, they can satisfactorily show the leukemia-related effects of leukemic microvesicles. Moreover, they can serve as evidence of disease progression by affecting adjacent normal cells in the leukemic bone marrow microenvironment.
Future in vivo studies are required to evaluate the quality of these affected healthy HSCs in normal or probably malignant hematopoiesis.
Footnotes
Acknowledgements
The authors thank the Iranian Blood Transfusion Organization and Atieh Hospital for providing cord blood samples. They also acknowledge Nemazi Hospital and Taleghani Hospital for providing them with the bone marrow aspirates of patients. They wholeheartedly thank the Stem Cell Technology Research Center for all technical equipment. F.R. did all the experiments, prepared the manuscript, and analyzed the data; M.S. generated the hypothesis, designed part of the study, and discussed the data; D.M. and M.H.K. designed part of the study; S.A.K.A. and M.R. provided clinical samples and discussed the data; M.I.S. and J.K. provided clinical samples.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This project was supported by Tarbiat Modares University and Shiraz University of Medical Sciences.
