Abstract
The mutations of epidermal growth factor receptor are detected in gastric cancer, indicating its suitability as a target for receptor tyrosine kinase inhibitors, as well as a marker for clinical outcome of chemotherapeutic treatments. However, extraction of quality tumor tissue for molecular processes remains challenging. Here, we aimed to examine the clinical relevance of urinary cell–free DNA as an alternative tumor material source used specifically for monitoring epidermal growth factor receptor mutations. Therefore, 120 gastric cancer patients with epidermal growth factor receptor mutations and 100 healthy controls were recruited for the study. The gastric patients also received epidermal growth factor receptor inhibitor treatment for a serial monitoring study. Paired primary tumor specimens were obtained with blood and urine samples, which were taken at a 1-month interval for a duration of 12 months. We found that urinary cell–free DNA yielded a close agreement of 92% on epidermal growth factor receptor mutation status when compared to primary tissue at baseline, and of 99% epidermal growth factor receptor mutation status when compared to plasma samples at different time points. Thus, our data suggest that urinary cell–free DNA may be a reliable source for screening and monitoring epidermal growth factor receptor mutations in the primary gastric cancer.
Keywords
Introduction
Gastric cancer is the second most common cancer worldwide, while abnormal activation of epidermal growth factor receptor (EGFR) and its ligands has been associated with progression of gastric cancer and poor prognosis. 1 Thus, immunological or small molecule pharmacological inhibition of EGFR has been applied. For example, a synergistic action of Gefitinib, a small molecule inhibitor of EGFR tyrosine kinase, and Paclitaxel have been used to treat gastric cancer cells expressing high levels of EGFR. 2
One important question in the therapy targeting EGFR is detection of EGFR mutations in gastric cancer patients during the treatment of the disease. Tyrosine kinase inhibitors (TKIs) are used as the first line treatment for gastric patients with positive activating EGFR mutations, among which the exon 19 deletions and L858R point mutation appear to comprise the majority of all activating EGFR mutations.3–5 Nevertheless, the analysis on the EGFR mutations frequently fails to obtain satisfactory results, largely due to the difficulties in collecting quality biopsy specimens. Hence, it is extremely critical to find alternative methods for obtaining non-biopsy tumor samples to analyze the EGFR mutations.
Recent reports have demonstrated that a novel technique is to retrieve disease information through extraction and analysis of circulating tumor DNA (ctDNA) in the peripheral blood plasma.6–8 This technology is applied in a less invasive approach compared to conventional tissue biopsies, and has been used in the diagnosis of gastric cancer.6–8 However, the application of ctDNA in the dynamic diagnosis for EGFR mutations in gastric patients before/after chemotherapy has not been studied. Moreover, alternative samples other than primary tissue and blood plasma for detection and profiling of EGFR mutations will further improve the current diagnosis of gastric patients, especially in the long term after chemotherapy.
In the current study, we aimed to examine the clinical relevance of urinary cell-free DNA as an alternative tumor material source used specifically for monitoring EGFR mutations. Therefore, 120 gastric cancer patients with EGFR mutations and 100 healthy controls were recruited for the study. The gastric patients also received EGFR inhibitor treatment for a serial monitoring study. Paired primary tumor specimens were obtained with blood and urine samples, which were taken at a 1-month interval for a duration of 12 months. We found that urinary cell–free DNA yielded a close agreement of 92% on EGFR mutation status when compared to primary tissue at baseline, and of 99% EGFR mutation status when compared to plasma samples at different time points. Thus, our data suggest that urinary cell–free DNA may be a reliable source for screening and monitoring EGFR mutations in primary gastric cancer.
Methods and materials
Ethical approval
All human and animal studies have been approved by the ethics committee of the First Affiliated Hospital of Zhengzhou University and have therefore been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
Patients
All persons gave their informed consent prior to their inclusion in the study. A total of 120 gastric cancer patients who were treatment naive and underwent treatment at our clinics were recruited for this study. Sample extractions were conducted via procedures approved by the institutional review board. A total of 100 healthy volunteers provided blood and urine specimens as controls. Patients who were selected in the trials were all positive for the EGFR mutation L858R, which was discovered in the molecular profiling of the primary tissue. All patients received treatment with Gefitinib.
Study design and sample procession
The first peripheral blood and urine specimens of gastric cancer patients were obtained immediately prior to the start of Gefitinib treatment. The tissue biopsy samples were obtained at mean time for EGFR mutations. Thereafter, sample extractions were performed at a 1-month interval for a duration of 12 months. Urinary samples of 50 mL were obtained each time together with 3 mL of peripheral blood.
Detection of EGFR ctDNA
Blood and urine samples were processed within 3 h of extraction to avoid loss of the cell-free DNA integrity. In order to extract blood plasma, centrifugation of whole blood specimens was performed for 8 min at 1200g. A repeat centrifugation step was done on the supernatant to ensure the removal of any remaining contaminating cells. For urine specimens, a single high-speed centrifugation step of 15,000g for 8 min was done and 8 mL of supernatant was recovered. Cell-free DNA from both urine and plasma were purified using the Qiagen’s QIAamp Circulating Nuclei Acid kit (Qiagen, Valencia, CA, USA), according to the manufacturer’s instructions. Elutes were stored at −20°C prior to the EGFR molecular test. DNA integrity was examined and DNA content was quantified using Nanodrop 1000 (Thermo Scientific, Rockford, IL, USA).
The EGFR mutation profiling of all samples was carried out using droplet digital polymerase chain reaction (PCR) (ddPCR, QX200 ddPCR system, Bio-Rad Laboratories Inc., Nutley, NJ, USA). Probes/primers for L858R were obtained from Bio-Rad’s PrimePCR™ ddPCR™ Mutation Assays (Bio-Rad Laboratories Inc.), according to the manufacturer’s instructions. Briefly, 20µL reaction mix was mixed with the cell-free DNA samples from both plasma and urine specimens. Thermocycling conditions were as follows: 96°C for 8 min followed by 40 cycles of 93°C for 30 s and 57°C for 1 min. Thereafter, an enzyme deactivation step at 100°C for 10 min was performed.
Statistical analysis
The correlation between urinary/plasma ctDNA with primary tissue biopsies was analyzed using the Cohen’s kappa coefficient. Comparisons of cell-free DNA with healthy negative controls were performed using unpaired Student’s t test. Receiver operating curves (ROC) and the area under the curves (AUC) were examined for evaluating the clinical value of urinary cell-free DNA. Categorical variables presented in the study were represented as mean ± standard deviation. All statistical analyses were performed using the GraphPad Prism 6.0 (GraphPad Software, Inc., La Jolla, CA, USA).
Results
Quantification of the amount of ctDNA in plasma and in urine
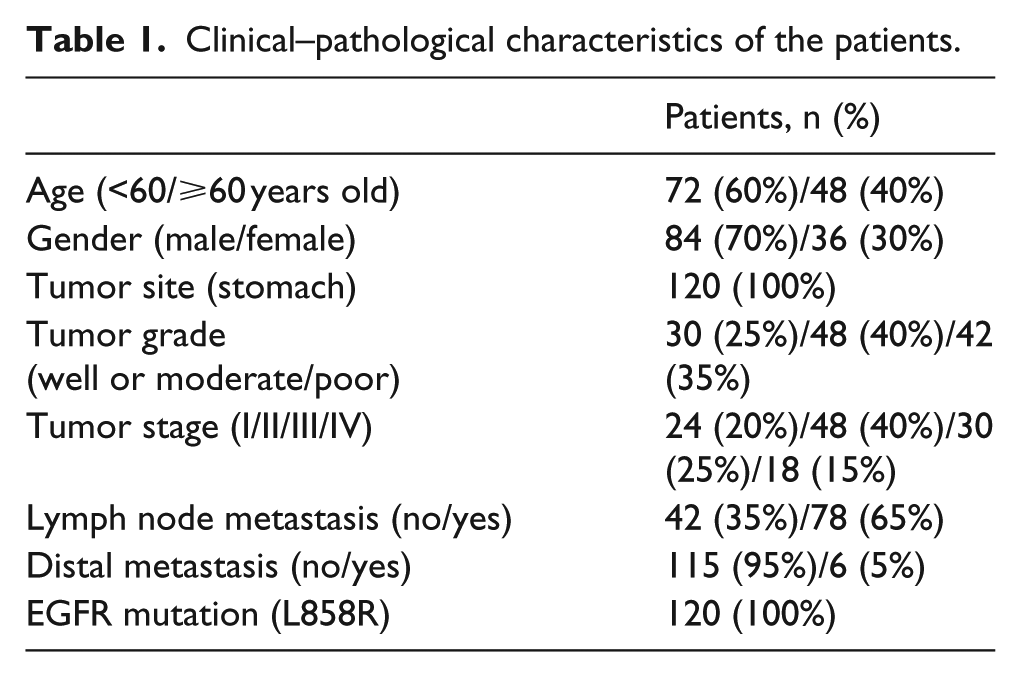
First, 120 biopsy gastric cancer tissue samples together with matched peripheral blood and urine specimens were analyzed. The characteristics of the patients are shown in Table 1. All the patients who were selected in the current study had L858R point mutation. The amount of ctDNA in urine specimens was analyzed, showing that the cell-free DNA that was obtained from either 3 mL plasma or 10 mL urine from the patients had no significant difference in the absolute amount (p > 0.05, Figure 1). However, the amount of ctDNA from the patients was significantly higher than healthy controls, using either 3 mL plasma (16.5 ± 3.3 ng vs 3.8 ± 1.5 ng, p < 0.05, Figure 1) or 10 mL urine (17.1 ± 3.6 ng vs 3.7 ± 1.8 ng, p < 0.05, Figure 1).
Clinical–pathological characteristics of the patients.
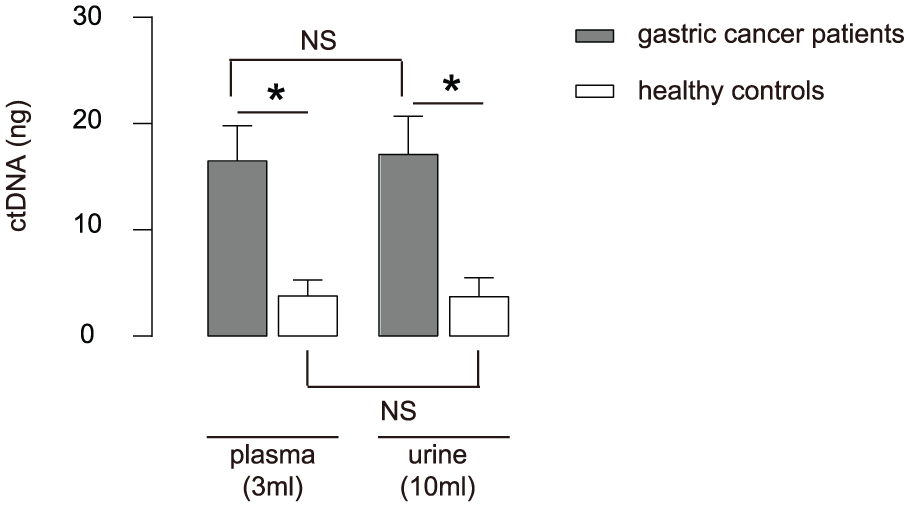
Quantification of the amount of ctDNA in plasma and in urine: comparison of the amount of urinal and plasma ctDNA between gastric cancer patients (n = 120) and healthy controls (n = 100).
Detection of EGFR mutations in plasma and urinary ctDNA before chemotherapy
We then analyzed the sensitivity for EGFR mutation detection, using either plasma ctDNA or urine ctDNA. First, we compared sensitivity using ctDNA from the tissue biopsy samples and from urinary ctDNA samples, and found the overall concordance rate was 92% (Figure 2(a)). Next, we compared sensitivity using ctDNA from the plasma samples and from urinary ctDNA samples, and found the overall concordance rate was 99% (Figure 2(b)).

Detection of EGFR mutations in plasma and urinary ctDNA before chemotherapy: EGFR concordance measurements for 120 gastric cancer patients and 100 healthy controls. (a) EGFR mutation status agreement between primary tissue specimens and urinary ctDNA. (b) EGFR mutation status agreement between plasma and urinary ctDNA. (c and d) ROC analysis of urinary (c) and plasma (d) ctDNA.
The ROC analysis for urinal cell-free DNA showed an area under the curve (AUC) of 0.98 for healthy individuals and gastric cancer patients (Figure 2(c)). AUC using plasma cell–free DNA was 0.97 for healthy individuals and gastric cancer patients (Figure 2(d)). There was no statistical significance in the AUC values using urine and plasma ctDNA (p > 0.05).
Dynamic detection of EGFR mutations in urinary ctDNA during chemotherapy
We then tracked the dynamic changes of cell-free DNA quantity and the mutational status in a serial test during Gefitinib treatment. Specimens were extracted at a 1-month interval for a duration of 12 months. Throughout the follow-up, we did not detect significant differences in the amount of ctDNA in the plasma and in the urine (Figure 3), and we did not detect differences in the EGFR mutations in the plasma and in the urine. At the early phase of treatment, we observed an immediate drop of the quantity of ctDNA, but again the drop was similar in the plasma and in the urine (Figure 3). Together, our data suggest that urinary cell-free DNA may be a reliable source for screening and monitoring EGFR mutations in primary gastric cancer.

Dynamic detection of EGFR mutations in urinary ctDNA during chemotherapy: serial monitoring of ctDNA in urine and in plasma from the gastric cancer patients after Gefitinib treatment. Specimens were extracted at a 1-month interval for a duration of 12 months. Throughout the follow-up, no significant differences in the amount of ctDNA in the plasma and in the urine were detected.
Discussion
Gastric cancer cells highly express EGFR, suggesting that the prevalence of such mutations in gastric cancer patients could have a clinical relevance. Numerous studies have identified the need to accurately and rapidly track the dynamic changes in EGFR mutational profile during cancer treatment. 9 Here, our study shows the feasibility to use urinary cell–free DNA in this endeavor. Most importantly, our results suggest that urinary cell–free DNA is both prognostic and predictive of treatment response and disease outcome. This finding may potentially complement the current disease treatment when the patient develops drug resistance. 9
The gold standard for EGFR molecular profiling of gastric cancer is primary tissue biopsy. However, some patients with poor general body condition may not be able to receive repeated surgical treatments and we may fail to obtain quality specimens. Thus, biopsies using plasma ctDNA or urinary ctDNA may be used as a substitute. Although plasma specimens have been shown to provide a reliable source of ctDNA for this purpose, the repeated sampling from the patients’ vessels may be daunting. On the contrary, urinary ctDNA is obtained in a non-invasive manner, which renders it an ideal body fluid for examination. Urine tests are commonly applied, in which the urine samples are routinely used for measurements of a variety of health-related parameters in many diseases. Here, we show that the ctDNA in the urine is directly linked to the disease status of gastric cancer patients with EGFR mutations who are undergoing chemotherapeutic treatment. Our results indicate a high concordance matching of the primary tissue EGFR molecular profiles. In addition, it is found that the quantity of ctDNA from the urine was comparable to those from the plasma at a lower concentration. AUC comparisons with plasma and urinary cell-free DNA showed no significant difference in sensitivity and specificity.
Prior studies have determined the use of urinary ctDNA as a marker for diagnosis of prostate cancer, 10 bladder cancer, 11 and colorectal cancer. 12 However, to the best of our knowledge, this study is the first investigation to show the usefulness of urinary cell–free DNA in monitoring of gastric cancer patients with EGFR mutations during chemotherapy. The serial monitoring of urinary cell–free DNA is expected to benefit the gastric cancer patients who undergo chemotherapeutic treatment and to improve the current gastric cancer treatment.
Footnotes
Acknowledgements
Xiu-Qin Shi and Wen-Hua Xue were equal contributors to this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
