Abstract
Pseudogenes were once considered to be genomic fossils without biological function. Interestingly, recent evidence showed that a lot of pseudogenes are transcribed in human cancers, and their alterations contribute to multiple cancer development and progression. It is apparent that many pseudogenes transcribe noncoding RNAs and contribute to the role noncoding genome plays in human cancers. On this basis, some pseudogene transcripts are currently ranked among the classes of long noncoding RNAs. In this study, we identified a new pseudogene-derived long noncoding RNA termed SFTA1P by analyzing the microarray data of non–small cell lung cancer from Gene Expression Omnibus datasets. We found that SFTA1P expression was significantly decreased in non–small cell lung cancer tissues compared with normal tissues in non–small cell lung cancer microarray data. Moreover, decreased SFTA1P expression is only correlated with lung adenocarcinoma patients’ poor survival time but not with lung squamous cell carcinoma patients’ survival. In addition, gain-of-function studies including growth curves, migration, invasion assays, and in vivo studies were performed to verify the tumor suppressor role of SFTA1P in non–small cell lung cancer. Finally, the potential underlying pathways involved in SFTA1P were investigated by analyzing the SFTA1P-correlated genes in The Cancer Genome Atlas lung adenocarcinoma and normal tissues RNA sequencing data. Taken together, these findings demonstrate that pseudogene-derived long noncoding RNA SFTA1P exerts the tumor suppressor functions in human lung adenocarcinoma. Our investigation reveals the novel roles of pseudogene in lung adenocarcinoma, which may serve as a new target for lung adenocarcinoma diagnosis and therapy.
Introduction
Lung cancer is one of the leading causes of cancer-related death worldwide with a 5-year overall survival rate as low as 15%. 1 In spite of current advances in clinical and experimental oncology and molecular targeting therapy for lung cancer, the prognosis of lung cancer remains dissatisfying.2,3 For lung cancer, non–small cell lung cancer (NSCLC) accounts for 80% of all lung cancer, and the major two types are lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). 4 Interestingly, LUAD and LUSC differ greatly in many aspects; 5 hence, it is critical to explore the different molecular basis between LUAD and LUSC.
In the past decade, fast advance in bioinformatics and deep sequencing technique has led to the discovery that only less than 3% human genomes are protein coding genes, while the others are noncoding genes including microRNA (miRNA) genes, long noncoding RNA genes, and pseudogenes.6,7 More importantly, ENCODE project and GENCODE annotation have revealed that these noncoding genome elements play essential roles in normal physiology as well as, when disorder, in human diseases.8,9 As a major subject class of the noncoding family, pseudogenes have been considered as nonfunctional “junk genes” or “genomic fossil” for long time due to their harbor mutations that abrogate their transcription or translation.10,11 Fortunately, the recent improvement of genome-wide platforms has revealed that many pseudogenes are transcriptionally active, and their disorder-transcribed RNAs contribute to several human diseases including cancer.12–16 For example, Kalyana-Sundaram et al. 17 developed a bioinformatics pipeline and identified 2082 pseudogene transcripts, among them, 218 expressed only in cancer samples and a breast cancer–specific pseudogene-ATP8A2-ψ was further validated. In addition, Han et al. 18 detected 9925 pseudogene transcriptions in seven cancer types from The Cancer Genome Atlas (TCGA) RNA-seq data, and many pseudogene transcripts are tissue and/or cancer specific. Despite the abundance of pseudogenes identification in the human cancers, the pathophysiological roles of pseudogenes in LUAD remain poorly understood.
Recently, multilayered functions of pseudogene-derived lncRNAs have been reported in several cancers. The PTEN pseudogene, PTENP1, possesses a regulatory function of PTEN through acting as a decoy for PTEN-related miRNAs and competing for these miRNAs, while loss of PTENP1 released these miRNAs, which instead targeted PTEN and reduced its protein levels.19,20 In this study, we identified a new pseudogene-derived lncRNA, termed SFTA1P, which is located in 10p14 and 693 nt in length. The expression pattern and biological function of SFTA1P in cancers are not reported until now, and we found that SFTA1P was significantly down-regulated in LUAD tissues and cells through analyzing the microarray data and RNA sequencing data from Gene Expression Omnibus (GEO) and TCGA. Furthermore, the relationship between SFTA1P and LUAD patients’ clinical features and survival was analyzed and gain-of-function assays were performed to investigate the roles of SFTA1P in LUAD tumorigenesis.
Materials and methods
Tissue specimens’ collection
We obtained 60 paired LUAD and adjacent non-tumor lung tissues from patients who underwent surgery at The Second Affiliated Hospital of Nanjing Medical University from 2011 to 2012. These patients were diagnosed with LUAD (stages I, II, and III) based on histopathological evaluation. The patients did not receive any local or systemic treatment before operation. The ethics committee of The Second Affiliated Hospital of Nanjing Medical University approved the study protocol.
Cell lines
Seven NSCLC cell lines (A549, H1975, SPC-A1, PC9, H1299, H1650, and SK-MES-1) and the normal bronchial epithelial cell line, HBE, were obtained from the Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences (Shanghai, China). A549, H1650, H1975, and H1299 cells were cultured in Roswell Park Memorial Institute (RPMI)-1640, and HBE, SK-MES-1, PC9, and SPC-A1 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco-BRL, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Gibco, Carlsbad, CA, USA), 100 U/mL penicillin sodium, and 100 mg/mL streptomycin sulfate at 37°C in a humidified air atmosphere containing 5% CO2.
RNA extraction and quantitative polymerase chain reaction assays
Total RNA was extracted with TRIzol reagent from tissue samples or cells according to the manufacturer’s instructions. A total RNA of 1 µg was reverse transcribed in a final volume of 20 µL using random primers under standard conditions with the PrimeScript RT Reagent Kit (TaKaRa, Dalian, China). To analyze SFTA1P expression levels, we used SYBR Premix Ex Taq (TaKaRa), following the manufacturer’s instructions. Primer sequences for SFTA1P and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) are as follows: SFTA1P F, 5′ CAGCATTCCAGGTGGGCTTT 3′, R, 5′ CCTTGTTTGGCTTACTCGTGC 3′; GAPDH F, 5′ GGGAGCCAAAAGGGTCAT 3′, R, 5′ GAGTCCTTCCACGATACCAA 3′. Quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) was performed in triplicate on an ABI 7500, and data were calculated using the comparative cycle threshold (CT) (2−ΔΔCT) method.
Cell transfection
The human SFTA1P complementary DNA (cDNA) sequence was, respectively, synthesized and subcloned into the pCDNA3.1 vector (GeneChem, Shanghai, China) to generate the pCDNA-SFTA1P vector and for ectopic expression in LUAD cells. Empty vector was used as a control. A549 and PC9 cells were transfected with pCDNA-SFTA1P vector or empty vector using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA), and the cells were incubated for 48 h before use in assays.
Cell migration and invasion assays
Transwells (Corning, Tewksbury, MA, USA, 8.0-µm pores) were used to measure LUAD migration and invasion ability. For migration assays, 3 × 104 cells were suspended in 300 µL of DMEM or RPMI-1640 containing 1% fetal bovine serum and transferred to the upper chamber. For the invasion assays, 1 × 105 cells in 1% fetal bovine serum medium were placed into the upper chamber of an insert coated with Matrigel (Sigma–Aldrich, St Louis, MO, USA). Medium containing 10% fetal bovine serum (FBS) was added to the lower chamber. After incubation for 24 h, the cells that had migrated or invaded through the membrane were stained with methanol and 0.1% crystal violet, imaged, and counted using an IX71 inverted microscope (Olympus Corporation, Tokyo, Japan).
Protein analysis
After 48 h of transfection, cells were lysed using a lysis buffer which contained the mammalian protein extraction reagent RIPA (Beyotime, Beijing, China), a protease inhibitor cocktail (Roche, Basel, Switzerland), and phenylmethylsulfonyl fluoride (PMSF) (Roche). Cell protein lysates that contained 50-µg protein were electrophoresed on 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), transferred onto 0.22-mm nitrocellulose (NC) membranes (Sigma–Aldrich), and incubated with specific antibodies. Specific bands were detected by enhanced chemiluminescence (ECL) chromogenic substrate and quantified by densitometry (Quantity One software; Bio-Rad Laboratories, Hercules, CA, USA). GAPDH antibody was used as control; anti-EZH2, MMP2, and MMP9 (1:1000) were purchased from Cell Signaling Technology, Inc. (Danvers, MA, USA).
In vivo metastasis assay
The 4-week female BALB/c nude mice were maintained under specific pathogen-free conditions and manipulated according to protocols approved by the Shanghai Medical Experimental Animal Care Commission. SPTA1P or empty vector stably transfected A549 cells were harvested. For metastasis assay, suspended cells were injected into the tail veins of mice, which were sacrificed 7–8 weeks after injection. The lungs were removed and photographed, and visible tumors on the lung surface were counted. This study was carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The protocol was approved by the Committee on the Ethics of Animal Experiments of the Nanjing Medical University.
Statistical analysis
All statistical analyses were performed using SPSS 20.0 software (IBM). The significance of differences between groups was estimated by the Student’s t-test, Wilcoxon test, or chi-square test. Kendall’s Tau-b and Pearson correlation analysis were conducted to investigate the correlation between SFTA1P and other gene expressions. Two-sided p values were calculated, and a probability level of 0.05 was chosen for statistical significance.
Results
SFTA1P is down-regulated in LUAD tissues
To investigate the pseudogene-derived lncRNA alterations in NSCLC, we first analyzed the profiles of NSCLC patient (GSE18842) from GEO and found that SFTA1P was most down-regulated in NSCLC tissues compared with normal tissues (Figure 1(a)). Meanwhile, another two datasets (GSE30219 and GSE19188) were used to confirm the result in GSE18842, and the results showed that SFTA1P was also significantly down-regulated in both the datasets (Figure 1(b)). In addition, we analyzed the SFTA1P expression level in LUAD, LUSC, and normal tissues RNA sequencing data from TCGA, and also, we found that SFTA1P was also significantly down-regulated in both LUAD and LUSC tissues compared with normal tissues (Figure 1(c)). Interestingly, the survival analysis using TCGA data showed that decreased SFTA1P was only associated with LUAD patients’ shorter survival time but not with LUSC patients’ survival (Figure 2(a)). These findings indicate that decreased SFTA1P may involve in LUAD development and progression.

SFTA1P expression is down-regulated in NSCLC tissues: (a) hierarchical clustering analysis of the lncRNAs and pseudogene-derived lncRNAs that were expressed differentially (GSE18842, fold change > 2; p < 0.05) between NSCLC samples and paired non-tumor samples, (b) SFTA1P expression was analyzed in GSE30219 and GSE19188 datasets from GEO, and (c) SFTA1P expression levels were analyzed in LUAD, LUSC, and normal tissues from TCGA RNA sequencing data.
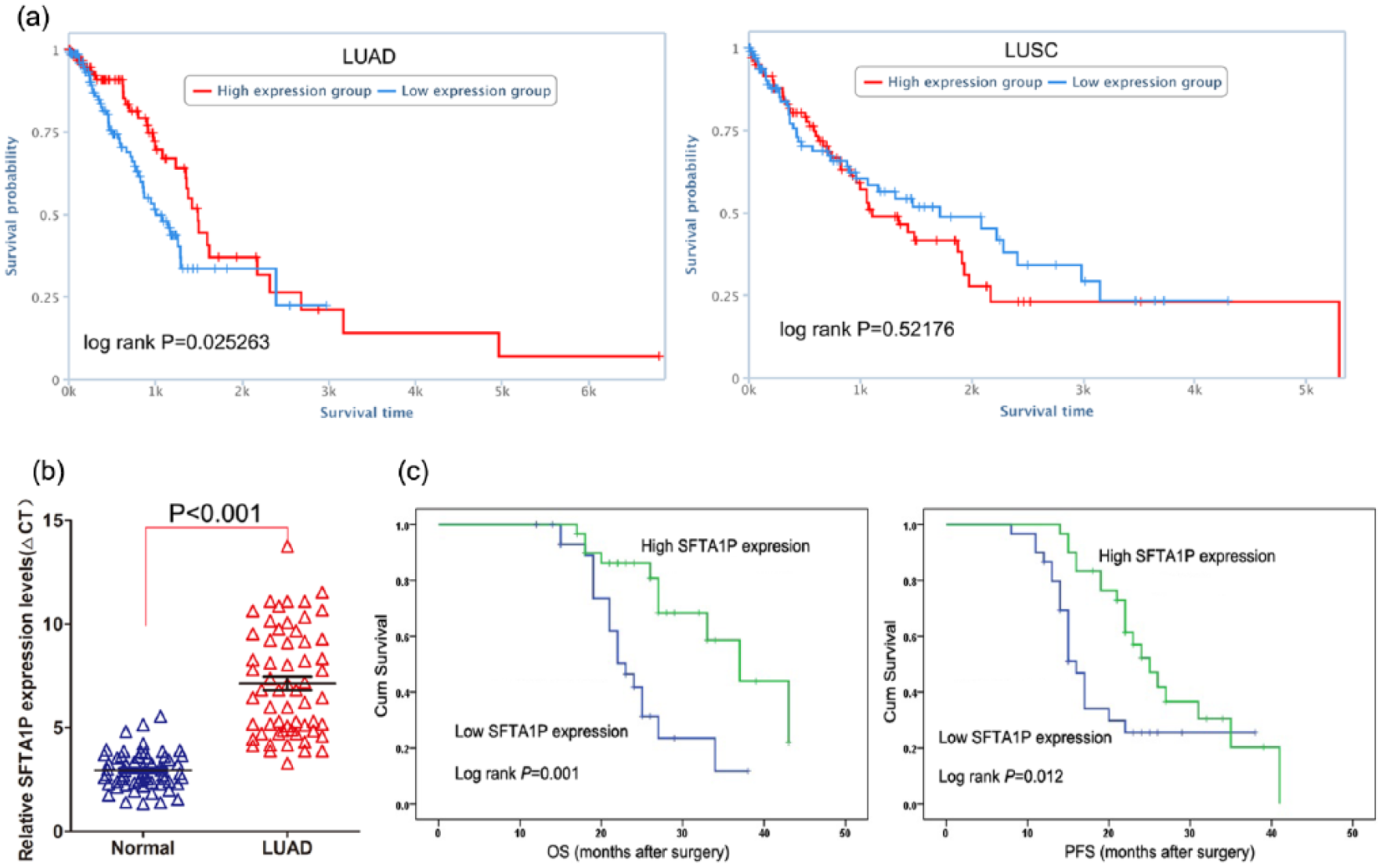
Decreased SFTA1P expression is associated with LUAD patients’ shorter survival: (a) the relationship between SFTA1P expression and LUAD or LUSC patients’ survival was analyzed using TCGA data, (b) SFTA1P expression was analyzed by qPCR in LUAD and normal tissues (n = 60), and (c) Kaplan–Meier survival analysis of OS and DFS in LUAD patients (p < 0.001 for both OS and DFS) based on SFTA1P expression.
Decreased SFTA1P is correlated with LUAD patients’ poor prognosis
To further determine whether SFTA1P was decreased in LUAD tissues, a total of 60 paired LUAD tissues were evaluated for SFTA1P expression using qRT-PCR. The results showed that SFTA1P was significantly down-regulated in 50/60 LUAD tissue samples (fold change > 2) (Figure 2(b)). To investigate the relationship between SFTA1P levels and LUAD patients’ clinicopathologic feature, the patients were divided into two groups: the high-SFTA1P group (n = 30, fold change ⩾ median) and the low-SFTA1P group (n = 30, fold change ⩽ median). Statistical analysis revealed that decreased SFTA1P expression is correlated with advanced pathological stage (p = 0.004) and lymph node metastasis (p = 0.035). However, SFTA1P expression was not associated with other factors including gender (p = 0.598) and age (p = 0.436) in LUAD (Table 1).
Correlation between SFTA1P expression and clinicopathological characteristics of LUAD patients (n = 60).

LUAD: lung adenocarcinoma; TNM: tumor node metastasis.
Overall p < 0.05.
Next, Kaplan–Meier survival analysis was performed to explore the relationship between SFTA1P expression and LUAD patient survival. The results showed that the overall and progression-free survival (PFS) rate over 3 years for the low-SFTA1P group is shorter than that in high-SFTA1P group (Figure 2(c)). Furthermore, univariate survival analysis showed that tumor size, lymph node metastasis, tumor nodes metastasis (TNM) stage, and SFTA1P expression level could be viewed as prognostic factors (Table 2). Other clinicopathological features including sex and age were not statistically significant prognostic factors. Moreover, multivariate Cox regression analyses showed that expression of SFTA1P (p = 0.019), along with lymph node metastasis (p = 0.001), was independent prognostic factor for LUAD patients (Table 2).
Univariate and multivariate analysis of over-survival in LUAD patients (n = 60).

LUAD: lung adenocarcinoma; HR: hazard ratio; 95% CI: 95% confidence interval; TNM: tumor node metastasis.
Overall p < 0.05.
Effect of SFTA1P on LUAD cell proliferation
To investigate the biological function of SFTA1P in LUAD cells, we first evaluated the expression of SFTA1P in various LUAD cell lines using qRT-PCR. As shown in Figure 3(a), SFTA1P expression was down-regulated in all LUAD cell lines compared with HBE. Then, SFTA1P expression was up-regulated in A549 and PC9 cells by transfection with pCDNA-SFTA1P vector, and the results showed that SFTA1P expression was significantly increased in A549 and PC9 cells compared with control (Figure 3(b)). To assess the effect of SFTA1P on LUAD cells phenotype, we performed gain-of-function assays. 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assays showed that SFTA1P overexpression had no effect on the growth of A549 and PC9 cells compared with control cells (Figure 3(c)).
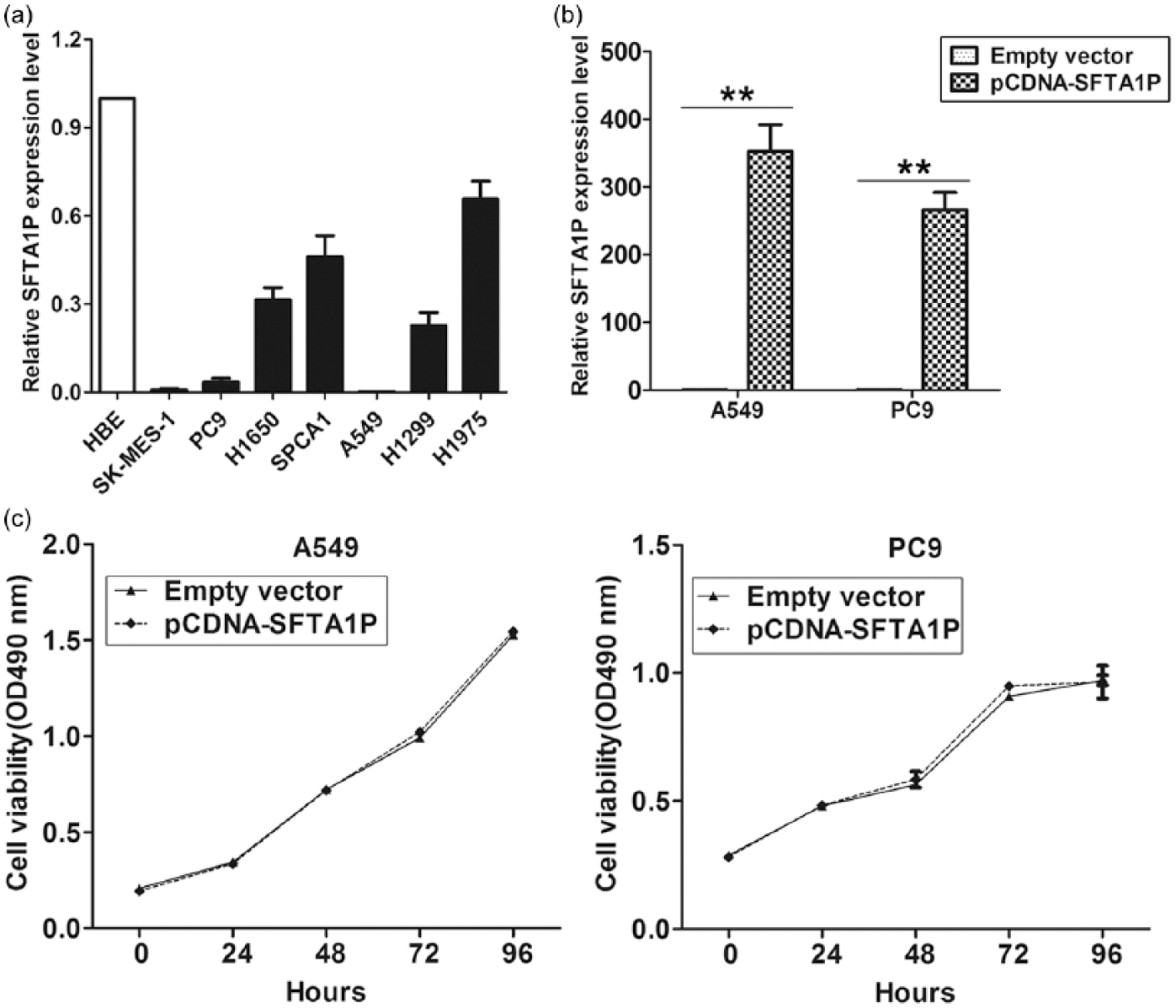
The effect of SFTA1P on LUAD cells proliferation: (a) SFTA1P expression was analyzed by qPCR in LUAD, LUSC, and normal cell lines, (b) SFTA1P expression was analyzed by qPCR in A549 and PC9 cells transfected with pCDNA-SFTA1P or empty vector, and (c) MTT assays were performed to determine the cell viability for pCDNA-SFTA1P or empty vector–transfected A549 and PC9 cells.
SFTA1P inhibits LUAD cells’ migration and invasion in vitro
Cancer cells’ migration, invasion, and metastasis are significant aspects of LUAD progression, which may further result in poor prognosis and short survival for LUAD patients. Therefore, we investigated the effect of SFTA1P on LUAD cells’ migration and invasion by performing transwell assays. The results showed that SFTA1P overexpression impeded LUAD cells’ migration and invasion compared with controls (Figure 4). These results suggest that SFTA1P had tumor-suppressive function that could inhibit LUAD cells’ migration and invasion.
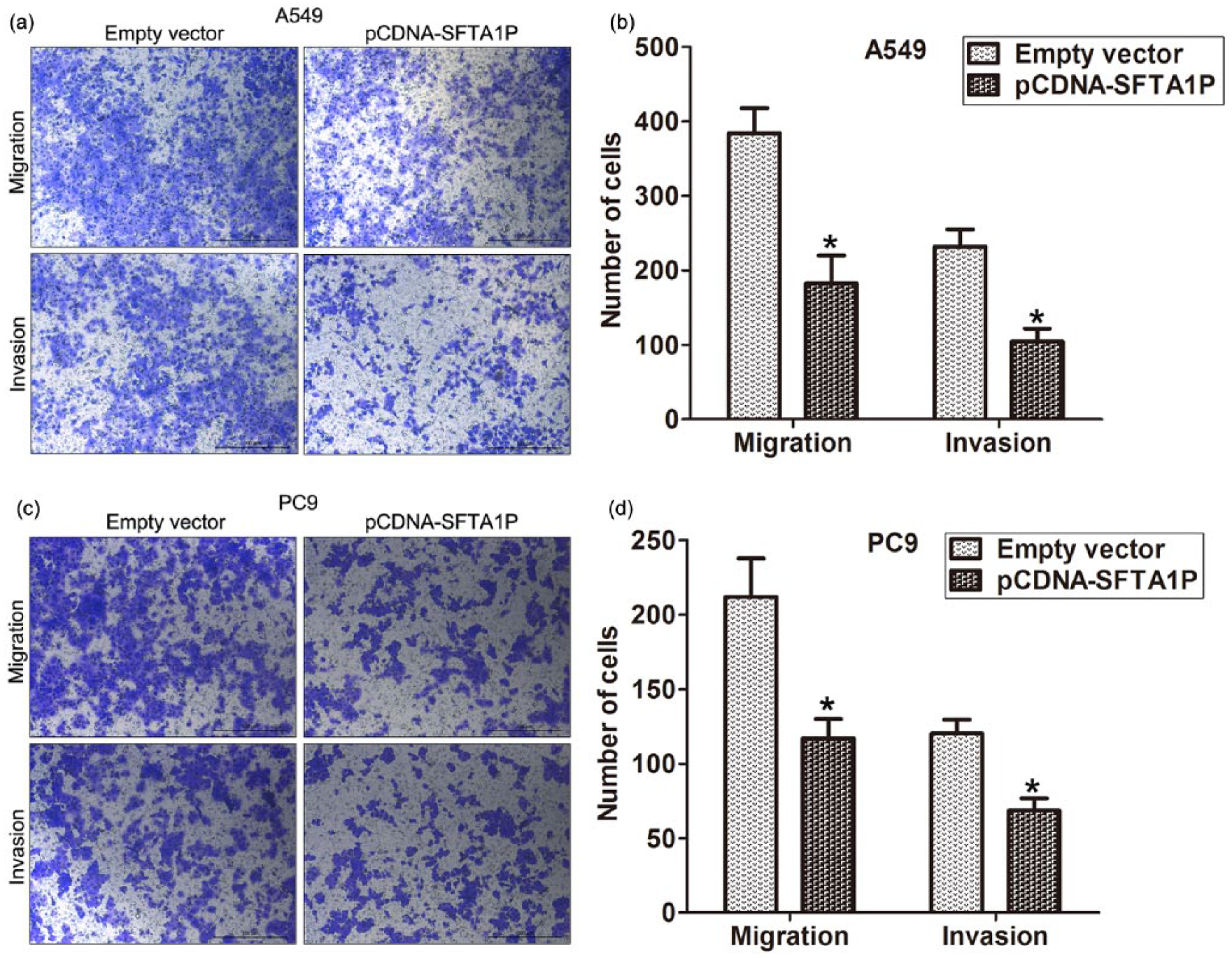
SFTA1P overexpression inhibits LUAD cells migration and invasion: (a and b) transwell assays were used to investigate the changes in migratory abilities of SFTA1P overexpression cells and (c and d) transwell assays were used to investigate the changes in invasive abilities of SFTA1P overexpression cells.
SFTA1P suppresses LUAD cell metastasis in vivo
To investigate the effects of SFTA1P on the metastasis of LUAD cells in vivo, A549 cells stably transfected with pCDNA-SFTA1P were injected into nude mice. Metastatic nodules on the surface of lungs were counted after 8 weeks. The results showed that overexpression of SFTA1P led to a reduction in the number of metastatic nodules compared with those in the control group (Figure 5(a) and (b)). This difference was further confirmed following examination through hematoxylin and eosin (HE) staining of lung sections (Figure 5(c)). The in vivo findings complemented the results of functional in vitro studies involving SFTA1P.

SFTA1P overexpression inhibits LUAD cells metastasis in vivo: (a) analysis of an experimental metastasis animal model was performed by injecting A549 cells stably transfected with pCDNA-SFTA1P into nude mice, (b) the numbers of tumor nodules on lung surfaces are shown, and (c) visualization of the entire lung and HE-stained lung sections.
Potential target genes of SFTA1P in LUAD
To explore the potential targets that are involved in SFTA1P tumor suppressor function in LUAD, we performed correlation analysis in TCGA LUAD tissues sequencing data. As shown in Figure 6(a) and (b), SFTA1P expression is positively related to tumor suppressor DLC, HOPX, and RHOBTB2 expression, while negatively related to oncogene EZH2, FOXM1, and PKL1 expression. These findings indicated that SFTA1P could function as tumor suppressor in LUAD cells might through regulating these important oncogenes or tumor suppressors expression. What is more, Western blot analysis was performed to confirm the expression level of potential target genes of SFTA1P. Compared with control cells, SFTA1P up-regulation significantly reduced EZH2 protein (Figure 6(c)). Besides, SFTA1P overexpression could also decrease the expression level of MMP2 and MMP9 (Figure 6(d)).

Potential targets involved in SFTA1P tumor suppressor function: (a and b) the relationship between SFTA1P and DLC, HOPX, RHOBTB2, EZH2, FOXM1, or PKL1 was analyzed in LUAD tissues using TCGA data and (c and d) the expression level of EZH2, MMP2, and MMP9 protein using Western blot analysis.
Discussion
In the past decade, the function of noncoding elements in human genome has been highlighted due to the implementation of GENCODE project, coming along with lots of studies that have revealed the key roles of microRNAs and lncRNAs in almost all aspects of biology.21–23 In addition, their alterations have been found to contribute to human disease, especially cancers.24,25 Conversely, although some studies documented the pseudogenes’ participation in competing endogenous RNA (ceRNA) interactions and contribution of tumorigenesis, it is difficult to study pseudogenes on a large scale due to their high homology to parental genes that could not be distinguished easily. Recently, benefits from the development of bioinformatics pipeline and a handful of investigations have evidence indicating that thousands of pseudogenes are transcribed and highlighting the involvement of pseudogenes and their derived RNAs during pathogenesis of human diseases, such as cancer.12,16,17 To date, a few intriguing reports have revealed the involvement of pseudogenes in human cancer development and regulation of target genes, by functioning as endogenous competitors for miRNA, RNA-binding protein (RBP), or translational machinery and generating endogenous small-interference RNA.20,26,27
Karreth et al. found the in vivo proto-oncogenic function of murine B-Raf pseudogene through regulation of B-Raf by decoying miR-134, miR-543, and miR-653. Similarly, its human ortholog-BRAFP1 also elicits oncogenic activity as ceRNAs that elevate BRAF expression and mitogen-activated protein kinase (MAPK) activation. 28 Moreover, Wang et al. identified a novel tumor-suppressive pseudogene, termed AOC4P, whose expression was significantly down-regulated in hepatocellular carcinoma (HCC) samples. Decreased AOC4P expression is correlated with poor prognostic outcomes, and AOC4P overexpression significantly reduced cell proliferation, migration, and invasion through binding to vimentin protein and promote its degradation. 29 In this study, we also identified a new novel tumor-suppressive pseudogene named SFTA1P in LUAD, which was consistent with Zhao et al.’s 30 data. We found that SFTA1P expression is significantly down-regulated in NSCLC tissues, but its decreased expression only correlated with LUAD patients’ survival time. Furthermore, the level of SFTA1P is confirmed in a cohort of 60 paired LUAD and normal tissues, and relationship analysis showed that decreased SFTA1P expression is associated with LUAD patients’ poor prognosis and shorter survival time. Gain-of-function assays showed that SFTA1P overexpression could impair LUAD cells’ migration, invasion, and metastasis but had no effect on LUAD cells proliferation. Our findings indicate that SFTA1P may be an important regulator of LUAD metastasis and progression, which serves as a prognostic marker for LUAD patients.
Generally, pseudogenes involve in regulation of cancer cells phenotype mostly via affecting their parental genes expression. In this study, we explored SFTA1P potential underlying targets through analyzing its related genes in LUAD tissues. SFTA1P expression is negatively related to EZH2 and FOXM1 and positively related to HOPX in LUAD tissues. EZH2 and FOXM1 are important oncogenes that promote cancer cell migration and metastasis,31,32 while HOPX exerts tumor-suppressive function through Ras-induced senescence in human lung cancer. 33 These findings suggested that SFTA1P might involve in LUAD cells invasion and metastasis via influencing those genes expression. In summary, we first identified a new tumor-suppressive pseudogene-derived lncRNA SFTA1P in LUAD which may be potential therapeutic targets. Although no pseudogenes have been well characterized in LUAD, the research of pseudogenes is expanding quickly. Therefore, integration of pseudogenes and their derived lncRNA into LUAD biology will deepen our understanding of the mechanisms of this deadly disease, and some specific pseudogenes may be translated into clinical applications for diagnosis, prognosis, or treatment of LUAD patients.
Footnotes
Acknowledgements
The authors apologize to all researchers whose relevant contributions were not cited due to space limitations. H.Z., Y.X., and R.X. contributed equally to this work and should be regarded as joint first authors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by National Natural Scientific Foundation of China grants 81602013 (to Fengqi Nie) and 81602003 (to Xuefei Shi).
