Abstract
Long noncoding RNAs play an important role in various biological processes, including tumorigenesis. FOXC1 (Forkhead box C1) is a member of the Forkhead box family of transcription factors and plays a crucial role in nasopharyngeal carcinoma. In this study, a novel long noncoding RNA (FOXCUT) located upstream of FOXC1 was investigated in 42 nasopharyngeal carcinoma patients. Our analysis revealed that the expression levels of FOXCUT and FOXC1 in nasopharyngeal carcinoma tissues were significantly higher than those observed in chronic nasopharyngitis tissues and that FOXCUT expression was positively correlated with FOXC1 expression. Additionally, knockdown of FOXCUT significantly inhibited proliferation and migration of nasopharyngeal carcinoma cell lines and resulted in downregulated expression of the matrix metalloproteinase 7 and matrix metalloproteinase 9, as well as vascular endothelial growth factor A and β-catenin. Our findings suggested that FOXCUT expression contributed to the development and progression of nasopharyngeal carcinoma by targeting FOXC1 and that FOXCUT might be useful as a potential nasopharyngeal carcinoma biomarker and therapeutic target.
Introduction
Nasopharyngeal carcinoma (NPC) is a type of head and neck carcinoma that is very common in Southern China and Southeast Asia. It was estimated that >60,000 people were diagnosed with NPC in the past year, constituting a major public health problem in China. 1 Although most primary NPC patients are sensitive to radiotherapy and chemotherapy, there remain a significant percentage of NPC recurrence and distant metastasis. 2 The poor prognosis of patients with NPC is largely the result of the tumor recurrence and distant metastasis after therapy. Therefore, it is necessary to explore the molecular basis of NPC to determine promising therapeutic strategies.
Forkhead box C1 (FOXC1) is a member of the Fox transcription-factor family and is overexpressed in multiple malignant tumors and involved in both tumor development and healthy biological processes. 3 Previous studies showed that FOXC1 overexpression correlated with poor clinical prognosis in melanoma and hepatocellular carcinoma4,5 and with poor clinical outcomes in cancers, such as non-small cell lung cancer, breast cancer, and pancreatic ductal adenocarcinoma.6–8 Recent data implicated FOXC1 in the induction of epithelial–mesenchymal transition (EMT) and regulation of cell proliferation and migration in NPC; 9 however, the underlying mechanisms associated with NPC tumorigenesis remain elusive.
Long noncoding RNAs (lncRNAs) describe RNA molecules >200 nucleotides in length and without protein-coding capacity. Although many lncRNAs are expressed at low levels in cells and tissues, lncRNAs play important roles in various biological processes.10,11 Additionally, lncRNAs cooperate with adjacent coding sequences to form functional lncRNA–mRNA pairs involved in tumorigenesis and cancer progression.
FOXCUT is a novel lncRNA (NR_125804.1) located on chromosome 6p25 and upstream of the FOXC1 promoter. Previous studies showed that the FOXCUT–FOXC1 regulatory network is associated with tumorigenesis and cancer progression in oral squamous cell carcinoma and breast cancer;12,13 however, the function of the FOXCUT–FOXC1 pair in NPC has not been described. In this study, we investigated the expression levels and clinicopathological characteristics of the FOXCUT–FOXC1 pair in NPC and explored the role of FOXCUT in cell proliferation and migration associated with NPC to evaluate its efficacy as a potential therapeutic target.
Materials and methods
Clinical tissue samples
A total of 42 freshly biopsied NPC tissue samples and 30 chronic inflamed nasopharyngeal tissue samples were collected from a Hospital in Shenzhen, China, from January 2015 to June 2016. NPC diagnosis was histopathologically confirmed, and none of the patients received therapy prior to the biopsy. The tissues from each subject were immersed overnight in RNAlater reagent (Qiagen, Hilden, Germany) at 4°C and stored at −70°C. The study was approved by the hospital’s ethics committee, and written informed consent was obtained from all patients prior to biopsy. The clinical characteristics are described in Table 1.
Correlation between the expression of FOXCUT and FOXC1 features in NPC.
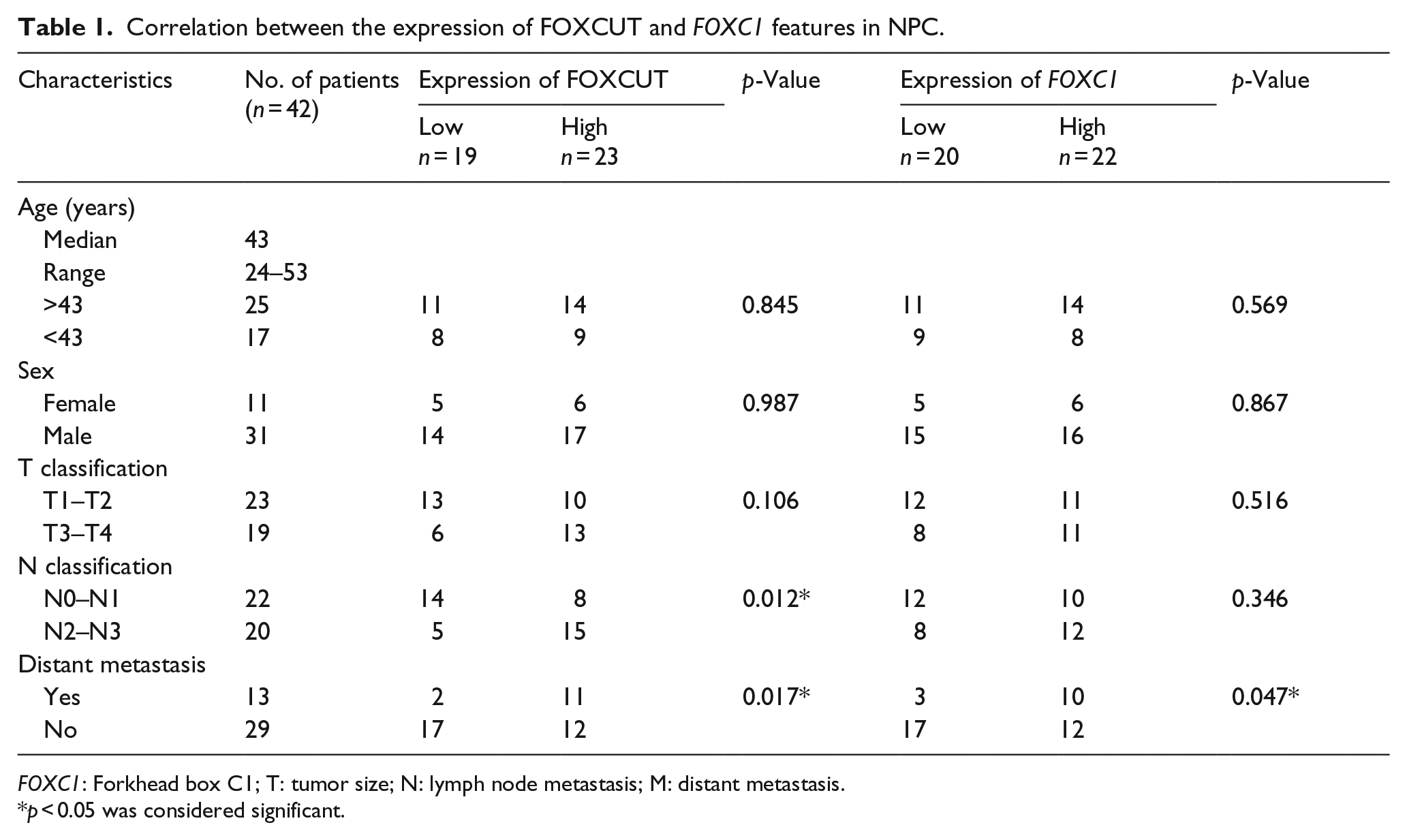
FOXC1: Forkhead box C1; T: tumor size; N: lymph node metastasis; M: distant metastasis.
p < 0.05 was considered significant.
Cell lines and cell culture
Human NPC cell lines, including CNE1, CNE2, 5-8F, 6-10B, and SUNE-1, were obtained from the Laboratory of Otorhinolaryngology Head and Neck Surgery, South Medical University, Guangzhou, China, and cultured at 37°C under 5% CO2 in RPMI-1640 medium (Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum (FBS; Invitrogen). The human immortalized nasopharyngeal epithelial cell line NP69 was obtained from Shenzhen University and cultured in keratinocyte serum-free medium (Invitrogen).
RNA isolation and real-time polymerase chain reaction
Total RNA was isolated from fresh tissues or cultured cells using TRIzol reagent (Takara, Otsu, Japan) and reverse transcribed to complementary DNA (cDNA) using the Reverse Transcription System (Promega, Madison, WI, USA) according to the manufacturer’s instructions. Quantitative real-time polymerase chain reaction (PCR) analysis was performed on an ABI PRISM 7900 sequence-detection system using a SYBR GreenER qPCR mix kit (Takara) and done in triplicate. The primer sequences are shown in Table 2. Relative RNA expression level was calculated using the 2−ΔΔCt method. The PCR conditions were 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 34 s.
Primer sequences for quantitative real-time PCR.

FOXC1: Forkhead box C1; GAPDH: glyceraldehyde 3-phosphate dehydrogenase; MMP: matrix metalloproteinase; VEGF-A: vascular endothelial growth factor A.
Small interfering RNA transfection
The small interfering RNA (siRNA) sequences, including one negative control siRNA (si-NC) which was nonhomologous to any human genome sequence, two FOXC1 siRNA sequences, and two lncRNA FOXCUT siRNA sequences, were obtained from GenePharma (Shanghai, China). The target sequences are presented in Table 3. 5-8F cells were plated at a density of 5 × 104 cells per well in six-well plates for 24 h to achieve 50% confluency prior to transfection. siRNAs were transfected using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions. After 48 h, cells were subjected to further experiments. All data were from at least three independent experiments and are presented as mean ± standard deviation (SD).
Target sequences for siRNA.

FOXC1: Forkhead box C1.
Western blot analysis
At 48-h post-transfection, 5-8F cells were lysed with radioimmunoprecipitation assay buffer and boiled for 5 min. Protein samples were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to a polyvinylidene fluoride membrane (Thermo Fisher Scientific, Portsmouth, NH, USA). Bovine serum albumin (5%) in Tris-buffered saline with Tween-20 (TBS-T) was used to block the membrane. The membranes were incubated overnight at 4°C with primary antibodies for FOXC1 (Proteintech, Wuhan, China), β-actin (Proteintech, Chicago, IL, USA), matrix metalloproteinase 7 (MMP7) (Santa Cruz Biotech, Dallas, TX, USA), MMP9 (Santa Cruz Biotech), vascular endothelial growth factor A (VEGF-A; Abcam, Cambridge, UK), and β-catenin (Abcam). After washing three times with TBS-T, the membranes were incubated with secondary antibodies (Proteintech). Immunoreactive signals were visualized using an EasySee Western Blot Kit (TransGen Biotech, Beijing, China), and protein bands were quantified using Quantity One 4.5.0 software (Bio-Rad, Hercules, CA, USA). The experiment was done in triplicate.
Cell proliferation assay
Following transfections, Cell Counting Kit-8 (CCK-8) assay kits (Beyotime Institute of Biotechnology, Shanghai, China) were used to measure cell proliferation according to the manufacturer’s protocol. 5-8F cells were seeded into 96-well plates, and cell transfections were performed, followed by incubation for 0, 24, 48, or 72 h and the addition of 10 µL CCK-8 reagent to each well. Absorbance was measured at 450 nm using iMARK™ Microplate Absorbance Reader (Bio-Rad), and each experiment was performed at least three times.
Cell migration assay
Cell migration assays were performed using the xCELLigence system (Roche, Basel, Switzerland) according to the manufacturer’s instructions. The system is a real-time migration monitoring device with E-plate 16. The experiment was performed on NPC cells transfected with FOXCUT siRNA. After a 24-h transfection, 5 × 104 cells were seeded in serum-free media in the upper chamber, with the lower chambers containing 10% FBS as a chemoattractant. Obtained data were plotted as cell relative-migration index versus time and analyzed using RTCA software (Acea Biosciences, Miami, FL, USA). Each experiment was performed in triplicate, and the results are presented as mean ± SD.
Wound-healing assay
Wounds were scratched in six-well plates using pipette tips after cells had reached ~90% confluence. Photographs were obtained using an Olympus microscope (Olympus, Tokyo, Japan) at 0, 24, and 48 h after siRNA transfection. The experiments were performed in triplicate.
Statistical analysis
A chi-square test or Fisher’s exact test was used to analyze statistical differences in clinical characteristics, and Student’s t test was used to analyze differences between two groups. All data were analyzed using SPSS version 21.0 (SPSS, Inc., Chicago, IL, USA) and GraphPad Prism version 5.0 (GraphPad Software, San Diego, CA, USA). Data are expressed as mean ± SD, and a p <0.05 was considered statistically significant.
Results
FOXC1 and FOXCUT are overexpressed in both NPC cells and biopsied NPC tissues
We detected the relative expression levels of FOXC1 and FOXCUT in different NPC cell lines, including 5-8F, CNE2, CNE1, SUNE-1, and 6-10B, and noncancerous control NP69 cells. As shown in Figure 1(a) and (b), FOXC1 and FOXCUT expression in all NPC cell lines was significantly upregulated as compared with levels observed in NP69 cells, with the highest expression levels observed in 5-8F cells. Therefore, 5-8F cells were chosen for subsequent analysis. Furthermore, quantitative PCR was conducted to evaluate FOXC1 and FOXCUT expression in tissue samples from 42 NPC patients and 30 patients with chronic nasopharyngitis. We observed consistently higher expression of FOXC1 and FOXCUT in NPC tissues relative to levels in chronic inflamed nasopharyngeal tissues (Figure 1(c) and (d)). These results demonstrated that the expression of FOXC1 and FOXCUT was frequently upregulated in both NPC cell lines and tissues.

FOXC1 and FOXCUT expression is frequently upregulated in NPC cell lines and tissues. Expression of (a) FOXC1 and (b) FOXCUT was detected by quantitative PCR in NPC cell lines and noncancerous control NP69 cells. Expression of (c) FOXC1 and (d) FOXCUT was detected by quantitative PCR in NPC and inflamed tissue samples.
Correlations between FOXCUT and FOXC1 expression and clinicopathological characteristics associated with NPC
To reveal potential correlations between FOXCUT/FOXC1 expression and clinicopathological characteristics in NPC patients, according to the mean value of relative FOXCUT and FOXC1 expression, the 42 NPC patient samples were divided into two groups: those exhibiting high expression levels of FOXCUT (n = 23)/FOXC1 (n = 22) and those exhibiting low expression levels of FOXCUT (n = 19)/FOXC1 (n = 20). As presented in Table 1, upregulated FOXCUT expression was associated with advanced lymph node classification (p = 0.012) and distant metastasis (p = 0.017), whereas upregulated FOXC1 expression was associated with distant metastasis (p = 0.047). Additionally, high levels of FOXCUT and FOXC1 expression were correlated with metastatic NPC tissues (p = 0.017 and p = 0.047, respectively; Table 1). However, statistical analyses showed no significant correlation between FOXCUT/FOXC1 expression and age or sex.
FOXCUT siRNA knockdown decreased FOXC1 mRNA levels in 5-8F cells
To explore relationships between FOXC1 and FOXCUT expression, in silico assays were performed using the UCSC gene browser (http://genome.ucsc.edu). Our findings indicated that FOXCUT is located at 6p25 and transcribed from the upstream region of the FOXC1 promoter (Figure 2(a)), suggesting potential interactions between FOXCUT and FOXC1 in NPC.

LncRNA association with adjacent protein-coding transcripts. (a) FOXCUT is an lncRNA located upstream of the FOXC1 promoter. (b) FOXC1 expression was significantly attenuated following FOXC1 siRNA transfection (*p < 0.05 as compared with the si-NC group). Notably, transfection with FOXC1 siRNAs had no effect on FOXCUT expression levels based on RNA expression analysis. (c) FOXC1 protein levels decreased following FOXC1 siRNA transfection. (d) FOXCUT and FOXC1 levels were significantly attenuated following FOXCUT siRNA transfection. (e) FOXC1 protein levels decreased following FOXCUT siRNA transfection.
To further investigate potential correlations between FOXC1 and FOXCUT expression, we used RNA interference to silence FOXC1 expression in 5-8F cells. Our results showed that FOXC1 expression was inhibited by FOXC1 siRNAs si-FOXC1-1 and si-FOXC1-2, with si-FOXC1-1 exhibiting higher degrees of inhibitory efficiency (Figure 2(b)). Western blot analysis shows FOXC1 protein expression to be reduced following si-FOXC1 transfection (Figure 2(c)). Notably, FOXC1 siRNAs had no effect on FOXCUT expression in 5-8F based on RNA expression analysis (Figure 2(b)).
Furthermore, FOXCUT expression was attenuated by ~60% following transfection with FOXCUT siRNA, with co-suppression of FOXC1 mRNA levels by ~50% (Figure 2(d) and (e)). These results suggested that FOXC1 mRNA levels are potentially regulated by the lncRNA FOXCUT.
FOXCUT knockdown inhibits the proliferation and migration of 5-8F cells
After establishing siRNA efficacy, we investigated the effects of FOXCUT and identified the function of FOXC1. CCK-8 assays demonstrated that FOXCUT silencing resulted in markedly lower tumor cell proliferation (Figure 3(a)). Additionally, wound-healing assays demonstrated that the migratory potential of FOXCUT-silenced cells was significantly reduced when compared with scrambled si-NC-treated NPC cells (Figure 3(b)). Furthermore, real-time cell-migration assays verified these phenotype changes in 5-8F cell lines (Figure 3(c)).

Effects of FOXCUT knockdown on the proliferation and migration of 5-8F cells. (a) Cell proliferation in the si-NC group was greater relative to that observed in the si-FOXCUT group. (b) Wound-healing assay showing that FOXCUT knockdown reduced the migration of 5-8F cells. (c) Cell migration assay indicating that the si-FOXCUT group exhibited decreased migration as compared with that observed in the control group.
FOXCUT knockdown decreases levels of VEGF-A, β-catenin, and MMPs
β-Catenin is one of the most important EMT-regulated gene in NPC cells, and β-catenin and MMP levels, as well as expression of the angiogenesis factor VEGF-A, were identified as significant NPC-related markers,9,14,15 with MMP expression and mesenchymal markers correlating with FOXC1 expression in cancer.16,17 To determine the role of FOXCUT in NPC, NPC cells exhibiting high levels of FOXCUT expression were subjected to FOXCUT siRNA knockdown. Quantitative PCR and western blotting results confirmed reduced expression of MMP7, MMP9, VEGF-A, and β-catenin following FOXCUT silencing (Figure 4(a) and (b)). Therefore, we hypothesized that FOXCUT is involved in regulating NPC cell growth by targeting FOXC1.

MMP, VEGF-A, and β-catenin expression following FOXCUT knockdown by siRNA in 5-8F cells. (a, b) Western blot and quantitative PCR analysis of MMP7, MMP9, VEGF-A, and β-catenin indicated their downregulation following FOXCUT silencing.
Discussion
NPC is among the most malignant head and neck carcinomas and exhibits a remarkably distinctive ethnic and geographic distribution. 18 In recent years, genetic disorders in NPC have been widely studied, but most of previous studies regarding NPC-associated genes focused on protein-coding genes. Increasing studies have demonstrated that lncRNAs are involved in the development and progression of malignant tumors. 19 Better understanding of the expression patterns and functions of NPC-associated lncRNAs is crucial to develop new treatment strategies and diagnostic biomarkers for NPC.
This study is the first to identify co-upregulation of FOXCUT and FOXC1 expression upregulated in clinical NPC tissues and cultured NPC cell lines relative to levels measured in chronic inflamed tissues and human immortalized nasopharyngeal epithelial cells. Furthermore, high levels of FOXCUT expression were correlated with lymph node metastasis and distant metastasis, whereas high levels of FOXC1 expression were correlated with distant metastasis in NPC patients. These results indicated that FOXCUT and FOXC1 were both involved in NPC progression and suggested that they could potentially be useful in a marker panel by integrating scores from multiple markers for diagnosing NPC.
The FOXC1 protein belongs to the Forkhead transcription-factor family and is a key regulator of diverse cellular functions. FOXC1 is abnormally overexpressed in various types of malignant tumors, including those associated with ovarian cancer, breast cancer, and primary prostate cancer.8,20,21 Previous studies of NPC-related genes primarily focused on the protein-coding gene FOXC1, and here, we performed bioinformatics analysis to identify a novel lncRNA (FOXCUT) located upstream of the FOXC1 promoter region. This finding led to our hypothesis that FOXCUT and FOXC1 constitute an lncRNA–mRNA pair involved in NPC progression.
Through RNA interference assays, we discovered that FOXCUT knockdown reduced FOXC1 mRNA levels. By contrast, the FOXC1 knockdown did not have a similar effect on FOXCUT expression. These findings supported an interactive role between lncRNA FOXCUT and FOXC1 and were similar to previous reports related to squamous cell carcinoma. 12
Increasing evidence implicates lncRNAs in NPC tumorigenesis, progression, and metastasis. The lncRNA CDKN2B antisense RNA 1 (ANRIL) promotes cancer progression, 22 whereas X-inactive specific transcript (XIST) was reported as a potential marker and therapeutic target in NPC. 23 However, the expression and function of FOXCUT associated with NPC had not been elucidated. Here, we showed that FOXCUT knockdown decreased NPC cell proliferation and migration. This is the first report of a role for this lncRNA in NPC development, and our findings indicated potential roles as both a biomarker in NPC and an important functional regulator in cancer similar to the results associated with lnc00974 interactions with KRT19 in hepatocellular carcinoma. 24
Recent studies demonstrated that MMPs play fundamental roles in the migration and proliferation of tumor cells. 25 Additionally, high levels of MMP expression are associated with NPC development. Yang et al. 26 reported that MMP2 and MMP9 promote NPC metastasis, and Hsu et al. 27 showed that MMP9 enhances human NPC cell invasion. Here, we investigated whether these MMPs were also associated with FOXCUT-mediated NPC progression. Following FOXCUT knockdown, we observed reduced expression of MMP7 and MMP9, implying that MMPs are involved in FOXCUT–FOXC1-mediated functions related to NPC cell proliferation and migration.
Previous studies demonstrated FOXC1 involvement as an EMT-related gene and that suppression of FOXC1 expression attenuates EMT progression. 9 One of the signals initiating an EMT is the canonical Wnt pathway, whose stimulation triggers the translocation of β-catenin to the nucleus. 28 Abnormal activation of β-catenin contributes to cancer development through the EMT. 29 Meanwhile, EMT also promoted angiogenesis by modulating pro-angiogenic factors, such as VEGF-A. 30 Our results revealed that FOXCUT downregulation repressed the expression of β-catenin and VEGF-A, with these findings indicating possible roles for FOXCUT in EMT progression and tumor angiogenesis in NPC through regulating FOXC1 expression.
In summary, our results revealed that FOXCUT and FOXC1 are overexpressed in NPC cells and are associated with distant metastasis, thereby potentially representing markers indicative of poor prognoses for NPC patients. Furthermore, we described functions related to the novel lncRNA FOXCUT involving the targeting of FOXC1 in NPC cells. These results suggested that FOXCUT might be acted as tumor promoter provides a potential diagnostic marker and novel therapeutic target for NPC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The study was approved by the ethics committee of the Affiliated Shenzhen Bao’an Hospital of Southern Medical University (2016010631).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grants 81470071 and 81171652), the Clinical Science and Technology Projects of Jiangsu Province (grant BL2014072), and National Key Clinical Program (no. 2014ZDZK003).
Informed consent
Written informed consent was obtained from all patients prior to biopsy.
