Abstract
Background
Pharmacogenomics aims to optimise drug therapy based on genetic makeup, but traditional clinical trial design faces challenges with complexity, cost and data integration.
Objectives
This study explores integrating generative artificial intelligence (AI), specifically large language models (LLMs) like Llama3 8B, Mistral 7B v0.3 and Phi-3 Mini 3.8B, into pharmacogenomics clinical trial design through Retrieval-Augmented Generation frameworks and local knowledge bases to address the challenges.
Materials and Methods
We conducted a comparative analysis of LLMs, evaluating the accuracy, relevancy, response time and operational efficiency with a case study that assessed LLMs’ capacity to address key trial design elements. The LLMs were locally run using an RTX 4080 mobile graphics card and Intel Core i9-13980HX central processing unit, with Open-WebUI employed.
Results
Our results show that Llama3 8B and Phi-3 Mini 3.8B both achieved an accuracy and relevancy score of 0.92 and 0.89, showcasing their underscore of advanced capabilities in delivering both accurate and contextually relevant outputs. More thorough results showed that Phi-3 Mini 3.8B excelled in efficiency and scalability, while Llama3 8B provided greater contextual depth.
Conclusion
This study indicates that generative AI offers transformative potential in pharmacogenomics clinical trials, enhancing efficiency and outcomes. However, challenges such as potential bias and the need for further validation remain. Addressing these limitations and advancing multimodal AI capabilities will further support inclusive and effective trial designs.
Keywords
Introduction
Pharmacogenomics is a rapidly evolving field that aims to optimise drug therapy based on an individual’s genetic makeup. Previous studies have shown that genetic factors can influence drug response, and that personalised medicine based on pharmacogenomic data has the potential to improve patient outcomes, and optimising clinical trial design for pharmacogenomics studies is crucial for improving patients’ outcomes. 1 However, traditional approaches in designing clinical trials for pharmacogenomics studies can be very challenging, sometimes leading to sub-optimal results.
Recent advancements in artificial intelligence (AI) have opened new avenues for innovation in pharmacogenomics. Among the most promising AI technologies are large language models (LLMs), a subset of generative AI. 2 These models have demonstrated remarkable capabilities in processing and generating human-like text based on vast data sets. They can analyse extensive biomedical literature, generate hypotheses and simulate complex biological processes, making them powerful tools for enhancing clinical trial design. 3
One innovative approach is deploying Retrieval-Augmented Generation (RAG) to retrieve relevant information and use LLMs for decision-making and data analysis. 4 By leveraging RAG, researchers can efficiently access and integrate various clinical trial data sets, including previous clinical studies and relatable guidelines. LLMs can revolutionise clinical trial design in pharmacogenomics by enabling more precise and efficient planning and execution. They can assist in generating pharmacogenomics clinical trial data sets that reflect various genetic variations, optimising inclusion and exclusion criteria for patient populations and designing adaptive trials that can adjust in real-time based on interim results. 5 Furthermore, LLMs can facilitate streamlining protocol development and improve overall trial design to ensure they are more targeted and less prone to failure.
Studies by Wang et al. and Jin et al. have explored leveraging language models to enhance and address the complexity of clinical trial designs.6, 7 While they innovatively used LLMs in their research, their methodologies did not incorporate the RAG framework and were limited by the medical knowledge in their training data. Furthermore, their findings did not specifically address pharmacogenomics trial innovations and focused solely on selecting relevant patient populations for clinical trials. 7
To address these limitations, this article proposes a case study to test the effectiveness of generative AI integrated with local knowledge bases (LKBs) to support decision-making in designing pharmacogenomics clinical trials, thereby assessing its practical impact and gathering insights for future improvements. We will examine how combining RAG and LLMs can streamline the design process by interacting with researchers and optimising the development of more efficient and effective pharmacogenomics clinical trial designs. Additionally, we will discuss the current state of LLM applications in this field, highlight key benefits and potential challenges and provide a forward-looking perspective on future developments.
To provide a comprehensive understanding of these developments, the following section offers an overview of existing literature and current research in the field.
Review
Pharmacogenomics is a crucial element of personalised medicine. It focuses on studying genetic variations among individuals to tailor the efficacy and safety of medications. 8 Pharmacogenomics is essential for rational drug development and prescribing practices, as it considers how an individual’s genotype affects their response to medications. Combining clinical pharmacology with genomics predicts drug responses based on genetic differences. 9 Pharmacogenomics helps optimise treatment plans and improve patient outcomes in various populations. The overarching aim of this approach is to advance personalised medicine, tailoring treatments to minimise side effects, enhance drug effectiveness and ensure precise dosing strategies. 1
Pharmacogenomics in Clinical Trials
Pharmacogenomics plays a crucial role in clinical trials for drug development by categorising patients according to their genetic markers. This method seeks to enhance drug effectiveness, optimise the design of clinical trials and tailor treatments to individual requirements. It influences all phases of drug development and tracks adverse effects after the drug is on the market. 10 Pharmacogenomics in clinical trials is essential for pinpointing patients who are most likely to respond positively to a particular drug. This strategy can greatly enhance the effectiveness and safety of medications by enabling treatments to be tailored to individual genetic factors. 11 It significantly improves therapeutic outcomes by choosing the most effective drugs and ideal dosages for each patient, thus minimising adverse drug reactions and allowing for more precise and accurate treatment dosing. Moreover, pharmacogenomics accelerates the drug approval process and facilitates the faster introduction of new medications to the market by identifying patient groups most likely to benefit from these treatments. 12
Despite its potential, the traditional approach to designing pharmacogenomic studies presents significant challenges. Analysing the vast and complex data sets required for these studies can be overwhelmingly intricate and prohibitively expensive. The sheer volume and diversity of genetic data and the need for advanced computational resources result in substantial financial and logistical burdens. 13 A study by Bienfait et al. highlighted common challenges in pharmacogenomics, such as the lack of population diversity, a complex regulatory environment and genetic variability in drug metabolism. 14 Ko and Gim further identified barriers, including the complexity of integrating and standardising extensive genomic data sets, designing patient-specific trials with appropriate eligibility criteria and managing ethical considerations. 15
Additionally, Arbitrio et al. noted that traditional methods struggle with detecting rare genetic variants, complicating patient population selection and stratification. 16 Developing precise criteria and methods for these processes could improve the relevance and applicability of study results. However, small and heterogeneous study populations limit the generalizability of findings, potentially resulting in unreliable outcomes and affecting overall efficacy and safety profiles. These challenges can lead to significant hurdles and potential failures in pharmacogenomics research. To overcome these obstacles, comprehensive and robust study design strategies are crucial.
Role of AI in Clinical Trials
Recent breakthroughs in AI-powered healthcare research and practice suggest that AI could transform the traditional approach to designing clinical trials. Generative AI can be a promising tool to streamline clinical trials, which could optimise trial protocols by facilitating the identification and recruitment of suitable participants, leading to shorter trial durations and reduced costs.
17
Additionally, AI-powered patient selection has the potential to limit exposure to ineffective treatments and improve the generalisability of trial results, ultimately enhancing the quality of medical research.
5
A study exploring machine learning to predict and prevent early termination of clinical trials suggests that AI can optimise trial design and resource utilisation using a comprehensive local data set such as
Hence, leveraging generative AI’s capabilities might significantly allow researchers to improve clinical trial design, efficiency and success. For example, generative AI can enhance patient selection and stratification. 19 This empowers researchers to design targeted recruitment strategies, enrolling only patients who are more likely to benefit from the specific therapy under investigation, thereby improving trial efficiency and reducing exposure to potentially ineffective treatments. Furthermore, generative AI could optimise trial design and protocols. 17 They could uncover trends and associations in these large data sets, leading to the development of more robust and efficient trial protocols that minimise risks and maximise the chances of success.
Generative AI could also manage and disseminate the volume of data associated with clinical trials by automating tasks like data summarisation and report generation. 17 However, ensuring interpretability and transparency in these models is paramount. Researchers must understand the rationale behind patient selection, stratification and predicted outcomes to foster trust in AI-driven recommendations and facilitate informed decision-making. Transparency can be achieved through careful model design and explanations for generated outputs. 20
Generative AI and LLMs
Generative AI, a subfield of AI, focuses on creating models that can generate new content, whether it be text, images, music or other forms of data. It primarily leverages machine learning techniques to learn patterns from existing data and generate new, similar content. 21 Over recent years, one of the most prominent advancements in generative AI has been the development and proliferation of LLMs. These models, built on vast data sets and extensive computational power, have revolutionised natural language processing by producing human-like text, facilitating translation, answering questions and even creating content autonomously. 22
In the past year, the development of LLMs has been particularly notable, marked by the rapid adoption and integration of these models into various applications and industries. However, despite their impressive capabilities, LLMs are limited by the scope of their training data. They are inherently constrained by the information they have been exposed to, which can result in gaps when dealing with highly specialised or nuanced topics. 23 This limitation underscores the need for more advanced approaches to enhance their utility and adaptability.
Transformer Architecture
The foundation of modern LLM is the transformer architecture, introduced by Vaswani et al. in 2017. 24 This architecture departed from previous recurrent models by utilising self-attention mechanisms, allowing it to weigh the importance of different words in a sentence irrespective of their position. This innovation enabled parallel data processing, significantly enhancing the efficiency and scalability of training large models. Transformers are the backbone of many state-of-the-art LLMs, facilitating the handling of vast amounts of data and complex language structures with unprecedented accuracy and coherence. 25
Prompt Engineering
Prompt engineering is an emerging technique crucial for optimising the performance of LLMs. It involves crafting specific inputs or ‘prompts’ to effectively guide the model’s output. By carefully designing prompts, users can steer the model to generate desired responses, enhancing the utility of LLMs in various applications such as content creation, customer support and research assistance.26, 27 Prompt engineering has become an essential skill for maximising the capabilities of LLMs, ensuring that the generated content is accurate, relevant and aligned with user expectations.
Multimodal
Multimodal models represent the next frontier in generative AI, aiming to simultaneously integrate and process multiple data types, such as text, images and audio. These models can generate more comprehensive and contextually enriched outputs by leveraging diverse data sources. For instance, a multimodal model can understand and describe an image, generate a corresponding caption, or even create an image based on a textual description. Integrating multiple modalities enhances the model’s ability to understand and interact with the world more human-likely, opening up new possibilities for applications in fields such as creative arts, education and interactive technologies. 28
RAG Frameworks
RAG frameworks represent a significant advancement in the capabilities of LLMs. 29 These frameworks enhance the performance of LLMs by incorporating an information retrieval component, which allows the model to access and utilise a vast repository of knowledge to produce original and informative outputs. This integration leads to more accurate and contextually relevant responses, as the retrieval mechanism ensures that the generated content is grounded in up-to-date and pertinent information. 4 Incorporating generative AI within RAG frameworks serves to refine both the retrieval and generative processes, creating a synergistic effect that boosts the system’s overall efficacy.
By leveraging the strengths of both retrieval and generation, RAG frameworks can dynamically access external knowledge bases and synthesise new information in real-time, leading to outputs that are not only creative but also highly informed and reliable. 30 The collaborative nature of RAG frameworks involves different experts handling various tasks, each optimising a specific aspect of the information processing pipeline. This division of labour ensures that the retrieval component focuses on gathering the most relevant and accurate information while the generative component concentrates on producing coherent and contextually appropriate text. The result is a more robust and efficient framework that significantly enhances the quality of outputs generated by LLMs, making them more suitable for various applications, from customer service and technical support to content creation and academic research. 31
Integrating AI with LKBs
LKBs play a crucial role by serving as repositories for local knowledge, encompassing scientific literature, clinical databases and other relevant data sets specific to a particular domain or region. 32 These repositories are essential for maintaining a comprehensive and up-to-date collection of information that can be readily accessed and utilised in various applications. Unlike standalone LLMs, RAG utilises both the internal training data of the LLMs and leverages external information retrieved from LKBs. This hybrid approach allows RAG to overcome the critical limitations of LLMs, which often suffer from outdated or incomplete information due to the static nature of their training data. 4 RAG can effectively access and leverage relevant data from LKBs with sophisticated retrieval methods in generative models. This improves the accuracy and relevance of the generated responses and enriches query semantics by grounding them in real-world knowledge.
Integrating AI with LKBs enables a dynamic and interactive knowledge management system that can adapt to new information and evolving user needs. It facilitates a more informed decision-making process by ensuring the most pertinent and up-to-date information is readily available. This approach also promotes a more holistic understanding of complex issues by synthesising information from diverse sources, thereby supporting a more comprehensive and nuanced perspective.33, 34
Ethical and Practical Considerations in Using Generative AI for Designing Pharmacogenomics Clinical Trials
The use of generative AI in pharmacogenomics clinical trials raises significant ethical and practical issues that must be addressed to ensure these technologies are used responsibly and effectively. Its implementation necessitates a nuanced approach to navigate the ethical and practical complexities. 35 A key concern lies in the inherent susceptibility of AI models to biases. In pharmacogenomics trials, these biases can manifest as underrepresenting specific genetic groups. This can lead to skewed trial results and recommendations that fail to generalise across diverse populations. 36
Transparency and interpretability are crucial for fostering trust in AI-driven decision-making within trial design. Researchers and clinicians must understand the rationale behind AI-generated recommendations to implement them confidently. 37 This necessitates developing AI models with built-in mechanisms to explain their decision-making processes. Regulatory compliance and ethical oversight are equally critical. Integrating AI into pharmacogenomics trials demands adherence to regulatory guidelines and ethical standards. Regulatory bodies increasingly emphasise transparency, accountability and patient safety regarding AI in healthcare. 38 Achieving compliance requires a collaborative effort between AI developers, clinical researchers, ethicists and regulatory authorities.
Furthermore, the protection of patient data is paramount. Pharmacogenomics deals with highly sensitive genetic information, where any misuse could result in privacy breaches and discrimination.39, 40 Robust data security measures are essential, including stringent anonymisation techniques and strict adherence to regulatory standards like the General Data Protection Regulation (GDPR) 41 and the Health Insurance Portability and Accountability Act (HIPAA).41, 42 Researchers must prioritise secure data storage and restrict access to authorised personnel only. Additionally, AI models should be designed to prevent inadvertent disclosure of personal information.
Methodology
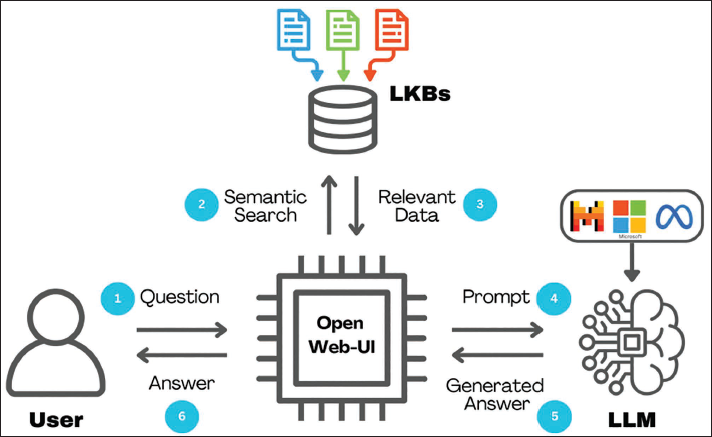
Based on the review above, the methodology of this research study integrates cutting-edge AI models with advanced implementation techniques to explore the potential of generative AI in transforming pharmacogenomics clinical trial design. By employing a combination of state-of-the-art generative AI approaches and comparative analysis methods, the study aims to enhance decision-making processes in designing pharmacogenomics clinical trials by providing recommendations based on the answers generated by integrating LKBs and LLMs detailed in Figure 1. Notably, this study does not contain any studies with human or animal participants.
Comparative Analysis of AI Models
The comparative analysis methodology employed three distinct AI models: Llama3 8B, 43 Mistral 7B v0.3 and Phi-3 Mini 3.8B.43–45 These models were selected due to their status as leading advancements in generative AI, with demonstrated strengths in producing coherent and contextually appropriate text. Additionally, they were chosen for their ability to operate with LKBs, which ensures data privacy by allowing models to run locally, an important consideration given the ethical requirements of handling sensitive data. By comparing multiple models, the study aims to identify the strengths and weaknesses of each, offering valuable insights into their applicability in real-world scenarios. The accuracy and relevancy metrics were evaluated using the Recall-Oriented Understudy for Gisting Evaluation (ROUGE) scores and Cosine similarity methods. 46 These approaches quantify the alignment between AI-generated content and reference texts, assessing both the precision of word overlap (ROUGE) and the semantic similarity of text embeddings (Cosine similarity) from the AI-generated answers compared to the LKBs employed in this research. Meanwhile, the power consumption, response time and cost data were obtained from the AI software.
The comparison criteria were defined as follows:
Accuracy: The correctness of the AI-generated responses when compared to established clinical databases and guidelines. Relevancy: The degree to which the generated responses are pertinent to the questions. Power consumption: The energy efficiency of each model during operation has implications for both environmental impact and operational costs. Response time: The speed at which each model processes queries and generates answers is crucial for real-time decision-making. Costs: The financial expenditure of running each model, including computational resources and potential licensing fees.
Implementation
To implement the AI models locally, we utilised Ollama, a tool designed to load LLMs efficiently on local machines. The AI models were run locally to address the ethical considerations and data protection for future development. We employed Open-WebUI as the deployment framework for the AI models, leveraging its user-friendly interface and robust features. Open-WebUI was selected due to its open-source nature, ease of use and built-in RAG integration. Open-WebUI supports both open-source models and closed-source LLMs, such as GPT-4, providing flexibility and extensibility in our research. The hardware configuration for running these models included a 32-GB RAM setup, an RTX 4080 mobile graphics card and an Intel Core i9-13980HX central processing unit (CPU).
Overall System Framework of Integrating Local Knowledge Bases (LKBs) and Large Language Models (LLMs) for Artificial Intelligence (AI)-generated Answers.
Case Study
The case study involved posing sample questions to support pharmacogenomics clinical trial decision-making for each AI model. These sample questions were designed to cover various aspects of trial design and the AI-generated responses were compared against existing clinical trial databases to evaluate their accuracy and relevancy. By incorporating specialised LKBs from previous open-source clinical trial data from
The rationale for each question chosen is the following:
Question 1: To ensure that the trial population is representative of diverse genetic backgrounds, which is essential for generalisable and equitable research outcomes. Question 2: To establish appropriate outcome measures for evaluating treatment success in a highly prevalent and complex condition like depression. Question 3: To ensure that control groups are appropriately selected to maintain scientific rigour while safeguarding the well-being of young participants. Question 4: To ensure that trials are designed with a comprehensive understanding of the diverse multi-regional factors, promoting ethical integrity and regulatory compliance. Question 5: To provide insights into how the AI models identify specific genetic data relevant to drug safety. Question 6: To evaluate the AI models’ ability to identify biomarkers used in oncology-personalised medicine to improve patient outcomes.
By systematically addressing these questions, the case study aimed to explore the practical impact of integrating generative AI with specialised knowledge bases on the efficiency and effectiveness of designing pharmacogenomics clinical trials. The findings provide valuable insights into the potential benefits and limitations of using advanced AI technologies in this critical area.
Results
The Supplementary Material of this article details the answers provided by the AI models to the generated questions within the case study. Based on our methodology testing results, our case study data analysis offers a comprehensive evaluation of the performance of the three AI models used, revealing nuanced insights into their strengths and areas for improvement.
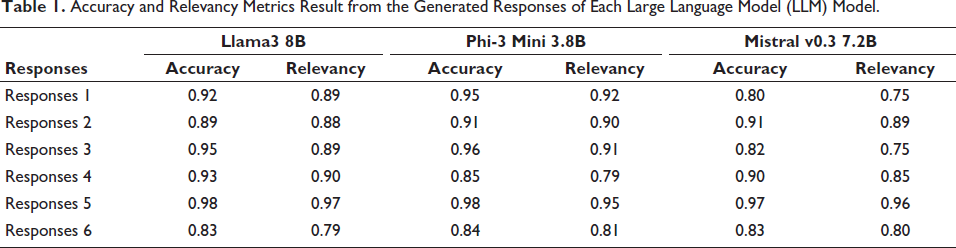
Accuracy and Relevancy Metrics Result from the Generated Responses of Each Large Language Model (LLM) Model.
Comparison of Average Accuracy and Relevancy Scores for Different Large Language Model (LLMs).
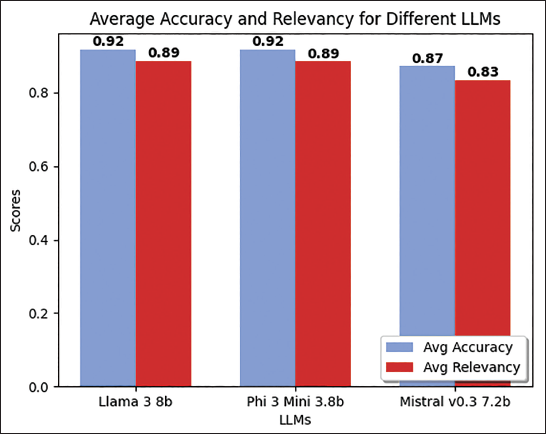
Accuracy and relevancy are paramount in determining the effectiveness of AI models in generating appropriate and precise responses. The results, as shown in Table 1 and Figure 2, indicate that Llama3 8B and Phi-3 Mini 3.8B both achieved an accuracy score of 0.92, showcasing their superior capability in producing correct answers. Mistral v0.3 7B, with a slightly lower accuracy score of 0.87, still demonstrates robust performance. Relevancy, which measures the contextual appropriateness of the responses, also favoured Llama3 8B and Phi-3 Mini 3.8B, each scoring 0.89. Mistral v0.3 7B, with a relevancy score of 0.83, indicates a need for further refinement to enhance its contextual understanding. These findings underscore the advanced capabilities of Phi-3 Mini 3.8B and Llama3 8B in delivering both accurate and contextually relevant outputs, which are critical for the complex nature of clinical trial design in pharmacogenomics.
(A) Character Counts: The Verbosity of Responses Generated by Each Large Language Model (LLM) Across Various Questions; (B) Cost of Energy Consumption: The Energy Costs Associated with Running Each LLM for Different Questions; (C) Response Time: The Response Times of the LLMs for Various Questions; (D) Energy Consumption: The Energy Consumption of Each LLM When Answering Different Questions.
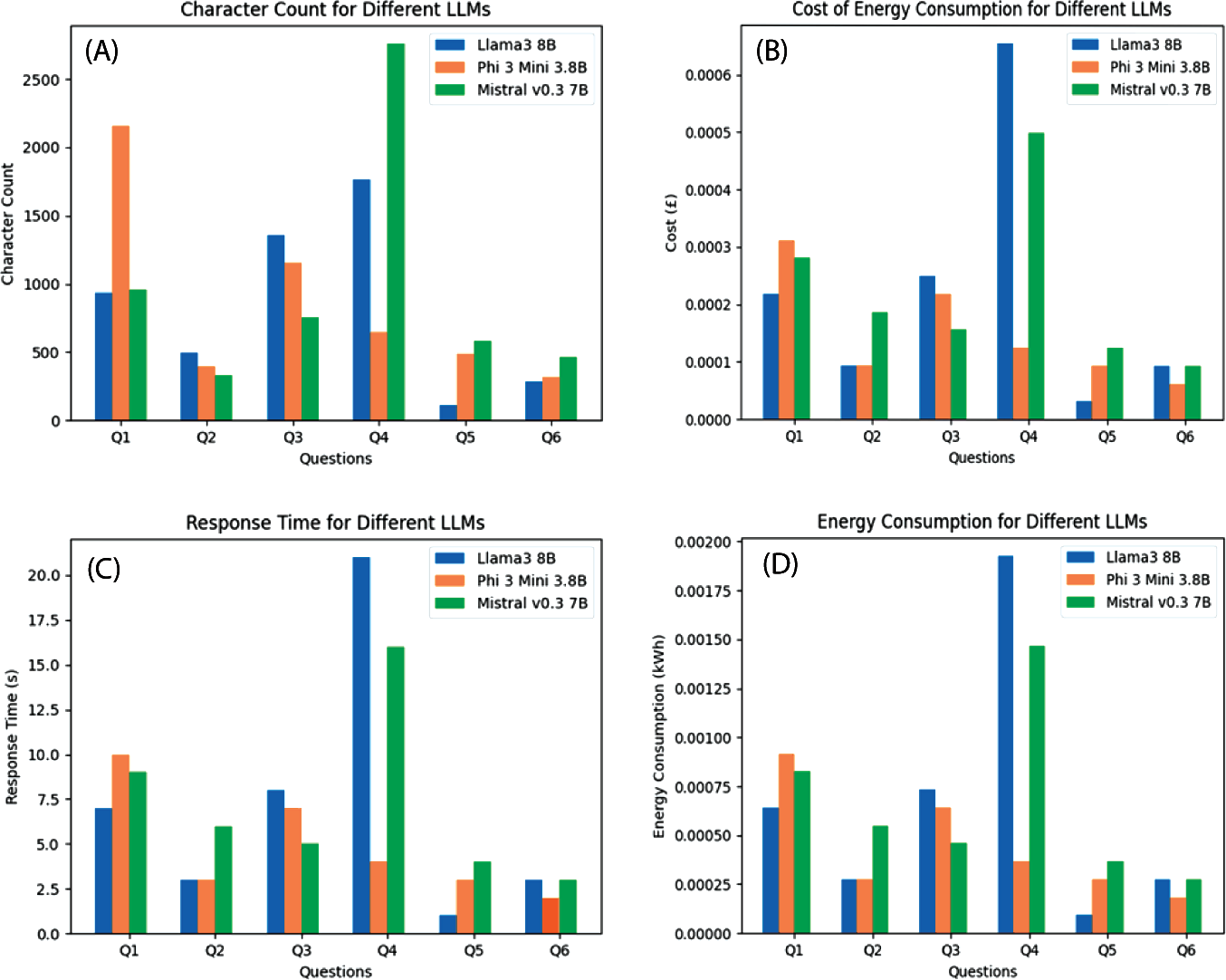
Character counts provide insight into the verbosity and detail level of the responses generated by the AI models. The analysis in Figure 3 shows that Mistral v0.3 7B often produced the highest character counts, particularly for complex questions, indicating a tendency to provide more elaborate and detailed responses. While more concise, Llama3 8B and Phi-3 Mini 3.8B maintained a balance between brevity and informativeness. This balance is crucial for clinical applications requiring concise yet comprehensive information. The character count data suggests that future refinements should aim to optimise the verbosity of responses to ensure they are detailed enough to be informative but concise enough to be practical for clinical use.
Discussion
The balance between accuracy, relevancy and character counts is vital for the practical application of AI models in clinical trial design. High accuracy and relevancy scores are essential for ensuring the outputs are correct and contextually appropriate. However, the character count analysis reveals that overly verbose responses can be cumbersome, whereas overly concise responses may lack the necessary details. Thus, the optimal model for clinical trial design should strike a balance, delivering precise and relevant information concisely to enhance usability and efficiency. This study highlights the potential of integrating generative AI into clinical trial design, demonstrating how advancements in AI can revolutionise the efficiency and effectiveness of these processes.
The power consumption comparative results, as shown in Figure 3, a critical factor when considering the deployment of AI models on a large scale within clinical settings, reveal that Phi-3 Mini 3.8B has the lowest power consumption, making it the most energy-efficient model among the three. This efficiency translates into lower operational costs and a smaller environmental footprint, which are significant advantages for sustainable AI deployment. In contrast, Llama3 8B and Mistral v0.3 7B exhibit higher energy consumption, highlighting a potential area for improvement to make them more viable for extensive use in resource-constrained environments.
The response time comparison results in Figure 3 also showed that Phi-3 Mini 3.8B demonstrates the shortest average response time, indicating its ability to process and generate answers rapidly. This speed is essential for dynamic clinical trial environments where quick iterations and adjustments are necessary. On the other hand, although their performance is still within acceptable ranges for less time-sensitive tasks, Llama3 8B and Mistral v0.3 7B possessed longer response times. The combined analysis of power consumption and response time emphasises the superior operational efficiency of Phi-3 Mini 3.8B. Its lower energy usage and faster response times make it highly suitable for large-scale and real-time applications in clinical trial design. Conversely, while Llama3 8B and Mistral v0.3 7B are still effective, their higher power consumption and slower response times suggest that they may be better suited for scenarios where these factors are less critical.
In terms of cost efficiency, the results in Figure 3 show that Phi-3 Mini 3.8B emerges again as the most cost-effective model, offering the lowest average cost per operation. This cost advantage, coupled with its superior power consumption and response time performance, makes Phi-3 Mini 3.8B an economically viable option for large-scale clinical trial designs. Llama3 8B and Mistral v0.3 7B, while effective, incur higher operational costs, which may limit their widespread adoption in cost-sensitive scenarios.
When considering the accuracy and relevancy of the responses, both Llama3 8B and Phi-3 Mini 3.8B achieve high scores, underscoring their capability to generate precise and contextually appropriate answers. However, the character count analysis indicates that Mistral v0.3 7B, despite producing more detailed responses, often exceeds practical verbosity limits. This highlights the importance of balancing detail with conciseness to ensure informative and user-friendly responses. The comparative analysis across all evaluated factors positions Phi-3 Mini 3.8B as the overall best performer. It excels in accuracy, relevancy, power consumption, response time and cost efficiency, demonstrating a balanced and robust performance. Although similar in many respects, Llama3 8B and Mistral v0.3 7B do not achieve the same level of balanced excellence. This highlights the significant advancements made by smaller LLMs, which can now deliver high performance across multiple dimensions.
Moreover, based on our results, small language models like Phi-3 Mini have shown that they can provide accurate and relevant results while optimising operational costs and power consumption. Future developments will likely focus on improving these models’ capabilities through fine-tuning and integrating them more deeply into clinical workflows. This will enable more precise patient stratification, adaptive trial protocols and streamlined data management, ultimately leading to more efficient and effective clinical trial designs. Future improvements should focus on refining verbosity and further enhancing the models’ performance to fully harness the potential of generative AI in revolutionising clinical trial processes, ensuring they are more efficient, cost-effective and sustainable.
Conclusion
This article represents the pioneering exploration of integrating generative AI with pharmacogenomics clinical trial design. Utilising the latest generative AI models, Mistral v0.3 7B, Phi-3 Mini 3.8B and Llama3 8B, this study demonstrates that AI can optimise trial design decision-making through more precise patient stratification, adaptive trial protocols and streamlined data management. However, several limitations are acknowledged. The potential for bias in AI-generated recommendations is a concern, as models can inherit and amplify biases in the data. Additionally, the current methodology requires further validation with larger and more diverse data sets to ensure the robustness and generalisability of the results.
Building on the results, future work should prioritise enhancing smaller language models to match or surpass the performance of larger models. Additionally, future research should aim to develop and test multimodal LLMs that incorporate vision capabilities to handle diverse data types, such as genomic sequences and medical imaging. This advancement will enable a more holistic analysis and a deeper understanding of patient data, paving the way for more precise and effective clinical decision-making. Moreover, integrating and scaling AI models with existing clinical trial databases and local genetic data sets will facilitate more accurate and contextually relevant AI-generated insights tailored to specific populations and regional healthcare needs. Developing suitable prompt engineering techniques will be critical, guiding the AI to produce more targeted and useful outputs for clinical trial design.
Exploring ethical considerations, such as ensuring transparency and interpretability of AI models, protecting patient privacy and complying with regulatory standards is also essential. Addressing these issues will foster trust and encourage the adoption of AI-driven approaches in designing pharmacogenomics clinical trials. Our findings advocate for continued advancements in AI integration to address limitations and enhance clinical trial design, ultimately promoting more efficient and effective pharmacogenomics trials.
Footnotes
Abbreviations
AI: Artificial intelligence; CPU: Central processing unit; EMA: European Medicines Agency; FDA: Food and Drug Administration; GDPR: General Data Protection Regulation; HIPAA: Health Insurance Portability and Accountability Act; ICH: International Council for Harmonisation; LKB: Local knowledge base; LLM: Large language model; SmPC: Summary of Product Characteristics; RAG: Retrieval-Augmented Generation; ROUGE: Recall-oriented understudy for gisting evaluation; RTX: Ray Tracing Texel Extreme (Specific Reference to GPU Hardware).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Ethical Approval
This study utilised publicly available data that did not require individual informed consent or ethical approval.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Informed Consent
There are no human subjects in this article; therefore, informed consent is not applicable.
