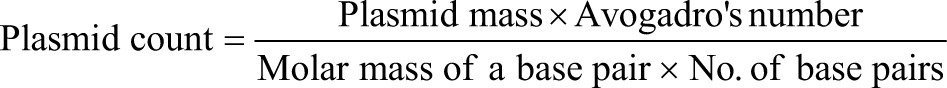Abstract
Background:
Antimicrobial resistance (AMR) remains a formidable global health threat. Conventional strategy of developing new antibiotics is costly and unsustainable. Thus, innovative approaches for resensitising bacteria using clustered regularly inter-spaced short palindromic repeats (CRISPR) technology are sought.
Aims and Objectives:
The aim of this study was to review existing technologies and approaches with the aim to identify the most feasible and practical approach to solve AMR globally.
Methods:
It is a narrative review of CRISPR research to identify a practical medical solution for combating AMR. By considering biological, social and pharmaceutical factors, feasible solutions and their limitations were identified. Next, CRISPR solutions were examined, then filtered based on strict medical expectations. A suitable cargo type and delivery mechanism will be established. Finally, the feasibility from a practical and economic perspective is considered.
Results:
Among the Cas effector modules examined, Cas12k was identified as the most promising candidate. Notably, Cas12k possesses a small protein size and lacks trans-cleavage activity, enhancing its specificity for targeted bacterial strains. Additionally, its naturally inactivated endonuclease domain further contributes to its precision. In line with this investigation, DNA plasmid was identified as the optimal cargo storage, while lipid nanoparticles were identified as the optimal delivery method.
Conclusion:
This is the first time an end-to-end consideration to AMR was considered for CRISPR-based strategies. This review offers valuable insights into a promising solution to addressing the AMR crisis. Such knowledge is essential to guide the development of novel AMR solutions.
Keywords
Introduction
Before the twentieth century, bacterial infections often resulted in death. The discovery of antibiotics by scientists had immediate success in saving lives.[1] As an evolutionary response, bacteria evolved specialised antibiotic resistance genes (ARGs). Global mortality attributable to antimicrobial resistance (AMR) in 2019 was around 1.27 million.[2]
Bacteria develop ARGs as a result of human overuse and misuse of antibiotics, for example, personal care products, pesticides, livestock growth promotion, over-prescribing antibiotics for any general infection, disease prevention in livestock etc.[3-6] ARGs can transfer to other bacteria via manure, fertiliser, groundwater, surface runoffs, hospital wastewater, viruses, clouds and through our internal microbiomes.[7-10] Bacteria spread ARGs using three horizontal gene transfer (HGT) methods: (a) ‘transformation’ by soaking up external DNA from the environment (b) ‘transduction’ via bacteriophages and (c) ‘conjugation’ using plasmids.[11] This has led to the proliferation of multidrug-resistant bacteria.
Public health organisations like the World Health Organisation (WHO), Centers for Disease Control and Prevention (CDC) and Gates Foundation are building awareness to reduce the inappropriate use of antibiotics. Governments are enforcing regulations to control the indiscriminate use of antibiotics (e.g., in 2006, use of antibiotics for livestock growth was banned in the European Union [EU]).[12] These social issues are tough to tackle and slow to bear fruits.
Small- and medium-sized enterprises (SMEs) are major contributors to antibiotics development globally, accounting for 81% of antibacterial programmes.[13] Although a driving force in innovation, SMEs are struggling to finance their efforts. Four SME biotechs, Achaogen, Tetraphase Pharmaceuticals, Aradigm and Melinta Therapeutics, which recently gained U.S Food and Drug Administration (FDA) approval for novel antibiotics and even began selling them went bankrupt.[13] Big companies including Novartis, Sanofi, AstraZeneca and GSK are moving away from antimicrobial research.[13]
To address the multi-faceted AMR problem explained above, this narrative review explores the scientific knowledge from clustered regularly inter-spaced short palindromic repeats (CRISPR) research since 1993 [Figure 1].
Flowchart showing inclusion and exclusion criteria for literature search
Medical Strategy for Solving AMR
An effective strategy to solve the crisis is to resensitise bacteria to antibiotics. ARGs are primarily found in bacterial plasmids and are spread through conjugation. Knocking out the ARGs in the infecting bacteria could weaken them and disease may subside. It is possible to cure the infection by employing this resensitising medicine as an adjuvant antibiotic therapy and administering an antibiotic for which the ARGs were destroyed. Alternative approaches using the trans-cleavage property of CRISPR-Cas systems to kill bacteria[14] may be lethal to humans in vivo.[15] Instead, the approach of resensitising bacteria is beneficial because it eliminates the need to spend billions to invent new antibiotics. Resensitising the bacteria will also reduce the necessary dose of antibiotic to be prescribed to cure the infection.
Comparison of Existing CRISPR Solutions for Targeting ARGs
CRISPR is a naturally occurring bacterial immune defence mechanism which precisely identifies and chops up viral DNA or RNA to protect the bacteria from viruses.[16] CRISPR Class-1 systems are many-component systems consisting of multiple effector modules that are complicated for scientists to use with specificity in a heterogeneous cell.[16] Class-2 systems, on the other hand, have single multi-domain proteins that are relatively easier to engineer. Most current CRISPR research is focused on Class-2 systems. However, Class-1 with its cascade of effector modules is very abundant in nature and 90% of all bacteria and archaea that employ CRISPR use Class-1.[17]
A naturally occurring CRISPR Class-2 system contains three components. The first part contains two CRISPR-associated genes Cas1 (nuclease/integrase) and Cas2 (endoribonuclease), which help bacteria adapt to new viruses.[18] The second part has clusters of these spacers, which are regularly interspaced by short palindromic repeats, called the CRISPR locus, used as templates to identify viral genomes.[18] The third part is the effector module – crRNA runs interference with the virus by cleaving its DNA or RNA. crRNAs can be engineered in labs to create precise gene cutting tools.[18]
CRISPR technology is advancing at a tremendous pace. Below are the characteristics of the three major Class-2 Types-II, V and VI that are relevant to solve the AMR problem.
CRISPR Edits – Cas9 (Class-2, Type II), Cas12 (Class-2, Type V), Cas13 (Class-2, Type VI)
Cas9 and Cas12 effector modules cleave double stranded DNA. Both cleave a strand by seeking out the Protospacer Adjacent Motif (PAM), aligning the protospacer sequence, and finally cleaving. The prokaryote’s natural repair processes fix breaks by homologous recombination or by non-homologous recombination.[19] This cleaving and repair process continues multiple times until an indel error occurs.[19] This indel knocks out the gene, but theoretically a functioning silent or conservative missense mutation could still happen.
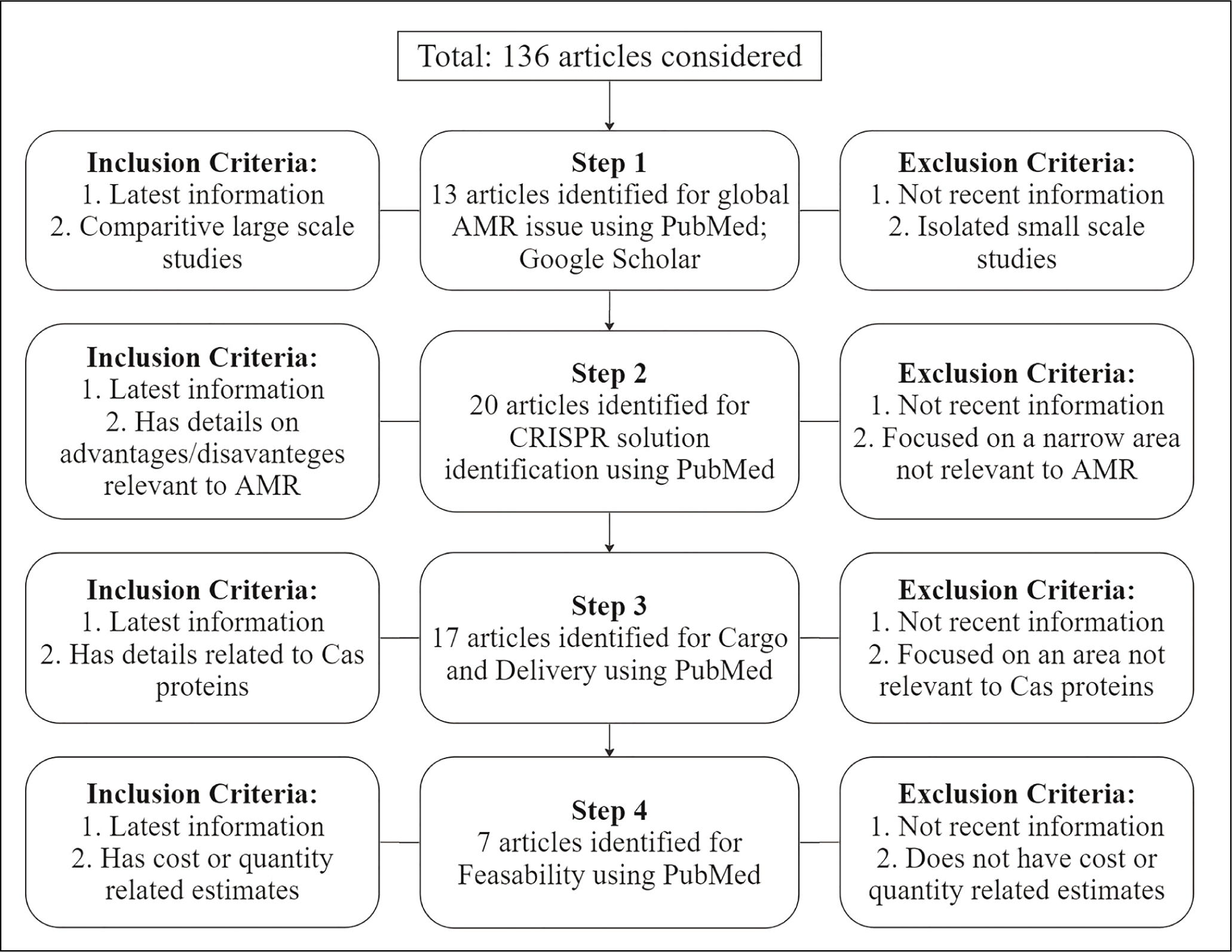
Cas9 does a straight double stranded cut, whereas Cas12 cuts the two strands in a staggered fashion [Figure 2], leading to different lengths between the two strands of DNA.[20] Out of both effector modules Cas12 acts faster than Cas9.[21] Cas12a cleavage behaviour leads to over 50% of edits being over six base pair (bp) deletions, while Cas9 generates small indels of one or two bp.[20] This Cas12 action is desirable to quickly knockout ARGs. Cas12’s fast action is utilised in diagnostic applications like HOMESv2, DETECTR and E-CRISPR.[22]
The mechanism through which Cas12k performs cleavage. Cas12k uses the crRNA for a DNA double-strand break. It cuts at different sites for both strands leading to a staggered cut
ARGs are primarily in the DNA, but Cas13 action is on RNA. Additionally, Cas13 also exhibits a non-specific trans-cleavage activity.[23] This makes Cas13 unsuitable for solving AMR.
Target Cleavage Efficiency and Specificity
An important difference in the actions of Cas9 and Cas12 is that Cas9 targets GC rich targets, while Cas12 effectors are suited to AT rich targets.[24] Studies show that compared with Cas9, Cas12a shows a lower frequency of off-target events.[25,26] When a cleavage activity occurs in the target location, it’s called a cis-cleavage, and a mistake in cis-cleavage is known as an off-target cleavage. Cas12 also exhibits non-target trans-cleavage. This action amplifies the detection signal and is beneficial for diagnostic applications.[27] However, in vivo applications have found this to be fatal.[15] Most of the Cas12 effector proteins discovered – Cas12a, Cas12b, Cas12c, Cas12d, Cas12e, Cas12f, Cas12g, Cas12h, Cas12i, Cas12j – exhibit trans-cleavage activity. Cas12k is the only known Cas12 effector that has not shown any trans-cleavage activity.[27]
Evaluating Cas12k
Transposons or ‘jumping genes’ are genes that move around in the prokaryote’s DNA. Cas12k (V-K, C2c5 or V-U5) is a transposon-encoded CRISPR-Cas variant. It is small in size, efficient, specific, sensitive and does not have trans-cleavage activity.[28] In bacteria, V-K systems provide a biological advantage for the host transposons by directing integration into other mobile genetic elements, to enhance transposon mobility and to minimise the damage to the host.[28] The bacteria Scytonema hofmanni natively carries CRISPR-associated transposase (ShCAST). A group led by BROAD institute showed that ShCAST can be used to mediate efficient and precise genome insertions in Escherichia coli.[28]
The Cas12k protein consists of only 639 amino acids.[29] This is less than half the size of Cas9 at 1368 amino acids,[30] giving Cas12k a clear advantage of being flexible enough to fit for multiple delivery methods. Its small size also allows space for other components of the CRISPR-Cas system to fit into one plasmid.
Compared with Cas9 or Cas12a, Cas12k has a naturally inactivated endonuclease domain, allowing it tight control over the domain’s cutting function which is advantageous for transcriptional regulation.[29] By designing sgRNAs targeting the genome so that the Cas12k–sgRNA complex binds to the target gene, transcriptional elongation of the gene can be blocked and gene expression can be downregulated.[29]
Inserting stop codons using base editing to precisely convert four codons (CAA, CAG, CGA and TGG) into stop codons silences genes.[31] Cas12k has been used to insert DNA downstream of the protospacer.[28] So, theoretically speaking, to knockout ARGs and resensitise antimicrobial-resistant bacteria, one can precisely insert stop codons TAA, TGA or TAG within ARG sequences using Cas12k. This is another benefit for Cas12k.
Medical Application: Drug Evolvability, Specificity and Non-immunogenicity of Cas12k
The Cas12k system has high drug evolvability. They offer several potential upsides compared with other similar systems because of their compact size, easy programmability and unidirectional integration behaviour.[32] Cas12k has very few off-target effects, making it have a high specificity of 98.1%.[32]
The gRNAs composition will highly affect the immunogenicity by initiating an innate immune response. The 5’-triphosphate gRNAs trigger the innate immune response leading to cytotoxicity but 5’-hydroxylated gRNAs do not.[33] So, 5’-hydroxylated gRNA for the CRISPR-Cas system should be produced to ensure the least possible cytotoxicity after delivery. No research has yet been published on the immunogenicity of Cas12k.
Evaluating CRISPR Drug Delivery Mechanisms
There are many drug delivery methods. Knowing the type of cargo, the space it takes in the capsule, the time it takes before breaking down and its feasibility will be important in assessing how useful they will be for solving AMR.
Cargo Types
Delivery Methods
Viral
Lentiviral (LV) vectors can accommodate all CRISPR components.[36] LVs have the advantage of low prevalence of pre-existing immunity and low immunogenicity in the general human population.[37,38] LVs ensure sustained expression as they integrate randomly into host genomes; however, there is risk of insertional mutagenesis. The potential transduction of antigen-presenting cells by lentiviruses has been observed to stimulate a robust immune reaction against the transgene product.[37]
Adenovirus (AdV) readily accommodates the CRISPR-Cas machinery. Its non-integrative nature mitigates the risk of off-target effects, rendering it a prevalent choice in clinical trials.[34] However, exposure to AdVs throughout an individual’s lifespan often results in the development of neutralising antibodies, triggering acute immune responses. This leads to tissue viral vector inactivation and tissue inflammation.[33-35,39]
Adeno-associated viruses (AAVs) stand as prominent viral delivery platforms, esteemed for their minimal cytotoxicity, replication deficiency, capability to infect both quiescent and dividing cells and diverse serotypes facilitating targeted tissue infection.[34] Furthermore, their exemplary safety profile has garnered regulatory approval for clinical applications. But AAVs have a limited payload (<4.7 kb) and can also activate the immune system.[33-35,40-43]
Phage therapy has emerged as a compelling therapeutic avenue in combating AMR. Recent years have witnessed a proliferation of studies exploring phage therapy.[44] Nonetheless, employing phages as delivery vectors for the CRISPR-Cas system has not yet been approved by the FDA.[45] It has several limitations, including inability to target multiple hosts, development of immunity against the phage and safety concerns around the natural evolution of the phage beyond its intended function.[33,34,46]
Physical
Physical delivery techniques include electrical, thermal and mechanical methods. Permeating the cell membrane to deliver the CRISPR cargo via these methods is not practical in vivo. For in vitro applications, these methodologies are straightforward and yield efficient editing outcomes with minimal constraints on cargo size. However, when applied in vivo these techniques may significantly impact cell viability or induce tissue damage. Therefore, any physical delivery method is not viable for delivery into the human body.[34]
Chemical
There are several chemical delivery methods but most will not work efficiently as delivery vectors for the CRISPR-Cas system.
Gold nanoparticles (AuNPs) can induce a non-specific inflammatory response and has potential toxicity at high concentrations in vivo.[34] The DNA nanoparticle delivery system utilises a yarn-like DNA nanoclew (NC), suitable for applications both ex vivo and in vivo.[34] The DNA nanoparticles are yet to be thoroughly researched and need further investigations on immunogenicity, safety and efficacy of treatments.[34,47] Cell-penetrating peptides (CPPs) do not confer protection against protease degradation nor ensure cell type-specific targeting.[34] Virus-like particles (VLPs) production necessitates cell culturing, incurs relatively high costs and they offer poor stability for RNA molecules.[34,48] Covalently encapsulating polymer nanoparticles (PNPs) offers less protection against immune responses and nuclease degradation while non-covalent PNP encapsulation exhibits reduced loading capacity and packaging efficiency, necessitating elevated dosages, thereby leading to cytotoxicity.[34,49]
Lipid nanoparticles (LNPs) possess the capacity to shield the cargo from nuclease degradation and immunological responses, with the flexibility to adapt their compositions for efficient in vivo delivery. Having small particle size, uniformity, stability and targeted delivery are imperative prerequisites for the clinical utilisation of LNPs in vivo.[34]
Out of all of these options for viral, physical and chemical delivery methods, LNPs are the most suitable option for use in humans. LNPs are also easy to prepare and administer.[34] LNPs are the most extensively studied non-viral vectors for nucleic acids and some FDA-approved vaccines also use LNPs.[35,50] LNPs have very low immunogenicity and further research is ongoing to regulate its immunogenicity.[50]
Feasibility of the Solution from a Practical and Economic Perspective
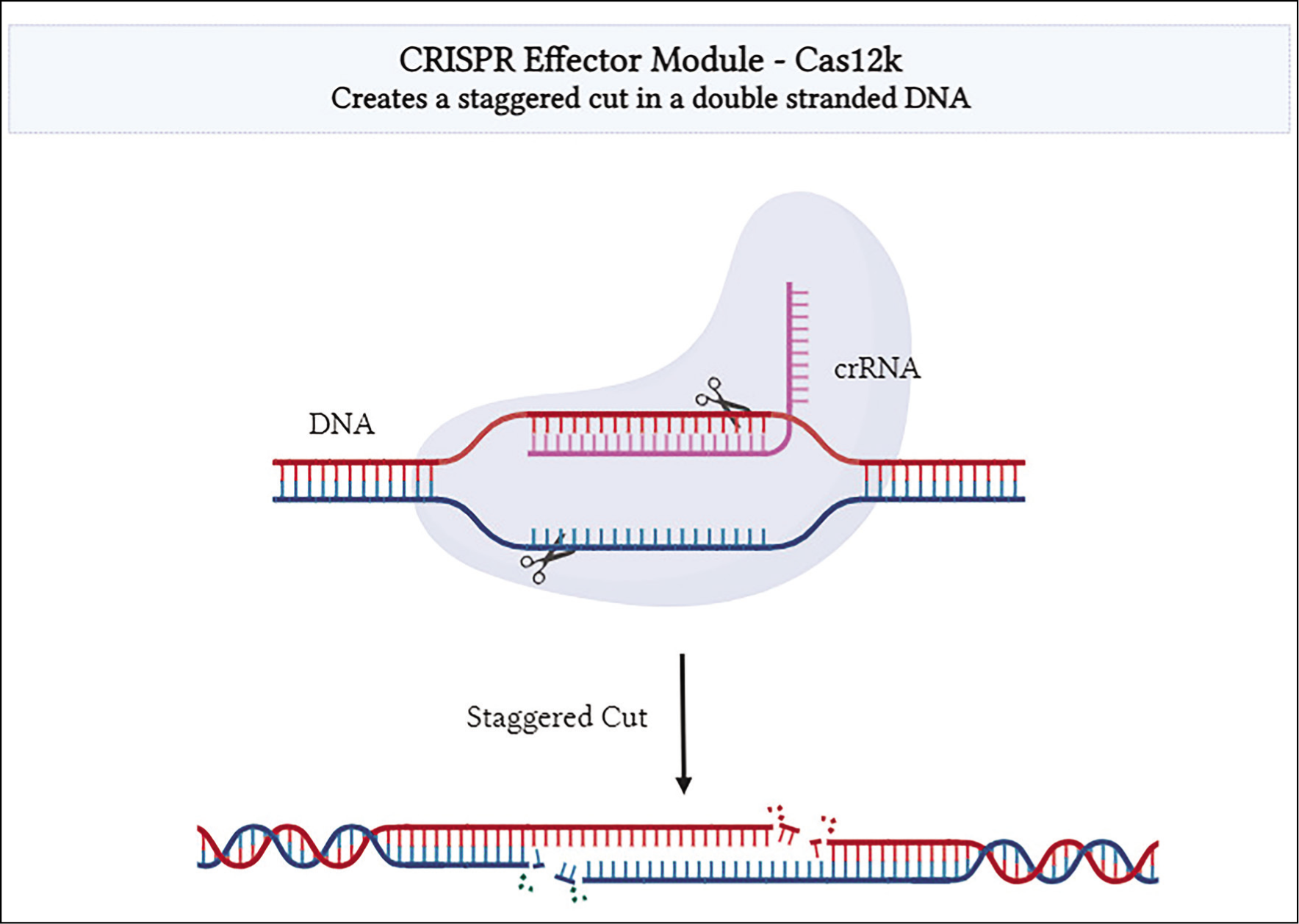
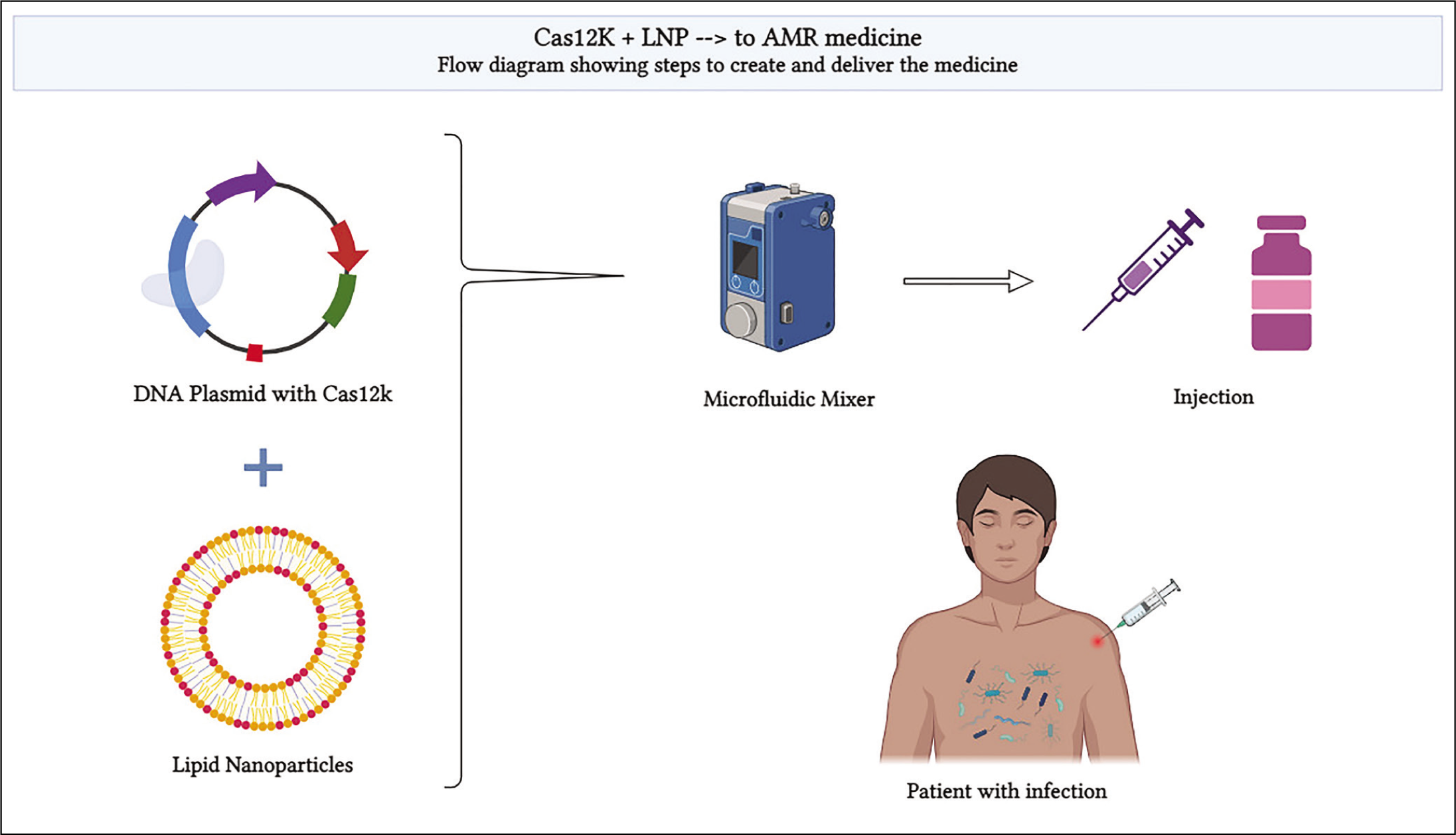
By considering the process of the DNA plasmid production, the price of one injection will be determined [Figure 3]. Count of bacteria in the blood of a sick person is typically <10 centrifugal units (CFU) per mL of blood.[51] The average human has 5000 mL of blood,[52] so a gross estimate would be 50,000 CFU of bacteria in blood. Here I assume 20 transfer units (TU) of cargo per CFU of bacteria will be used. This is to ensure that even if some of the cargo may get segregated before entering the cell and some may receive a defective CRISPR system at least one perfectly working system will be present in every bacteria.[53] From 20 TU per CFU an estimated 1×10 6 plasmids are required. Producing 4 µg of dry plasmid will cost an estimated $0.65 USD / bp.[54] Assuming an 8.4 kb plasmid size to integrate a large piece of DNA, for single-step insertion,[55] the cost for one order will be $5460. The average molar mass of a DNA bp is 660 grams/mole.[56] Using Equation 1 an estimated 4.3449×10 11 plasmid copies will be present for 4 µg of dry plasmid. This will be enough for 4,34,490 injections. Hence, the cost of one injection with enough plasmids is $0.012566. The cost of producing one dose of a mRNA vaccine from scratch encased by LNPs is $2.7.[57] Assuming the same process will be used for storing DNA plasmids the cost would not increase. So, the total production cost for one vaccine will be $2.72. The price may increase based on the cost of researching, testing the vaccine through preclinical trials, putting it through clinical trials for FDA approval and the final distribution of the vaccine.
The Cas12k production and delivery process. LNPs and the DNA plasmid will be mixed in a microfluidic mixer so the LNPs encase the DNA plasmid. It will then be stored in injection vials, transported and administered to the patient with infection
Conclusion
Cas12k emerges as a standout candidate to combat AMR due to its exceptional efficiency in targeting resistant bacterial strains. It consists of only 639 amino acids and has no trans-cleavage activity. Cas12k possesses an inherent advantage in transcriptional regulation, owing to its naturally inactivated endonuclease domain. This attribute enhances its utility in precisely modulating gene expression, thereby increasing its therapeutic potential against AMR.
The versatility of Cas12k extends to its delivery mechanisms, wherein it can be efficiently transported using LNPs, offering a cost-effective means of dissemination. Additionally, its genetic blueprint can be encapsulated within a DNA plasmid, encoding both the guide RNA (gRNA) and the Cas12k protein, facilitating ease of storage and distribution.
Each vaccine of Cas12k is estimated to be priced at $2.72; this is around the same price of an RNA vaccine making it highly affordable. However, it’s crucial to note that this estimate may fluctuate based on the expenses incurred during the research, preclinical and clinical trials, as well as the final distribution logistics.
In theory, this solution can work on any bacteria as long as the ARG can be identified. However, a limitation of this study is that it is based on knowledge from existing research, and CRISPR is a relatively new field where the scientific literature does not have sufficient depth in certain areas. For example, further investigation is required to ascertain the adequacy of a single injection versus the necessity of a daily dosage over the course of a week. Additional research is required to evaluate the feasibility of accommodating multiple distinct ARGs within a single plasmid for combating multidrug-resistant bacteria. Further investigation is also needed to find out the precise dosage requirements for addressing blood, lung, skin, urinary tract, ear-nose-throat and ocular infections in humans. Finally, ethical and legal considerations will also need to be addressed before initiating clinical trials.
Supplemental Material
Supplemental material for this article is available online.
Footnotes
Acknowledgements
Maurice Cheung was my mentor, who helped keep me on schedule, helped identify various publishing avenues and critically reviewed the study proposal. Tan Wen Hui critically reviewed the study proposal.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author received no financial support for the research, authorship and/or publication of this article.
Credit author statement
HSN conducted the research and wrote the manuscript.
Data availability
Not applicable.
Use of artificial intelligence
Artificial intelligence was not used in the preparation of this manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.