Abstract
Background
Liver cancer is a complex disease, and the upregulation of Nrf2, a key molecule in the Keap1/ARE signaling pathway, is expected to become a new method for liver cancer and prevention. Curcumin is a natural antioxidant, and polydopamine nanoparticles, as a new type of nano-drug carrier, have excellent biocompatibility and drug-loading performance and can improve the targeting and efficacy of drugs.
Objectives
This study aims to explore the effect of polydopamine nanoparticle-encapsulated curcumin on upregulating the expression of Nrf2 through the Keap1/ARE signaling pathway, thereby improving liver damage in mice with liver cancer and providing new strategies and basis for the treatment and prevention of liver cancer.
Materials and Methods
After the H22 cell culture and liver cancer mouse model were established, they were divided into groups for the study of the mechanism of action of the prepared polydopamine nanoparticles-coated curcumin (CUR-PDA NPs), and the effect of CUR-PDA NPs expression on inhibition of cell proliferation and migration, promotion of apoptosis, and delaying liver cancer progression was studied through animal experiments. This process is related to the Keap1/ARE signaling pathway and Nrf2 expression. The mechanism is further verified by Add-Nrf2, Nrf2-shRNA, Keap1-shRNA, and other methods.
Results
CUR-PDA NPs upregulate the expression of Nrf2 through the Keap1/ARE signaling pathway to improve liver injury in mice with liver cancer. There is an interaction between the expression of CUR-PDA NPs and Keap1. Increasing the expression of Keap1 can inhibit the activation of Nrf2, thereby inhibiting the antioxidant and detoxification mechanisms, resulting in reduced cell survival rate, increased apoptosis rate, reduced migration, and reduced levels of IL-1β and TNF-α.
Conclusion
CUR-PDA NPs upregulates the expression of Nrf2 through the Keap1/ARE signaling pathway to improve liver injury in mice with liver cancer. This provides new strategies and a basis for liver cancer treatment and prevention.
Introduction
Although there are many treatment methods for liver cancer, the recurrence rate is high, resulting in a low cure rate (Rich et al., 2017). It is very important to find more effective treatment methods. After extensive research, it was found (Riordan & Williams, 2000) that curcumin showed potential anti-cancer activity in cancer treatment. Curcumin has powerful antioxidant properties that can neutralize free radicals, reduce oxidative stress, and protect cells from oxidative damage, thereby helping to inhibit the growth of cancer cells (Ikeda, 2019). In addition, curcumin can inhibit the growth of cancer cells by regulating cell cycle proteins, reducing DNA synthesis, and other ways (Perrone et al., 2015). Studies have shown that curcumin can activate Nrf2 and promote the expression of antioxidant enzymes, thereby reducing oxidative stress damage to cells and protecting cells from damage (Simioni et al., 2018). Although curcumin can be used in the treatment of cancer, the current mechanism of action in liver cancer remains to be studied.
Abnormal activation of Nrf2 may be related to tumorigenesis and growth (Slattery et al., 2015). In addition, Nrf2 serves as a key regulator of cellular anti-oxidative stress response. When cells are oxidatively stressed, Nrf2 will be activated and enter the nucleus, resulting in the transcription of anti-oxidative stress genes in cells and reducing inflammatory responses and inflammatory factors, which helps to maintain cellular homeostasis and reduce inflammatory damage (Wang et al., 2019). Furthermore, in hepatocellular carcinoma (HCC) cells, the activation of Nrf2 is associated with tumor proliferation and survival. Studies have found (Liu et al., 2016) that when oxidative stress aggravates, the activity of Keap1 is inhibited, which will cause Nrf2 to break away from the inhibition of Keap1, thereby prompting Nrf2 to enter the nucleus and bind to ARE, initiating the expression of antioxidant genes, thus enhancing the antioxidant capacity of cells (Loboda et al., 2016). The Keap1/ARE signaling pathway regulates the expression of a series of antioxidant enzymes and detoxifying enzymes in liver cells. These enzymes help remove free radicals and harmful oxidative substances, maintain the oxidative balance in cells, and thus protect liver cells from oxidative damage (Tu et al., 2019). Moreover, by interfering with the Keap1/Nrf2/ARE signaling pathway, it can inhibit the proliferation of liver cancer cells, induce apoptosis, inhibit angiogenesis, and enhance chemotherapy sensitivity (Toma et al., 2011). Although the Keap1/ARE signaling pathway and Nrf2 play an important role in liver cancer, whether curcumin can play a therapeutic role in liver cancer by upregulating the expression of Nrf2 through the Keap1/ARE signaling pathway remains to be further studied.
Studies have found (Franson et al., 1991) that curcumin is easily metabolized and eliminated in the body, resulting in low bioavailability and low solubility in water, thus affecting its absorption in the body. The use of polydopamine nanoparticles can increase the solubility of curcumin and extend the residence time of curcumin in the body, thus improving its bioavailability and making it work more effectively (Hayes et al., 2015). In addition, by using nanoparticle encapsulation, the dose of curcumin can be released gradually, reducing potential toxic side effects (Ma et al., 2019). Based on this, this study will focus on the intervention role of the drug loading system in liver cancer, through understanding the specific mechanism of action, in order to provide a basis for the clinical treatment of such patients, at the same time, the successful application of this drug loading system will also be conducive to the full development of mesoporous silica nanoparticles in the medical field, which is of great significance for the clinical application of traditional Chinese medicine.
Materials and Methods
Experimental Materials
Mouse-derived liver cancer cell line H22 (Shanghai Guandao); CUR (specification: 100 g, purity: ≥98%, molecular formula: C42H72O14) Ita Biotech; rabbit anti-mouse Keap1, Nrf2, ARE (all purchased from Beijing Biorebo Technology); secondary antibody (Shanghai Uniview Biotechnology); scanning electron the microscope (scanning electron microscope, TEM) was purchased from Shanghai Carl Zeiss, and the inverted microscope (Shanghai Leica Microsystems).
H22 Cell Culture and the Establishment of Liver Cancer Mouse Models
The culture was based on 37°C, 5% CO, and cells were cultured and passaged in a cell culture incubator. Collect cells in the logarithmic growth phase and use trypan blue to evaluate their survival rate. The cell survival rate is required to exceed 95%. Then, use phosphate buffered saline (PBS) to adjust the cell concentration to 1 × 107 cells per mL to facilitate animal experiments: 0.2 mL of H22 cell suspension was injected subcutaneously in the right armpit of mice.
Mice that were successfully modeled were included in the model group. The mice were orally administered with 50 mg kg−1 CUR and were designated as the CUR group. The mice were orally administered with the same dose of PDA NPs and polydopamine nanoparticles-coated curcumin (CUR-PDA NPs) and were designated as the PDA NPs group and CUR-PDA NPs group, respectively. Also set Keap1-shRNA, Add-Keap1, CUR-PDA NPs+Keap1-shRNA, CUR-PDA NPs+Add-Keap1, Add-Nrf2, Nrf2-shRNA, CUR-PDA NPs+Add-Nrf2, and CUR-PDA NPs+ Nrf2-shRNA group.
Preparation and Characterization of Nanoparticles
Preparation of CUR-PDA NPs
Add an appropriate amount of CUR to distilled water and stir to fully dissolve to obtain a uniform CUR solution with a concentration of 20 mg/mL; then place the PDA solution in an ultrasonic cleaner to remove insoluble impurities and bubbles to form a uniform PDA Precursor solution, transfer the cleaned solution to a water bath, keep the temperature at 70°C–80°C, and continue stirring to further diffuse the PDA molecular chains and form nanoparticles. Slowly dissolve the CUR solution in the PDA precursor solution, and mix them well by stirring well to obtain a uniform CUR-PDA solution. Unreacted substances were removed by ultracentrifugation, and finally, the nanoparticle samples were dried to obtain composite samples of CUR-PDA NPs.
Nanoparticle Characterization
Dip a small amount of the nanoparticle dispersion liquid and evenly drop it on the conductive adhesive. After natural drying, spray gold for 1 min. Then use TEM to observe the shape and structure of the nanoparticles, and analyze its potential with dynamic laser scattering.
Cell Viability Detected by Cell Counting Kit (CCK)
The transfected liver cancer cells were seeded into a 60-well plate at 1 × 103 (200 µL). After 4 h, the cell culture medium was aspirated, and DMED medium containing 10% CCK-8 was added and incubated in the dark for 1 h. The growth of the cells on days 1, 2, 3, 4, and 5 was measured with a CCK-8 kit (Wuhan Jinkairui), and the growth curve was measured. Operate strictly in accordance with the standards of the CCK-8 manual. The absorbance value of each small well at a wavelength of 570 nm was measured by the enzyme labeling method, and the cell viability was calculated.
Transwell Detection of Cell Migration
Add 100 µL (200 µg/mL) Matrigel (Shanghai Yisheng) into the chamber. Next, liver cancer cells of each group will be prepared to a concentration of 1 × 105 cells/mL, and then these cells will be seeded. Put the serum-free culture medium containing SDF-1α into the lower chamber of Transwell (Wuhan Jinkarui), incubate for 12 h, take out and wash, fix with paraformaldehyde, stain with crystal violet, absorb the floating color, observe and incubate at 565 Detect absorbance at nm. A conventional microscope was used to observe the cell invasion after 0, 24, and 48 h of culture, and the images were analyzed, and corresponding calculations were performed.
Apoptosis
After each group of liver cancer cells was cultured for 24 h, the supernatant was first discarded, then washed with PBS solution, and then 0.25% trypsin was added for digestion, and centrifuged to collect the cell pellet. Wash the cells three times with PBS solution, and then add 500 µL of Binding Buffer solution (cell density is 1 × 106 cells/mL). Then, add 5 µL of Annexin V-FITC (Wuxi Puhe) under dark conditions and incubate. Next, 5 µL of propidium iodide (PI) was added, and the incubation was continued for 10 min under the same conditions. Finally, the apoptosis of cells in each group was detected by flow cytometry.
Enzyme-linked Immunosorbent Assay (ELISA) Detection of IL-1β, TNF-α Levels
Dissolve the anti-IL-1β or TNF-α antibody in the buffer, add it to a 96-well microplate, let stand at 4°C, pour off the antibody solution, and then wash the well walls with the buffer. The treated samples are added to the wells of the enzyme plate and incubated at room temperature for a period of time. The samples in the wells are poured out and specific anti-IL-1β and TNF-α detection antibody solutions are added to bind them to the wells. IL-1β and TNF-α binding on the pore walls. Add the substrate solution to cause the enzyme-labeled secondary antibody to produce a color reaction, and use a microplate reader or other photometer to measure the optical density of each well to determine the concentration of IL-1β and TNF-α in the sample.
Serum Biochemical Analysis
Blood samples were collected from the vein or tail vein of mice with liver cancer, left to stand at room temperature for about 30 min, waiting for the blood to coagulate, and centrifuged to separate the serum. Use the serum biochemical analysis instrument and the corresponding chemical kit, and operate according to the operating instructions of the kit. Measurements of ALT, AST, and TBil concentrations were recorded.
Polymerase Chain Reaction (PCR) Detection of Keap1, Nrf2 Gene Expression
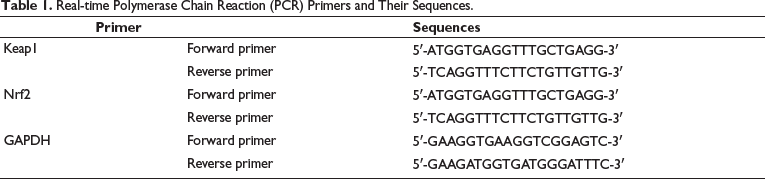
MKN28 cells were ground and homogenized with liquid nitrogen, total mRNA was extracted, and reverse transcribed into cDNA using TaqManTM advanced miRNA cDNA synthesis tool. Analysis was carried out by real-time PCR system analysis (Shanghai Unicom), and GAPDH was used as an internal reference. Relative expression levels were estimated using the 2−∆∆Ct method. Table 1 lists the primers and the sequence of the primers.
Real-time Polymerase Chain Reaction (PCR) Primers and Their Sequences.

Western Blotting (WB) Analysis of Protein Expression
Total protein was extracted from MKN28 cells by TriZol reagent, and its concentration was detected. The electrophoresed total protein was heated to 100°C and incubated for 5 min and then electrophoresed using SDS-polyacrylamide gel (120 V, 100 min). Transfer the separated proteins to a PVDF membrane. Membranes were blocked with 5% skim milk and incubated overnight at 4°C with rabbit anti-mouse Keap1, Nrf2, ARE, and GAPDH. The next day, incubate and wash the membrane, repeat three times, and incubate with a secondary antibody (1:10,000) for 1 h (37°C). Wash with PBS, repeat three times, and observe ImageJ images.
Statistical Analysis
The data obtained in the above experiments were analyzed using SPSS26.0 and GraphPad Prism software. If there is no special requirement, p < 0.05 is used as the inspection standard.
Results
The Mouse Model of Liver Cancer was Successfully Constructed and the Composite Material of CUR-PDA NPs was Prepared
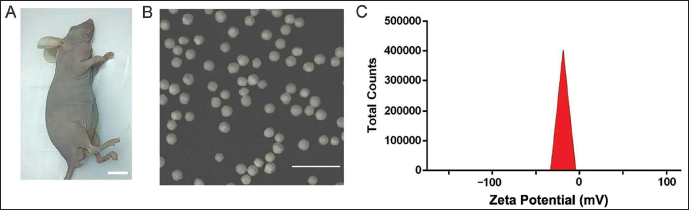
The presence of peanut-sized masses in the armpits of mice indicates that the modeling was successful (Figure 1A). No mice died during the research modeling period, and the modeling success rate was 100%.
The TEM image shows that the particles of the prepared CUR-PDA NPs composite material are round spheres with uniform size and normal shape (Figure 1B). The Zeta potential is −14.3 mV (Figure 1C), indicating that the nanoparticles are relatively stable with low dispersion.
Characterization of Polydopamine Nanoparticles-coated Curcumin (CUR-PDA NPs) Composites. The Presence of Peanut-sized Masses in the Armpits of Mice (Scale Bar = 1 cm) (A); TEM Image of CUR-PDA NPs (Scale Bar = 1 µm) (B); Zeta Potential Distribution of CUR-PDA NPs (C).
CUR-PDA NPs have a positive effect on the proliferation and migration of liver cancer cells and delay the progression of liver cancer. This process is related to the Keap1/ARE signaling pathway and Nrf2 expression.
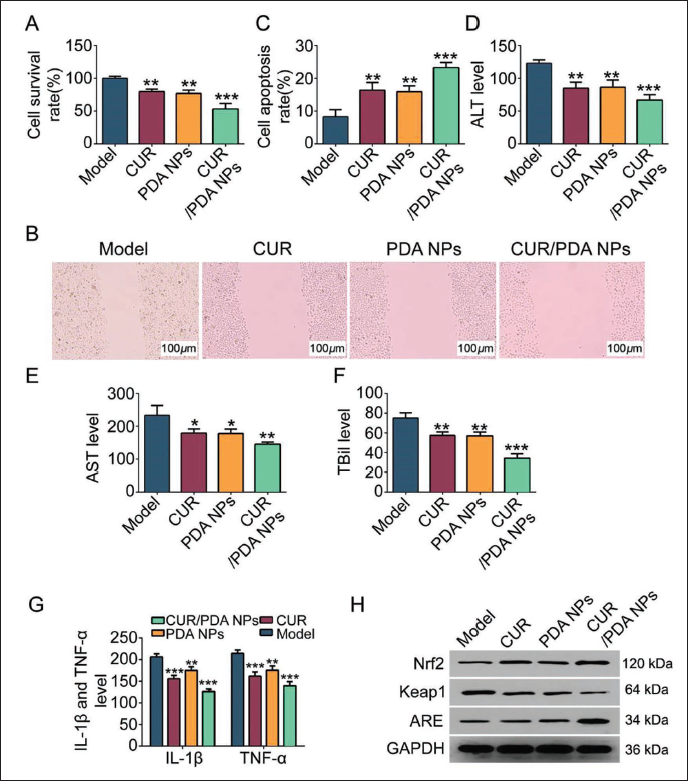
In order to explore the impact of CUR-PDA NPs on the progression of liver cancer, we observed the biological behavior of mouse liver cancer cells and found that CUR, PDA NPs, and CUR-PDA NPs intervention The cell proliferation and migration decreased significantly (Figure 2A and B), and the apoptosis increased significantly (Figure 2C); at the same time, the levels of ALT, AST, and TBil in the blood samples of mice with liver cancer showed a downward trend (vs. model group, p < 0.05, Figure 2D–F), suggesting that the liver function of the mice was improved; the levels of IL-1β and TNF-α also showed an abnormal decrease (vs. model group, p < 0.05, Figure 2G), and the above indicators were measured by CUR-PDA. The NPs group showed the most significant improvement.
In addition, the results of WB detection showed that under the intervention of CUR, PDA NPs, and CUR-PDA NPs, the expression of Keap1 in the Keap1/ARE signaling pathway was downregulated, and the expressions of ARE and Nrf2 were upregulated (Figure 2H), indicating that CUR-PDA NPs can inhibit the proliferation of liver cancer cells. The process of migration, promoting apoptosis, and delaying the progression of liver cancer may be related to changes in Keap1/ARE signaling pathway and Nrf2 expression.
Polydopamine Nanoparticles-coated Curcumin (CUR-PDA NPs) Can Inhibit the Proliferation and Migration of Liver Cancer Cells and Promote Their Apoptosis, Delaying the Progression of Liver Cancer. (A) Cell Counting Kit (CCK) Detects Cell Viability; (B) Transwell Detects Cell Migration; (C) Flow Cytometry Detects Cell Apoptosis; (D–F) Serum Biochemical Analysis Detects ALT, AST, and TBil Levels; (G) Enzyme-linked Immunosorbent Assay (ELISA) Detects IL-1β and TNF-α; (H) Western Blotting (WB) Detects the Expression of Keap1/ARE and Nrf2; Compared with the Model Group. **p < 0.01, ***p < 0.001.
CUR-PDA NPs can inhibit the proliferation and migration of liver cancer cells and promote their apoptosis, thereby delaying the progression of liver cancer via inhibiting Keap1 and upregulating ARE expression.
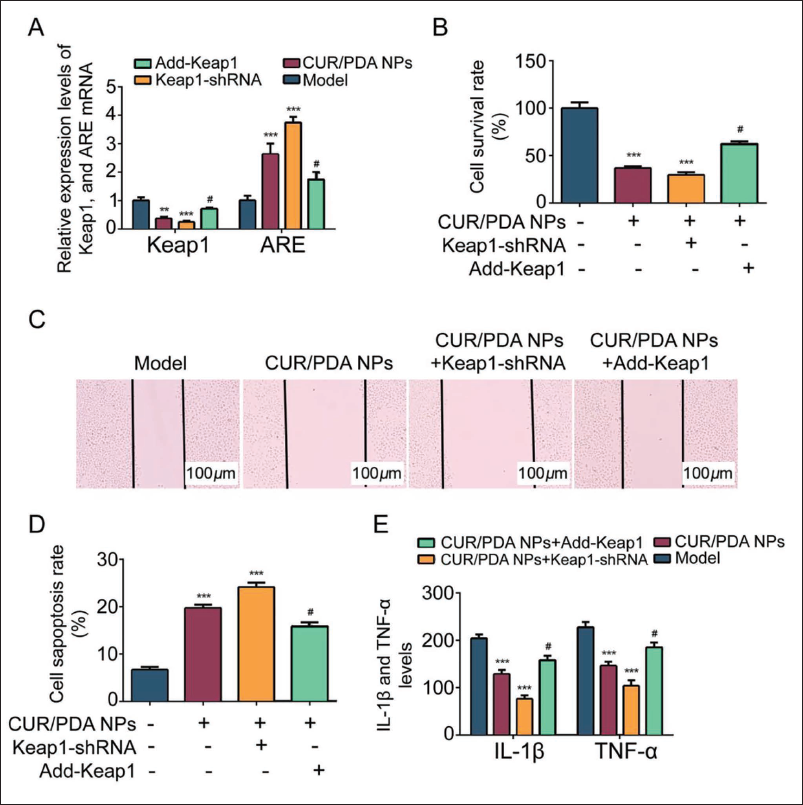
Further testing the regulatory effect of CUR-PDA NPs on Keap1/ARE, it was found that Keap1 was downregulated and ARE was upregulated in liver cancer cells in the CUR-PDA NPs group, which was consistent with the trend of the Keap1-shRNA group, suggesting that CUR-PDA NPs can mediate downregulation of Keap1 and upregulation of ARE (Figure 3A). After CUR-PDA NPs+Keap1-shRNA intervention, cell survival, and migration were inhibited, and the levels were the lowest (vs. the other three groups, Figure 3B and C), and the cell apoptosis rate was the highest (Figure 3D); IL-1β, TNF-α. The level also showed to be at the lowest level (vs. the other three groups, p < 0.05, Figure 3E). When CUR-PDA NPs were combined with Add-Keap1, the above trend was reversed.
Polydopamine Nanoparticles-coated Curcumin (CUR-PDA NPs) Can Inhibit the Proliferation and Migration of Liver Cancer Cells, Promote Apoptosis, and Delay the Progression of Liver Cancer, Which is Related to the Activation of the Keap1/ARE Signaling Pathway. (A) Polymerase Chain Reaction (PCR) Detects Keap1 and are mRNA; (B) Cell Counting Kit (CCK) Detects Cell Survival Rate; (C) Transwell Detects Cell Migration; (D) Flow Cytometry Detects Cell Apoptosis; (E) Enzyme-linked Immunosorbent Assay (ELISA) Detected IL-1β and TNF-α Levels; Compared with the Model Group, ***p < 0.001; Compared with CUR-PDA NPs Group, #p < 0.05.
CUR-PDA NPs Can Inhibit the Proliferation and Migration of Liver Cancer Cells and Promote Their Apoptosis Via Upregulating the Expression of Nrf2
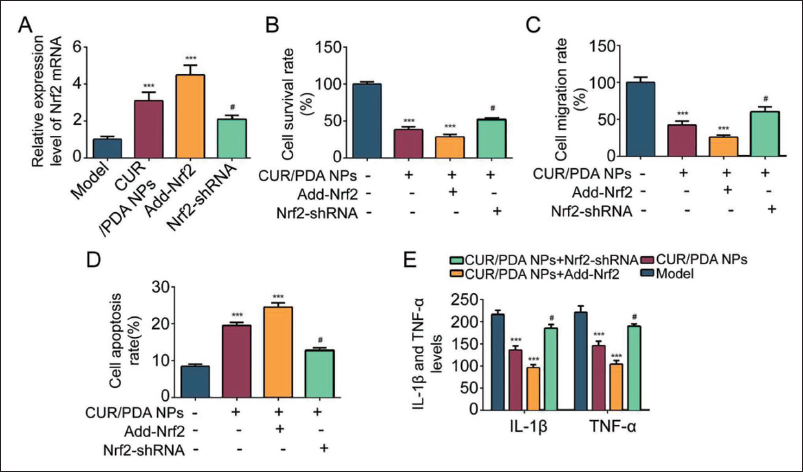
Further testing the regulatory effect of CUR-PDA NPs on Nrf2, it was found that Nrf2 was upregulated in liver cancer cells in the CUR-PDA NPs group, which was consistent with the trend in the Add-Nrf2 group, suggesting that CUR-PDA NPs can mediate the upregulation of Nrf2 (Figure 4A). After CUR-PDA NPs+Add-Nrf2 intervention, cell survival, and migration were inhibited, with the lowest levels (vs. the other three groups, Figure 4B and C), and the highest cell apoptosis rate (Figure 4D); IL-1β, TNF-α. The level also showed to be at the lowest level (vs. the other three groups, p < 0.05, Figure 4E). When CUR-PDA NPs were combined with Nrf2-shRNA, the above trend was reversed.
Polydopamine Nanoparticles-coated Curcumin (CUR-PDA NPs) Can Inhibit the Proliferation and Migration of Liver Cancer Cells and Promote Their Apoptosis, Delaying the Progression of Liver Cancer is Related to the Upregulation of Nrf2 Expression. (A) Polymerase Chain Reaction (PCR) Detects Nrf2 mRNA; (B) Cell Counting Kit (CCK) Detects Cell Survival Rate; (C) Transwell Detects Cell Migration; (D) Flow Cytometry Detects Cell Apoptosis; (E) Enzyme-linked Immunosorbent Assay (ELISA) Detects IL-1β, TNF-α; Compared with Model Group, ***p < 0.001; Compared with CUR-PDA NPs Group, #p < 0.05.
CUR-PDA NPs Upregulate the Expression of Nrf2 Through the Keap1/ARE Signaling Pathway to Improve Liver Injury in Mice with Liver Cancer
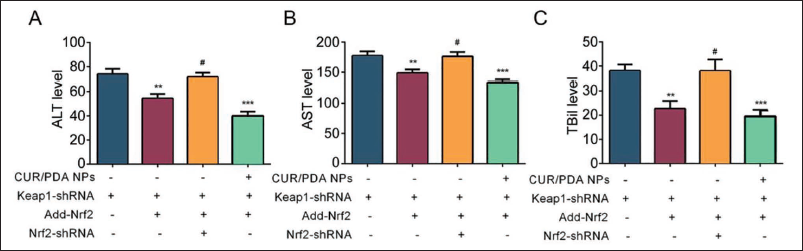
Under the intervention of Keap1-shRNA+Add-Nrf2, the levels of ALT, AST, and TBil in the blood samples of liver cancer mice showed a downward trend (vs. Keap1-shRNA group, p < 0.05, Figure 5A–C). On this basis, adding after Nrf2-shRNA intervention, the above levels were reversed, indicating that the Keap1/ARE signaling pathway can upregulate the expression of Nrf2. Moreover, the CUR-PDA NPs+Keap1-shRNA+Add-Nrf2 group had the lowest levels of ALT, AST, and TBil, suggesting that CUR-PDA NPs can improve liver damage in mice with liver cancer by inhibiting the expression of Keap1 and upregulating the expression of ARE and Nrf2.
Study on the Effect of Polydopamine Nanoparticles-coated Curcumin (CUR-PDA NPs) on Improving Liver Injury in Mice with Hepatocellular Carcinoma by Upregulating the Expression of Nrf2 Through the Keap1/ARE Signaling Pathway. (A) Serum Biochemical Analysis Detected ALT, AST, and TBil Levels; (B) Compared with Keap1-shRNA Group, **p < 0.01, ***p < 0.001; (C) Compared with the Keap1-shRNA Group Compared with the +Add-Nrf2 Group, #p < 0.05.
Discussion
Studies have shown (Peng et al., 2018) that curcumin inhibits the activation of IKK and prevents IKK from phosphorylating IκB protein, thereby inhibiting the activity of NF-κB, thereby inhibiting the growth and invasion of cancer cells and reducing the occurrence of inflammatory reactions (Maghsoudi et al., 2017). In addition, curcumin can also inhibit the phosphorylation of STAT3 by inhibiting the activation of JAK/STAT, regulating the expression of Bcl-2, VEGF, and MMP genes, thereby inhibiting the growth and metastasis of tumor cells (Peng et al., 2013). In our cell experiments, we first found that CUR can reduce cell survival rate, greatly increase apoptosis rate, and reduce cell migration (Zhang et al., 2019). Under the intervention of CUR-PDA NPs, the results showed more consistent changes. This shows that CUR-PDA NPs have a more significant impact on cells than CUR. This may be because CUR-PDA NPs are more easily absorbed and utilized by cells. It has better stability and can maintain effective concentrations in the body for a longer period of time, so it has a more lasting and significant effect on cells, and therefore has a more significant effect on cells (Yang et al., 2021). Compared with CUR, CUR-PDA NPs may have a broader mechanism of action (Yang et al., 2020). In addition, both CUR and CUR-PDA NPs can reduce the levels of ALT, AST, TBil, IL-1β, and TNF-α (Itoh et al., 2003). It shows that CUR-PDA NPs have active anti-proliferation and migration effects of liver cancer cells. Further, WB testing found that CUR-PDA NPs increased the expression of ARE, decreased Keap1, and increased the expression of Nrf2. ARE is one of the downstream target genes of Nrf2, and its increased expression further suggests the activation and expression of Nrf2. It is suggested that CUR-PDA NPs can inhibit the expression of Keap1, thereby releasing its inhibitory effect on Nrf2, and further promoting the activation and expression of Nrf2. The process of CUR-PDA NPs inhibiting liver cancer cells is related to Keap1/ARE signaling pathway and Nrf2 expression, which can reduce the inflammatory response of liver cancer cells.
Studies have found that (Li & Kong, 2009; Velichkova & Hasson, 2005) when cells suffer from oxidative stress, the combination of Keap1 and Nrf2 weakens, Nrf2 is stabilized and enters the nucleus, and activates the expression of ARE-initiated genes SOD, GST, and others, thereby enhancing the cell’s resistance to oxidative stress. When the expression of Keap1 is downregulated, Nrf2 is stabilized and accumulated, which triggers the excessive activation of the oxidative stress response, leading to DNA damage, cell cycle abnormalities, apoptosis induction, and others in cancer cells, and plays a direct toxic effect on cancer cells (de la Vega et al., 2018). In this study, we used Keap1-shRNA and Add-Keap1 and found that under the intervention of CUR-PDA NPs+Keap1-shRNA, cell survival rate decreased, apoptosis rate increased, migration decreased, and IL-1β, TNF-α. The levels showed a significant decrease in all, indicating that activation of Nrf2 can inhibit cell growth and migration and promote apoptosis while reducing the levels of inflammatory factors IL-1β and TNF-α. While CUR-PDA NPs+Add-Keap1 can reverse this result, it can be seen that there is an interaction between the expression of CUR-PDA NPs and Keap1. Increasing the expression of Keap1 can inhibit the activation of Nrf2, thereby inhibiting the antioxidant and detoxification mechanisms, resulting in reduced cell survival rate, increased apoptosis rate, reduced migration, and reduced levels of IL-1β and TNF-α. It also shows that CUR-PDA NPs can inhibit the proliferation and migration of liver cancer cells, promote apoptosis, and delay the progression of liver cancer, which is related to the regulation of the Keap1/ARE signaling pathway, specifically by downregulating the expression of Keap1 and thereby increasing the expression of ARE. It shows that CUR-PDA NPs have a positive effect on the proliferation and migration of liver cancer cells. Delaying the progression of liver cancer is related to inhibiting Keap1 and upregulating ARE expression. These findings are important for a deeper understanding of the anti-liver cancer mechanism of CUR-PDA NPs and its Nrf2-based treatment methods.
Studies have reported (Gérard & Goldbeter, 2014) that activation of Nrf2 can promote the proliferation and cell cycle progression of cancer cells by downregulating negative cell cycle regulatory proteins such as p21 and p27 or upregulating cyclin D1, CDK4/6, and others; and Nrf2 is inhibited by Keap1 in the cytoplasm mediated ubiquitination degradation when stimulated by compounds, intracellular Nrf2 will be released and react with cis-acting elements. IL-4 can induce the expression of Nrf2 and increase the levels of antioxidant enzymes, thereby inhibiting oxidative stress and inflammatory responses. In addition, Nrf2 can inhibit the inflammatory response induced by IL-1β and reduce the interaction of factors such as IL-6 and TNF-α to exert the synergistic effect of Nrf2 and IL-4, thereby participating in the excessive proliferation and uncontrolled growth of cancer cells. Cell cycle regulation. Nrf2 interacts with STAT3 to inhibit STAT3 transcriptional activity, thereby reducing the expression of IL-6 levels. In addition, it can also upregulate the expression of miR-155, inhibit the production of TNF-α, and reduce the expression of IL-6, thus interfering with tumor cells. It interacts with immune cells to reduce the sensitivity of tumors to immune responses, downregulates the expression of TNF-α, IL-1β, and IL-8, and inhibits tumor metastasis and invasion. In this study, under the combined intervention of CUR-PDA NPs and Add-Nrf2, the results were consistent with those of CUR-PDA NPs+Keap1-shRNA, while CUR-PDA NPs+Nrf2-shRNA reversed this phenomenon. This suggests that the activation of Nrf2 is crucial for the efficacy of CUR-PDA NPs, and the effect of Keap1 may be achieved only through the activation of Nrf2. This verified that the effect of CUR-PDA NPs on inhibiting the proliferation and migration of liver cancer cells, promoting apoptosis, and delaying the progression of liver cancer is related to upregulating the expression of Nrf2. Therefore, the combined use of CUR-PDA NPs and Add-Nrf2 can effectively treat the disease, while the use of CUR/PDA NPs and Nrf2-shRNA may reduce the therapeutic effect.
Conclusion
In summary, CUR-PDA NPs improve liver injury in mice with liver cancer by upregulating the expression of Nrf2 through the Keap1/ARE signaling pathway. It provides us with new ideas and methods for the treatment of liver cancer and also provides a reference for us to better understand the cellular defense system and drug delivery mechanism.
Footnotes
Abbreviations
CUR-PDA NPs: Polydopamine nanoparticles-coated curcumin; Keap1: Kelch-like ECH-associated protein 1; Nrf2: Nuclear factor erythroid 2-related factor 2.
Acknowledgments
The authors gratefully acknowledge the Xinchang County People’s Hospital laboratory for providing the necessary equipment for this study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Ethnic Committee of Xinchang County People’s Hospital.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
