Abstract
Background
Miao medicine Radix Wikstroemia indica (RWI) is derived from the root of W. indica (L.) C. A. Mey. which has an analgesic effect, but its mechanism is not clear. Because of the toxicity of RWI, the research group used the “Sweat soaking method” to process RWI in the early stage to reduce its toxicity. Therefore, this study explored the analgesic effect and mechanism of processed RWI through network pharmacology and metabolomics.
Purpose
To explore the analgesic effect and therapeutic mechanism of RWI processed by “Sweat soaking method”.
Materials and Methods
The torsion experiment was carried out with acetic acid. The metabolomic analysis of serum samples was carried out based on 1H-NMR technology, and the intersection targets of RWI and pain diseases were screened by network pharmacology for gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis.
Results
RWI has an analgesic effect and is related to metabolites such as 4-pyridoxic acid, l-glutamic acid, and agmatine. It is involved in arginine and proline metabolism, arginine biosynthesis, and alanine, aspartate, and glutamate metabolism. In network pharmacology, there were 404 common targets between RWI and pain diseases, and eight core targets were screened, including SRC, STAT3, and HSP90AA1. GO functional enrichment analysis found that RWI had effects on molecular processes such as protein phosphorylation and response to xenobiotic stimulus, cell composition such as receptor complex and membrane raft, and molecular functions such as enzyme binding. KEGG pathway enrichment analysis obtained 193 pathways. Arginine proline metabolism and nitrogen metabolism are involved in the same pathway as metabolomic analysis.
Conclusion
RWI has an analgesic effect, and its therapeutic mechanism mainly involves arginine and proline metabolism.
Introduction
Radix Wikstroemia indica (RWI) was first recorded in “Herbs Medicine of Lingnan,” which is derived from the dried root of W. indica (L.) C. A. Mey, and was often used in the Guizhou Miao area. At present, studies have shown that RWI has an analgesic effect (Geng et al., 2006; Lu et al., 2013). The research group previously studied that RWI had hepatorenal toxicity. Therefore, on the basis of respecting the traditional processing methods of Miao folk Traditional Chinese Medicine, the research group optimized the use of artificial sweat to replace the traditional “sweat soaking method” (carrying medicinal materials on the back of the body, using sweat and body warm medicine) to reduce the toxicity of RWI.
Network pharmacology analysis uses the method of network analysis to explore the interrelationship between biological effectiveness and soreness of each node by establishing a drug-target-disease network. Furthermore, this analysis is used to study the mechanism of multi-target and multi-pathway of drugs, which is consistent with the mechanism of multi-component, multi-pathway, and multi-target of traditional Chinese medicine. It has been widely used in the study of the mechanism of traditional Chinese medicine (Bi et al., 2019; Chen & Chen, 2021; Liu & Wang, 2019; Xie et al., 2019).
As a newly developed “omics” discipline, metabolomics can comprehensively monitor the changes of metabolites in the body after external stimulation (Guo et al., 2012). It is based on high-throughput detection and multivariate statistical analysis methods. By analyzing the differences in endogenous metabolites in the body, it seeks biomarkers and related metabolic pathways, thereby revealing the regulatory mechanism of the overall state of the organism after traditional Chinese medicine treatment.
The 1H-NMR technique has the advantages of mature and stable technology, simple and fast sample processing, short detection time, fast analysis speed, high recognition, good reproducibility, and high throughput (Li et al., 2018; Wang et al., 1987; Zhang et al., 2012). In this experiment, 1H-NMR metabolomics technology was used as a detection method to analyze the analgesic metabolic pathway of processed RWI, which provided the basis for the clinical use of RWI.
Materials and Methods
Medicine and Reagent
RWI was purchased from Yulin Yinfeng International Chinese Medicine Port, Yuzhou District, Yulin City, Guangxi Zhuang Autonomous Region (Batch No.20160115), it was identified as the root of W. indica (L.) C. A. Mey. by Professor Li Wei from Guizhou University of Traditional Chinese Medicine. Aspirin Enteric-coated Tablets (Batch No. BJ55569) from Bayer Pharmaceutical Healthcare Co., Ltd. Glacial acetic acid (Batch No. 20200909) was purchased from Tianjin Beichen Fangzheng Reagent Factory.
Animal Materials
SPF KM mice were obtained from Changsha Tianqin Biotechnology Co., Ltd., No.: SCXK (Hunan) 2019–0013, body weight 19 ± 1 g, half male and half female. The animal experiment was approved by the Ethics Committee of Guizhou University of Traditional Chinese Medicine (No. 20210163).
Processing of RWI and Preparation of its Extract
RWI powder was sprayed with 1% synthetic perspiration and stirred evenly. After moistening, it was dried in an oven for 24 hours at 37°C ± 0.5°C for 30 days to obtain RWI-processed products (synthetic perspiration components:
The 5 kg processed RWI powder was added to 14 times 70% ethanol for percolation. All the percolated liquid was combined for concentration and drying to obtain ethanol extract.
Serum Metabolomic Study
Grouping and Administration of Mice
Forty-eight KM mice were divided into the control group, model group, positive group, and high, medium, and low dose groups of RWI. The positive group was given aspirin enteric-coated tablets (0.2 g/kg). The model group was given an equal volume of 1% CMC solution. The doses of RWI high, medium, and low dose groups were 0.52, 0.26, and 0.13 g/kg, respectively. The control group was not treated. Each experimental group was administered continuously for seven days, once a day, and the administration method was gastric irrigation.
Establish a Mouse Pain Model by Acetic Acid Writhing Method
After the final administration for 1 hour, the mice were intraperitoneally injected with 0.6% glacial acetic acid (10 mL/kg). The latency time of the torsion reaction and the number of writhing within 15 minutes were observed and recorded. The analgesic inhibition rate of the drug was calculated to observe the analgesic effect of the drug.
Inhibition rate (%) = (Average number of torsion in the model group − Average number of torsion in the drug group)/Average number of torsion in the model group
Sample Collection
One hour after the end of the experiment, the mice in each group were taken blood from the abdominal aorta, and the serum was separated after centrifugation for 1H-NMR metabolomic analysis.
NMR Sample Pretreatment
The serum samples were centrifuged (13,000 r/min, 15 minutes) after thawing. The supernatant (180 µL) was taken into a vial containing 200 µL PBS buffer. The vial was sealed with a sealing membrane and taken out after pre-freezing at –80°C, and quickly transferred to the vacuum freeze dryer for drying. The freeze-dried serum samples were taken out, and 500 µL of D2O and 100 µL of TSP (1 mg/mL) were added. The supernatant was transferred to the NMR tube for testing.
1H-NMR Detection
Serum samples were detected by nuclear magnetic resonance spectrometer using CPMG and LED impulse sequence (instrument number: Bruker 500 MHz AVANCE III HD), and water peaks were suppressed by pre-saturation. Detection parameters: the spectrum width is 8,000 Hz, the number of sampling points is 64 k, and the cumulative number is 64 times. The nuclear magnetic spectrum of the sample was obtained by superconducting pulse Fourier transform.
Data Treatment
The phase correction and baseline correction of the spectrum were performed using Mest ReNova-9.0.1 software, and the signal peak of 0.00 ppm was used as the standard calibration. The spectrum in the range of δ0.4–6.0 was segmented and integrated according to 0.04 ppm of each segment. The δ4.6–5.0 integral segment was removed to exclude water peaks, and finally, the area normalization was performed. The data were exported in the form of Script: NMR CSV Matrix (*.csv *.txt) file.
The data were input into the MetaboAnalyst 5.0 website, principal component analysis (PCA) and orthogonal partial least squares discriminant analysis (OPLS-DA) were used for multivariate statistical analysis. The accuracy of the OPLS-DA model was evaluated by 100 repeated permutation tests. Differential metabolites were found by VIP value, t-test, and FC analysis. The screening conditions were VIP > 1, p < 0.05, FC ≥ 2, or FC ≤ 0.5.
Metabolic Pathway Analysis
The Kyoto Encyclopedia of Genes and Genomes (KEGG) database was used to analyze the metabolic pathways of differential compounds and the analgesic metabolic pathways of RWI based on differential metabolites were found through analysis.
Network Pharmacology Research
Screening of Active Components and Potential Targets of RWI
The effective compounds of RWI were screened by the TCMSP website, and the screening conditions were OB % ≥ 30 and DL ≥ 0.18. The SMILES number of the compound was searched using the PubChem database, and the gene name of the target was queried using the SwissTargetPrediction and UniProt databases. In addition, the components of RWI were found through literature, and the chemical structure of the components was searched by PubChem database and uploaded to Swiss ADME for prediction. The predicted results were screened. The screening principle was that the gastrointestinal absorption was high, and at least two results in the five principles of drug-likeness (Lipiski, Ghose, Veber, Egan, and Muegge) were “Yes.” The screened components were searched for target genes through the SwissTargetPrediction database.
Pain Target Screening
The related targets in OMIM, DisGeNET, and Gene Cards databases were collected with “pain” as the keyword. The disease-related targets obtained from the above three databases were summarized, and duplicate data were deleted to obtain disease targets.
Component-disease Intersection Target Acquisition
Venny 2.1.0 website was used to draw the Venn diagram of drug targets and soreness targets, and the same targets were screened.
Protein-protein interaction (PPI) Network was Established to Screen Core Targets
The intersection targets were input into the STRING website to establish the PPI network diagram. The confidence level was set to 0.9. Cytoscape software was used to visualize the PPI network diagram in TSV format. Then, topological analysis was performed using the “CytoNCA” plug-in, and the core targets were selected with Degree, Closeness, and Betweenness as quantitative indicators.
Gene Ontology (GO) Function and KEGG Pathway Analysis
The intersection targets were input into the DAVID website for GO function analysis and KEGG pathway analysis, and then the top results were selected according to the p value and visualized on the micro-bioinformatics platform.
Construct a “drug-component-target-disease-pathway” Network Diagram
Cytoscape software was used to construct a network diagram of “RWI-component-intersection target-pain-pathway.” Each node in the diagram represented RWI, component, intersection target, main signal pathway, and pain, and the edge represented the interaction between them. We can more intuitively observe the target of RWI analgesia and the corresponding network pathway.
Results
Observation on Writhing Behavior of Mice Induced by Acetic Acid
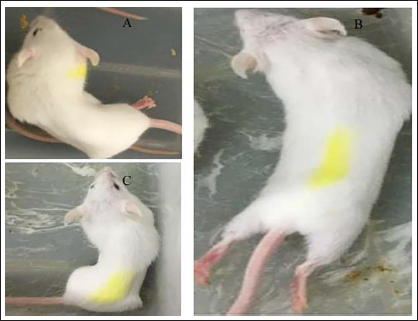
After injection of glacial acetic acid, mice showed writhing behaviors such as abdominal contraction, body distortion, and hindlimb extension, as shown in Figure 1.
Behavioral Observation of Acetic Acid-induced Writhing in Mice.
The Effect of RWI on Acetic Acid-induced Writhing Behavior in Mice
Compared with the model group, the positive group, and each dose group of RWI could obviously decrease the number of torsion in mice (p < 0.01), but the effect of the RWI administration group was not as good as that of the positive group. The writhing inhibition rate of the positive group was 50.27%, and the writhing inhibition rate of the high, medium, and low dose groups of RWI was 40.49%, 32.79%, and 28.42%, respectively, the inhibition rate of RWI increased with the increase of concentration. In addition, the positive group and each administration group could obviously prolong the writhing latency time of the mouse (p < 0.01), as shown in Table 1.
The Effect of RWI on Acetic Acid-induced Writhing Behavior in Mice.
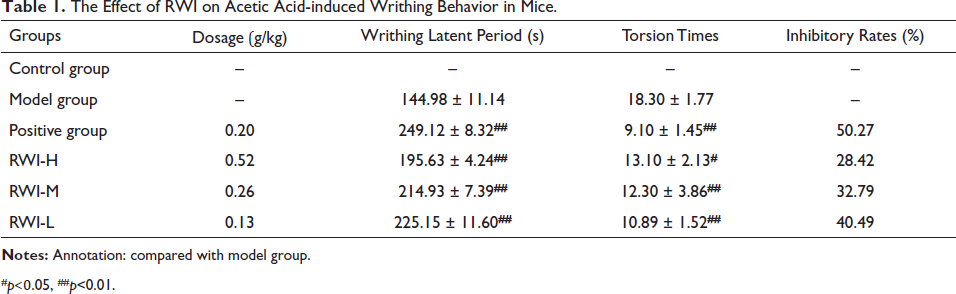
#p<0.05, ##p<0.01.
Nuclear Magnetic Data Export and Pretreatment of Serum Samples
The NMR spectrum is shown in Figure 2. The detection data table is input into MetaboAnalyst 5.0 software, and all detected ppm chemical shifts are retained. Smooth data screening conditions: the robust estimate such as interquartile range, the following parameters should be met:
1H-NMR Spectra of Serum Sample.
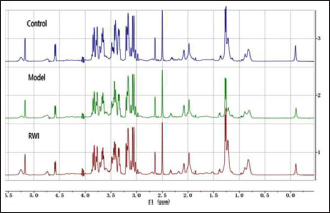
Less than 250 variables: 5% will be filtered; Between 250 and 500 variables: 10% will be filtered; Between 500 and 1,000 variables: 25% will be filtered; Over 1,000 variables: 40% will be filtered, Normalization by sum was used as the condition for sample standardization, and the data normalization condition was mean-centered and divided by the standard deviation of each variable. The data were corrected according to the above criteria, as shown in Figure 3.
Mouse Serum Before and After Correction Figures. (A) Before Correction, (B) After Calibration.
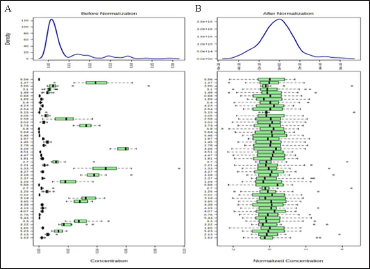
Multivariate Statistical Analysis
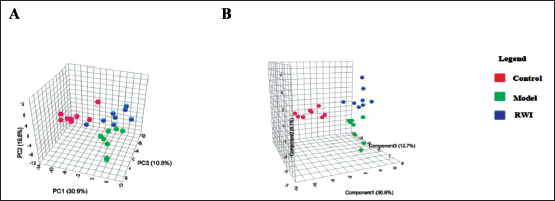
Multivariate statistical analysis was performed on the serum sample data of the control, model, and RWI groups. In the PCA score plot, the data of each group were basically separated. In the PLS-DA score plot, the data of each group were further separated, as shown in Figure 4. It shows that the metabolism of mice in each group has changed. Combined with the experimental data of acetic acid writhing, it showed that the abnormal metabolism caused by pain in mice was improved after administration. The evaluation parameters of the OPLS-DA model of the control group and the model group were R2Y: 0.947 and Q2: 0.827. The evaluation parameters of the OPLS-DA model of the model group and the RWI group were R2Y: 0.977 and Q2: 0.737. The evaluation parameters of the OPLS-DA model of the control group and the RWI group were R2Y: 0.862 and Q2: 0.608, indicating that the established model has a good interpretation rate, fitness, and predictive ability (Figure 5).
PCA and PLS-DA were used to Analyze the 3D Scores of the Model Group, Control Group, and RWI. (A) PCA Analysis Diagram, (B) PLS-DA Analysis Diagram.
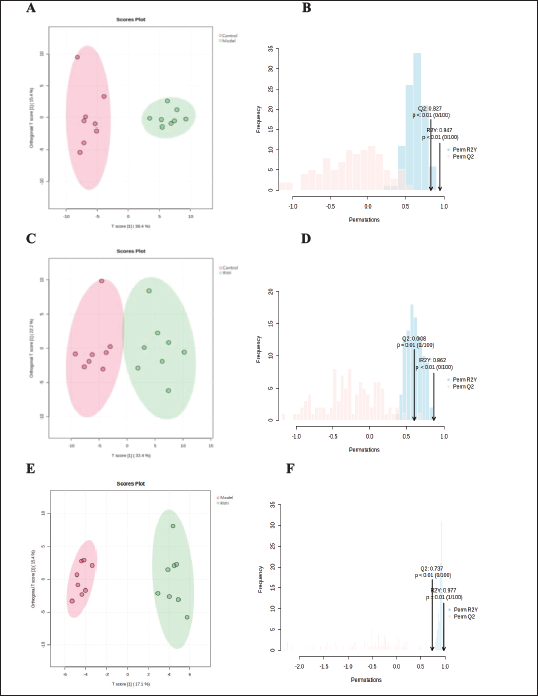
OPLS-DA Score Plot and Model Permutation. (A and B) Control versus Model (C and D) Control versus RWI (E and F) RWI versus Model.
Screening of Differential Metabolites
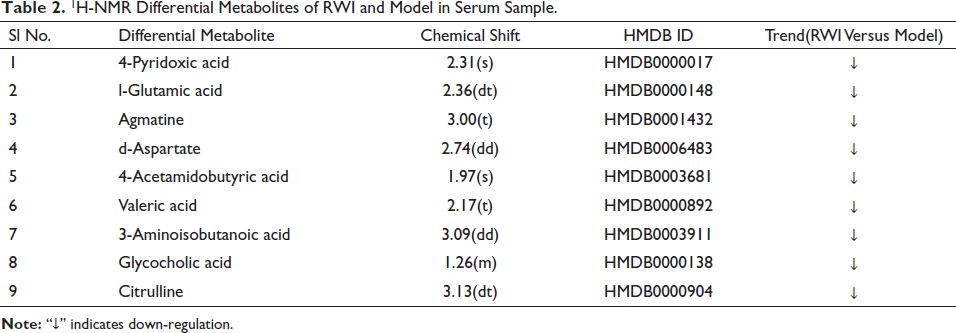
Based on the VIP value > 1, FC value ≥ 2 or ≤ 0.5, and p < 0.05 of the OPLS-DA model, nine differential metabolites between the model group and the RWI group were screened out. They were 4-pyridoxic acid,
1H-NMR Differential Metabolites of RWI and Model in Serum Sample.

Metabolic Pathway Analysis
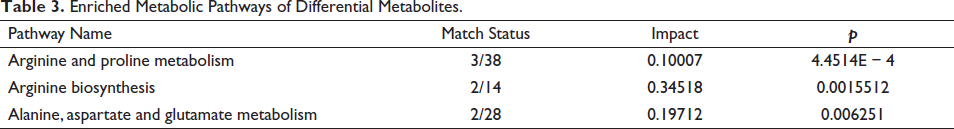
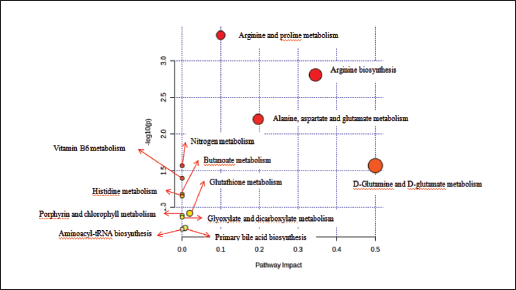
The differential metabolites were analyzed the pathways were predicted by MetaboAnalyst website, and 13 metabolic pathways were obtained. Three important metabolic pathways, Arginine and proline metabolism, Arginine biosynthesis and Alanine, aspartate, and glutamate metabolism, were screened by Impact > 0.1, p < 0.05, as shown in Table 3, Figures 6 and 7.
Enriched Metabolic Pathways of Differential Metabolites.
Significantly Changed Metabolic Pathways and Links Between Pathways.
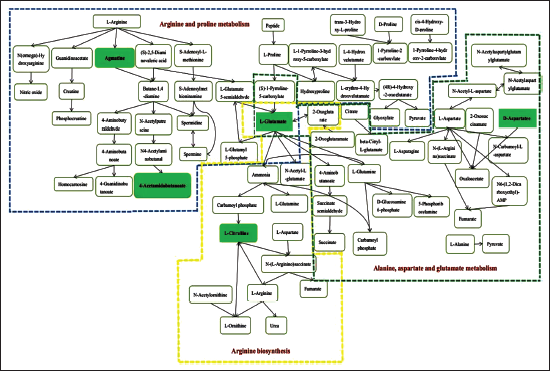
Path Diagram.
Active Components and Targets of RWI
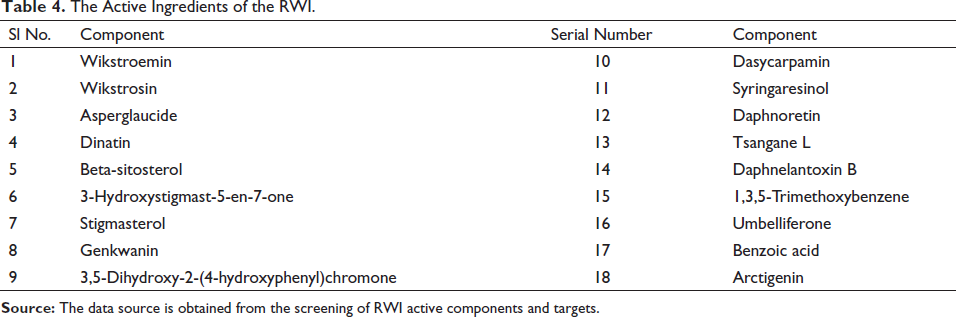
Through the TCMSP website, 10 effective active compounds of RWI were screened out with OB % ≥ 30 and DL ≥ 0.18 as the standard, and then eight active components were screened out by referring to the literature and using the five principles of drug-like, as shown in Table 4. PubChem, SwissTargetPrediction, and UniProt databases were used to search for component targets, and 639 target genes were obtained after deleting duplicate targets.
The Active Ingredients of the RWI.
Disease Target
The disease targets obtained from the Gene Cards website were screened according to the relevance score greater than the median. After screening twice, 3,555 disease targets with a score ≥ 1.783 were finally selected. Disease targets were queried in the Gene Map of OMIM database, and 16 disease targets were obtained. The DisGeNET database obtained 1,554 disease targets. The targets in the three databases were summarized, the repeated parts were deleted, and 4,294 disease targets were obtained.
Component-disease Intersection Target
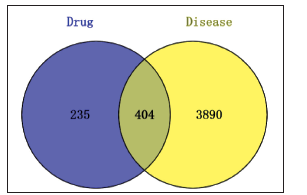
Venn diagram of components and soreness targets was made by using Venny 2.1.0 online website. It was observed that there were 404 common targets, and the results are shown in Figure 8.
Venn Diagram of Drug and Disease.
PPI Network Construction
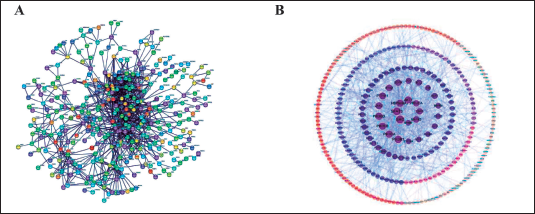
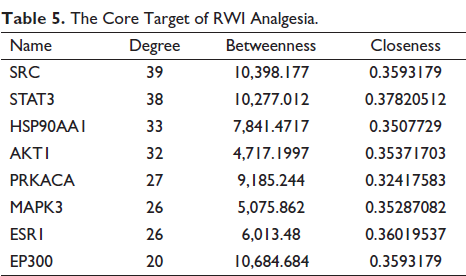
The PPI network was established by inputting the common targets of components and diseases into the STRING website, and 403 nodes and 1,005 connecting lines were obtained, in which the nodes represent proteins and the edges represent the connections between proteins, as shown in Figure 9 (A). Cytoscape software was used to visualize the PPI network. The larger the diameter of the point and the deeper the color, the more proteins interact with the node and the higher the degree value. The thicker the edge and the darker the color, the stronger the interaction between the two proteins, as shown in Figure 9 (B). By calculating the values of degree, closeness, and betweenness, the first 20 targets with the largest values were screened out, and the intersection was taken to obtain eight intersection targets, which were taken as the core targets, as shown in Table 5.
PPI Network Diagrams. (A) PPI Network of Potential Targets of RWI Analgesia, (B) The Visual PPI Network Diagram of the Potential Targets of RWI Analgesia was Established.
The Core Target of RWI Analgesia.
GO Function and KEGG Pathway Analysis
GO analysis mainly included BP, CC, and MF analysis. Through GO analysis, 1,628 results were obtained, and 1,306 important items were screened out according to p ≤ 0.05. There were 953 items related to BP, including protein phosphorylation, response to xenobiotic stimulus, and protein autophosphorylation. There were 138 items related to CC, including the plasma membrane, a receptor complex, and an integral component of the presynaptic membrane. There were 215 items related to MF, involving enzyme binding, protein tyrosine kinase activity, and identical protein binding. Draw histograms for the first 10 pre-BP analysis, CC analysis, and MF analysis (Figure 10 (A)).
GO Function and KEGG Pathway Enrichment Analysis Diagrams. (A) GO Enrichment of RWI Analgesic Target, (B) KEGG Pathway Enrichment Map.
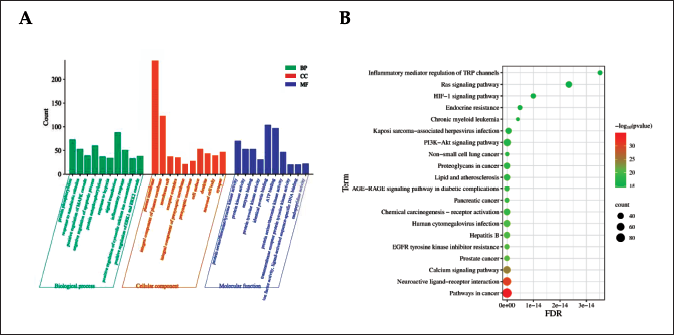
A total of 193 pathways were obtained by KEGG pathway enrichment analysis, and 182 important pathways were screened out with p ≤ 0.05, including cancer pathways, prostate cancer, and EGFR tyrosine kinase inhibitor resistance. The top 20 pathways with relatively small p values and the most enriched targets were selected to draw a bubble diagram (Figure 10 (B)). The darker the color of the point, the higher the credibility, and the larger the point, the more genes are contained.
Construction of “drug-component-target-disease-pathway” Network Diagram
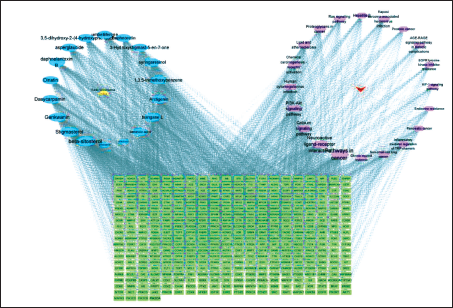
The diagram of “RWI-components-Intersection Targets-Pain-Pathway” was constructed, as shown in Figure 11. There are 444 nodes and 2,083 edges in the figure. The blue ellipse represents the RWI component, the yellow triangle represents RWI, the red arrow represents the pain, the purple hexagon represents the signal pathway, the green rectangle represents the intersection target, and the edge represents the interaction between the several. The larger the point, the more important, and the greater the analgesic effect.
“RWI-component-intersection target-pain-pathway” Network Diagram.
Network Pharmacology and Metabolomics Analysis
By combining network pharmacology with metabolomics, two common pathways were obtained, arginine and proline metabolism and nitrogen metabolism. Combined with the above research, it is shown that the metabolism of arginine and proline for the “sweat soaking method” has processed the main pathway involved in the analgesia of RWI.
Discussion
Pharmacodynamic studies have shown that the “sweat soaking method” after processing the RWI has an analgesic effect, by regulating 4-pyridoxine,
NO plays a similar role as many neurotransmitters and modulators in the central and peripheral nervous systems and can participate in the transmission and modulation of nociceptive sensory information.
Glutamate is an important excitatory neurotransmitter in the central nervous system. N-methyl-
Through network pharmacology research, 18 components of RWI were screened out, and 639 component target genes, 4,294 disease targets, and 404 intersection targets were obtained. Through PPI network construction, eight core targets of RWI analgesia were obtained, which were SRC, STAT3, HSP90AA1, AKT1, PRKACA, MAPK3, ESR1, and EP300, respectively. Among them, SRC kinase is coupled to the sites of NMDA receptor subunits. Phosphorylation of these sites will enhance the endocytosis of NMDA receptors and mediate the CS process. SRC is activated by the NMDA receptor to become an additional source of inward calcium current, which enhances NMDA receptor activity and reduces pain threshold (Childers & Baudy, 2007; Dahl & Keane, 2012; Katsura et al., 2006; Salter & Pitcher, 2012). Pain information is initially integrated from peripheral receptors to the dorsal horn of the spinal cord, and is transmitted to the higher center of the brain to form a conscious pain. When the peripheral nerve is injured, the microglia and astrocytes in the spinal dorsal horn are sequentially activated. Once the microglia are activated, a number of proinflammatory cytokines such as IL-6 will be released. Under the action of cytokines such as IL-6, the phosphorylation level of JAK is increased, and STAT is further activated. Phosphorylated STAT molecules affect the expression of related molecules. The phosphorylation of STAT3 is closely related to the activation and proliferation of astrocytes. Down-regulation of STAT3 phosphorylation in astrocytes can significantly alleviate the occurrence of pain (Todd, 2010; Wang et al., 2015;).
Through GO analysis, it was found that the target gene can play an analgesic role by affecting molecular processes such as protein phosphorylation, response to xenobiotic stimulus, cell composition such as receptor complex and membrane raft, and molecular functions such as enzyme binding.
Through KEGG, it was found that gene enrichment-related pathways included EGFR, AGE-RAGE, PI3K-Akt, and some cancer-related pathways. The imbalance of the EGFR-mediated signaling pathway is a key factor in pain. Therefore, regulating EGFR overexpression and blocking the transmission of pathways can relieve pain. The activation of the AGE-RAGE pathway can cause the release of many inflammatory mediators and the increase of oxygen-free radical production, which affects the function of the central nervous system, and aggravates cognitive dysfunction. Inflammatory cytokines are recognized as important biological molecules that promote the occurrence and development of pain. Therefore, RWI can achieve an analgesic effect by affecting the AGE-RAGE signaling pathway (Chhipa et al., 2019). The PI3K/Akt pathway can regulate the growth and survival of neuronal cells. Akt activation can activate its downstream important target protein mTOR and apoptosis-related proteins to regulate the PI3K/Akt/mTOR pathway, which is related to the regulation of the analgesic effect of neuropathic pain. Therefore, pain can be alleviated by regulating the PI3K/Akt pathway (Jiang & Shao, 2022; Liu & Li, 2020; Zhang, 2018).
Conclusion
In this study, it was found that RWI had an analgesic effect after the “sweat soaking method” processing. Metabolomics studies have shown that the analgesic mechanism of RWI is related to arginine and proline metabolism and arginine biosynthesis. At the same time, network pharmacology studies have shown that RWI can act on SRC, STAT3, HSP90AA1, AKT1, and other targets, and participate in cancer pathways and calcium signaling pathways to achieve the purpose of analgesia. The combination of the two methods shows that RWI plays an analgesic role mainly through arginine and proline metabolism.
Abbreviations
GO: Gene ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes; OPLS-DA: Orthogonal partial least squares discriminant analysis; PCA: Principal component analysis; PPI: Protein-protein interaction; RWI: Radix Wikstroemia indica.
Footnotes
Acknowledgments
The author expresses her gratitude to the College of Pharmacy of Guizhou University of Traditional Chinese Medicine for the experimental technology and site.
Author’s Contribution
Caiyao Han and Guo Feng are involved in the processing of raw medicinal materials and the extraction of alcohol extracts, animal experiments, data processing, and article writing. Chenchen Ren and Wei Li participated in the experimental design and data collection. Wen Liu and Gang Liu guided the experiment. Xueli Song, Kexin Ma, and Yan Lei collated the data. Zhengyan He, Tingting Liu, Jinxin Hou, and Ju Zhang assisted the experiment.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
Participants in this experiment received informed consent and ethical approval.
Funding
This work was supported by the National Natural Sciences Foundation of China (No. 81760766, 82060767), the Guizhou Province Science and Technology Foundation Project (Grant no. Guizhou Scientific Basis ZK[2022] General 461), the Guiyang College of TCM Doctor Startup Fund Project (Grant no. Guizhongyi Doctor Fund [2017] 1), the National training program for innovative backbone talents for TCM (Grant no. Zjjh [2019] 128), the “Thousand” level Innovative Talents Project in Guizhou Province (Grant no. Qrlf [2020] 4), and the Fund project of Guizhou administration of traditional Chinese Medicine (Grant no. QZYY-2020-083).
