Abstract
Background
Hyperlipidemia is a common precursor to cardiovascular diseases, necessitating effective intervention. This study explores the potential of a bioactive fraction derived from Kalanchoe pinnata leaf, a less-explored natural source, for its antihyperlipidemic properties.
Objectives
The primary objectives of this research are to isolate and assess the bioactive fraction from Kalanchoe pinnata leaves extract for its antihyperlipidemic attributes using a murine model. The study also aims to characterize the fraction and investigate its interactions with hyperlipidemia-related proteins.
Materials and Methods
Kalanchoe pinnata leaves were extracted and fractionated via flash chromatography. Isolated fractions underwent phytochemical, antioxidant, and enzyme inhibition assays in vitro. In vivo, selected fractions were tested in Triton WR-1339-induced hyperlipidemic mice. Fraction F2, which exhibits the most potent activity, was characterized using HPTLC MS/MSn chromatography. Molecular docking studies explored interactions between F2 compounds and hyperlipidemia-related proteins.
Results
Fraction F2 displayed significant in vivo antihyperlipidemic activity, with observable morphological changes in mice. HPTLC MS/MSn identified four major compounds in F2, while docking studies revealed potential interactions with hyperlipidemia-related proteins.
Conclusion
This study underscores F2’s potential as a potent antihyperlipidemic agent sourced from Kalanchoe pinnata leaves. The identified compounds hold promise for further research in hyperlipidemia therapy development, emphasizing the therapeutic potential of natural Kalanchoe pinnata compounds for managing hyperlipidemia and related cardiovascular conditions.
Introduction
Hyperlipidemia, a term that denotes elevated levels of lipids in the bloodstream, has emerged as a significant global health concern. This condition is characterized by the excessive presence of cholesterol and triglycerides, two vital lipid components, in the blood. Hyperlipidemia has garnered worldwide attention due to its high prevalence and its well-established association with several life-threatening diseases, most notably cardiovascular disorders (Benjamin, 2022; Kader et al., 2023; Singh & Nain, 2023).
An epidemiological panorama reveals its ubiquity, transcending geographical, cultural, and socioeconomic boundaries. The prevalence of hyperlipidemia varies across regions and is influenced by dietary patterns, lifestyle choices, genetic predispositions, and other environmental factors. This condition finds its roots in the perturbations of lipid metabolism, which can manifest as either primary (genetic) or secondary (acquired) forms.
Hyperlipidemia’s notoriety primarily stems from its well-established link to cardiovascular diseases (CVDs). Elevated levels of low-density lipoprotein cholesterol (LDL-C), often termed “bad cholesterol,” contribute significantly to the development of atherosclerosis—atherosclerotic plaques in arteries that can lead to coronary artery disease, myocardial infarction (heart attack), and stroke. Additionally, hyperlipidemia can engender the formation of lipid deposits in other vascular beds, posing risks to peripheral arteries and potentially leading to conditions like peripheral artery disease (Mainieri et al., 2023).
Beyond cardiovascular concerns, hyperlipidemia is also implicated in pancreatitis, a painful and sometimes life-threatening inflammation of the pancreas. High levels of triglycerides, another lipid constituent, are a common culprit in this context. Furthermore, the metabolic syndrome, characterized by a cluster of cardiovascular risk factors including obesity, high blood pressure, and insulin resistance, is closely entwined with hyperlipidemia (Hou et al., 2014; Kiseleva et al., 2000).
The pharmacological armamentarium for managing hyperlipidemia is vast and multifaceted. Statins, a class of drugs that inhibit the enzyme HMG-CoA reductase, represent the cornerstone of lipid-lowering therapy. These drugs effectively reduce LDL-C levels, leading to a decreased risk of cardiovascular events. However, they are not devoid of side effects, with potential concerns ranging from muscle pain (myalgia) to hepatotoxicity, necessitating vigilant monitoring.
Fibrates, another class of lipid-lowering agents, primarily target elevated triglyceride levels. Despite their efficacy, fibrate use can be associated with gastrointestinal disturbances and a heightened risk of gallstones. Bile acid sequestrants, yet another group of drugs, aid in reducing LDL-C levels by binding to bile acids in the intestine, preventing their reabsorption. However, their palatability is often a hurdle for patient adherence, and they may lead to constipation and malabsorption of fat-soluble vitamins.
Natural products, derived from plant sources, have been a focal point of research in the quest for alternative treatments for hyperlipidemia. These substances often contain bioactive compounds with lipid-lowering properties, making them attractive candidates for therapeutic exploration (David et al., 2015; Singh & Pattnaik, 2023; Zhang et al., 2018).
“Kalanchoe pinnata,” a traditional remedy with a long history of use in certain cultures, has gained prominence as a potential natural product for hyperlipidemia management. Derived from specific plant sources, it offers the prospect of lipid reduction without the adverse effects associated with conventional medicines. Preliminary studies suggest that “Kalanchoe pinnata” may contain compounds that modulate lipid metabolism, potentially lowering both LDL-C and triglyceride levels (El Abdellaoui et al., 2010).
Our study represents a pioneering endeavor to comprehensively investigate the efficacy of “Kalanchoe pinnata” in the treatment of hyperlipidemia. Employing a rigorous scientific approach, we aim to elucidate the mechanisms through which this natural product exerts its lipid-lowering effects. We will conduct randomized, controlled clinical trials, meticulously monitoring lipid profiles and assessing the safety profile of “Kalanchoe pinnata.”
This groundbreaking research seeks to bridge the gap between traditional remedies and evidence-based medicine, potentially offering a safe, effective, and accessible alternative for hyperlipidemia management. The profound implications of our study may reshape the landscape of lipid-lowering therapy, addressing not only the global prevalence of hyperlipidemia but also the associated burden of cardiovascular diseases and other related conditions.
Our hypothesis posits that “Kalanchoe pinnata,” hitherto unexplored in the context of hyperlipidemia, possesses lipid-modulating properties that can significantly lower LDL-C and triglyceride levels. We anticipate that this natural product may operate through distinct mechanisms, distinct from conventional medications, minimizing side effects while providing efficacious lipid control.
Through rigorous scientific investigation, we endeavor to validate this hypothesis and contribute invaluable insights into the realm of natural products as a novel approach to hyperlipidemia management. The outcomes of our study could herald a transformative shift in the paradigm of hyperlipidemia therapy, offering new hope for countless individuals grappling with this global health challenge.
In conclusion, this study represents a logical continuation of our previous research, which encompassed comprehensive in vitro assays, including phytochemical assessment (total phenolic content, total flavonoid content, and total steroidal content), in vitro antioxidant studies (DPPH assay, H2O2 assay, NO2 assay, and ABTS assay), and in vitro enzyme inhibition assays (alpha amylase, alpha glycosidase, and pancreatic lipase). Notably, our earlier investigations identified the most potent fraction, F2, characterized by the presence of key compounds such as sitosterol, quercetin, stigmasterol, and gallic acid (Singh et al., 2023).
In the current study, we aim to elucidate the potential relationship between the isolated F2 fraction and hyperlipidemia, a pressing global health issue. Our objective is to further delineate the lipid-lowering properties of F2, thereby providing valuable insights into the therapeutic potential of natural products in the management of hyperlipidemia. This research continuity aligns with our commitment to advancing scientific knowledge and addressing critical health challenges through rigorous empirical exploration.
Materials and Methods
Collection, Authentication, and Isolation of Bioactive Guided Fractions
It is imperative to acknowledge that the essential steps encompassing the collection, authentication, and isolation of bioactive guided fractions have been meticulously conducted and previously reported in our comprehensive research article. In our prior publication, we have elucidated in detail the methodologies employed in the collection of plant materials from their natural habitats, the rigorous authentication process to ensure species specificity, and the systematic isolation of bioactive guided fractions through various extraction and purification techniques (Singh et al., 2023).
In vitro Phytochemical Assessment, Antioxidant Assay, and Enzyme Inhibition Assay
In our previously published work, we judiciously explicated the methodologies deployed for in vitro phytochemical assessment, encompassing the precise quantification of total phenolic content, total flavonoid content, and total steroidal content. Furthermore, we conducted a rigorous inquiry into the antioxidant potential of our bioactive fractions through a suite of sophisticated in vitro assays, including DPPH assay, H2O2 assay, NO2 assay, and ABTS assay. Additionally, our investigative ambit extended to in vitro enzyme inhibition assays, wherein we meticulously evaluated the inhibitory effects on alpha-amylase, alpha-glycosidase, and pancreatic lipase activities (Singh et al., 2023).
These foundational assays, underscored by their methodical execution and their outcomes, have been scrupulously documented, ensuring utmost transparency and scientific rigor in our research framework. In the present study, we undertake a seamless progression from these well-established methodologies to scrutinize the potential implications of our isolated fraction in the context of hyperlipidemia. This scientific continuum underscores our unwavering commitment to advancing the frontiers of scientific knowledge and proactively addressing pivotal health challenges through an empirically grounded approach.
In vivo Investigation
Animal Procurement
In the executed experimental series, the chosen animal model consisted of eight-week-old Swiss albino mice. These mice were sourced from the Animal House facility within the Department of Pharmaceutical Sciences and Technology at Birla Institute of Technology, Mesra. The procurement and utilization of these animals were carried out in compliance with IAEC approval number 1972/PH/BIT/108/20/IAEC.
The mice were group-housed, with each enclosure accommodating six individuals. Their housing environment was meticulously regulated, maintaining a 14-hour light and 10-hour dark cycle, with precise control over temperature (23 ± 3°C) and humidity parameters (60 ± 10 %). Moreover, the mice were provided ad libitum access to standard rodent pellet diets and water, ensuring their consistent exposure to standardized environmental conditions throughout the experimentation (Elmowafy et al., 2022).
Experimental Framework and Triton Induction in Animals
Male mice, with a mean weight approximating 20 ± 2 grams, underwent a five-day acclimatization phase during which they were granted unrestricted access to both water and standard rodent chow. Subsequently, these animals were categorized into five distinct groups (normal treated group, standard treated group, Triton-induced group, fraction F1 treated group, and fraction F2 treated group), each comprising six subjects, and were assigned individualized identification codes systematically. The administered dosages of the test compounds (F1 and F2) and standard (Fenofibrate) were accurately standardized to 80 and 13 mg/kg body weight, respectively. These substances were expertly formulated into an oral suspension employing CMC-Na as the vehicle. This clarification has been added to the manuscript for precision and is highlighted for easy reference, achieving a final concentration of approximately 0.5%, which was administered via the oral route in a saline vehicle over one week.
On the sixth day of this regimen, excluding the control group, an intravenous parenteral administration of an aqueous Triton WR-1339 solution, dosed at 400 mg/kg, was precisely delivered through the tail vein. After 24-hour interval, blood specimens were judiciously retro-orbitally collected and subsequently employed for the comprehensive assessment of various lipid parameters. On the terminal day of the experimental protocol, the animals were ethically euthanized in strict accordance with IAEC protocol to facilitate subsequent in-depth histopathological analyses (Elmowafy et al., 2022; Lee et al., 2006).
HPTLC MS MSn Analysis
It is pertinent to acknowledge that the comprehensive characterization of compounds through high-performance thin-layer chromatography mass spectrometry (HPTLC MS/MSn) for the most potent fraction, denoted as F2, has been diligently conducted and meticulously detailed in our previously published research article (Singh et al., 2023).
In silico Molecular Docking and Simulation Study
In pursuit of a comprehensive understanding of the interactions between the identified compound from our HPTLC MS/MSn analysis and specific protein targets, an in silico molecular docking and simulation study was conducted. The proteins selected for this study were PDB IDs 3QNT, 3MWD, and 1B09, each chosen for its relevance to our research objectives and downloaded from the RCSB protein bank (Chu et al., 2010; Pandit et al., 2000; Pepys et al., 2006).
The structural data of these proteins were retrieved from the Protein Data Bank (PDB), providing a sound basis for our computational investigations. The identified compound, derived from our HPTLC MS/MSn analysis, was subjected to molecular docking simulations with these protein structures. The purpose of this study was to explore and predict the potential binding modes, affinities, and interactions between our compound of interest and the selected protein targets (Chandran et al., 2022). This computational approach, which integrates molecular docking and simulation techniques, allowed us to gain valuable insights into the molecular-level interactions that may underlie the compound’s efficacy in the context of hyperlipidemia. The results obtained from these in silico studies contribute significantly to the elucidation of the compound’s pharmacological mechanisms and its potential as a therapeutic agent for hyperlipidemia (Cheng et al., 1996; Jorgensen et al., 1996).
ADME Study
Schrodinger’s QikProp module was utilized to ascertain the ADME features. When comparing the specific molecule’s attributes with those of 95% of the currently marketed drugs, QikProp delineated boundaries. The utilization of QikProp aids in averting false positives in high-throughput screening assays, given its recognition of 30 distinct reactive functional groups. It evaluates the potential of analogs based on Lipinski’s rule of five, an imperative aspect for ensuring a drug-like pharmacokinetic profile through rational drug design (Pires et al., 2015).
Results and Discussion
Extraction and Isolation of Bioactive Guided Fractions
It is imperative to note that in our earlier study, as documented by Singh et al. (2023), a meticulous process of extraction and isolation was conducted, resulting in the isolation of a total of four potent bioactive fractions. The detailed procedures and outcomes of this extraction and isolation process have been comprehensively reported in the study by Singh et al. (2023).
These four bioactive fractions, each characterized by their unique composition and properties, hold significant promise and form the foundation of our continued research endeavors. Building upon the knowledge garnered from their isolation, we aim to delve deeper into the therapeutic potential and mechanisms of action of these fractions, particularly in the context of hyperlipidemia. This research continuity underscores our commitment to advancing scientific knowledge and addressing critical health challenges through an evidence-based approach.
In vitro Phytochemical Assessment, Antioxidant Assay, and Enzyme Inhibition Assay
It is essential to acknowledge that in our prior research, comprehensive in vitro phytochemical assessment, antioxidant assay, and enzyme inhibition assay were diligently conducted. The detailed methodologies and results of these critical analyses have been exhaustively documented in our earlier study.
In vivo Assay
Impact of Bioactive Fraction F2 on Biochemical Indices
Based on the above findings, it was observed that fractions F1 and F2 exhibited the most potent activity in overall in vitro studies. Consequently, these fractions were selected for further in vivo studies. Upon a comprehensive assessment of biochemical indices, it becomes evident that all experimental cohorts subjected to therapeutic intervention exhibited substantial reductions in crucial lipid parameters, notably encompassing total cholesterol, triglycerides, low-density lipoprotein (LDL), and very-low-density lipoprotein (VLDL) levels. This reduction in lipid levels is in stark contrast to that in the Triton-induced control groups, which experienced elevated lipid levels.
Notably, fraction F2 consistently emerged as the most efficacious among the experimental groups, manifesting robust activity relative to the untreated control cohort. These compelling and consistent findings collectively underscore the substantial potential of fraction F2 as a promising agent for mitigating hyperlipidemia and related metabolic disorders. As such, these results warrant further extensive investigations and clinical scrutiny to ascertain its therapeutic efficacy in the clinical context as shown in Table 1.
Effect of Bioactive Fractions (F1 and F2) on Biochemical Parameters.
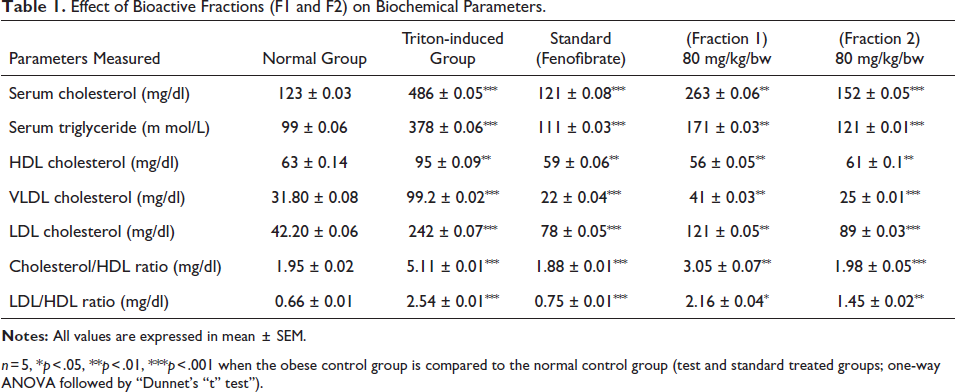
n = 5, *p < 0.05, **p < 0.01, ***p < 0.001 when the obese control group is compared to the normal control group (test and standard treated groups; one-way ANOVA followed by “Dunnet’s “t” test”).
Histopathological Analysis
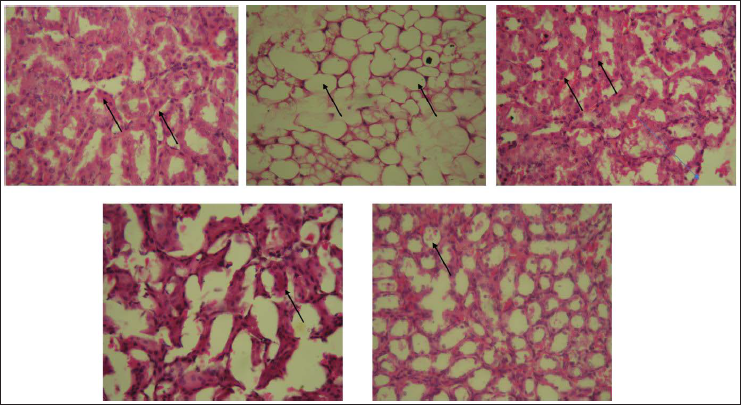
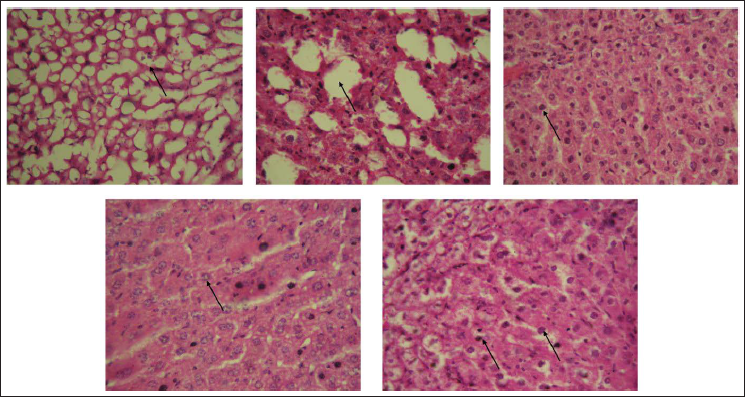
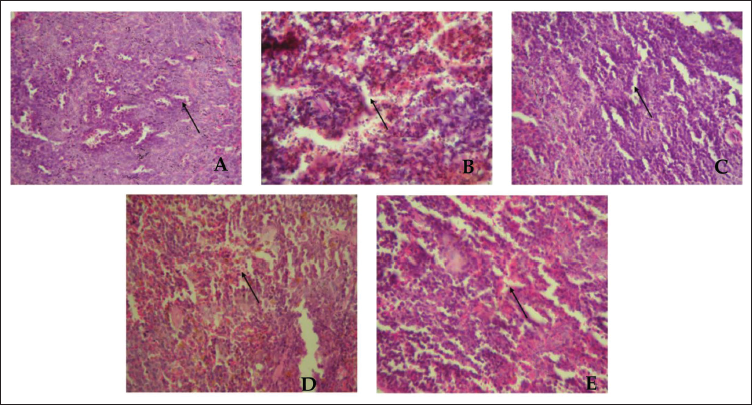
Histopathological analysis of Triton WR-1339-induced mouse models revealed that the Triton-induced group exhibited significant hepatocyte ballooning, extensive steatosis, and inflammation in liver tissue, as shown in Figure 1, along with damage in kidney, pancreas, and artery tissues, as shown in Figures 2–4. In contrast, the fraction F2 treated group showed marked reductions in hepatocyte ballooning and lipid accumulation in liver tissue compared to the Triton-induced group, and it also demonstrated significant improvements in kidney, pancreas, and arterial tissues. These results suggest that fraction F2 had a particularly significant and promising therapeutic effect in mitigating hyperlipidemia-induced damage across multiple organs, warranting further investigation.
Kidney’s Histology of Mice Treated with Triton WR-1339-induced Hyperlipidemia and Treated Groups. Representative Figures Were Stained with H × E with the Original Magnification of 40×. Arrows Indicate the Nephron Integrity. (A) Normal Control Group. (B) Triton WR-1339-induced Group. (C) Standard (Orlistat) Treated Group. (D) Fraction F1 Treated Group. (E) Fraction F2 Treated Group.
Liver’s Histology of Mice Treated with Triton WR-1339-induced Hyperlipidemia and Treated Groups. Representative Figures Were Stained with H × E with the Original Magnification of 40×. Arrows Indicate the Hepatic Steatosis. (A) Normal Control Group. (B) Triton WR-1339-induced Group. (C) Standard (Orlistat) Treated Group. (D) Fraction F1 Treated Group. (E) Fraction F2 Treated Group.
Pancreas Histology of Mice Treated with Triton WR-1339-induced Hyperlipidemia and Treated Groups. Representative Figures Were Stained with H × E with the Original Magnification of 40×. Arrows Indicate the Islet’s Integrity. (A) Normal Control Group. (B) Triton WR-1339-induced Group. (C) Standard (Orlistat) Treated Group. (D) Fraction F1 Treated Group. (E) Fraction F2 Treated Group.
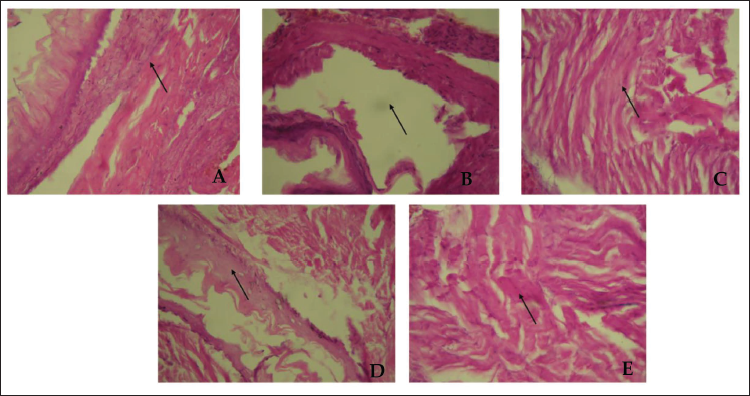
Artery’s Histology of Mice Treated with Triton WR-1339-induced Hyperlipidemia and Treated Groups. Representative Figures Were Stained with H × E with the Original Magnification of 40×. The Arrow Indicates the Endothelial Health and Arterial Wall Thickness. (A) Normal Control Group. (B) Triton WR-1339-induced Group. (C) Standard (Orlistat) Treated Group. (D) Fraction F1 Treated Group. (E) Fraction F2 Treated Group.
HPTLC MS/MSn Analysis
Under the meticulous scrutiny of HPTLC MS/MSn analysis, our investigation has unveiled the presence of four major compounds within the sample. These compounds, identified as sitosterol, stigmasterol, thiamine, and quercetin, have been unequivocally identified and their structural characteristics delineated. The compounds were identified via cross-referencing the obtained spectral data with Chemspider. It is noteworthy that these findings have been extensively detailed and documented in our previously published research article, thus serving as a foundational reference point for our ongoing research endeavors.
Molecular Docking Study
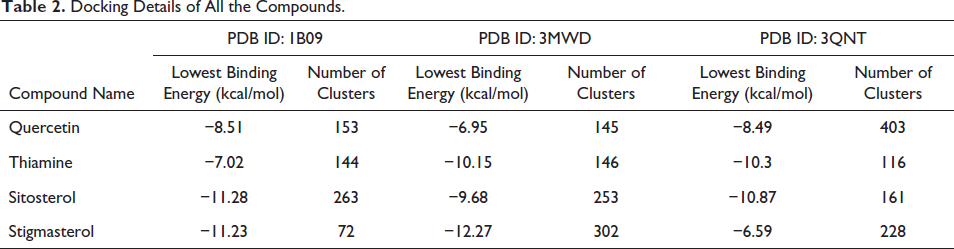
The docking simulation was performed with a total of 500 runs using the GA algorithm. Following the completion of the docking process, the identified compounds (quercetin, thiamine, sitosterol, and stigmasterol) were screened based on the lowest binding energy exhibited by the conformer with the largest cluster. An analysis of the resulting log file (.dlg file) revealed that sitosterol displayed the most favorable docking score when interacting with proteins human C-reactive protein complex (1B09) and NPC1L1 (3QNT). On the other hand, stigmasterol exhibited the best docking score when interacting with truncated human ATP-citrate lyase (protein 3MWD). To obtain the coordinates for the best conformer of the ligands, we employed an in-house utility Python script called “write_largest_cluster_ligand.py,” which is available within MGLTool 1.5.7. Subsequently, protein–ligand complexes were generated and subjected to molecular dynamics (MD) simulation, as shown in Table 2.
Docking Details of All the Compounds.
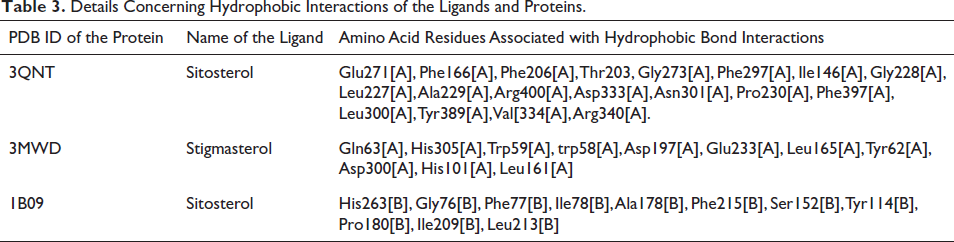
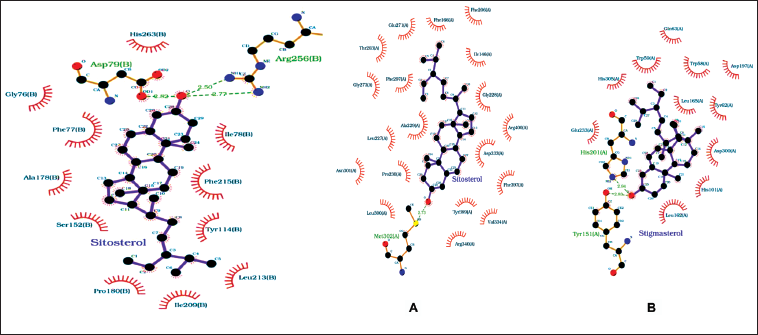
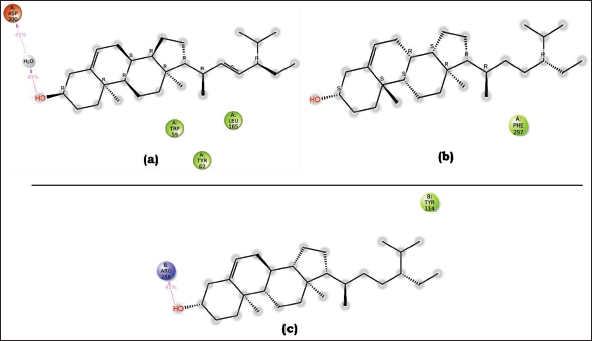
The interaction diagrams depicting the binding between 2D protein and ligands were generated and examined using LigPlot+ v.2.2.8. The ligands, namely, stigmasterol and sitosterol, exhibited favorable hydrogen bonding and hydrophobic interactions with their respective proteins across three complexes. In the case of human C-reactive protein complexes (PDB ID: 1B09), the hydroxyl group (-OH) present in sitosterol formed a single hydrogen bond with the sulfur atom of Met302[A], with a bond length of 2.73 Å. As for ATP-citrate lyase (PDB ID: 3MWD), the hydroxyl (-OH) group of stigmasterol participated in hydrogen bonding with the nitrogen atom in the imidazole ring of His201[A] and with the oxygen atom attached to the benzene ring of Tyr151[A], exhibiting bond lengths of 2.94 Å and 2.83 Å, respectively. In NPC1L1 (PDB ID: 3QNT), the hydroxyl group of sitosterol engaged in two hydrogen bonding interactions with two amino groups present in Arg256[B], displaying bond lengths of 2.50 and 2.77 Å. Additionally, the hydroxyl group formed another hydrogen bond with one of the hydroxyl groups of Asp79[B], with a bond length of 2.82 Å. The ligands also established multiple hydrophobic interactions with their corresponding proteins, as shown in Table 3 and Figure 5.
Details Concerning Hydrophobic Interactions of the Ligands and Proteins.

Interaction of the Best-docked Ligand (A) Sitosterol with Protein 3QNT, (B) Stigmasterol with 3MWD, and (C) Sitosterol with 1B09 Proteins.
Molecular Dynamics Simulation Analysis
Molecular dynamics simulation studies were carried out for the three PLCs:
Complex a: Stigmasterol—3MWD PLC Complex b: Sitosterol—1B09 PLC Complex c: Sitosterol—3QNT PLC
RMSD Analysis
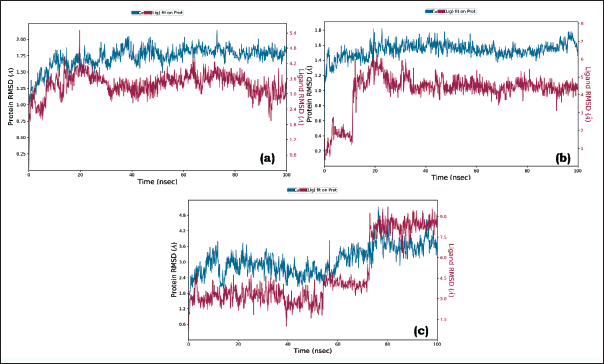
RMSD analysis of the protein–ligand complex reveals whether a simulation has achieved equilibrium. RMSD of the protein in the presence of ligand should converge at a constant value of 1–3 Å. However, for small biomolecules, minor fluctuations in this range are perfectly acceptable. Moreover, if the protein/ligand undergoes a large conformation change, there might be deviations in RMSD values, but it should converge at a constant value for a particular simulation. Herein for complex a, the conformation revealed significant RMSD values in the range 1.0–1.8 Å, which displays the stability of the protein (backbone and c-alpha). Similarly, for complex b, the RMSD values were in the range of 1.2–1.6 Å. For complex c, RMSD converged at a value of 2.4–3.6 Å. Though there were fluctuations in all the PLCs, these minor deviations are acceptable. This observation indicates that the protein, in the presence of our external ligand, maintained its conformation without significant changes. Consequently, it effectively accommodated the ligand within the active site. The ligand RMSD metric is a measure for assessing the stability of the ligand within the protein and its binding pocket. In the simulation of complex a, the RMSD values initially ranged from 1.2 to 1.2 Å (0–50 ns). Following this, there were minor fluctuations, after which RMSD converged to a constant value of 3.0–3.6 Å (50–100 ns). For complex b, the ligand was stable throughout the simulation with an RMSD value of 4–5 Å (15–100 ns). There were minor fluctuations with a slight deviation in RMSD from 0 to 15 ns. For complex c, the ligand was stable throughout the simulation with an RMSD value of 3 Å (0–55 ns), 3.5 Å (55–75), and 8.5–9 Å (75–100). Therefore, all our PLCs have demonstrated effectiveness in binding to the active site and interacting with the critical amino acid residues involved. Overall, all the PLCs were found to be stable. There were slight deviations for complexes b and c though all the confirmations were maintained constantly throughout the simulation time.
Because possible effective inhibitors should be able to bind strongly to the enzyme and create stable non-dynamic complexes, the dynamicity property provides a valid criterion to evaluate the efficiency of a proposed inhibitor. These findings are consistent with the images in Figures 6 A, B, and C, where the RMSD values can be visually observed.
Root Mean Square Deviation (RMSD) Graph of (A) Stigmasterol-3MWD PLC, (B) Sitosterol-3QNT PLC, and (C) Sitosterol-1B09 PLC.
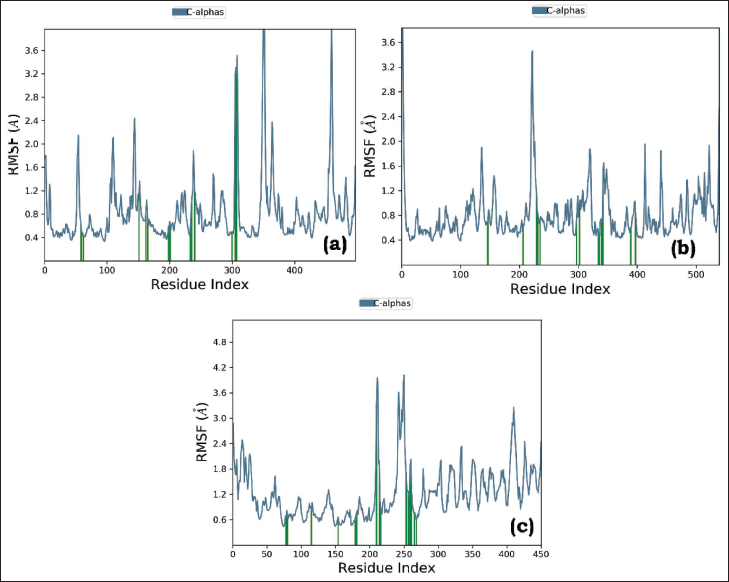
RMSF stands for root mean square fluctuation, a measure used in molecular simulations to assess the flexibility and dynamics of atoms or residues within a biomolecular system. It quantifies the average deviation or fluctuation of the positions of atoms or residues from their mean positions throughout the simulation. The RMSF values are typically represented as a plot or graph to visualize the regions of the molecule that experience significant fluctuations during the simulation. A higher RMSF value indicates greater flexibility or movement, while a lower RMSF value suggests relative stability or restricted motion. Figure 7 A, B, and C displays the regions of the protein that experience the most significant fluctuations during the simulation. The peaks on the plot generally correspond to the tails (N- and C-terminals) of the protein, which tends to fluctuate more than other parts of the protein. The alpha helices and beta strands, which are secondary structural elements, are usually more rigid than the unstructured regions of the protein and therefore fluctuate less compared to the loop regions. Herein, there were minor scale fluctuations throughout the course of the simulation in all PLCs. The protein residues that interact with the ligand are marked with green-colored vertical bars, and they signify the ligand contacts.
Root Mean Square Fluctuation (RMSF) Graph of (A) Stigmasterol-3MWD PLC, (B) Sitosterol-3QNT PLC, and (C) Sitosterol-1B09 PLC.
Throughout the simulation, the protein’s interactions with the ligand can be monitored and analyzed. These interactions can be classified into different types, summarized, and depicted in the plot provided in Figure 8 A, B, and C. The protein–ligand contacts are categorized into four main types: hydrogen bonds, hydrophobic interactions, ionic interactions, and water bridges. Hydrogen bonds play a vital role in maintaining the secondary structures of proteins. These bonds contribute to the stability of protein folding, aiding in the maintenance of alpha helices, beta sheets, and other secondary structural elements. Additionally, the presence of hydrogen bond donors and acceptors is a crucial factor in passive diffusion across cell membranes. This process is of utmost importance in the absorption and distribution of drugs within the body, highlighting the significance of hydrogen bonds in pharmacokinetics. As per the graph presented in Figure 8, a value of 0.4 suggests that for 40% of the simulation time, the specific interaction is maintained. However, values over 1.0 are possible as some protein residues may make multiple contacts of the same subtype with the ligand.
Plot (Stacked Bar Charts) of Protein Interactions with the Ligand Supervised Throughout Molecular Dynamics Simulation of (A) Stigmasterol-3MWD, (B) Sitosterol-3QNT PLC, and (C) Sitosterol-1B09 PLC (green: H-bonds, lavender: hydrophobic, pink: ionic, and blue: water bridges).
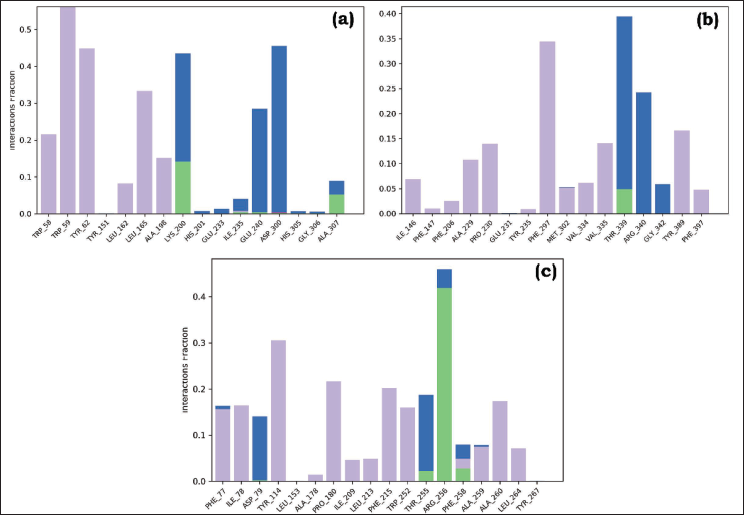
As illustrated in Figures 8 A, B, and C, all PLCs maintained hydrogen bond interactions. The number of H-bonds established with the amino acid residues of the protein was: complex a (Lys200: 15% and Ala307), complex b (Thr339: 5%), and complex c (Arg256: 45%). The simulation demonstrated more hydrophobic contacts, ionic interactions, and water-mediated linkages in all PLCs. These suggest that all the PLCs remained stable throughout the simulation, and the system’s backbone fluctuations were modest.
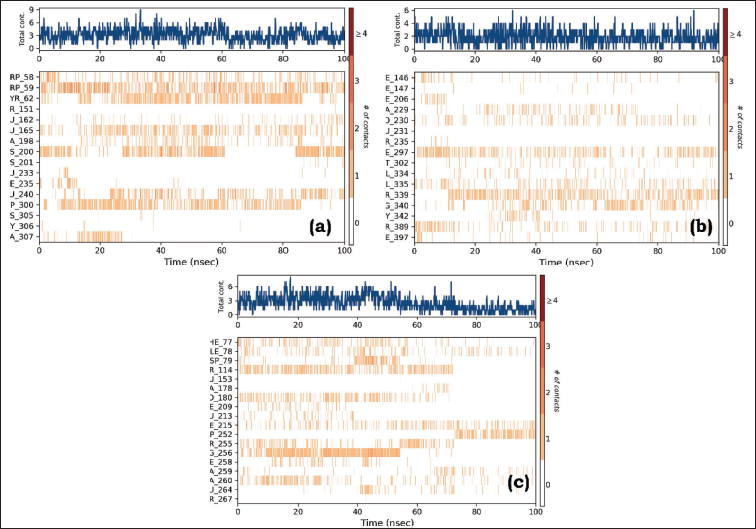
Protein–ligand contacts are an essential aspect of simulating molecular interactions between proteins and ligands as they represent their mode of interaction with each other. The timeline representation illustrates the interactions and contacts (H-bonds, hydrophobic, ionic, and water bridges) summarized in Figure 9. The top panel overviews the total number of specific contacts formed between the protein and the ligand throughout the trajectory. In the bottom panel, each trajectory frame indicates the residues that interact with the ligand. Residues that establish multiple specific contacts with the ligand are depicted in a darker shade of orange, as indicated by the color scale on the right side of the plot. Herein, it is evident that all PLCs maintained contact with the proteins. This signifies that complex b is more stable than complex a.
Specific Contacts Made by the Protein with the (A) Stigmasterol-3MWD PLC, (B) Sitosterol-3QNT PLC, and (C) Sitosterol-1B09 PLC Throughout the Trajectory. (Dark color indicates more specific contact with the ligand.)
Detailed ligand interaction analysis in molecular dynamics simulations provides essential information for understanding binding mechanisms, guiding rational drug design, predicting binding free energies, and gaining mechanistic insights into ligand–target interactions. Herein, the selected trajectory (0.00 through 100.00 ns) and the interactions that persist for more than 30.0% of the simulation time have been highlighted. It is worth noting that it is possible to observe interactions exceeding 100% because certain residues may have multiple interactions of a single type with the same ligand atom, leading to cumulative interaction times beyond the duration of the simulation. The detailed interactions observed are shown in Figure 10.
A Schematic Representation of Detailed Ligand Atom Interactions (A) Stigmasterol-3MWD PLC, (B) Sitosterol-3QNT PLC, and (C) Sitosterol-1B09 PLC with the Protein Residues.
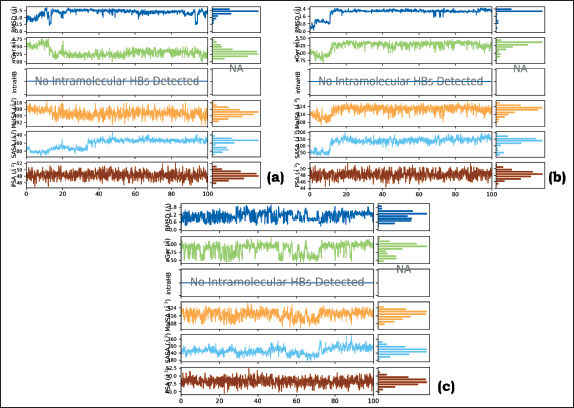
Several ligand properties are of utmost importance in the context of drug design and molecular interactions. These properties include ligand RMSD, radius of gyration (rGyr), intramolecular hydrogen bonds (intraHB), molecular surface area (MolSA), solvent accessible surface area (SASA), and polar surface area (PSA). A glimpse of these properties is displayed in Figure 11.
Representation of the Ligand Properties (a) Stigmasterol-3MWD PLC, (b) Sitosterol-3QNT PLC, and (c) Sitosterol-1B09 PLC.
Post-simulation MM-GBSA Analysis
The calculated binding free energies (∇G) for the PLCs are −78.30 ± 5.85, −82.50 ± 6.66, and −62.20 ± 5.35 kcal/mol. The results indicate favorable binding, with the second PLC demonstrating the highest affinity at −82.50 kcal/mol.
ADME Result
Using the QikProp tool of the Schrodinger software, we examined 10 physical descriptors and pharmaceutically significant features of some natural moieties. Major descriptors provided here are necessary for predicting the drug-like properties of compounds. Four compounds were screened, and among them, two exhibited identical docking scores. Consequently, two chemicals were selected based on this outcome. According to Lipinski’s rule of five, each structure had drug-like qualities and significant values for the properties’ analysis. The first three characteristics are based on Lipinski’s rule of five and include solubility (QPlogS) > 7; partition coefficient between octanol and water (logPo/w), which ranges from −2 to −6.5; and molecular weight (mol.MW), which is 500. Blood–brain barrier passage was shown by the brain/blood partition coefficient (QplogBB) measure, which is necessary for inhibition. The apparent MDCK cell permeability in nm/s was predicted by the QPPMDCK. The blood–brain barrier is thought to be well mimicked by MDCK cells, and the higher the value of QPPMDCK, the more permeable the cell is. All the proposed compounds displayed acceptable ADME properties, as shown in Table 4.
ADME Analysis of the Top Docked Compounds.
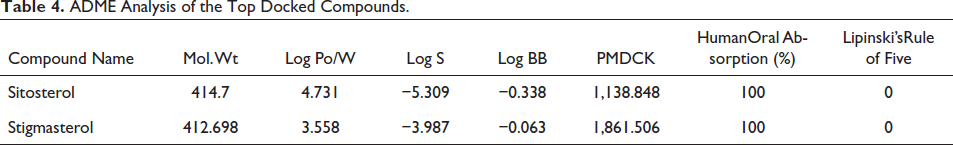
Conclusion
In sum, the rigorous process of extraction and isolation, as expounded upon in the antecedent study, has culminated in the isolation of four potent bioactive fractions from Kalanchoe pinnata leaves extract. These bioactive fractions, each endowed with singular compositions and attributes, proffer substantial potential for subsequent scientific inquiry into the realm of hyperlipidemia and its associated metabolic perturbations.
The in vitro phytochemical assessment, antioxidant assay, and enzyme inhibition assay have bestowed invaluable insights into the bioactive potential intrinsic to these fractions. Notably, fraction F2 has consistently evinced remarkable efficacy in abating pivotal lipid parameters, encompassing total cholesterol, triglycerides, LDL, and VLDL levels, in Triton-induced hyperlipidemic murine models. These observations underscore the substantive potential of fraction F2 as a propitious therapeutic moiety for the amelioration of hyperlipidemia.
Histopathological scrutiny corroborates the therapeutic promise of fraction F2, as it has exhibited noteworthy ameliorations in hepatocyte ballooning, lipid accrual, and tissue damage across multiple organs, encompassing the hepatic, renal, pancreatic, and arterial domains. HPTLC MS/MSn analysis has divulged the existence of four principal compounds within the sample matrix: sitosterol, stigmasterol, thiamine, and quercetin. Molecular docking investigations have elucidated that sitosterol and stigmasterol have engendered favorable binding interactions with specific proteins implicated in hyperlipidemia.
Molecular dynamics simulations have imparted insights into the stability and interactions manifesting within the protein–ligand complexes, substantiating the sustained constancy of these assemblages. The RMSD, RMSF, and protein–ligand contact analyses harmonize with the stability of these complexes, auguring their potential as efficacious inhibitory agents.
Post-simulation MM-GBSA analysis has divulged a propitious binding free energy for the protein–ligand complexes; further ADME analysis has adduced that the preeminent docked compounds, sitosterol and stigmasterol, evince attributes befitting pharmacological agents, encompassing solubility, partition coefficient, and molecular weight, thus positioning them as auspicious candidates for subsequent endeavors in drug development.
In synthesis, the compendious scientific findings at hand illuminate the auspicious potential of these bioactive fractions, notably fraction F2, as propitious candidates for the genesis of therapeutic agents, en route to the rectification of hyperlipidemia and its allied metabolic aberrations. Subsequent clinical investigations and validation are required to ascertain their clinical efficacy and safety. This evidence-driven modus operandi underscores our steadfast dedication to the advancement of scientific acumen and the concomitant resolution of pressing health exigencies.
Abbreviations
MS: Mass spectrometry; b.w: Body weight; ADME: Absorption, Distribution, Metabolism, and Excretion; CMC-Na: Carboxymethyl Cellulose Sodium; MM-GBSA: Molecular Mechanics/Generalized Born Surface Area; LDL: Low-density Lipoprotein; HDL: High-Density Lipoprotein; VLDL: Very Low-density Lipoprotein.
Footnotes
Acknowledgments
The authors are thankful to the Department of Pharmaceutical Sciences and Technology and Central Instrument Facility of Birla Institute of Technology, Mesra, Ranchi, for providing all the necessary chemicals, equipment, and analytical tools for the study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The animals used in the experiment were approved by institutional animal ethical committee of Birla Institute of Technology, Mesra, Ranchi, with the approval number of 1972/PH/BIT/108/20/IAEC.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
