Abstract
Background and Objectives
4-Methylumbelliferone (4-MU) is a coumarin compound that can be extracted from the medicinal plant with anti-cancer properties, Smilax china L. In recent years, studies have revealed its potential as an anti-tumor and anti-metastasis drug with promising effects in cancer treatment. Despite an increase in research on the metabolic patterns of tumor cells, no prior research has suggested that 4-MU inhibits tumor proliferation by blocking glycolysis. This thesis presents evidence that 4-MU binds to proteins involved in glycolysis, thus mediating its anti-tumor effects.
Materials and Methods
Network pharmacology, transcriptomics, and molecular docking were utilized to forecast the potential targets and probable pathways of 4-MU’s anti-cancer activity, and the affinity of 4-MU towards potential targets was discovered using microscale thermophoresis (MST) detection.
Results
The results of transcriptome analysis brought to light that the genes with differential expressions were primarily enriched in metabolic pathways, including glycolysis-related proteins. Using network pharmacology and molecular docking, our study identified Heat Shock Protein 90 Alpha Family Class A Member 1 (Hsp90AA1), mitochondria, phosphoglycerate kinase 2 (PGK2), glycerol-3-phosphate dehydrogenase (GPD2), and glucose-6-phosphate isomerase (GPI) as potential targets of 4-MU. The strong binding affinity between 4-MU and these proteins was confirmed by the MST assay.
Conclusion
The findings indicate that 4-MU can hinder glycolysis by binding to glycolysis-associated proteins such as Hsp90AA1, PGK2, GPD2, and GPI. This results in the prevention of the energy supply to the tumor tissue, which ultimately curbs tumor growth, thereby demonstrating its anti-tumor properties. These results conclude that 4-MU has the capacity to be a novel glycolysis inhibitor for cancer treatment. Moreover, the identification of these glycolysis-associated proteins as possible targets for cancer therapy offers new avenues for research in the field of cancer treatment, thus providing further valuable evidence for the anti-cancer mechanism of 4-MU.
Keywords
Introduction
Studies have confirmed that lactate is generally produced under hypoxic conditions, but even in the presence of sufficient oxygen, tumor cells will prefer to metabolize glucose rather than lactate to provide energy for themselves. This mode of metabolism is considered a sign of malignancy, and this process is known as aerobic glycolysis or the Warburg effect (Schwartz et al., 2017; Vaupel & Multhoff, 2021). Metabolically active tumor cells develop a tumor microenvironment rich in nutrient-depleting, hypoxic, acidic, and toxic metabolites that suppress the immune system. Glycolysis is accomplished in the cytoplasm through a sequence of enzyme processes. The abundant presence of enzyme correlations during glycolysis is a direct factor leading to tumor cell proliferation. Among them, Heat Shock Protein 90 Alpha Family Class A Member 1 (Hsp90AA1), phosphoglycerate kinase 2 (PGK2), glycerol-3-phosphate dehydrogenase (GPD2), and glucose-6-phosphate isomerase (GPI) are involved in promoting glycolysis and tumor cell proliferation. These enzymes may be potential biomarkers and targets for cancer diagnosis and treatment (Kathagen-Buhmann et al., 2016; Xu et al., 2018). Therefore, a growing amount of research is being conducted on the metabolic patterns of tumor cells. It has become one of the most effective anti-tumor methods to suppress glycolysis-related enzyme expression and down-regulate the capacity of glycolysis. Because the effect of inhibiting tumor cell proliferation was achieved.
4-Methylumbelliferone (4-MU) is a coumarin that can be isolated in significant quantities from the medicinal plant Smilax china L. with anti-cancer properties. In recent decades, studies have shown that 4-MU can also be used as an anti-tumor and anti-metastasis drug and has good effects on breast cancer (Hiraga et al., 2013; Karalis et al., 2019), hepatocellular carcinoma (Piccioni et al., 2015), lung cancer (Essa et al., 2022), and ovarian cancer (Vitale et al., 2021). Most studies have shown that 4-MU inhibits in vitro tumorigenicity and in vivo bone metastatic lesions by inhibiting the content of hyaluronic acid and glucose and the expression of matrix metalloproteinase-9 in the tumor microenvironment (Hiraga et al., 2013; Karalis et al., 2019; Nakamura et al., 2002). However, there is no report that 4-MU inhibits tumor proliferation by inhibiting glycolysis. The purpose of this study was to look into the impact of 4-MU on tumor cell proliferation by suppressing glycolysis-related enzyme expression and down-regulating the capacity of glycolysis.
Transcriptome is a discipline that studies gene transcription and transcriptome regulation at the overall level of tissues or cells. It is an RNA-based technology to study gene expression as a whole. The transcriptome is the sum of the total RNA that can be transcribed by a new cell. Otherwise, it is an important means of studying cell phenotype and function. The transcription process, in which DNA is used as a template to synthesize RNA, is the first step in gene expression and a key link in the regulation of gene expression. The transcriptome, unlike the genome, has temporal and spatial restrictions. Transcriptome studies look at the expression of all genes in a cell or tissue, thereby studying the changes that occur in a cell-living tissue under limited conditions.
Molecular docking technology is a drug design method based on the characteristics of receptors and the interaction between receptors and drug molecules (Galati et al., 2021). The interaction of small-molecule ligands with protein receptors is studied using theoretical simulations to predict binding patterns and affinities.
Microscale thermophoresis (MST) detects changes in the hydration layer, charge, and molecular size as well as the movement of molecules in a microscopic temperature gradient field. Structural/conformational changes in biomolecules make the hydration layer change; next, they can make trace changes in heat flow, which can be measured; and last, affinity can be determined (Entzian & Schubert, 2016; Seidel et al., 2013). To be gratified, small changes, such as protein phosphorylation or small molecules binding to targets, can be evaluated.
In this research, network pharmacology was used to preliminarily screen potential anti-tumor targets of 4-MU (Wang et al., 2022; Yue et al., 2017), and molecular docking technology was used to analyze the binding ability of 4-MU to potential targets. The MST assay was used to verify the affinity of 4-MU with proteins, find and identify potential protein targets, and provide reliable data support and new direction for the anti-tumor mechanism research of 4-MU.
Materials and Methods
Reagents and Proteins
4-MU (CAS: 90-33-5) was obtained from Shanghai Yuanye Bio-Technology Co., Ltd. (China), with a concentration of ≥98%. Hsp90AA1 (ab78425), PGK2 (ab123166), GPD2 (ab132692), and GPI (ab116799) were purchased from Abcam (Cambridge, UK).
Extract Total RNA from Cells
HepG2 cells purchased from the Cell Bank of the Chinese Academy of Sciences were cultivated with HepG2 cell-specific medium in a cell incubator containing 5% CO2 at 37°C (Prokel Life Science and Technology Co., Ltd.). HepG2 cells that are growing well were seeded in 6-well plates. After 24 h, by removing the original medium, add the original medium (control group) and drug-containing medium containing 50 µM 4-Mu (experimental group) into the 6-well plates.
After 24 h of culture, total RNA was extracted from both groups in accordance with the directions of the TruSeq™ RNA Sample Preparation Kit (San Diego, CA, USA). Total RNA concentration and purity were measured with NanoDrop 2000, and RNA integrity was checked. To build the sequencing library, 5 g of high-quality RNA samples (OD 260/280 = 1.8−2.2) were employed.
Transcriptome Sequencing
The mRNA was obtained by pairing magnetic beads containing Oligo (dT) with the ployA tail of the eukaryotic mRNA3′-terminal based on the A-T base. The mRNA was randomly cleaved into 300-dp fragments by the addition of fragmentation buffer, and single-stranded cDNA was synthesized by the addition of reverse transcriptase. As a result, the ultimate, stable double-stranded structure is formed.
To create the final library, the purified products from the ligation products, which were purified and fractionated, were submitted to PCR amplification.
Sequencing was performed on an Illumina HiSeq Xten/NovaSeq6000 platform.
Analyzing Differential Expression and Enriching Functional
In RNA-Seq analysis, gene abundance can be calculated by asking about the read count of clean read sequences located in genomic regions, which reflects the expression level of the gene. Higher abundance indicates higher gene expression levels. In this study, to distinguish differences in transcript and gene expression between different samples, RSEM was used to objectively examine the transcript and gene expression levels of distinct samples.
Through differential gene expression analysis, differentially expressed genes were identified by screening several genes, and then the differentially expressed genes’ specific functions were further studied.
Screening of Intersection Targets
Through the PubChem database (
The GeneCards database (
GO Function and KEGG Pathway Enrichment Analysis
The intersection targets obtained above were introduced into the OmicShare analysis platform, and the “Gene ontology (GO) enrichment analysis” and “Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis” projects were selected to conduct GO function and KEGG signaling pathway enrichment analysis on potential targets, with “Ensemble 104 or 51” and “Homo sapiens” as versions and species.
The p-values were arranged in increasing order, and then the top 20 signaling pathways were selected as critical pathways for analysis.
KEGG pathway enrichment analysis can show the importance of different signaling pathways in protein-protein relationship networks and is often employed to analyze drug action mechanisms in network pharmacology (Kanehisa et al., 2016). GO functional enrichment analysis is currently the most representative tool in the description of biological systems (Chen et al., 2017). The DAVID database enables annotation, visualization, and integration of cross-target GO function and KEGG signaling pathway gene discovery (Huang et al., 2007).
Based on the findings of the annotation of KEGG signaling pathways analysis, the pathways with more intersecting targets can be considered important regulatory pathways of 4-MU anti-cancer pathways. The intersection of 4-MU and tumors can be considered potential anti-tumor targets of 4-MU.
Protein–Protein Interaction
Protein–protein interaction (PPI) is essential for most biological functions and processes. To our knowledge, most proteins appear to initiate their functions by interacting with one another. PPI connection analysis aids in the study of disease molecular mechanisms from a systemic perspective and the discovery of new therapeutic targets.
Establishing a PPI network involves importing the cross-target genes into the STRING database (
Network Construction
Cytoscape 3.8.2 software was used to construct the “4-MU/Target/Pathway” network (Appleton et al., 2017). 4-MU, target genes, and pathways are shown by nodes in the network schematic representation, and edges represent the association between two nodes. The level of association between nodes was analyzed according to node scores.
Molecular Docking
The PDB database (
Protein Fluorescent Labeling
Protein labeling buffer was dissolved (or replaced) with samples of Hsp90AA1, PGK2, GPD2, and GPI. Each 100 to 500 µL of protein solution was mixed with 50 µg of fluorescent dye DyLight®488, and the reaction was carried out in the dark at room temperature for 1 h. At the end of the reaction, the mixture was purified using a Zeba rotary desalting column to get rid of the unreacted dye and added to phosphate-buffered saline (PBS) to substitute for the labeled protein. The samples were kept at 4°C after the protein concentration was determined.
Binding of the Labeled Target Protein with Compound Sample
PBS buffer with 10% dimethyl sulfoxide (DMSO) was used to reduce 4-MU to an ultimate concentration of 20 mM.
Hsp90AA1 fluorescent protein was diluted in PBS buffer containing 10% DMSO and 2% Tween 80. The solution concentration was then adjusted to achieve a fluorescence intensity of 400−1500 Fnorm. Sixteen PCR tubes, numbered 1−16, were used to add 20 µL of compound solution to tube 1 and 10 µL of PBS buffer containing 10% DMSO to tubes 2−16. After mixing with the liquid in tube 2, 10 µL of liquid from the complex solution in tube 1 was withdrawn and added to tube 3 for proportional dilution. After diluting the solution in every tube, 10 µL of liquid was aspirated and then discarded, resulting in a total amount of liquid of 10 µL in 1−16 tubes.
Each of the 1−16 tubes received a 10 L fluorescently labeled protein solution. After stirring well for 5 min, the system was equilibrated at room temperature. The volume of liquid in the 1−16 tubes remained at 20 µL. Fluorescently labeled proteins (PGK2, GPD2, and GPI) were handled in the same way.
MST Estimation with Fluorescence Scanning
The mixed solution in the PCR tube was aspirated into a standard capillary using the capillary and placed into the MST Monolith N.115 for fluorescence scanning. Using the capillarity, the combined solution in the PCR tube was sucked into a standard capillary and then scanned for fluorescence by placing it in the MST Monolith N.115.
Various samples were prepared under various assay settings, including test temperature, pH, and LED intensity for fluorescence scanning and MST measurement, and then the combination of 4-MU with Hsp90AA1, PGK2, GPD2, and GPI was detected.
Statistical Analysis
The Kolmogorov–Smirnov test was used to determine normality. Two-tailed comparisons between the two groups, the Student’s t-test, or a nonparametric test were utilized. The χ2 test was used to examine the AF induction rate. Statistical significance was defined as a p-value < 0.05. Means and the standard error of the mean (SEM) are presented for regularly distributed data, whereas medians and the interquartile range (IQR) are presented for non-normally distributed data. GraphPad Prism 8 was used for all statistical analyses.
Results
Differential Gene Analysis of RNA-Seq Transcriptome
Genes with statistically significant differences in expression (Log2FC(H_O/H) ≥ 2, p < 0.05) were screened by DEG-seq software, and the volcano plot and scatter plot of differentially expressed genes are shown in Figure 1A and B (Wu et al., 2020). Based on the differentially expressed genes, KEGG pathway and GO function enrichment analyses were done. KEGG pathways were mainly enriched in glycolysis/gluconeogenesis, central carbon metabolism of cancer, fructose and mannose metabolism, carbon fixation of photosynthetic organisms, and other metabolism—and energy-related pathways. GO function was mainly enriched in the processes of glucose catabolism, ethanol metabolism, nucleotide phosphorylation, and acetyl-coA biosynthesis, as shown in Figure 1C–E. The findings revealed that the pathways strongly affected by 4-MU were mostly related to glycolysis, which could be one of 4-MU’s anti-tumor strategies.
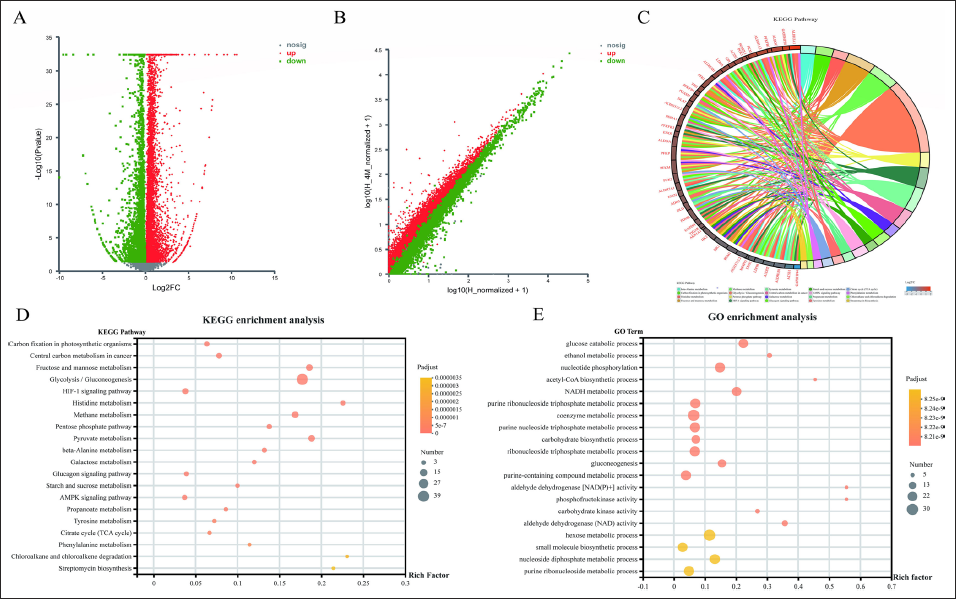
GO Function and KEGG Pathway Enrichment Analysis
A total of 111 target genes were obtained from 4-MU, and 810 cancer-related target genes were screened by setting a threshold score >10.00. Information on these 32 intersection targets is listed in Table 1.
Information on 32 Intersection Targets.
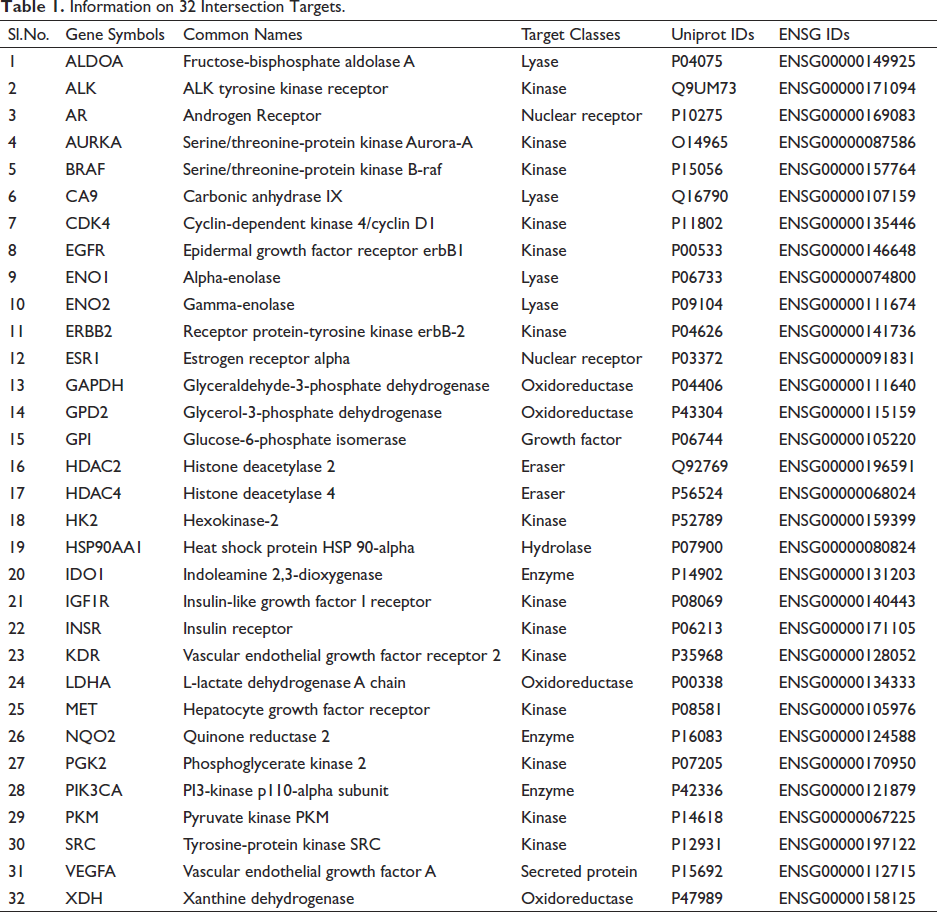
Following the import of the cross-target genes into the DAVID database, 61 pathways were obtained, and 57 potential pathways were screened (p < 0.05). Figure 2A depicts the top 10 anti-tumor mechanisms of 4-MU. Some genes are involved in the HIF-1 signaling pathway, such as Hsp90AA1, PGK2, GAPDH, and so on. Glycolysis/gluconeogenesis involves some genes, such as GPI, PGK2, GPD2, and so on. Proteoglycans in cancer involve some genes, such as GPI, PGK2, GPD2, Hsp90AA1, and so on. The cancer pathway involves some genes, such as Hsp90AA1, GPI, PGK2, GPD2, and so on. GO enrichment analysis yielded 244 GO entries, which were filtered according to the set value (p < 0.05) and ranked from small to large by p-value. The top 10 entries for biological process (BP), cellular component (CC), and molecular function (MF) were summarized, respectively, as shown in Figure 2. BP (Figure 2B) enrichment analysis mainly involved the glycolytic process, canonical glycolysis, positively regulated protein kinase B signaling pathway, and transmembrane receptor protein tyrosine kinase signaling pathway. CC (Figure 2C) enrichment analysis mainly involved receptor complexes, cytoplasmic, membrane, and cellular exosomes. MF (Figure 2D) enrichment analysis mainly involved protein tyrosine kinase activity, transmembrane receptor protein tyrosine kinase activity, protein serine/threonine/tyrosine kinase activity, and identical protein binding.
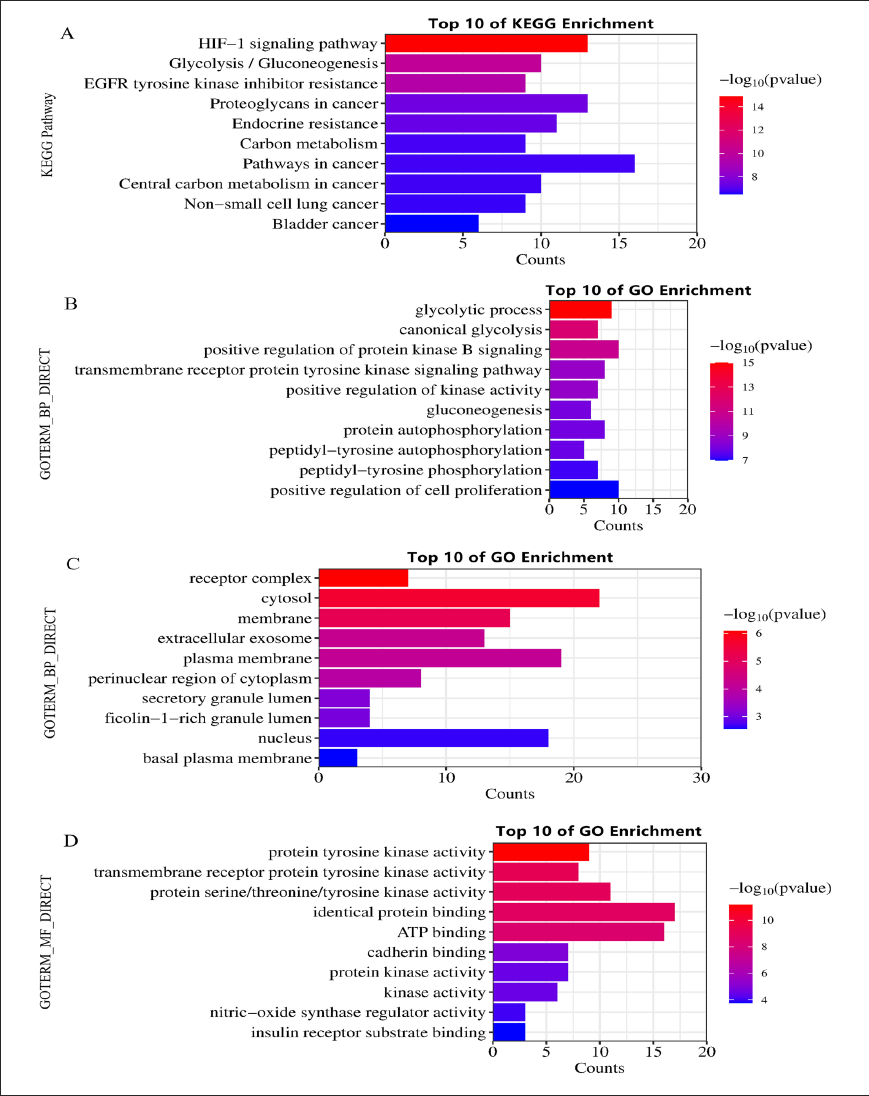
Network
The cross-target genes were integrated through the STRING database to generate an interaction network, which was analyzed using Cytoscape 3.8.2 software, as shown in Figure 3 and Table 2. Clearly, these proteins occupy central positions in the PPI network, such as Hsp90AA1, GPI, Receptor protein-tyrosine kinase erbB-2 (ERBB2), epidermal growth factor receptor (EGFR), insulin-like growth factor I receptor (IGF1R), SRC, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and PGK2. The results show that these proteins interact more closely.
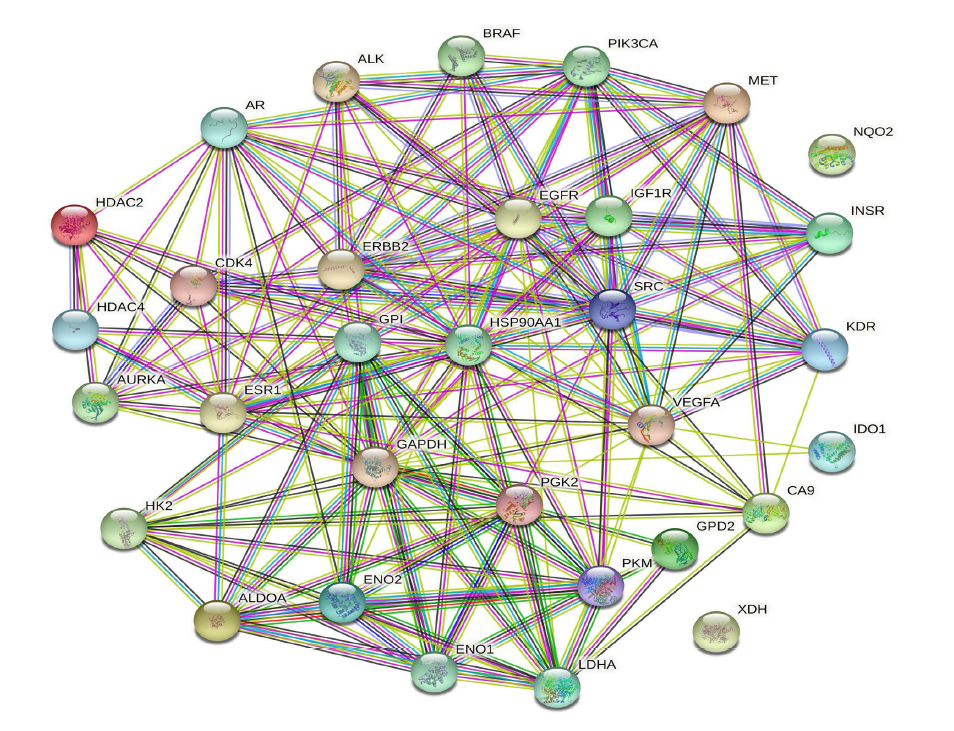
Degree of PPI and Network.
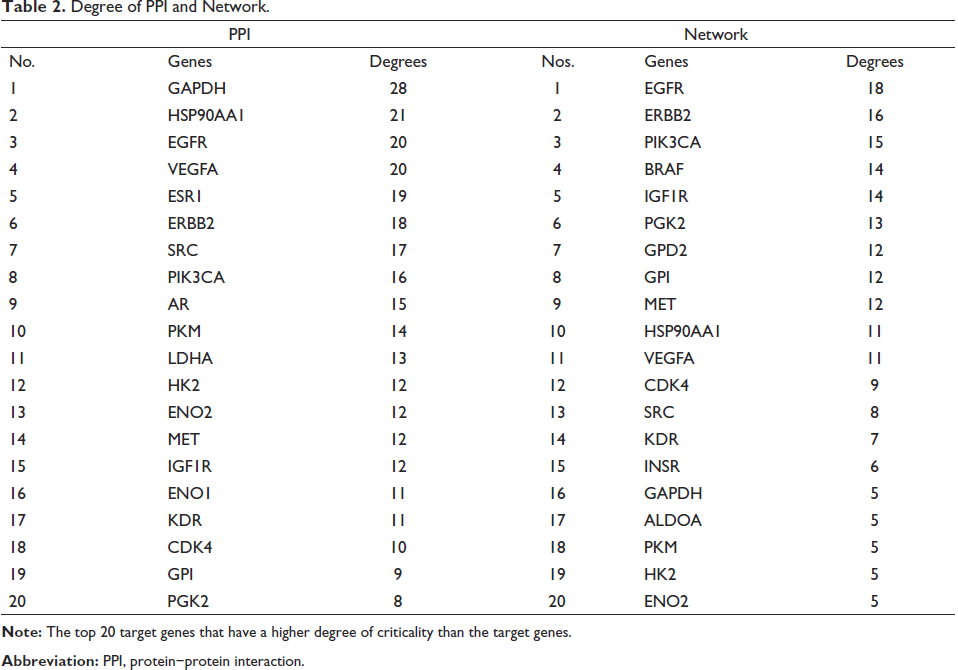
The 32 target genes, the top 20 signaling pathways, and 4-MU were imported into Cytoscape 3.8.2 software to construct the “4-MU/target/pathway” network, as shown in Figure 4. In the network diagram, blue nodes represent 4-MU, red nodes represent target genes, and yellow nodes represent pathways. The network contains a total of 53 nodes, including 32 target nodes, 20 path nodes, and 1 composite node, for a total of 230 edges. The network shows the interaction between nodes through edges, and the higher the correlation, the more concentrated the convergence of these edges, and the higher the degree score of the nodes. According to the degree of criticality of the target genes, the top 20 target genes are listed in Table 2 in the supplementary files.
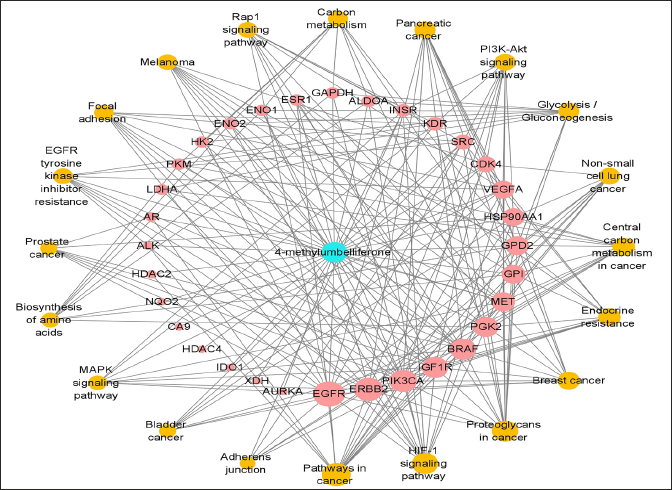
According to the results of PPI and network analysis, 10 proteins with strong interaction ability and a high interaction degree were selected. They were GPI (2C6C), GPD2 (6E8Y), PGK2 (2PAA), Hsp90AA1 (5NJX), BRAF (4MNF), PIK3CA (7R9V), EGFR (6DUK), MET (6I04), and IGF1R, respectively (5E1S), and ERBB2 (3WLW). This network indicates that 4-MU acts on multiple targets through multiple pathways to exert anti-tumor properties.
Molecular Docking
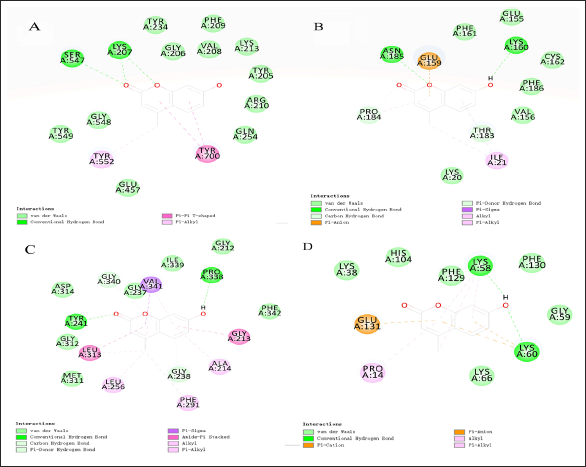
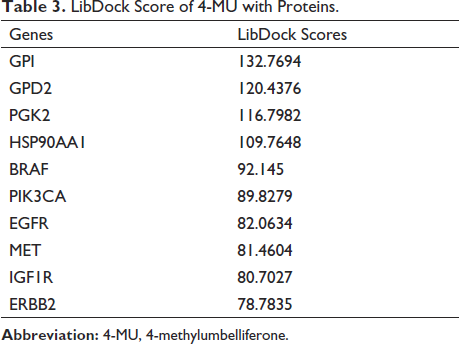
It is considered that the lower the binding energy between ligand and receptor, the higher the LibDock score, the higher the probability of interaction, and the stronger the potential activity of the compound. Ten target genes and 4-MU were uploaded to the Discovery Studio 2019 client for analysis. LibDock score ≥90 was used to analyze the docking results, and the results showed that Hsp90AA1, PGK2, GPD2, and GPI had high docking scores with 4-MU and strong binding activity. The docking results are shown in Figure 5 and Table 3.
LibDock Score of 4-MU with Proteins.
SD-test
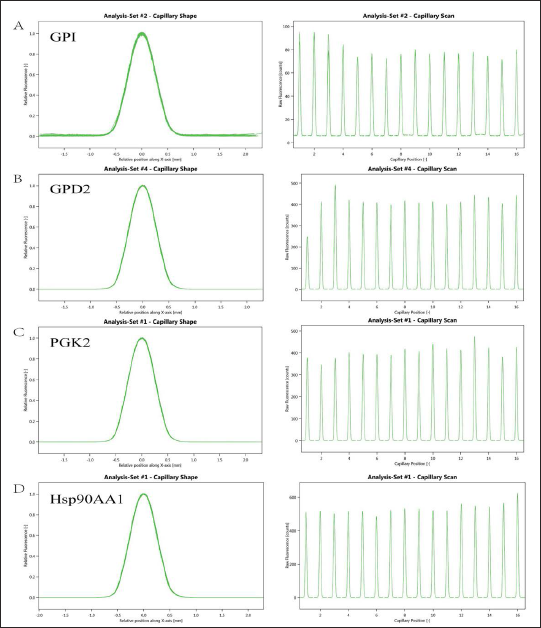
The SD-test was used to examine the effect of labeling on Hsp90AA1, PGK2, GPD2, and GPI and to exclude the effect on the protein from protein fluorescent labeling.
As illustrated in Figure 6, the SD-test findings revealed that the protein fluorescent tagging had minimal impact on each protein.
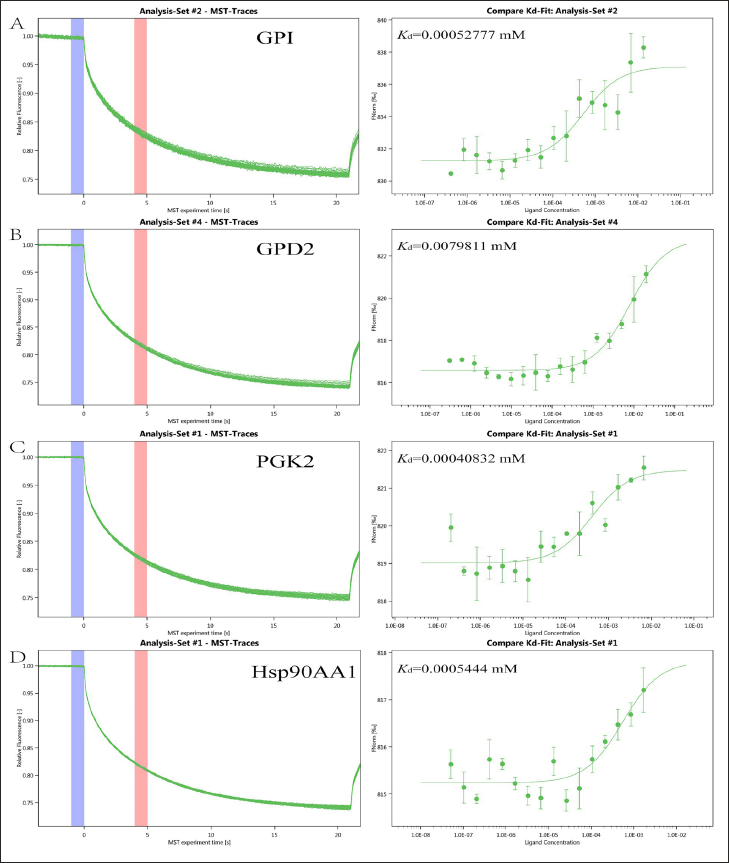
The Binding between 4-MU and Proteins
Following the application of various doses of 4-MU to the tagged target proteins as assessed by MST, the fluorescence signal’s change value during thermal stimulation of the temperature gradient field was multiplied by 1,000, which was the relative change value per thousand fluorescences. The dissociation constant Kd was calculated using the normalized fluorescence difference between the bound and unbound states. The results indicate that 4-MU binds tightly to Hsp90AA1, PGK2, GPD2, and GPI. These results are shown in Figure 7.
Discussion
In this study, we used bioinformatics methods and network pharmacology combined with molecular docking technology to explore the anti-tumor mechanism of 4-MU. Secondly, the biophysical technique MST experiment was used to verify the network pharmacology results, and the binding ability of 4-MU to the target proteins Hsp90AA1, PGK2, GPD2, and GPI was studied.
Based on previous research, Hepg2 cells were used for experiment decisions. Using 4-MU to administer the drug to the cells, RNA-seq sequencing of the affected cells was performed using Illumina second-generation sequencing technology. The result clearly revealed that under the influence of 4-MU, the cell changed at the gene level.
Through the induction, reorganization, and deep mining analysis of RNA-seq sequence discrepant data and differentially expressed genes of the medication administration group and the control group, the phenomenon that glycolysis and other metabolism-related pathways and related proteins were more concentrated was revealed.
The results of network pharmacology and virtual molecular interconnection showed that the glycolytic pathway had high comprehensive effect scores, so it was a necessity that its associated enzymes, such as Hsp90AA1, PGK2, GPD2, and GPI, had high comprehensive effect scores too, which supports the idea that the effect of 4-MU on the glycolysis level of tumor cells may be one of its anti-tumor mechanisms. Through the analysis of GO functional enrichment results, the cross-targets were mainly involved in BPs such as glycolysis, typical glycolysis, and gluconeogenesis; other CCs such as cytoplasm, membrane, and cell exosomes; and other MFs such as mainly regulated protein binding, ATP binding, and transmembrane receptor protein tyrosine kinase activity. GO function test results suggested that catabolism of small molecules and other glucose metabolism processes, as well as the cytoplasm or cell membrane, have the potential to be impacted by 4-MU and inhibit glycolysis. KEGG pathway enrichment results showed that 4-MU May play an anti-tumor effect by combining Hsp90AA1, PGK2, GPD2, and GPI, the core proteins of the glycolysis pathway. Key pathways such as the HIF-1 signaling pathway, tumor proteoglycan, and endocrine resistance also suggest that 4-MU combines with Hsp90AA1, PGK2, GPD2, and GPI to further influence the expression and activity of tumor-related proteins.
GPI is ubiquitous in most organisms and catalyzes the reversible isomerization of glucose-6-phosphate and fructose-6-phosphate during glycolysis, directly affecting glycolysis, gluconeogenesis, and glucose metabolism. When GPI is knocked down, oxygen consumption is restored and glycolytic activity is reduced. Downregulation of GPI expression can reduce cell migration and sensitize tumor cells to drugs (Lázár et al., 2021; Shao et al., 2020). In acute hepatitis, acute myocardial infarction, malignant tumor and other diseases, the serum GPI activity is significantly increased, and it is positively correlated with the severity of the disease. This means that the study of GPI has potential implications for the diagnosis and detection of many diseases. GPD2 is a key protein in aerobic glycolysis in tumor cells (Oh et al., 2023). GAPDH, which was converted from dihydroxyacetone phosphate by GPD2, produces nicotinamide adenine dinucleotide in the cytoplasm.
GPD2 activation changes its substrate affinity, preferentially converting GAPDH to DHAP and glycolysis catalysis, thereby promoting glioma cell proliferation (Li et al., 2022; Wu et al., 2022). PGK2 is an important protease in glycolysis, which not only catalyzes the conversion of glycerol-1,3-diphosphate to 3-phosphoglycerate but also produces ATP (Rogalski-Wilk & Cohen, 1997). PGK2 is secreted by a variety of tumor cell lines, such as pancreatic, breast, and colon cancer cells (Al Bawab et al., 2021; Ikemoto et al., 2003). PGK2 was also detected in the serum of patients with lung adenocarcinoma or pancreatic cancer, even though it was significantly higher than in normal controls.
Hsp90AA1 can bind to other proteins to form complexes, which in addition help transport proteins within and between cell membranes and additionally help with protein folding and stretching to regulate protein function. Hsp90AA1 can regulate protein function, which obtains the name “molecular chaperone” because of its involvement in the assembly of polymer complexes without changing protein structure (Zhang et al., 1999).
Hsp90AA1 can promote cellular effects, so the opposite of that is that Hsp90AA1 inhibitors can inhibit cellular action. It is inhibited by competitively binding to ATP. The failure of the binding between Hsp90AA1 with ATP forces Hsp90AA1 to change its conformation and its molecular chaperone function is inhibited, blocking the binding of Hsp90AA1 to ATP substantially eliminating signal transmission and energy exchange in tumor cells. So Hsp90AA1 inhibitors play an important role in inducing apoptosis in tumor cells (Li, 2004; Obermann et al., 1998). In accordance with the data above, as well as the analysis of Hsp90AA1, PGK2, GPD2, and GPI, these targets have the meaning of an in-depth study because of their significance in boosting aerobic glycolysis in tumor cells.
MST detection is a novel biomolecular interaction detection suitable for a variety of complex experimental environments (Young & Hartl, 2000). MST results showed that 4-Mu was associated with Hsp90AA1, PGK2, GPD2, and GPI. It has been reported that the fitted curve can be S-shaped or S-shaped specular. The chemistry, binding site, and molecular-structure changes produced by combining a protein with the sample of interest indicate the MST amplitude (Romain et al., 2020). These results suggest that protein structure changes such as activations of Hsp90AA1, PGK2, GPD2, and GPI can inhibit the level of glycolysis, leading to anti-tumor effects.
4-MU has the features of an excellent inhibitor of changes in protein conformation, inhibiting glycolytic levels by altering the conformation of Hsp90AA1, PGK2, GPD2, and GPI. In addition to glycolysis-related proteins, 4-MU May also inhibit tumor proliferation by affecting other proteins, which will be one of the focuses of future studies. This study found that 4-MU inhibited glycolysis levels, blocked the energy and information exchange of tumor cells, and inhibited the growth and division of tumor cells by combining with glycolysis-related proteins Hsp90AA1, PGK2, GPD2, and GPI, which provided a new direction for the anti-tumor research of 4-MU.
In the future, we will conduct some in vivo animal experiments and in vitro cell experiments based on this paper to further verify this mechanism and strive to find more possible pathways and targets.
Conclusion
These results suggest that 4-MU inhibits glycolysis by binding to glycolysis-related proteins Hsp90AA1, PGK2, GPD2, and GPI, blocking the energy supply of tumor tissue, and inhibiting tumor proliferation, and thus exerting anti-tumor effects. It suggests that 4-MU may be a novel inhibitor of glycolysis in tumor cells and that these glycolysis-related proteins could be possible novel cancer therapeutic targets, which further provides valuable evidence for the anti-cancer mechanism of 4-MU.
Summary
This paper demonstrates that 4-MU, which can be extracted from the medicinal plant with anti-cancer properties, Smilax china L., inhibits glycolysis by binding to glycolysis-related proteins Hsp90AA1, PGK2, GPD2, and GPI, blocking the energy supply of tumor tissue and inhibiting tumor proliferation, and thus exerting anti-tumor effects.
Availability of Data and Materials
The datasets used and/or analyzed in this study are available from the corresponding author upon reasonable request. The main supporting data can be found in the supplementary material to the article.
Abbreviations
4-MU: 4-Methylumbelliferone; GPI: Glucose-6-phosphate isomerase; GPD2: Glycerol-3-phosphate dehydrogenase, mitochondria; PGK2: Phosphoglycerate kinase 2; MST: Microscale thermophoresis; Hsp90AA1: Heat Shock Protein 90 Alpha Family Class A Member 1; KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: Gene ontology; PPI: Protein-Protein Interaction; PBS: Phosphate buffer saline; DMSO: Dimethyl sulfoxide; BP: Biological Process; CC: Cellular Component; MF: Molecular function; GAPDH: Glyceraldehyde-3-phosphate dehydrogenase.
Authors’ Contributions
S-TW designed and guided the experiments. Z-YL provided financial support, conducted the experiments, analyzed the data, and wrote the manuscript. QY, X-GC, Y-LZ, X-LP, LQ, and B-JZ searched for information and provided assistance in completing the experiment.
Footnotes
Acknowledgments
The authors are indebted to the Key Laboratory of Traditional Chinese Medicine Resources and Chemistry of Hubei Province for providing convenience in the experimental process. The authors are grateful to the editors and reviewers for their suggestions, which have greatly improved this manuscript. We declare that this work was done by the authors named in this article, and all liabilities pertaining to claims relating to the content of this article will be borne by the authors.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was supported by the Scientific Research Project of the Education Department of Hubei Province (Grant No. Q20212005).
Statement of Informed Consent and Ethical Approval
Necessary ethical clearances and informed consent were received and obtained, respectively, before initiating the study from all participants.
