Abstract
Background and Objectives
Both primary and secondary cancers require the involvement of glycolytic pathways. Cancer cell proliferation leads to the upregulation of glycolysis, which results in increased glucose consumption. For demonstrating that umbelliferone can effectively bind to several proteins involved in the glycolytic pathway, thereby inhibiting glycolysis and reducing cancer cell proliferation.
Materials and Methods
This study uses transcriptomics, network pharmacology, and molecular docking to predict the potential targets and possible pathways of umbelliferone against cancer and microscale thermophoresis (MST) to detect the affinity between umbelliferone and potential targets.
Results
Transcriptomic analysis revealed that differentially expressed genes were primarily associated with glycolytic and other metabolic pathways and proteins. According to network pharmacology and molecular docking results, glycolysis-related proteins such as glucose-6-phosphate isomerase (GPI), glycerol-3-phosphate dehydrogenase, mitochondrial (GPD2), phosphoglycerate kinase 2 (PGK2), and heat shock protein HSP-90 alpha (Hsp90AA1) are potential targets of umbelliferone against tumors. MST confirmed that umbelliferous lactone binds strongly to GPI, GPD2, and PGK2 but not to Hsp90AA1.
Conclusion
By binding to the glycolysis-related proteins such as GPI, GPD2, and PGK2, umbelliferone acts as an anti-tumor agent by inhibiting glycolysis, cutting off the energy supply to tumor tissue, and reducing tumor growth. It was suggested that umbelliferone might be a brand-new tumor glycolysis inhibitor and that these glycolysis-related proteins might be potential new targets for cancer therapy. This finding helped to establish a solid foundation for the anti-cancer action of umbelliferone.
Introduction
Although studies have shown that lactic acid is typically produced in anoxic environments, tumor cells will still prefer to metabolize glucose into lactic acid in order to fuel their own growth, even in environments with plenty of oxygen. The Warburg effect, or aerobic glycolysis, is the metabolic process that is thought to be indicative of malignant tumors (Schwartz et al., 2017; Vaupel et al., 2019). The immune system is suppressed by the tumor microenvironment that tumor cells with active metabolism will produce because it is full of nutrients consumed, hypoxia, acidity, and toxic metabolites. Glycolysis proceeds through a series of enzymatic reactions in the cytoplasm. A large number of related enzymes directly contribute to the growth of tumor cells during the glycolysis process. They include the heat shock protein HSP-90 alpha (Hsp90AA1), phosphoglycerate kinase 2 (PGK2), glycerol-3-phosphate dehydrogenase, mitochondrial (GPD2), glucose-6-phosphate isomerase (GPI), and GPD2. These enzymes might serve as potential biomarkers and treatment targets for tumors (Kathagen-Buhmann et al., 2018; Xu et al., 2017). As a result, increasing numbers of studies have concentrated on the metabolic profile of tumor cells. Inhibit the expression of glycolysis-related enzymes, thus reducing the level of glycolysis and ultimately stopping the proliferation of tumor cells, which is one of the most effective methods of anti-tumor therapy.
Umbelliferone is a coumarin that can be isolated in significant quantities from the anti-cancer plant vine pear root. It is now employed as an inhibitor of hyaluronic acid formation and has anti-inflammatory properties. Recent research has demonstrated that umbelliferone has positive effects on breast cancer (Ali et al., 2019; Kandil et al., 2016), patocellular carcinoma (Yu et al., 2015), prostate carcinoma (Shen et al., 2017), oral carcinoma (Vijayalakshmi & Sindhu, 2017), and more. It can also act as a tumor suppressor and anti-cancer cell metastasis agent. Most studies have shown that umbelliferone reduces the expression of matrix metalloproteinase 9 (Ali et al., 2019; Kandil et al., 2016; Karthikeyan et al., 2016), while decreasing the levels of hyaluronic acid and glucose in the tumor microenvironment and inhibiting tumorigenesis
The aim of this study was to investigate whether umbelliferone can inhibit the proliferation of tumor cells by inhibiting the expression and activity of glycolysis-related enzymes, thereby downregulating the level of glycolysis.
Transcriptomics is a discipline that studies gene transcription in cells at cellular level as well as transcription regulation. The study of gene expression at the RNA level is known as transcriptomics. The transcriptome, which is the total number of RNAs that can be transcribed from a living cell, is a useful tool for studying cellular phenotype and function. The transcriptional process of synthesizing RNA using DNA as a template is the first step in gene expression and a critical link in gene expression regulation.
A drug design approach based on receptor properties and interactions between receptors and drug molecules is known as molecular docking (Pinzi & Rastelli, 2019). Theoretical simulations are used to analyze the interaction between small molecules and protein receptors and to forecast their binding affinities and modes.
The directed movement of particles in a microscopic temperature gradient is known as microscale thermophoresis (MST). By monitoring changes in the trace heat flow brought on by modifications to the hydration layer, which are often brought on by modifications to the structure or arrangement of biomolecules, affinity can be identified (Jerabek-Willemsen et al., 2011; Wienken et al., 2010). It is possible to quantify even minute changes, like the phosphorylation of proteins or the binding of tiny compounds to their targets.
In this study, we first obtained almost all transcripts from control and administered groups of hepatocellular carcinoma (HCC) cells and then reacted to their expression levels. The potential targets and regulatory pathways of umbelliferone against HCC were then predicted using network pharmacology (Wang et al., 2022; Ye et al., 2016) and molecular docking. Finally, MST tests confirmed the presence of umbelliferone and glycolysis-related proteins, but the pharmacological activity associated with umbelliferone suppression of glycolysis in tumor cells was not. A series of related studies will be carried out in the future.
Materials and Methods
Reagents and Proteins
Umbelliferone (CAS: 93-35-6) was purchased from the Beijing Century Aoke Biology Research Co., Ltd. (China), with a concentration of ≥98%. Hsp90α (ab78425), GPI (ab116799), GPD2 (ab132692), and PGK2 (ab123166) were purchased from Abcam (Cambridge, UK).
Total RNA Extraction from Cells
HepG2 cells obtained from the Chinese Academy of Sciences cell bank were grown in a cell culture incubator at 37°C and 5% CO2 in HepG2 cell-specific media. After 24 h, the cultivated HepG2 cells were placed in 6-well plates and swapped between the original medium (the control group) and a drug-containing medium containing 50 mM umbelliferone (the drug treatment group). Following that, total RNA was extracted from both groups using the TruSeqTM RNA Sample Preparation Kit, and the concentration, purity, and integrity of total RNA were assessed using the NanoDrop 2000. Finally, 5 µg of good-quality RNA samples (OD 260/280 = 1.8–2.2) were chosen for library construction.
Transcriptome Sequencing
Transcriptome sequencing generally entails high-throughput sequencing of mature mRNA and ncRNA synthesized by polythymidine-affinity-purified RNA polymerase II transcripts (oligo-dT). Transcriptome sequencing, when compared to traditional microarray hybridization platforms, can detect the overall transcriptional activity of any species without the need to pre-design probes for known sequences, resulting in more accurate digitized signals, higher detection throughput, and a wider detection range, making it a powerful tool for in-depth transcriptome complexity research. To extract mRNA, we used magnetic beads and oligonucleotides (dT) with A-T bases at the 3ʹ-terminus of eukaryotic mRNA cells. By adding a fragmentation buffer, mRNA was randomly deconstructed into 300 DP pieces, and single-stranded cDNA was created by adding the reverse transcriptase to form a stable double-stranded structure. The products were purified and split with abstainers, and the selected products were amplified by PCR to obtain the final library, which was then analyzed by sequencing using HiSeq Xten/NovaSeq 6000 platforms (Wu et al., 2020).
Differential Expression Analysis and Functional Enrichment
In RNA-Seq analysis, the read counts of the clean read sequence found in the genome region—which reflected the level of gene expression—could be queried to determine the gene abundance. The degree of gene expression increases with abundance. In order to identify differences in transcripts and gene expression between various samples, RSEM was used in this study to quantitatively analyze the transcripts and gene expression levels of various samples. To find and separate differentially expressed genes from multiple genes and to thoroughly investigate the distinct functions of differentially expressed genes, a differential analysis of gene expression was conducted.
Prediction of Intersection Target Genes
The PubChem database (
GO Function and KEGG Pathway Enrichment Analysis
The Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathway enrichment analysis is frequently used in network pharmacology to examine the mechanism of drug action. It can highlight the significance of various signaling pathways in protein–protein relationship networks (Kanehisa et al., 2017). The most representative tool for the systematic description of biology at the moment is the gene ontology (GO) function enrichment analysis (Chen et al., 2017). The KEGG signaling pathway’s cross-targets and GO function can be annotated in the DAVID database for integrated gene discovery, visualization, and annotation (Huang et al., 2009). The pathway with the greatest number of cross-targets can be viewed as an essential regulatory pathway for the anticancer effects of umbelliferous lactones, according to an analysis of the KEGG signaling pathway annotation results.
Protein–Protein Interaction and Network Construction
The intersection targets were analyzed using the STRING website, with the species set to “
Molecular Docking
We obtained the 3D ligand files of umbelliferone from PubChem and the 3D structures of Hsp90AA1, GPI, GPD2, and PGK2 from the PDB Protein Data Bank. The interaction of umbelliferone with the four proteins was investigated using AutoDock software, and molecular docking was performed. The binding ability of the pharmacodynamic molecule to the target was assessed using the binding energy after successful docking, and binding forms with binding energies ≤ – 4 kcal mol–1 were included in the selection.
Protein Fluorescent Labeling
For protein fluorescent labeling, the protein samples such as GPI, GPD2, PGK2, and Hsp90α were dissolved in a protein labeling buffer. For 1 h, we mixed 100–500 µL of each protein solution with 50 µg of the fluorescent dye DyLight® 488. The mixture was purified using a Zeba Spin desalting column after the reaction to remove the unreacted dye and replace the labeled proteins with phosphate buffered saline (PBS) buffer. The protein concentration was calculated and kept at 4°C.
Binding of the Compound Sample with the Labeled Proteins
Umbelliferone was diluted to a final concentration of 20 mM in PBS buffer containing 10% dimethyl sulfoxide (DMSO). The fluorescently labeled protein Hsp90α was diluted in PBS buffer containing 10% DMSO and 2% Tween 80, and the solution concentration was adjusted so that the fluorescence intensity ranged between 400 and 1500 Fnorm. A total of 16 PCR tubes were labeled 1–16, and 20 µL of the compound solution was added to tube 1 and 10 µL of PBS buffer containing 10% DMSO to tubes 2–16. After mixing with the liquid in tube 2, 10 µL of liquid from the compound solution in tube 1 was removed and added to tube 3 for proportional dilution. After dilution of the solution in each tube, 10 µL of liquid in tube 16 was aspirated and discarded, leaving the volume of liquid in tubes 1–16 at 10 µL. A volume of 10 µL of fluorescently labeled protein solution was added to each of tubes 1–16, mixed thoroughly, and the reaction was allowed to equilibrate at room temperature for 5 min. GPI, GPD2, and PGK2 after fluorescence labeling were treated as above.
MST Determination
The mixed solution in the PCR tube was sucked into the standard capillary tube by the siphon phenomenon and placed in the MST Monolith NT.115 for fluorescence scanning. Different samples were set with different determination conditions, including LED intensity and test temperature for MST determination, to detect the binding between umbelliferone and GPI, GPD2, PGK2, and Hsp90AA1.
Results
RNA-Seq Transcriptome Differential Gene Analysis
Using DEG-seq software, the differentially expressed genes were checked for statistical significance (Log2FC(H_O/H) ≥ 2,
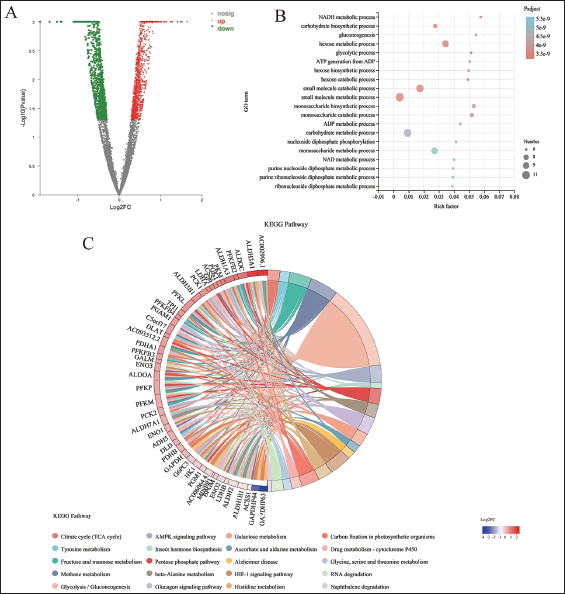
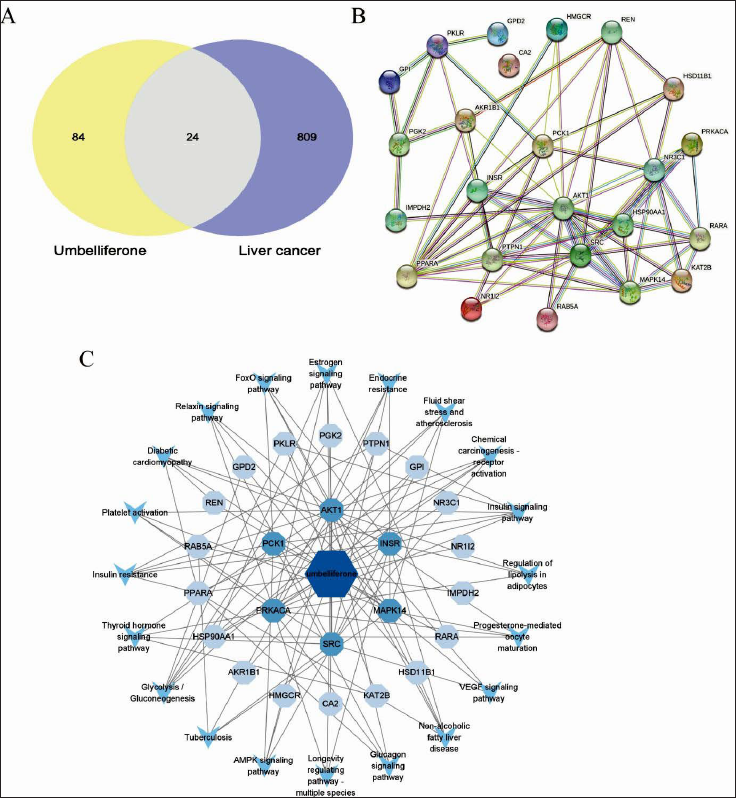
Protein–Protein Interaction and Network Construction
Based on the results of the transcriptome analysis, we performed a network pharmacology analysis. The PharmMapper database and GeneCards database were used to screen 24 potential drug targets and intersecting disease targets, as shown in Figure 2A. We used the STRING database to integrate the intersecting target genes to generate an interaction network, as shown in Figure 2B. The 24 target genes and 20 signaling pathways with umbelliferous lactones were imported into Cytoscape 3.9.1 software to construct an “umbelliferone/target/pathway” network, as shown in Figure 2C. In the network diagram, dark blue nodes represent umbelliferone, light blue nodes represent target genes, and sky blue arrow-shaped nodes represent pathways.
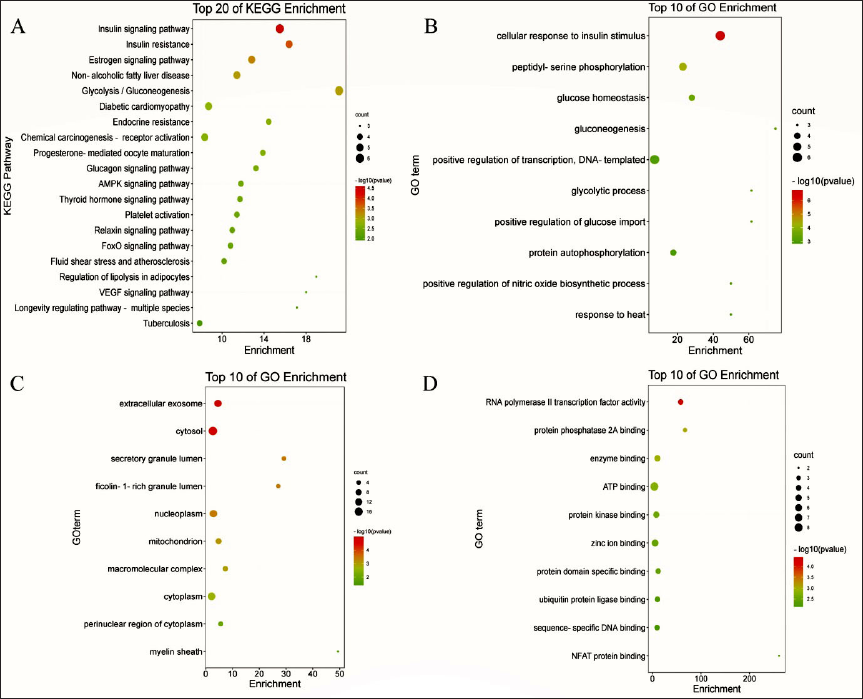
GO Function and KEGG Pathway Enrichment Analysis
We entered the intersecting targets into the DAVID database, resulting in 51 pathways and 42 putative pathways (
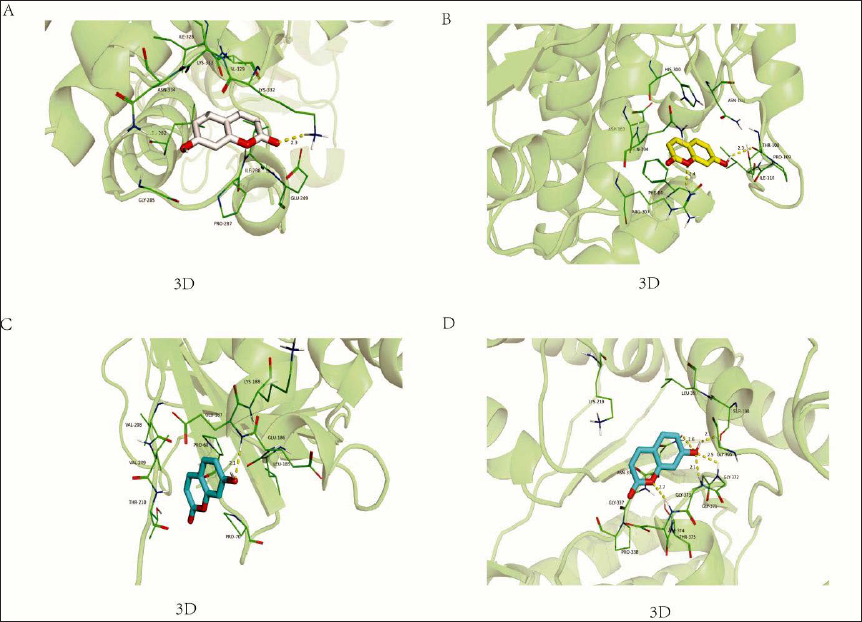
Molecular Docking Results
AutoDock was used to analyze the free binding energy and hydrogen bonding between the compounds and the targets. Interestingly, the docking binding energies of the compounds to all four targets were less than – 4 kcal/mol–1. The docking situation is shown in Figure 4, and the results of docking binding energy are shown in Table 1.
The Result of Docking Binding Energy.

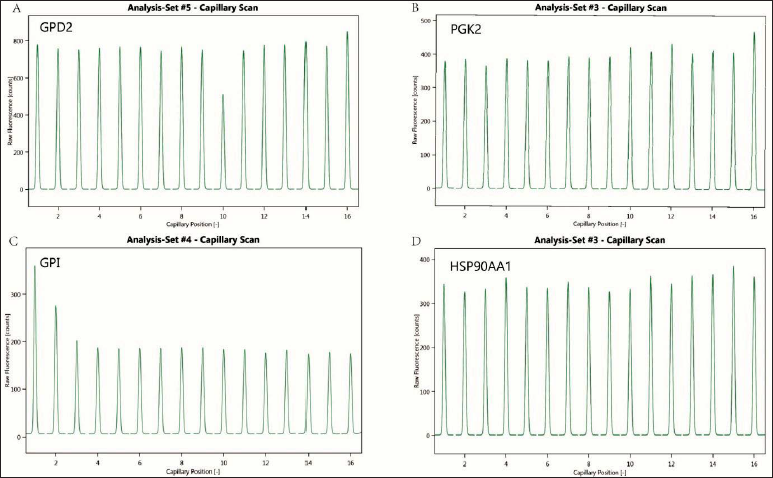
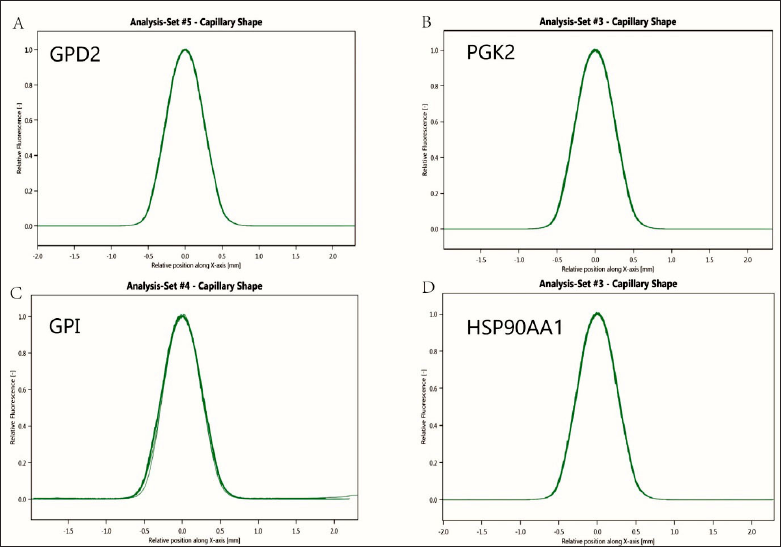
SD Test
Since protein fluorescent labeling may have an effect on proteins, we performed SD tests to examine the effect of labeling on GPI, GPD2, PGK2, and Hsp90α. SD test results showed that protein fluorescent labeling had little effect on each protein, as shown in Figures 5 and 6.
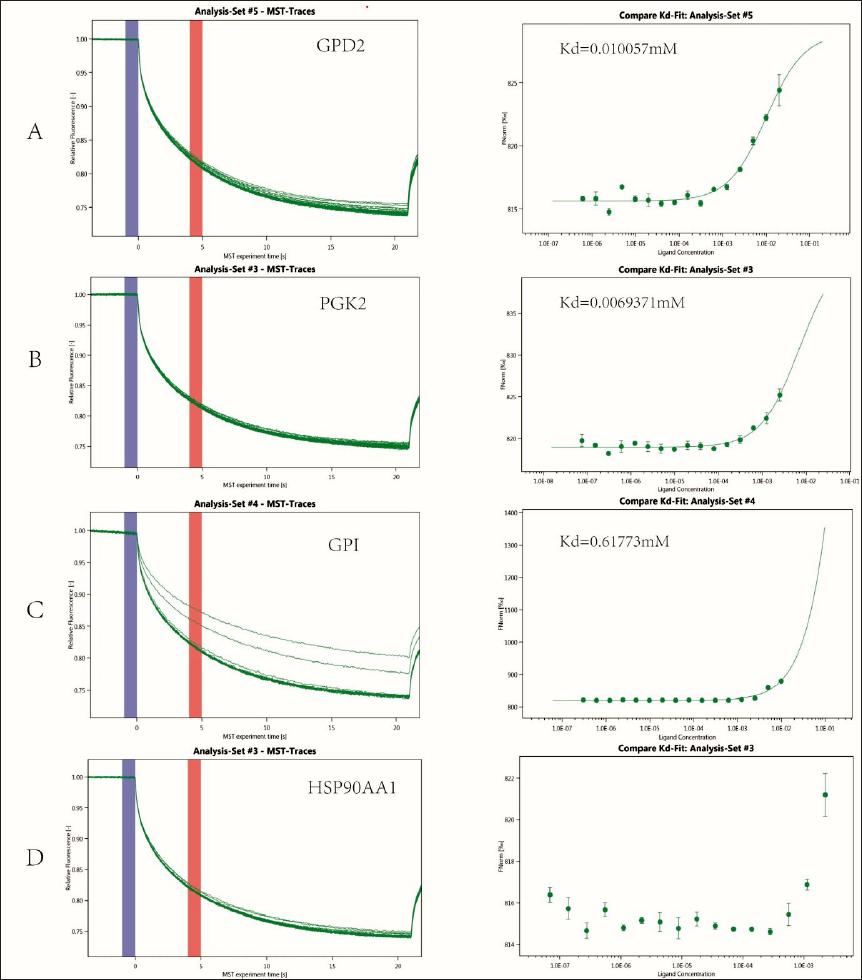
Binding of Umbelliferous Lactones to Proteins
After applying different concentrations to the labeled target proteins measured by MST, the change value of the fluorescence signal during heat excitation in the temperature gradient field was multiplied by 1000 to convert it into the relative change value of fluorescence per thousand times. The normalized fluorescence difference between bound and unbound states was used to calculate the dissociation constant
Discussion
In this study, network pharmacology in conjunction with molecular docking technology was used to investigate the antitumor mechanism of umbelliferone. The network pharmacology results were then confirmed using the geophysical technology MST experiment, and the binding affinity of umbelliferone for the target proteins such as GPI, GPD2, PGK2, and Hsp90AA1 was explored.
After the addition of umbelliferone, HepG2 cells were subjected to RNA-seq sequencing using Illumina’s second-generation sequencing technology. This revealed a more thorough and dynamic process of quantitative gene change caused by umbelliferone. Through deep mining analysis of the RNA-seq sequence data of the treatment group and the control group and the analysis of differentially expressed genes, it was found that glycolysis and other metabolic-related pathways and related protein enrichment were more concentrated. It was expected that umbelliferone may inhibit the proliferation of tumor cells by acting on glycolysis-related proteins and regulating the related pathways.
The influence of umbelliferone on the level of glycolysis in tumor cells may be one of the anti-tumor mechanisms, according to the findings of network pharmacology and virtual molecular interconnection, which revealed that the glycolysis pathway, and its related enzymes such as GPI, GPD2, PGK2, and Hsp90AA1 had a higher comprehensive effect score. Through the analysis of GO function enrichment results, intersection targets were mainly regulated transmembrane receptor protein tyrosine kinase activity, identical protein binding, ATP binding, and other MFs. These targets were primarily involved in the glycolysis process, canonical glycolysis, glucose, genesis, and other BPs involved in the cytosol, membrane, extracellular exosome, and other cell components. Umbelliferone may influence the cytosol or membrane, small molecule catabolism, and other glucose metabolism processes to inhibit glycolysis, according to the results of GO function. Umbelliferone may act as an anti-tumor agent by attaching to key glycolysis pathway proteins, such as GPI, GPD2, PGK2, and Hsp90AA1, according to the findings of the KEGG signaling pathway enrichment. It has also been suggested that umbelliferone may alter the expression and activity of tumor-related proteins by binding to GPI, GPD2, PGK2, and Hsp90AA1 through the HIF-1 signaling pathway, proteoglycans in the endocrine resistance, and other significant pathways.
With its ability to catalyze the reversible isomerization of glucose-6-phosphate and fructose-6-phosphate in glycolysis, GPI is present throughout most organisms and has a direct impact on glycolysis, glucose, and glucose metabolism. The oxygen consumption returned and the glycolysis activity decreased when GPI was inhibited. The ability of cells to migrate may be reduced, and tumor cells may become more responsive to treatment if GPI expression is downregulated (Chen et al., 2020; Rengaraj et al., 2012). The activity of GPI in the serum of patients was markedly elevated in many diseases, including acute hepatitis, malignant tumors, acute myocardial infarction, and more, and was positively correlated with the severity of the disease. As a result, the study of GPI has the potential to be important for the identification and diagnosis of numerous diseases. A crucial protein in tumor cells’ aerobic glycolysis is GPD2 (Mikeli et al., 2020). In the cytoplasm, GPD2 can transform the compound dihydroxyacetone phosphate (DHAP) into the enzyme glyceraldehyde-3-phosphate dehydrogenase (GAPD) and NADH. By preferentially converting GAPD to DHAP to catalyze glycolysis, GPD2 activation changes its affinity with substrates and encourages the growth of glioma cells (Lu et al., 2020; Zhao et al., 2021). In the glycolysis process, PGK2 is a crucial protease that can catalyze the formation of ATP as well as the conversion of glycerol-1,3-diphosphate to 3-phosphoglyceric acid (Wu et al., 1997). Many tumor cell lines, including pancreatic, breast, and colon cancer cells, secrete PGK2 (Jarrar et al., 2020; Yeh et al., 2008). Additionally, PGK2 was found in the serum of patients with lung adenocarcinoma, and it was also found to be significantly higher in the serum of a patient with pancreatic cancer than in healthy controls. Hsp90AA1 can form complexes with other proteins, not only accompanying protein transport within the membrane cell but also participating in protein folding and stretching. Hsp90AA1 is also involved in the assembly of polymer complexes and regulates the function of the protein without changing the protein structure, so it is called a “molecular chaperone protein” (Zuehlke et al., 2015). Hsp90AA1 inhibitors can competitively bind with ATP, forcing Hsp90AA1 to change its conformation and inhibit its molecular chaperone function, thus blocking the binding of Hsp90AA1 and ATP and effectively eliminating signal transduction of tumor cells, which plays an important role in inducing apoptosis of tumor cells (Prodromou et al., 1997; Srivastava, 2005). Based on the above research results and the analysis of GPI, GPD2, PGK2, and Hsp90AA1, these targets play an important role in promoting the aerobic glycolysis of tumor cells.
RNA-seq sequencing of HepG2 cells treated with umbelliferone was performed using Illumina second-generation sequencing technology to reveal the process of gene number changes under umbelliferone treatment. The in-depth mining and analysis of the RNA-seq sequence data of the treated and control groups, as well as the analysis of differentially expressed genes, revealed a more concentrated enrichment of metabolism-related pathways such as glycolysis and related proteins. This suggests that umbelliferone may inhibit the proliferation of cancer cells by acting on glycolysis-related proteins and regulating related pathways.
The results of network pharmacology and molecular docking showed that the combined effect of the glycolytic pathway and its related enzymes, such as GPI, GPD2, PGK2, and Hsp90AA1, scored high, suggesting that the effect of umbelliferone on the glycolytic level of cancer cells may be one of the anti-cancer mechanisms. The GO functional results suggest that umbelliferone may affect the cytoplasm or cell membrane and influence small molecule catabolism and other glucose metabolism. The enrichment results of the KEGG signaling pathway suggest that umbelliferone may bind core proteins, such as GPI, GPD2, PGK2, and Hsp90α, through the glycolytic pathway to exert anticancer effects.
MST is a general technique for quantifying intermolecular interactions based on the directed motion of molecules in a temperature gradient (Huang & Zhang, 2021). The MST results indicate that the selected compounds have potential molecular interactions with PGK2, GPD2, and GPI. According to Seidel et al. (2013), the fitted curves can be S-shaped or have mirror-image surfaces. The chemical reactions, binding sites, and conformational changes induced after binding lead to the standard sign of the MST amplitude. This suggests that the activation of PGK2, GPD2, and GPI induced by conformational changes may inhibit glycolysis and further produce anticancer effects.
Conclusion
In this study, the transcriptome results were used as a cue to predict the potential targets and regulatory pathways of umbelliferone anticancer drugs in combination with network pharmacology and molecular docking. The affinity between umbelliferone and glycolysis-related proteins was verified by the MST assay. Umbelliferone showed the characteristics of a typical structural change inhibitor, which could inhibit the level of glycolysis by changing the conformation of GPI, GPD2, and PGK2. Umbelliferone may affect other proteins in addition to those involved in glycolysis, which is one of the areas of research that will be focused on in the future. The results of this study show that umbelliferone binds to the glycolysis-related proteins such as GPI, GPD2, and PGK2; inhibit the level of glycolysis; block the energy supply of tumor tissue; inhibit the proliferation of tumor cells; and provide a new direction for the study of umbelliferone’s anti-tumor properties.
Footnotes
Summary
It was the first time to study the mechanism of umbelliferone anti-HCC cell glycolysis.
Umbelliferone could strongly bind to glycolysis-related proteins.
Umbelliferone could inhibit tumor cell glycolysis and thus inhibit the proliferation of cancer cells.
Abbreviations
BP: biological process; CC: cellular component; MF: molecular function; DMSO: dimethyl sulfoxide; KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: gene ontology; GPD2: glycerol-3-phosphate dehydrogenase, mitochondrial; PGK2: phosphoglycerate kinase 2; Hsp90AA1: heat shock protein HSP-90 alpha; GPI: glucose-6-phosphate isomerase; HCC: hepatocellular carcinoma; MST: microscale thermophoresis; PBS: phosphate buffered saline; PPI: protein–protein interaction; DHAP: dihydroxyacetone phosphate; GAPD: glyceraldehyde-3-phosphate dehydrogenase; NADH: nicotinic adenine dinucleotide.
Acknowledgments
The authors have indebted the Key Laboratory of Traditional Chinese Medicine Resources and Chemistry of Hubei Province for providing convenience in the experimental process. The authors are grateful to the editors and reviewers for their suggestions, which have greatly improved this paper. We declare that this work was done by the authors named in this article, and all liabilities pertaining to claims relating to the content of this article will be borne by the authors.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Statement of Ethical Approval and Informed Consent
Necessary ethical clearances and informed consent were received and obtained from all participants before initiating the study.
Funding
This research was funded by the Hubei University of Chinese Medicine Youth Program (Grant No. 2021ZZX008).
