Abstract
Background
Prosaikogenin F (PSF) has stronger anti-cancer bioactivity than saikosaponin A (SSA), however, it was hardly isolated due to its trace amount in the raw material of Radix Bupleuri (RB). In addition, the active chemical constituent was unstable under acidic conditions owing to a 13,28-epoxy-ether moiety at the D ring of its aglycone.
Objectives
This study was to develop an appropriate method for obtaining acid-sensitive PSF from SSA abundant in RB.
Materials and Methods
Enzymatic hydrolysis was employed and snailase was selected due to its good hydrolysis performance under nearly neutral circumstances. Hydrolysis conditions were then optimized by one-factor-at-a-time experimentation before response surface methodology (RSM) by Box-Behnken Design (BBD).
Results
The reaction system was constructed in Na2HPO4-NaH2PO4 buffer (pH 6.0) containing snailase/SSA (44:1) at 39°C, then the hydrolysis lasted for 12 h. Therefore, the highest conversion ratio of SSA was achieved at 100.0%.
Conclusion
The newly proposed method is eco-friendly for obtaining acid-sensitive PSF, which lays a solid foundation for its development to be an anti-cancer new drug.
Keywords
Introduction
Radix Bupleuri (RB) is the dried root of Bupleurum chinense DC. or Bupleurum scorzonerifolium Willd., and used in clinical practice for over two thousand years. Saikosaponin A (SSA) is a major bioactive chemical constituent in RB, exhibiting remarkable anti-inflammatory, anti-oxidation, and anti-viral effects (Du et al., 2018; Li et al., 2018; Li et al., 2020). The saponin is accordingly employed as a marker for the quality assessment of raw RB in prevailing Chinese Pharmacopeia. In a previous study, partial hydrolysis of glycosyl groups of a primary glycoside will result in a secondary saponin, which can be well absorbed in the gastrointestinal tract to largely enhance efficacy in vivo owing to their increased lipophilicity (Lu et al., 2022; Shen et al., 2019a; Upadhyaya et al., 2016). It was seen that strong bioactivity but low bioavailability of saikosaponins remains to be a conundrum, and efficient conversion of a primary saponin to the desired secondary saponin is a convenient approach to well resolve it.
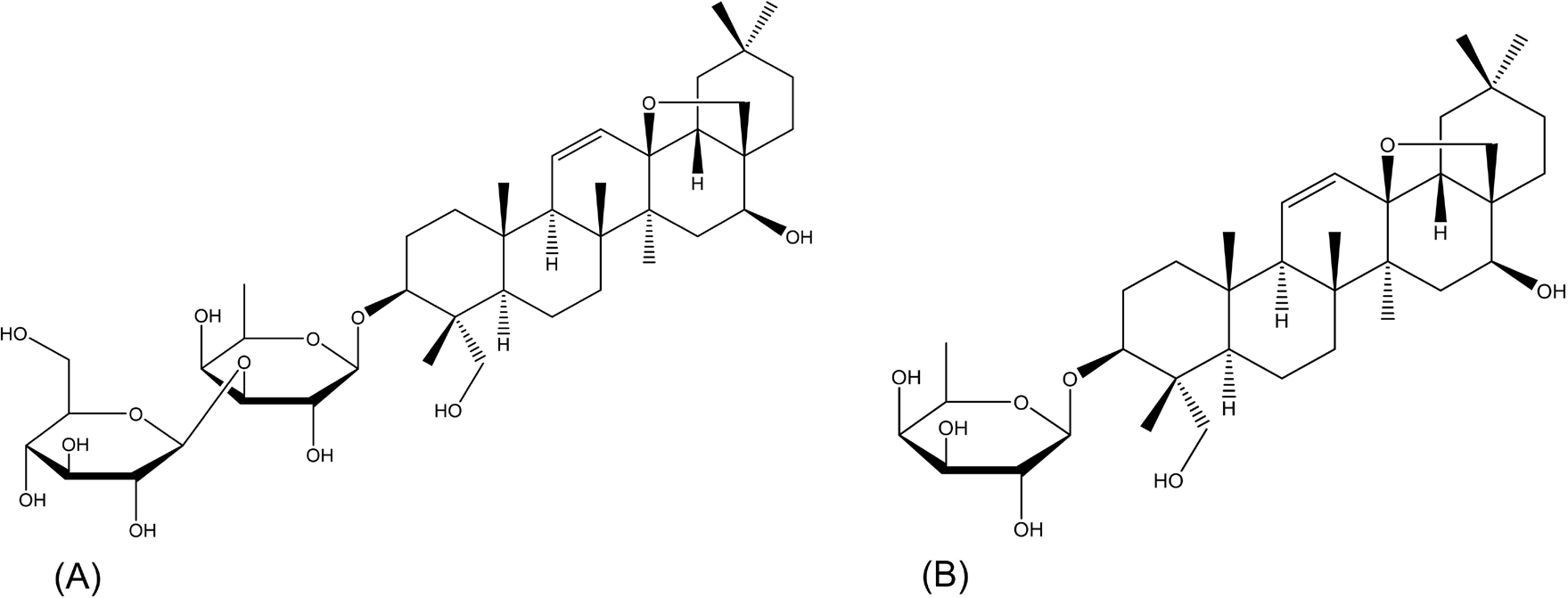
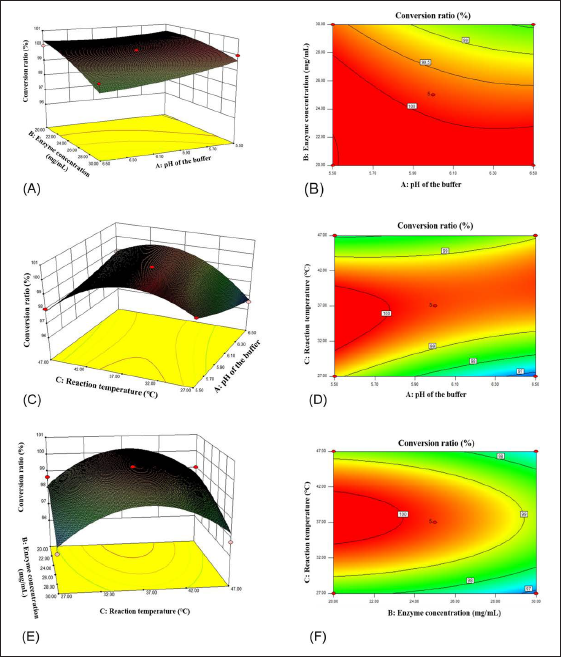
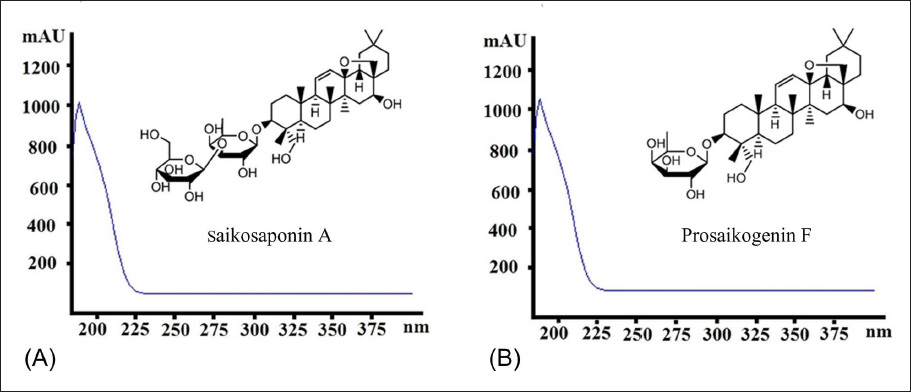
Enzymatic hydrolysis has shown well-recognized merits in great convenience and high specificity compared with acid hydrolysis and microbial transformation (Feng et al., 2019; Lu et al., 2023; Shen et al., 2019b,c). Among those primary saponins in raw RB, SSA (Figure 1A), a typical oleanane-type saponin featuring a unique 13,28-epoxy-ether moiety, is unstable under conventional acidic conditions for enzymatic hydrolysis. Prosaikogenin F (PSF, Figure 1B), an anti-cancer secondary glycoside of SSA, was hardly isolated due to its trace amount in raw material (Lee et al., 2022; Shi et al., 2020). To prevent the breakage of its epoxy-ether bond, the selection of a proper enzyme with high capacity is a vital step under strictly controlled pH conditions.
In this study, we aimed to establish a hydrolysis strategy for the convenient preparation of acid-sensitive PSF by enzymatic hydrolysis of SSA for the first time. Then, one-factor-at-a-time experimentation and response surface methodology (RSM) by Box-Behnken Design (BBD) were designed for optimizing the hydrolysis system. This research will lay a solid foundation for obtaining PSF based on “green chemistry” principles and be beneficial to the development of an anti-cancer new drug.
Materials and Methods
Reagents and Materials
SSA (Batch No.: 18041706, ≥98.0% by high-performance liquid chromatography-ultraviolet [HPLC-UV]) was isolated from the dried roots of B. chinense DC. and supplied by Chengdu Pufeide Biotechnology Co., Ltd. (China). HPLC grade Acetonitrile (ACN) (Batch No.: 2001101107M11) was the product of OmniGene LLC (United States) and other analytical grade reagents were bought from Sinopharm Chemical Reagent Co., Ltd. (China). β-glucosidase (activity: 100 U/g; Batch No.: 2020071806) and β-glucanase (activity: ≥20,000 U/g; Batch No.: 2020081501) were both bought from Jiangsu Ruiyang Biotechnology Co., Ltd. Cellulase (activity: ≥15,000 U/g; Batch No.: 20190719) were provided by Sinopharm Chemical Reagent Co., Ltd. (China). Snailase (Batch No.: S20200421) was supplied by Hefei BoMei Biotechnology Co., Ltd. (China).
HPLC Conditions
The content of SSA was determined by Agilent 1100 HPLC and the UV detection wavelength was 210 nm. 20 µL of standard solution or sample solution was injected for separation on a Kromasil C18 HPLC column (4.6 mm × 250 mm, 5 µm) kept at 35°C. Mobile phases, ACN (A) and water (B) were delivered at 1.0 mL/min constantly. The elution program was set up: 0 min, 40%A: 60%B; 20 min, 60%A: 40%B; 20.01 min, 40%A: 60%B; 25 min, 40%A: 60%B.
Preparation of Standard Solutions and Sample Solutions
SSA (12.60 mg) was dissolved in 100 mL methanol (MeOH) for preparing eight standard solutions (0.98−126 µg/mL). All solutions filtered through a 0.45 µm polytetrafluoroethylene membrane were injected for HPLC-UV analysis.
A certain volume of SSA standard solution and snailase solution was precisely mixed into a 1.5 mL tube containing Na2HPO4-NaH2PO4 buffer at a certain temperature for a period of time. Then, MeOH was transferred into the tube to terminate enzymatic hydrolysis, and the resulting mixture was centrifuged at 12000 rpm for 10 min. Before any injection for HPLC-UV analysis, the supernatant was filtered through a syringe membrane.
HPLC Method Validation
To investigate linearity, the prepared eight standard solutions were analyzed and the calibration curve of peak area (Y) versus the concentration of SSA (X) was drawn by linear regression using Microsoft Excel software (version 2016).
The highest, middle, and lowest concentrations of standard solutions and a sample solution were injected six consecutive times in a single day. In addition, the standard solution and the sample solution were analyzed on three consecutive days. Then, the intra-day and inter-day precision were calculated.
Six sample solutions were concurrently prepared and consecutively analyzed to evaluate the repeatability of the established HPLC method.
The standard solution and a freshly prepared sample solution were analyzed by the established HPLC method immediately, and then stored at room temperature. The determination of the solutions was performed again after 2, 4, 8, 16, 24, and 48 h.
The accuracy of the proposed HPLC method was evaluated by a recovery test. Three different amounts of standard solutions were added to the known amount of the sample solution. The recovery in percentage was then calculated by (detected amount-original amount)/added amount
Screening of Hydrolase
β-glucosidase, β-glucanase, cellulose, and snailase were compared in terms of the hydrolysis performance of SSA to its monosaccharide saponin, namely PSF. In detail, the reaction system was constructed by 5.0 µL of the substrate solution (5.00 mg/mL), 50.0 µL of the enzyme solution (1.00 mg/mL), and 145.0 µL of 0.20 M Na2HPO4-NaH2PO4 buffer in 1.5 mL tube. The pH of the buffer was adjusted to the optimal pH of each commercial enzyme, which was 5.0, 5.5, 5.0, and 6.5 individually. Constant incubation was carried out at 37.0°C water bath for 12 h. Then 200.0 µL of MeOH was transferred into the tube to terminate hydrolysis, and the resulting mixture was centrifuged at 12,000 rpm for 10 min. The filtered supernatant was subjected to HPLC-UV analysis. A proper enzyme was selected according to the conversion ratio of SSA.
Optimization of Enzymatic Hydrolysis Conditions
One-Factor-at-a-Time Experimentation
The effect of four factors, including enzyme concentration, pH of the buffer, reaction temperature, and hydrolysis duration, on the conversion ratio of SSA was investigated. In detail, 5.0 µL of SSA (5.00 mg/mL) was incubated with 50.0 µL of snailase solution (2.50−30.00 mg/mL) in 145.0 µL of 0.20 M Na2HPO4-NaH2PO4 buffer (pH 5.0−8.5). Then, hydrolysis was conducted at various temperatures (27°C−67°C) for different hydrolysis duration (2−18 h). In addition, a blank solution was prepared without snailase addition as the control.
Design of Response Surface Methodology
Enzymatic hydrolysis conditions were further optimized by RSM. In the present study, Design-expert software (version 8.0.6.1) was applied to design experiments and analyze data. A BBD with three variables was used to evaluate the interaction among them and ascertain their optimal values. According to the results from one-factor-at-a-time experimentation, three factors including pH of the buffer (A: 5.5−6.5), enzyme concentration (B: 20.00−30.00 mg/mL) and reaction temperature (C: 27°C−47°C) were investigated (Table A1). The non-linear quadratic model was obtained.
Calculations
The concentration of SSA was calculated from the plotted calibration curve. Then, the formula was employed for computing the conversion ratio of SSA (α):
where Ckb (µg/mL) represents the concentration of SSA in the blank sample solution; Crb (µg/mL) represents the concentration of unconverted SSA in the sample solution.
Structural Identification of the Product
After complete enzymatic hydrolysis, the resulting product was purified by a Hedera C18 HPLC column (20 mm × 250 mm, 5 µm) equipped with a Shimadzu LC-20AT HPLC. The eluate-containing target natural product was manually collected in a glass container and condensed to dryness. The chemical structure was elucidated by ESI+-MS (Thermo LXQ, USA) and NMR (Bruker Advance II 400 MHz, USA).
Results
HPLC Chromatograms
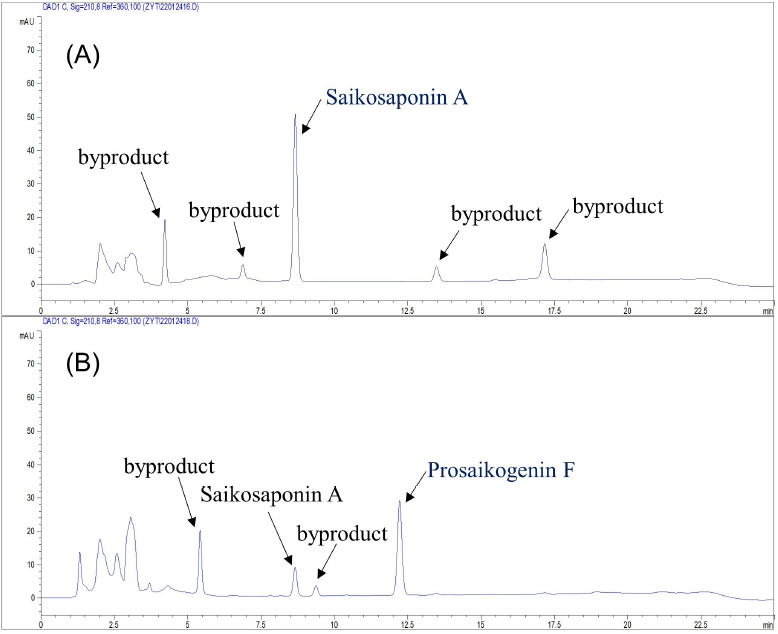
Typical HPLC chromatograms were shown in Figure A1, and the results showed that SSA and its hydrolysate could be well separated from the baseline (R > 1.5).
HPLC Method Validation
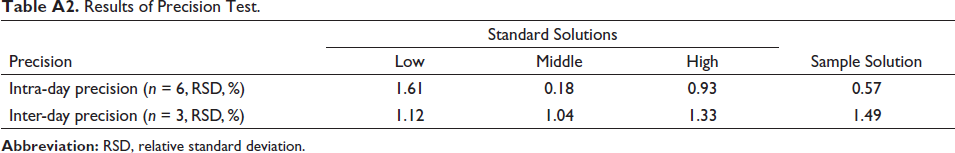
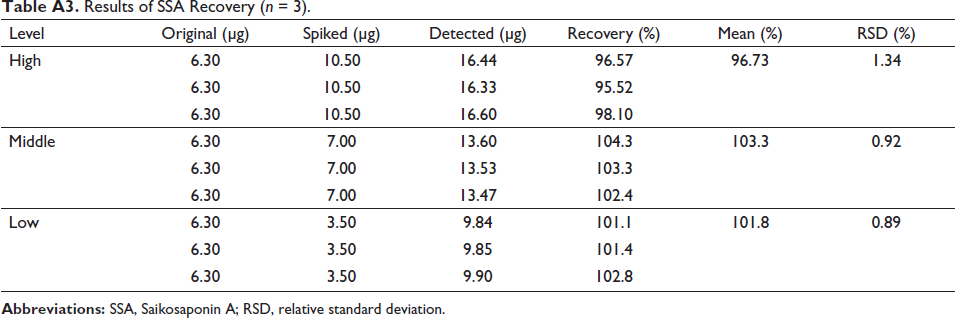
The calibration curve of SSA was drawn as Y = 6.5408X + 10.478 (r2 = 0.9991), indicating a good linear relationship in the range 0.98−126 µg/mL. Moreover, the intra-day and inter-day precision results were summarized in Table A2. The RSDs of the peak areas were all less than 2%, which showed that the analysis of SSA was precise. In the repeatability test, the RSD of the contents of SSA in the enzymatic hydrolysis reaction system was 1.88%, demonstrating good repeatability of the proposed analytical method. In the stability test, the RSDs of the peak areas were 1.33% and 1.03%, suggesting that the standard solution and the prepared sample solution were all stable at room temperature within 48 h. In addition, a good accuracy of the proposed HPLC method was showed in Table A3. The recoveries of SSA were 96.73%−103.3% with the RSDs ranging from 0.89% to 1.34%.
Screening of Hydrolase
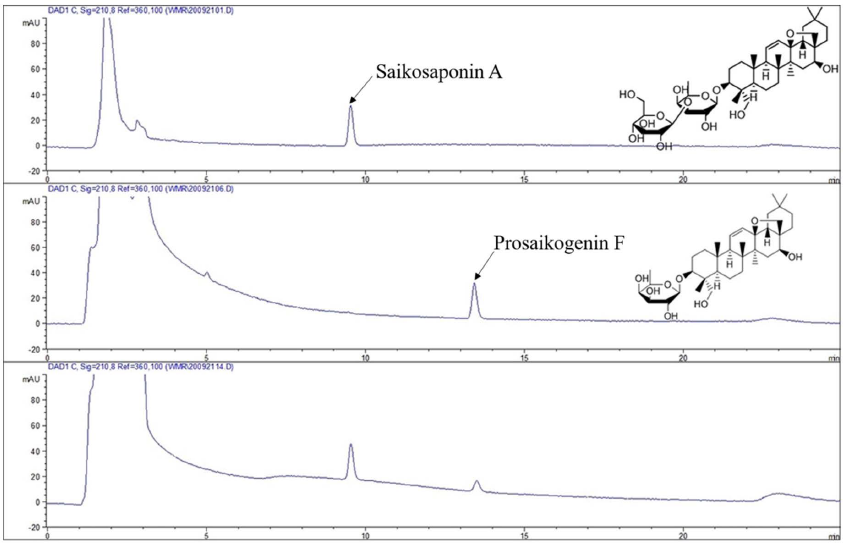
As illustrated in Figure A2, when the enzyme concentration was 1.00 mg/mL, the conversion ratio by snailase was much higher than those of the other three enzymes. It was also seen that the UV spectrum of the hydrolysate (PSF, Figure A3B) was consistent with that of the original primary glycoside SSA (Figure A3A) with λmax after hydrolysis by snailase at pH 6.0. As shown in Figure A4, the structure of PSF is unstable in the buffer of low pH conditions due to the breakage of the epoxy-ether bond. To keep the structural integrity of the resulting PSF, enzymatic hydrolysis should be carried out under a buffer of higher pH conditions. Consequently, snailase was selected for its best catalysis capability and its better hydrolysis performance in the buffer of higher pH circumstances.
One-Factor-at-a-Time Experimentation
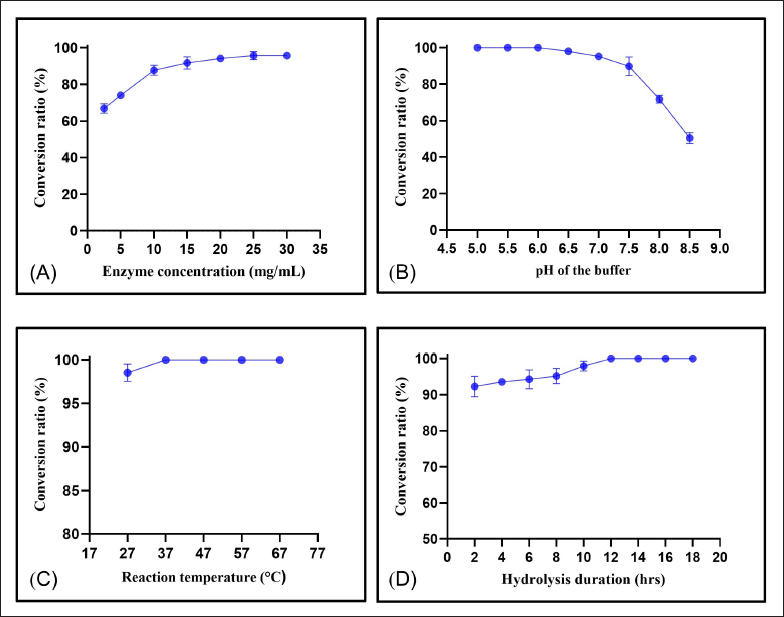
As illustrated in Figure 2, the conversion ratio of SSA gradually increased from 65.4% to 95.8% when the snailase concentration climbed to 25.00 mg/mL. The conversion ratio then remained at such a high level even though the concentration continued to increase. Therefore, the enzyme concentration of 25.00 mg/mL was chosen. Furthermore, 100.0% of SSA has been converted to PSF in the pH range 5.0−6.0, and the conversion ratio was 98.1% while the pH of the buffer was 6.5 and sharply dropped after the pH value was 7.0 or above. Given the structural integrity of PSF and the stronger robustness in the hydrolysis capability of SSA in less acidic circumstances, pH 6.0 was selected as the optimal hydrolysis condition. Moreover, the ratio ascended from 98.5% to 100% when the temperature of enzymatic hydrolysis was increased from 27°C to 37°C, and it kept constant at 100% after the temperature rose to 67°C. Accordingly, 37°C was selected as the optimal reaction temperature. In addition, the conversion ratio of SSA reached 100.0% after 12-h enzymatic hydrolysis. Hence, given the efficiency, 12 h was selected for subsequent complete enzymatic hydrolysis.
Optimization of the Conditions by RSM
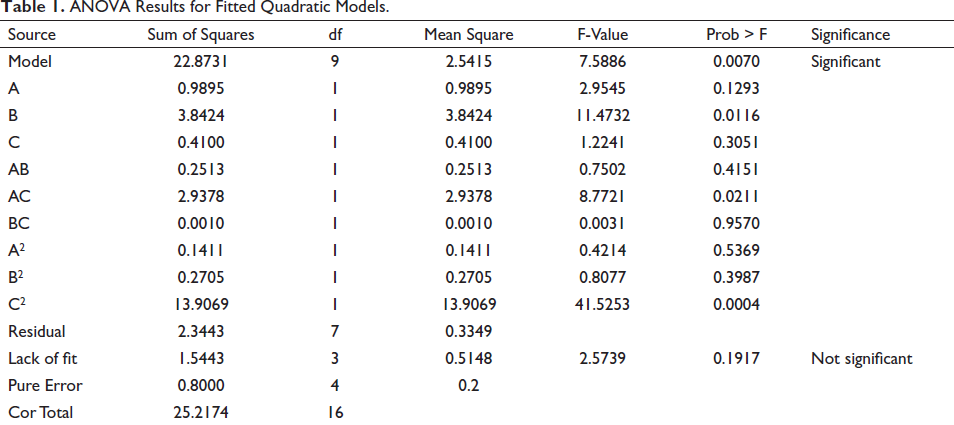
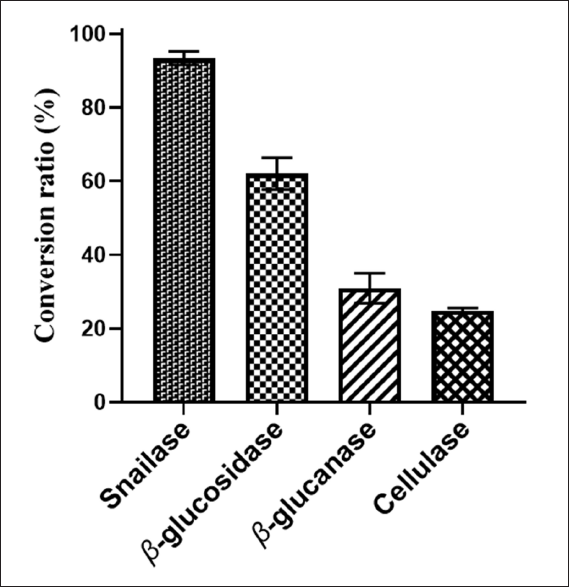
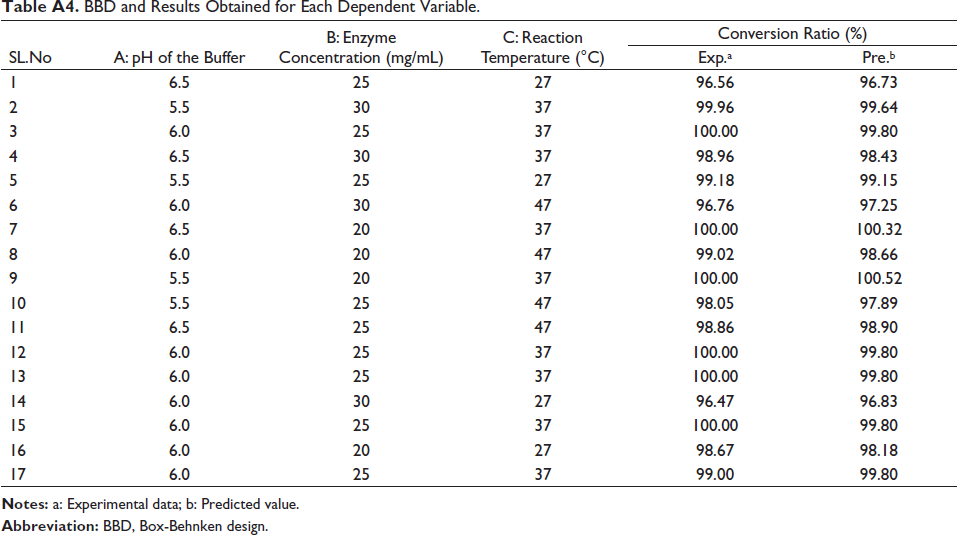
The experimental design matrix and responses were summarized in Table A4. According to Table 1, the estimated model was significant and the lack of fit test was not significant, so the model can be accurately used for the optimization of hydrolysis conditions. The fitted quadratic model was as follows:
ANOVA Results for Fitted Quadratic Models.
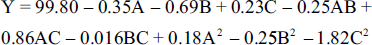
where Y represents the conversion ratio of SSA (%), A represents the pH of the buffer, B represents enzyme concentration (mg/mL), and C represents reaction temperature (°C).
Regression analysis showed that only 9.30% of the total variations were not able to explain by the quadratic model (R2 = 0.9070). RAdj2 value revealed the predicted values were in good agreement with the experimental values (RAdj2 = 0.7875). Moreover, the coefficient of variation was low (CV = 0.59), which indicated the reliability of the data. Meanwhile, the signal-to-noise ratio (Adeq Precision = 8.551) greater than 4 usually represented an adequate signal.
The conversion ratio was significantly affected by B-enzyme concentration (p < 0.05), followed by the A-pH of the buffer (p = 0.1293) and C-reaction temperature (p = 0.3051). The interaction between the pH of the buffer and reaction temperature exhibited the most significant effect (p < 0.05).
Besides, response surface plots and contour plots (Figure 3) showed the interactions between any two variables and their optimal ranges. To compare the predicted result with the practical value, three parallel experiments were conducted under the predicted hydrolysis conditions with a little modification (39°C, 22 mg/mL, pH 6.0). Finally, the conversion ratio of SSA was 100.0%, and the relative error was 0.60% compared with the predicted value of 100.6%. Thus, the model was reliable to predict the optimal hydrolysis conditions.
Structural Identification of the Product
The isolated product was identified as PSF and the chemical formula was C36H58O8.
ESI+-MS: m/z 641[M+Na]+, 657[M+K]+; 1H-NMR (400 Hz, DMSO-d6) δ: 0.55 (3H, s, H-24), 0.79 (3H, s, H-30), 0.82 (3H, s, H-25), 0.85 (3H, s, H-29), 0.91 (3H, s, H-27), 0.93 (3H, s, H-26), 4.16 (1H, d, H-1’), 5.31 (1H, dd, H-12), 5.78 (1H, d, H-11); 13C-NMR (100 MHz, DMSO-d6) δ: 38.2 (C-1), 25.5 (C-2), 80.3 (C-3), 43.0 (C-4), 46.5 (C-5), 17.2 (C-6), 31.1 (C-7), 41.7 (C-8), 52.5 (C-9), 35.8 (C-10), 131.9 (C-11), 130.7 (C-12), 83.4 (C-13), 45.2 (C-14), 35.4 (C-15), 63.1 (C-16), 46.3 (C-17), 51.7 (C-18), 37.5 (C-19), 31.5 (C-20), 34.3 (C-21), 25.2 (C-22), 63.2 (C-23), 12.8 (C-24), 18.6 (C-25), 19.8 (C-26), 20.7 (C-27), 71.4 (C-28), 33.8 (C-29), 24.1 (C-30), 105.2 (3-O-Fuc-C-1’), 72.2 (C-2’), 74.1 (C-3’), 71.6 (C-4’), 70.1 (C-5’), 17.0 (C-6’). These data were consistent with reported studies (Shi et al., 2020; Wang et al., 2021).
Discussion
PSF is a secondary triterpenoid glycoside of SSA and the latest research showed that PSF has an excellent anti-cancer effect on the cancer cells HCT 116 (Lee et al., 2022). Due to its trace amount in RB, it was hardly isolated from the raw material of RB directly. In the present study, we proposed to obtain PSF by enzymatic hydrolysis of SSA abundant in raw material for the first time. Snailase was selected due to its good hydrolysis performance under nearly neutral circumstances. What’s more, snailase is a complex of more than twenty enzymes, such as cellulase, β-D-glucosidase, amylase, and protease, which has gained increasing applications due to strong hydrolysis capability. As a natural enzyme, snailase also has some other advantages such as low cost and easy access. Furthermore, in our study, an optimal hydrolysis condition was determined for the eco-friendly and efficient preparation of PSF, which lays a firm foundation in industrial application. Enzymatic hydrolysis as green technology is becoming a hot area in natural products research nowadays. For instance, the content of compound K (C-K) is presently low in ginseng extracts, but it is better absorbed in the intestinal tract than other original ginsenosides. Hereby, some researchers made an effort to establish an efficient system for converting ginsenoside Rb1 to C-K by recombinant β-glucosidase hydrolysis (Kim et al., 2022).
Conclusion
In this study, a novel approach was developed for obtaining acid-sensitive PSF from SSA using snailase, by which the epoxy-ether ring of PSF remained stable as the process of enzymatic hydrolysis was performed at a high pH value. Moreover, the estimated model by BBD-RSM was able to well predict conditions for the highest conversion ratio. In conclusion, the newly developed approach was of good capacity to obtain anti-cancer PSF, and it is greatly feasible and has potential for industrial application.
Appendix
Experimental Parameters and Level for BBD.

Results of Precision Test.
Results of SSA Recovery (n = 3).
BBD and Results Obtained for Each Dependent Variable.
Footnotes
Summary
An appropriate approach was established for obtaining acid-sensitive prosaikogenin F from saikosaponin A for the first time. Enzymatic hydrolysis was employed and snailase was selected due to its good hydrolysis performance. Then, hydrolysis conditions were optimized by one-factor-at-a-time experimentation and RSM. The newly developed strategy lays a solid foundation for its development to be an anti-cancer new drug.
Abbreviations
RB: Radix Bupleuri; SSA: saikosaponin A; PSF: prosaikogenin F; RSM: response surface methodology; BBD: Box-Behnken design; HPLC-UV: high-performance liquid chromatography-ultraviolet; ACN: acetonitrile; MeOH: methanol; C-K: compound K.
Acknowledgments
Saikosaponin A was kindly provided by Chengdu Pufeide Biotechnology Co., Ltd. (City of Chengdu, Sichuan Province, PRC).
Authors’ Contributions
Yeting Zhu: Investigation, Formal analysis, Writing-original draft. Jiangsong Peng: Investigation, Formal analysis, Methodology. Yaqin Zhao: Investigation, Visualization, Validation. Mengru Wu: Methodology, Software. Suping Chen: Data curation, Validation. Jiali Shao: Validation, Formal analysis. Xubo Wang: Data curation. Guohua Xia: Conceptualization, Funding acquisition, Writing-review, and editing. Yuping Shen: Conceptualization, Funding acquisition, Supervision, Writing-review, and editing.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the National Natural Science Foundation of China (81873196), the Science and Technology Project of Changzhou City (CJ20220161), and the Science and Technology Planning Project of Zhenjiang City (SH2021057).
Statement of Informed Consent and Ethical Approval
Necessary ethical clearances and informed consent were received and obtained respectively before initiating the study from all participants.