Abstract
Immunoregulation and indoleamine 2,3-dioxygenase 1 (IDO1) play pivotal roles in the rejection of allogeneic organ transplantation. This study aims to elucidate the immune-related functional mechanisms of exosomes (Exos) derived from bone marrow–derived mesenchymal stem cells (BMSCs) overexpressing IDO1 in the context of allogeneic heart transplantation (HTx) rejection. A rat model of allogeneic HTx was established. Exos were extracted after transfection with oe-IDO1 and oe-NC from rat BMSCs. Exos were administered via the caudal vein for treatment. The survival of rats was analyzed, and reverse transcription qualitative PCR (RT-qPCR) and immunohistochemistry (IHC) were employed to detect the expression of related genes. Histopathological examination was conducted using hematoxylin and eosin (HE) staining, and flow cytometry was utilized to analyze T-cell apoptosis. Proteomics and RNA-seq analyses were performed on Exos. The data were subjected to functional enrichment analysis using the R language. A protein interaction network was constructed using the STRING database, and miRWalk, TargetScan, and miRDB databases predicted the target genes, differentially expressed miRNAs, and transcription factors (TFs). Exos from BMSCs overexpressing IDO1 prolonged the survival time of rats undergoing allogeneic HTx. These Exos reduced inflammatory cell infiltration, mitigated myocardial damage, induced CD4 T-cell apoptosis, and alleviated transplantation rejection. The correlation between Exos from BMSCs overexpressing IDO1 and immune regulation was profound. Notably, 13 immune-related differential proteins (Anxa1, Anxa2, C3, Ctsb, Hp, Il1rap, Ntn1, Ptx3, Thbs1, Hspa1b, Vegfc, Dcn, and Ptpn11) and 10 significantly different miRNAs were identified. Finally, six key immune proteins related to IDO1 were identified through common enrichment pathways, including Thbs1, Dcn, Ptpn11, Hspa1b, Il1rap, and Vegfc. Thirteen TFs of IDO1-related key miRNAs were obtained, and a TF-miRNA-mRNA-proteins regulatory network was constructed. Exosome miRNA derived from BMSCs overexpressing IDO1 may influence T-cell activation and regulate HTx rejection by interacting with mRNA.
Keywords
Introduction
With the progression of medical technology, organ transplantation has emerged as an efficacious therapeutic modality for addressing heart, kidney, and other organ failures. The principal impediment leading to the failure and mortality of organ transplantation is rejection 1 . Heart failure, stemming from cardiac dysfunction, stands as a chronic ailment with a heightened mortality rate. The pathogenic underpinnings of heart failure are intricate, encompassing factors such as hypertension, valvular and congenital heart diseases, and various risk factors 2 . Heart transplantation (HTx) stands as the exclusive recourse for end-stage heart failure treatment. However, it comes with the risk of rejection, which occurs when the recipient’s immune system recognizes the transplanted heart as foreign and mounts an immune response against it 3 . Allogeneic heart transplant rejection can be broadly categorized into acute rejection (AR) and chronic rejection 4 . AR is an inflammatory response in which T lymphocytes infiltrate the transplanted heart tissue. That triggers an immune response characterized by tissue damage, inflammation, and dysfunction of the transplanted heart 5 . While, chronic rejection typically occurs months to years after HTx and is a leading cause of long-term graft failure 6 . The main characteristic of chronic rejection is the development of a condition called chronic graft vasculopathy, which leads to narrowing due to the accumulation of immune cells, smooth muscle cells, and extracellular matrix within the vessel walls, leading to reduced blood flow and oxygen supply to the heart muscle 7 . As the incidence of heart failure escalates among patients, there is an increasing demand for HTx to augment both the duration and quality of life 8 . Nonetheless, the administration of immunosuppression consistently precipitates acute graft rejection in HTx 9 . Thus, it is imperative to scrutinize the molecular mechanisms contributing to the amelioration of HTx rejection.
Immunoregulation refers to the control and balance of the immune system to maintain homeostasis. It plays a crucial role in preventing excessive immune responses that can lead to harmful inflammation and tissue damage 10 . The innate immune response, instigated by inflammation, invariably induces organ dysfunction, thereby exacerbating allograft rejection 11 . Graft failure always leads death of HTx patients, while immune rejection is the main cause of graft failure, targeting the immune response is vital in HTx to recover myocardial injury 12 . Myocardial injury represents the foremost challenge in HTx, wherein injury to the myocardium, coupled with the activation of innate immunity, accentuates the impairment of heart function 13 . Consequently, the immune responses of T cells assume a pivotal role in HTx rejection. Indoleamine 2,3-dioxygenase (IDO) has been validated as an inducible enzyme with immune-regulatory activities, promoting tryptophan (Trp) catabolism 14 . Previous investigations have indicated that enhancing IDO activation can diminish Trp products, thereby attenuating T-cell activation and proliferation through the inhibition of dendritic cell (DC) function 15 . IDO has been affirmed to exert a protective function in organ transplantation by inhibiting rejection responses 16 . Studies demonstrated that the overexpression of IDO can safeguard lung allografts by suppressing T-cell responses and fortifying an antioxidant defense system 17 . In addition, Hill et al. 18 delineated the indispensable role of IDO expression in cardiac allograft rats governed by cytotoxic T lymphocyte–associated antigen 4 immunoglobulins (CTLA4Ig). CTLA4Ig is a vital factor in preventing allograft rejection and inducing tolerance 19 . IDO1 is primarily expressed in immune cells, such as DCs, macrophages, and monocytes. It plays a crucial role in immune tolerance and immune escape mechanisms by regulating the balance between immune activation and suppression20,21. Our previous research has also affirmed that exosomes (Exos) derived from overexpression IDO1 BMSCs enhance the immunotolerance of cardiac allografts 22 .
Bone marrow–derived mesenchymal stem cells (BMSCs) exhibit superior regulatory capabilities in immune responses, playing a pivotal role in medicinal therapies 23 . BMSC-secreted Exos play the main role in BMSC exerting therapeutic effects 24 , which contain various biologically active ingredient, that can interact with receptors to affect the signaling of target cells and are essential in immunomodulatory effects25,26. BMSC-derived Exos has been reported to exhibit inflammation regulation by inhibiting the nuclear factor κB (NF-κB) pathway and changing macrophages M2 polarization to exhibit immunomodulatory functions 27 . Notably, microRNAs (miRNAs) from BMSC paracrine factors, specifically Exos, hold promise in the treatment of cardiac injuries 28 . The confirmed potential of BMSC-derived Exos includes the promotion of cardiomyocyte proliferation, inhibition of reactive oxygen species (ROS), mitigation of cardiac injury, and preservation of cardiac function 29 . Exos, replete with active substances such as lipids, proteins, mRNA, long non-coding RNAs (LncRNAs), and miRNAs that convey information and regulate cellular functions30,31, assume a pivotal role in cardiac repair therapies 32 . LncRNA exerts function through sponging miRNAs to regulate the stability, translation, and translocation of mRNAs or is bound to proteins alone, for increasing the expression of target mRNA and involved in regulating transcriptional of nucleus and cytoplasm 33 . Several studies have underscored the significance of BMSC-derived Exos in conferring cardioprotection 34 . However, the mechanistic action of Exos derived from BMSCs overexpressing IDO1 in allogeneic HTx rejection remains elusive. This study aims to elucidate the immunomodulatory effects of miRNAs and proteins within Exos from BMSCs overexpressing IDO1, employing bioinformatics to unravel their role in allogeneic HTx rejection.
Materials and Methods
Animal and Experiment
Lewis and Wistar rats were procured from Hunan SJA Laboratory Animal Co., Ltd (No.99, Changchong Road, Longping Hi-Tech Park, Furong District, Changsha City, Hunan Province, China). The rats were maintained in a light-dark cycle at 22–24°C with ad libitum access to food and water, undergoing a 7-day adaptation period and a 12-hour fasting period prior to surgery. Wistar rats served as the donor group, while Lewis rats were designated as the recipients. BMSCs were extracted from the femur and tibia of Lewis rats, as per previously reported methods 35 . These cells were cultured in BMSC growth medium (Cyagen Biosciences, Sunnyvale, CA, USA) supplemented with 10% foetal bovine serum (FBS, Gibco, CA, USA), and P9 cells were selected for subsequent experiments.
Cell Transfection and Extraction of Exos
Overexpressed IDO1 lentivirus obtained from China Shanghai Genechem Co., Ltd. was used to transduce BMSCs via Lipofectamine 3000 reagent (Invitrogen Life Technologies, Carlsbad, CA, USA), following established procedures 22 . In short, mixed Lipofectamine 3000 reagent with overexpressed IDO1 lentivirus was in a transfection medium and incubated for 10 minutes at room temperature. After seeding the cells in the 6-hole plate under 5 × 104 cells/well and allowed to reach 60–80% confluency. Then, the transfection mixture was evenly applied dropwise to the cells, cultured in a 5% CO2 cell incubator at 37°C for 48 hours, and the transfection efficiency was detected by reverse transcription qualitative (RT-qPCR), shown in Supplemental Fig. 1.
After transfected BMSCs were collected, Exos were harvested using the Total Exosome Isolation Kit (4478359; Thermo Fisher Scientific, Waltham, MA, USA) in accordance with the manufacturer’s instructions. After processed BMSCs’ culture supernatant was collected, it was centrifuged at 2,000 × g for 30 minutes, followed by ultracentrifugation of 10,000 × g for 20 minutes for removing any cellular debris. Then, they were transferred to new tubes, and a Ribom™ Exosome isolation Reagent (Ribobio Co., Ltd, Guangzhou, China) (for cell culture media) was added and mixed well, at the 4°C refrigerator overnight. The next day, at a time, 2 mL was transferred to a new centrifuge tube, and centrifugation was performed at 15,000 × g for 30 minutes at 4°C repeatedly to extract all the mixed liquids, into pellet Exos. Subsequently, store the Exos. Transmission electron microscope (TEM, Hitachi, Japan, H7500) was employed to observe the exosome morphology, as shown in Supplemental Fig. 2.
Allogeneic HTx Model
All rats were anesthetized by isoflurane inhalation (1.5%–2%), as previously described 36 . First, after laparotomy, 50-U/L ice heparin saline solution (5 mL) at 4°C was injected through the inferior vena cava into Wistar rats. After 2 minutes, the abdominal aorta and inferior vena cava were cut for adequate bleeding. Subsequently, the thorax was opened, salt water for ice was cut, and a single 5-0 ligature was used to ligate the inferior and superior vena cava. The aortic and pulmonary arteries were cut, and the pulmonary vein of the heart was ligated and removed heart. Simultaneously, the abdomen of Lewis rats was opened, abdominal aorta and inferior vena cava were separated, and the heart from donor rats was implanted. The continuous single-point fitting method was employed to connect the aorta to the abdominal aorta and the pulmonary artery to the inferior vena cava. Analgesia was administered perioperatively using carprofen. Survival status was recorded daily, and mortality was calculated by Kaplan-Meier test. Exos (5 μg/mL) from each group were administered through the caudal vein 2 mL and categorized into the Model group (HTx rats), the BMSC group (HTx rats + Exos from BMSC), the NC group (HTx rats + Exos from transfected oe-NC BMSC), and the IDO1 group (HTx rats + Exos from transfected oe-IDO1 BMSC). The survival time of Lewis rats was analyzed using log-rank (Mantel-Cox) testing.
Reverse Transcription Qualitative PCR
Total RNA from rat heart tissues in each group was extracted using TRIzol reagent (Invitrogen, 15596026) and reverse transcribed to cDNA using the One Step Prime Script miRNA cDNA Synthesis Kit (Takara, Kyoto, Japan). RT-qPCR was performed using SYBR Green PCR Master Mix (Life Technologies, CA, USA) following the manufacturer’s protocols. The primers used for RT-qPCR are detailed in Table 1. The relative expression was calculated using the 2-ΔΔCt method, with GAPDH or U6 serving as the internal reference.
PCR Primer Sequence.

Histological Staining
HTx samples from each group were fixed in 4% paraformaldehyde and processed into paraffin slices. Sections were stained using the hematoxylin and eosin (HE) Stain Kit (G1120; Beijing Solebao Biotechnology Co., Ltd., China), and immunohistochemistry (IHC) was performed using the anti-IDO1 antibody (1:2,000; ab311847, Abcam, UK) to assess pathology and IDO1 expression.
Flow Cytometry
After concluding the experiment, spleens from each group were collected to isolate T lymphocytes. CD4+ T cells were extracted using the Stemcell 16052 StemSepTM Rat CD4+ T Cell Enrichment Kit (HY-001068; Shanghai Yes Service Biotech, Inc., China) following the product manual. The rate of programmed cell death was determined using the Annexin-V-FITC/PI apoptosis kit from Absin, a Chinese company, and assessed by FACScan flow cytometry.
Data Sources
Exos extracted through the methods outlined in “2.2 Cell Transfection and Extraction of Exosomes” were utilized as data sources. The BMSC group comprised Exos from normal BMSCs, the NC group included Exos from transfected oe-NC BMSC, and the IDO1 group involved Exos from transfected oe-IDO1 BMSC. Proteomics and RNA-seq analyses were conducted for further insights.
Extracted Protein and Trypsin Digestion
HTx samples from each group were collected, and Radio Immunoprecipitation Assay (RIPA) buffer (Sigma-Aldrich, St. Louis, MO, USA) with 1% protease and phosphatase inhibitor was added to extract total proteins. Following the previously mentioned method, the protein concentration was determined. A 100-mg protein solution underwent reduction with 90 μL of 10-mmol/L tris (2-carboxyethyl) phosphine (TCEP, Thermo Fisher) at 37°C for 30 minutes. Subsequently, 40 mmol/L iodoacetamide was added for alkylation in darkness for 40 minutes. Alkylated samples were then precipitated with cooled acetone at −20°C for 4 hours, followed by centrifugation at 10,000 × g/min at 4°C for 20 minutes. The protein samples were diluted in 50-mmol/L triethylammonium bicarbonate (TEAB), and trypsin/protein digestion was carried out twice 37 .
TMT Peptide Labeling and Mass Spectrometric Analysis
The three groups (BMSC group, NC group, and IDO1 group) were collected for proteomic analysis. Peptides were dissolved with 0.5m TEAB and labeled using the 6-label TMT kit (Thermo Fisher Scientific), following the kit’s instructions. Fractionation of peptides was performed using high-pH reverse high-performance liquid chromatography (HPLC) with an Agilent 300Extend C18 column (2.1 mm 100 mm, 3.5 μm, Agilent, USA). Peptide analysis was carried out using LTQ Orbitrap Elite, following the methods described by Stryiński et al. 38
Obtaining Differential miRNA by miRNA Chip
Total RNA from the three groups (BMSC group, NC group, and IDO1 group) was collected. The total RNA quality was assessed by formaldehyde denatured gel electrophoretic assay and purified using the mirVana purification miRNA Isolation Kit (AM1561). Dephosphorylation and labeling reactions were performed using the miRNA Complete Labeling and Hyb Kit from Agilent. After hybridisation, washing, and drying the slides, the hybrid image was obtained using the Agilent chip scanner (G2565CA). The Agilent Feature Extraction (v10.7) software was employed to analyze the hybrid images and extract the data.
Principal Component Analysis
Principal component analysis (PCA) represents a widely utilized algorithm for data dimensionality reduction. All samples from the BMSC group, NC group, and IDO1 group were included in the PCA analysis to visualize the degree of separation, which was accomplished with scatterplot3d package (v0.3-43) 39 .
Differentially Expressed Gene Analysis
Differentially expressed proteins (DEPs) between the IDO1 group and BMSC group (DEP1) and between the BMSC group and NC group (DEP2) were acquired using the limma
40
package (v3.44.3) with screening thresholds of|logFC| > 0.5 and
Obtained Immune-Related Genes
The study methods of Chen et al. 44 were referenced to obtain immune-related genes (IRGs) from the ImmPort and InnateDB databases, as detailed in Supplemental Table 1.
Functional Enrichment Analysis
The clusterProfiler
45
package (v1.58.0) was utilized for gene ontology (GO)
46
and Kyoto Encyclopedia of Genes and Genomes (KEGG)
47
enrichment analyses of DEPs and miRNAs, with significance determined at adj.
Protein Interaction Network
The protein-protein interaction (PPI) network was constructed using the STRING database (https://cn.string-db.org/) and visualized using Cytoscape (v 3.6.1) 48 , with subsequent screening of proteins for further analysis.
Target Gene Prediction and GeneMANIA Network Construction
Prediction of target genes for DEmiRNAs associated with IDO1 was performed using miRanda and miRDB databases. GeneMANIA was employed for predicting interactions, functional enrichment in networks, and pathways of key proteins related to IDO1. In addition, transcription factors (TFs) were predicted using the miRDB and miRWalk databases.
Statistical Analysis
All experimental data are presented as mean ± standard deviation (mean ± SD). GraphPad Prism 8 facilitated data analysis and plotting. T-tests were employed for comparisons between two groups, one-way analysis of variance (one-way ANOVA) for comparisons among groups, and two-way analysis of variance (two-way ANOVA) for pairwise comparisons between groups.
Results
Exos From Overexpression IDO1 BMSCs Can Effectively Relieve Allogeneic HTx Rejection in Rats
Initially, we established an allogeneic HTx model in rats, observing that the model group exhibited the shortest survival time after transplantation. While no significant difference was discerned between the BMSC group and NC group, the IDO1 group demonstrated a noteworthy extension in survival time (Fig. 1A). The expression of IDO1 was notably diminished in the model group, whereas the administration of Exos from BMSCs overexpressing IDO1 substantially elevated IDO1 expression, as validated by RT-qPCR and IHC (Fig. 1B, C). Histopathological analysis using HE staining revealed marked inflammatory cell infiltration, disorganization, diminished cardiomyocytes, and severe myocardial damage in the model group. The BMSC and NC groups exhibited similar outcomes, whereas the IDO1 group exhibited reduced inflammatory cell infiltration, a little vacuolation between myocardial cells and diminished myocardial damage (Fig. 1D). Flow cytometry analysis indicated a higher CD4 T-cell apoptosis rate in the BMSC group than the model group, with no significant difference from the NC group. Notably, the IDO1 group significantly promoted CD4 T-cell apoptosis (Fig. 1E). These findings collectively suggest that Exos derived from BMSCs overexpressing IDO1 enhance the survival time in allogeneic HTx in rats and mitigate rejection by inducing CD4 T-cell apoptosis.
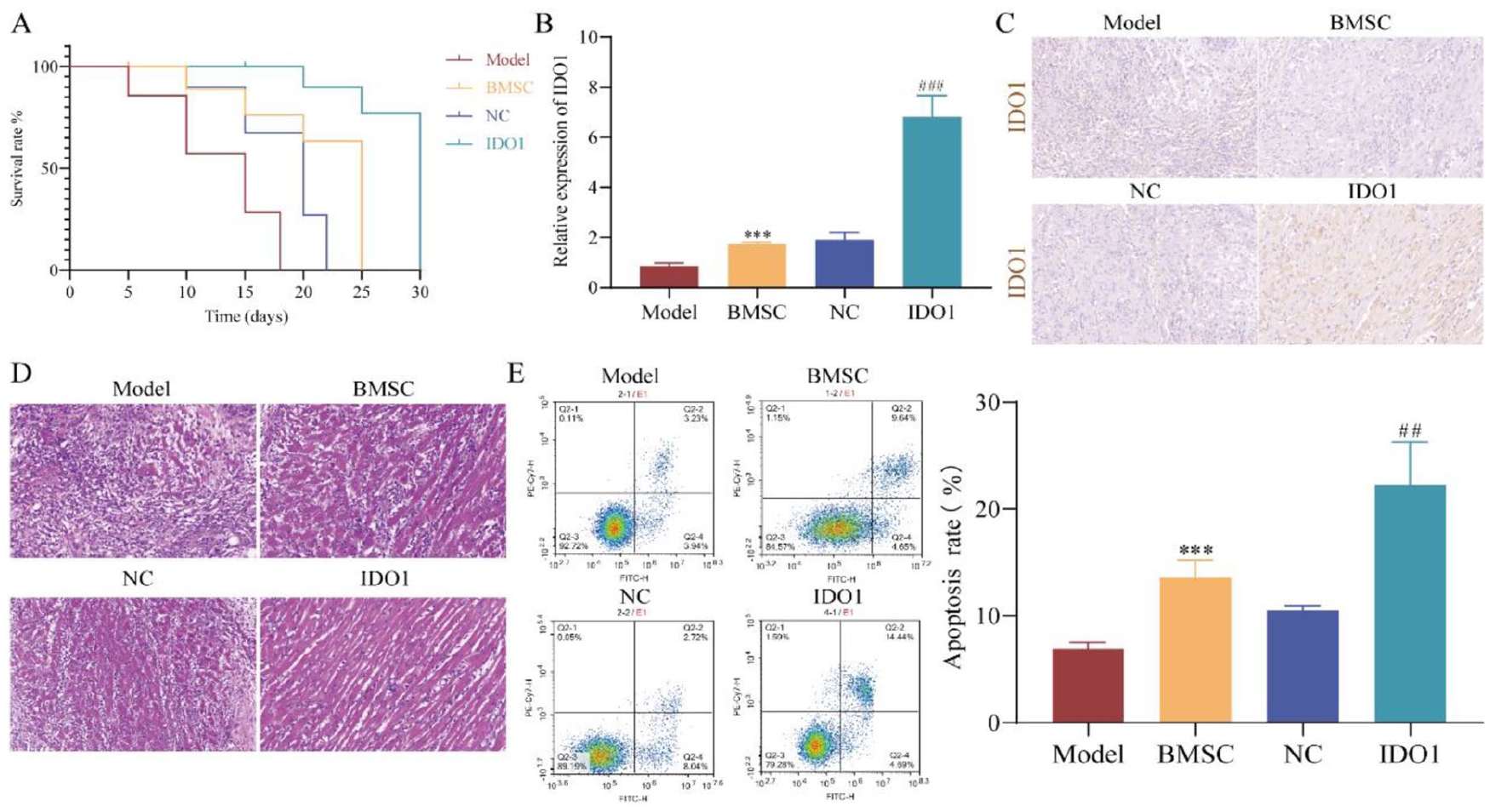
Exos from overexpression IDO1 BMSCs can effectively relieve allogeneic heart transplantation rejection in rats. (A) Kaplan-Meier analysis of the survival of rats. (B) RT-qPCR detected expression of IDO1. (C) IHC detected the expression of IDO1. (D) Histopathological examination by HE. (E) Flow cytometry analysis T-cell apoptosis. ***
Analysis of Immune-Related Differential Proteins in Exos of Rats BMSC
To elucidate the mechanism through which Exos from BMSCs overexpressing IDO1 alleviate HTx rejection in rats, we conducted proteomics analysis. PCA analysis revealed a distinct separation among the IDO1 group, BMSC group, and NC group, displaying no anomalies (Fig. 2A). Employing the limma package, we identified 169 differentially expressed proteins (DEP1) between the IDO1 and BMSC groups (Fig. 2B, C), as well as seven differentially expressed proteins (DEP2) between the NC and BMSC groups (Fig. 2D, E); detailed results are provided in Supplemental Table 2. Upon eliminating common proteins from DEP1 and DEP2, we obtained 162 DEPs (Fig. 2F), which intersected with 1,697 IRGs (Supplemental Table 1). Consequently, 13 immune-related differential proteins were identified as key proteins (Fig. 2G), encompassing C3, Ptx3, Hspa1b, Ntn1, Thbs1, Anxa2, Ctsb, Il1rap, Vegfc, Anxa1, Dcn, Ptpn11, and Hp.
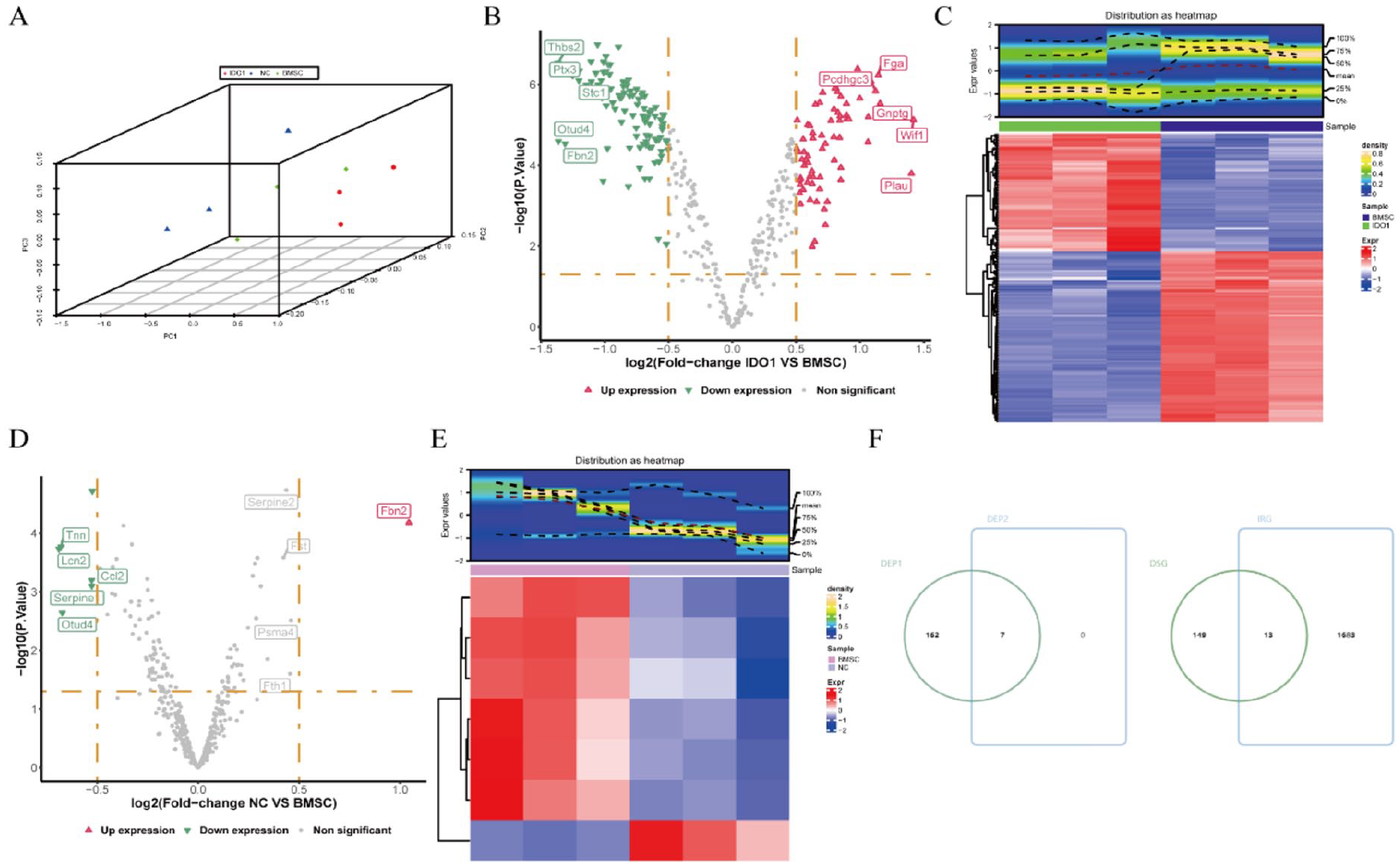
Analysis of immune-related differential proteins in exosomes of rats BMSC. (A) Proteomics dataset principal component analysis (PCA). (B and D) Volcano maps of differential protein expression: The x-axis depicts multiple log2FC of differential expression, while the y-axis represents −log10 (
Functional Enrichment Analysis and Expression Identification of Immune-Related Differential Proteins
Using the clusterProfiler package with an adjusted
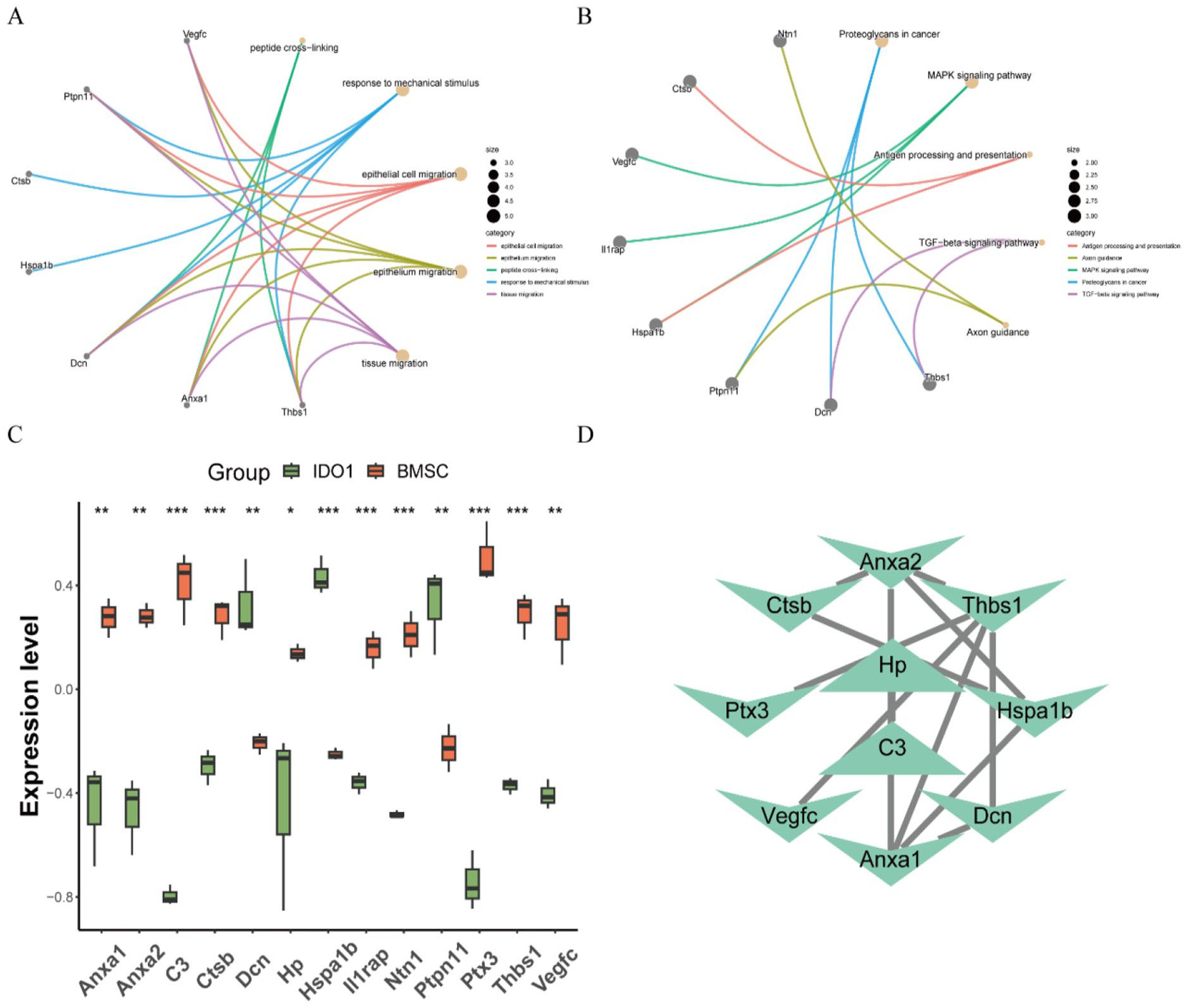
Functional enrichment analysis and expression identification of immune-related differential proteins. (A–B) GO enrichment and KEGG analysis of immune-related differential proteins. The connecting lines represent pathways and significantly enriched genes, with small dots on the left half denoting genes and large dots on the right half representing pathways. (C) Box plots of immune-related differential protein expression compared to the BMSC group. (*
In the KEGG pathway analysis, 10 functional pathways were enriched (Supplemental Table 4), encompassing proteoglycans in cancer, mitogen-activated protein kinase (MAPK) signaling pathway, antigen processing and presentation, transforming growth factor beta (TGF-beta) signaling pathway, and Axon guidance. The expression levels of immune-related differential proteins between the IDO1 and BMSC groups were evaluated using box diagrams, revealing the results shown in Fig. 3C. Relative to the BMSC group, the expression levels of Anxa1, Anxa2, C3, Ctsb, Hp, Il1rap, Ntn1, Ptx3, and Thbs1 were downregulated, while Hspa1b, Vegfc, Dcn, and Ptpn11 were upregulated. Utilizing the STRING database, a total of 10 nodes and 12 edges were obtained in the PPI network (Fig. 3D).
Analysis of Differential miRNAs in Rat BMSC Exos
We conducted a comprehensive analysis of differential miRNAs in Exos derived from BMSCs overexpressing IDO1. PCA analysis revealed distinct separation among the IDO1, BMSC, and NC groups, with no anomalies in the miRNA sequencing datasets (Fig. 4A). Using the DESeq2 package, we identified 425 differentially expressed miRNAs (DEmiRNA1) between the IDO1 and BMSC groups (Supplemental Table 5), as shown in Fig. 4B, C, and 380 differentially expressed miRNAs (DEmiRNA2) between the NC and BMSC groups (Supplemental Table 6), as depicted in Fig. 4D, E. Employing a significance threshold of
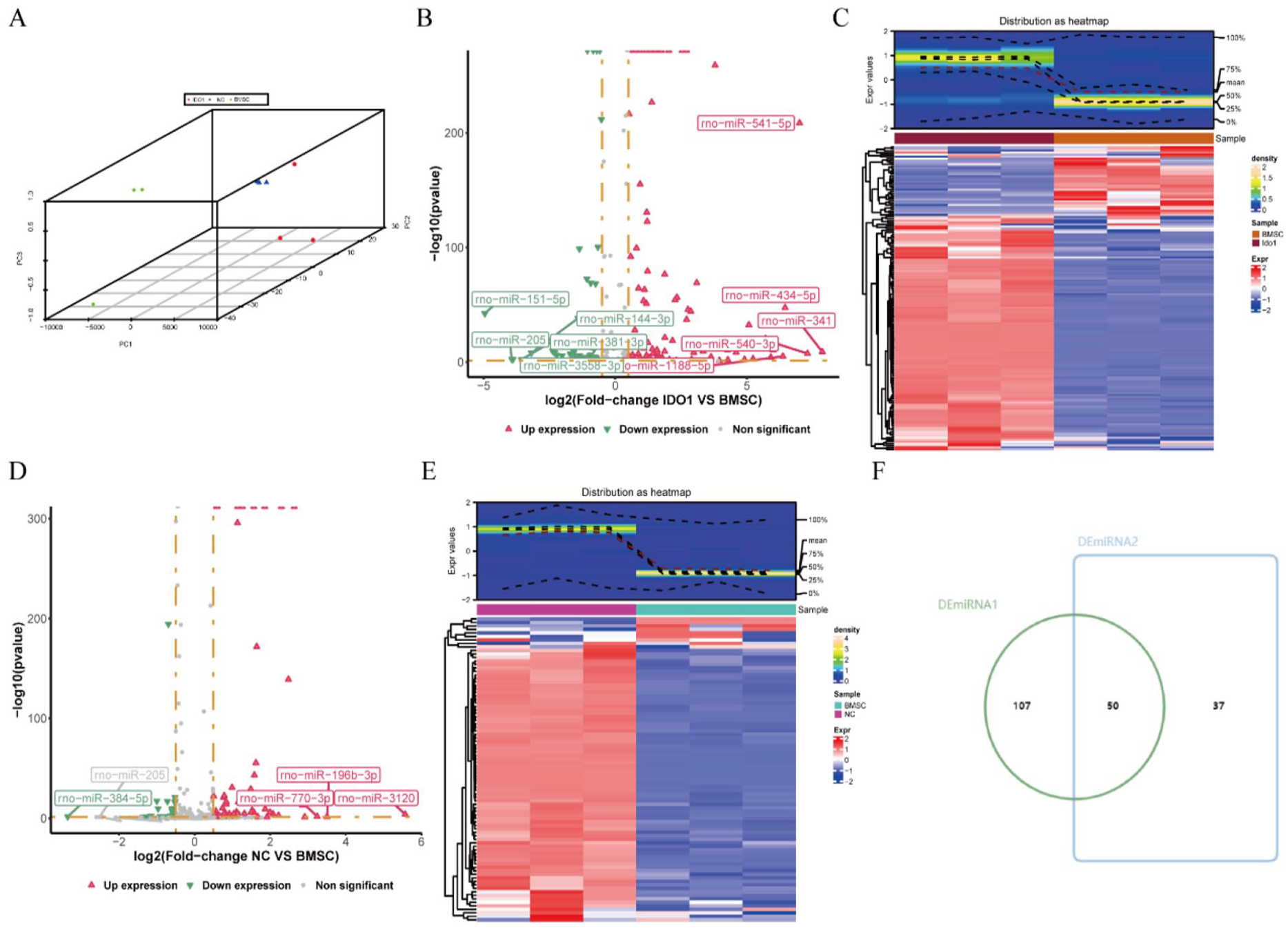
Analysis of differential miRNAs in exosomes of rat BMSCs. (A) PCA of miRNA-seq datasets. (B and D) Volcano plots of Differential miRNA expression. The abscissa represents the log2FC of differential expression, while the ordinate represents the −log10 (
Expression Identification, Target Gene Prediction, and Functional Enrichment Analysis of IDO1-Related Differential miRNAs
We assessed the expression levels of IDO1-related miRNAs between the IDO1 and BMSC groups in rat BMSC Exos using a box plot (Supplemental Table 10). Only miRNAs exhibiting significant differences were retained as IDO1-related differential miRNAs for subsequent analysis (Supplemental Table 11). The top 10 miRNAs are depicted in Fig. 5A, with all 10 showing downregulation compared to the BMSC group (Supplemental Table 12).
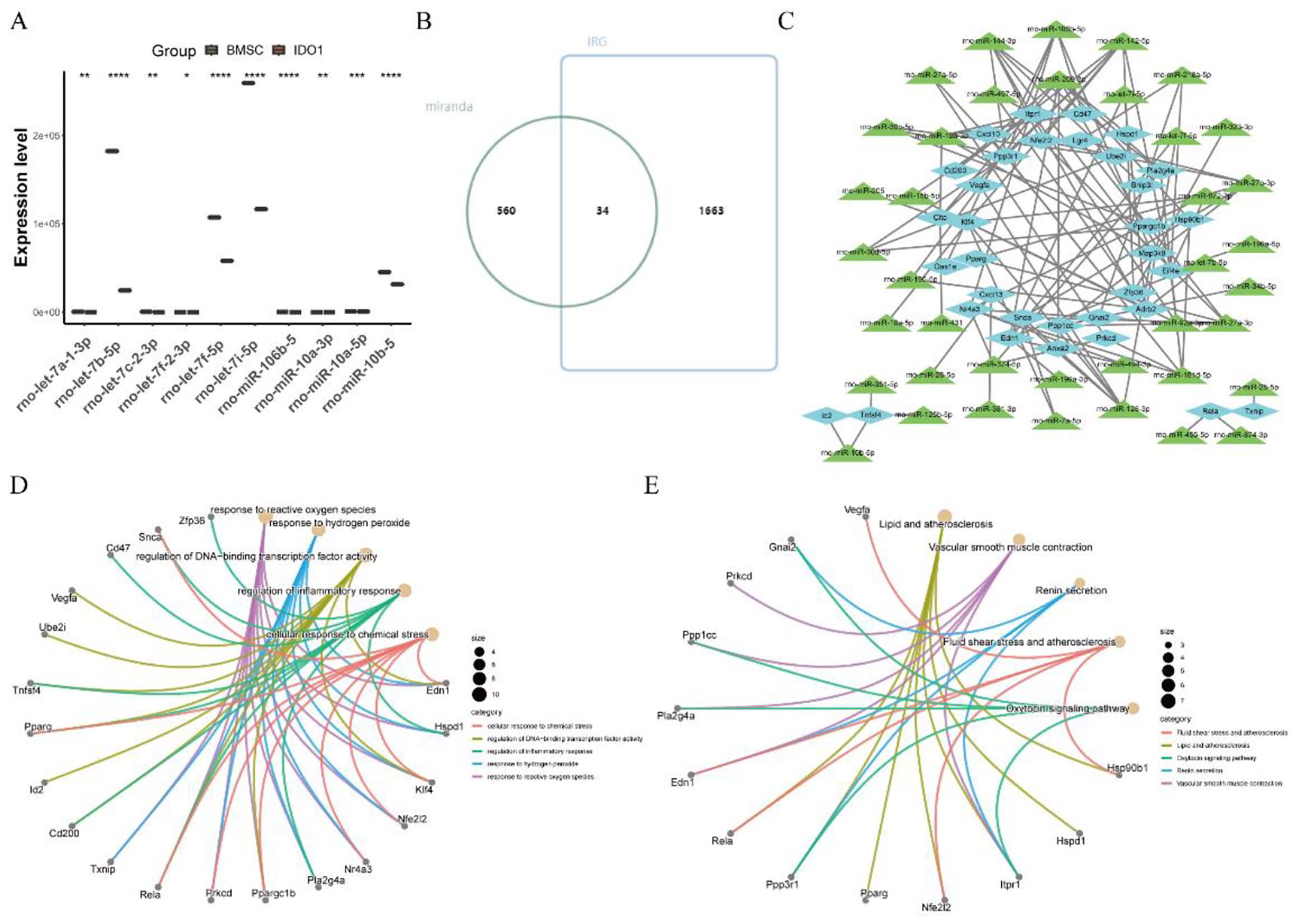
Expression identification, target gene prediction, and functional enrichment analysis of IDO1-related differential miRNAs. (A) Box plot illustrating the expression of IDO1-related differential miRNAs compared to the BMSC group.
Target genes (mRNAs) of IDO1-related differential miRNAs were predicted using miRanda and miRDB databases (Supplemental Table 13). The intersection of miranda and IRGs yielded 34 immune mRNAs related to IDO1 differential miRNAs (Supplemental Table 14), as illustrated in Fig. 5B. The target relationship between immune mRNAs of IDO1-related differential miRNAs and IDO1-related differential miRNAs is presented in Fig. 5C (see Supplemental Table 15 for detailed results).
Subsequently, KEGG and GO enrichment analyses were performed on these immune mRNAs of IDO1-related differential miRNAs. The top 5 results are shown in Fig. 5D, E. A total of 1,543 GO terms were obtained (Supplemental Table 16), encompassing 1,364 BP terms (e.g., response to ROS, response to hydrogen peroxide, regulation of DNA-binding TF activity, regulation of inflammatory response, and cellular response to chemical stress), 73 CC terms (e.g., secretory granule, protein serine/threonine phosphatase complex, phosphatase complex, midbody, and sperm plasma membrane), and 107 MF terms (e.g., transcription coregulator binding, RNA polymerase II-specific DNA-binding TF binding, cytokine activity, phosphatase binding, and transcription coactivator binding). In addition, 72 functional pathways were enriched in KEGG (Supplemental Table 17), including lipid and atherosclerosis, vascular smooth muscle contraction, renin secretion, fluid shear stress and atherosclerosis, and oxytocin signaling pathway.
Analysis and Identification of Key miRNAs and Key Proteins
Common enrichment pathways between IDO1-related immune proteins and immune mRNAs of IDO1-related miRNAs were selected as key pathways (Fig. 6A). These include proteoglycans in cancer, MAPK signaling pathway, Ras signaling pathway, and PI3K-Akt signaling pathway (Supplemental Table 18). Six IDO1-related immune proteins within these key pathways were identified as IDO1-related key proteins (Fig. 6B): Thbs1, Dcn, Ptpn11, Hspa1b, Il1rap, and Vegfc (Supplemental Table 19). Twenty-three IDO1-related miRNAs corresponding to immune mRNAs in key pathways were identified as key IDO1-related miRNAs (Fig. 6C): rno-miR-872-3p, rno-miR-142-5p, rno-miR-181d-5p, rno-miR-199a-5p, rno-miR-206-3p, rno-miR-218a-5p, rno-miR-455-5p, rno-miR-874-3p, rno-miR-128-3p, rno-miR-497-5p, rno-miR-92a-3p, rno-miR-15b-5p, rno-miR-195-5p, rno-miR-19b-3p, rno-miR-27a-3p, rno-miR-27b-3p, rno-miR-34b-5p, rno-miR-374-5p, rno-miR-144-3p, rno-miR-27a-5p, rno-miR-30b-5p, rno-miR-30d-5p, and rno-miR-106b-5p (Supplemental Table 20). Using GeneMANIA, we predicted the interaction and functional enrichment of IDO1-related key proteins in the network and pathways, revealing 20 interacting genes and 7 related pathways (Fig. 6D). We then predicted the TF of IDO1-related key miRNAs using miRDB and miRWalk databases, obtaining 13 TFs of IDO1-related key miRNAs. The regulatory network is shown in Fig. 6E (Supplemental Table 21), and the TF-miRNA-mRNA regulatory network (Supplemental Table 22) was constructed by combining the target relationships between IDO1-related immune mRNAs and miRNAs (Supplemental Table 15), as illustrated in Fig. 6F. Finally, we predicted the common miRNA of IDO1-related key proteins coding genes using miRWalk and miRDB databases (Supplemental Table 23) and visualized the network of key protein gene-miRNA relationships (Fig. 6G). The TF-miRNA-mRNA-proteins regulatory network was built by combining the TF-miRNA-mRNA regulatory network and the key protein gene-miRNA network (Fig. 6H).
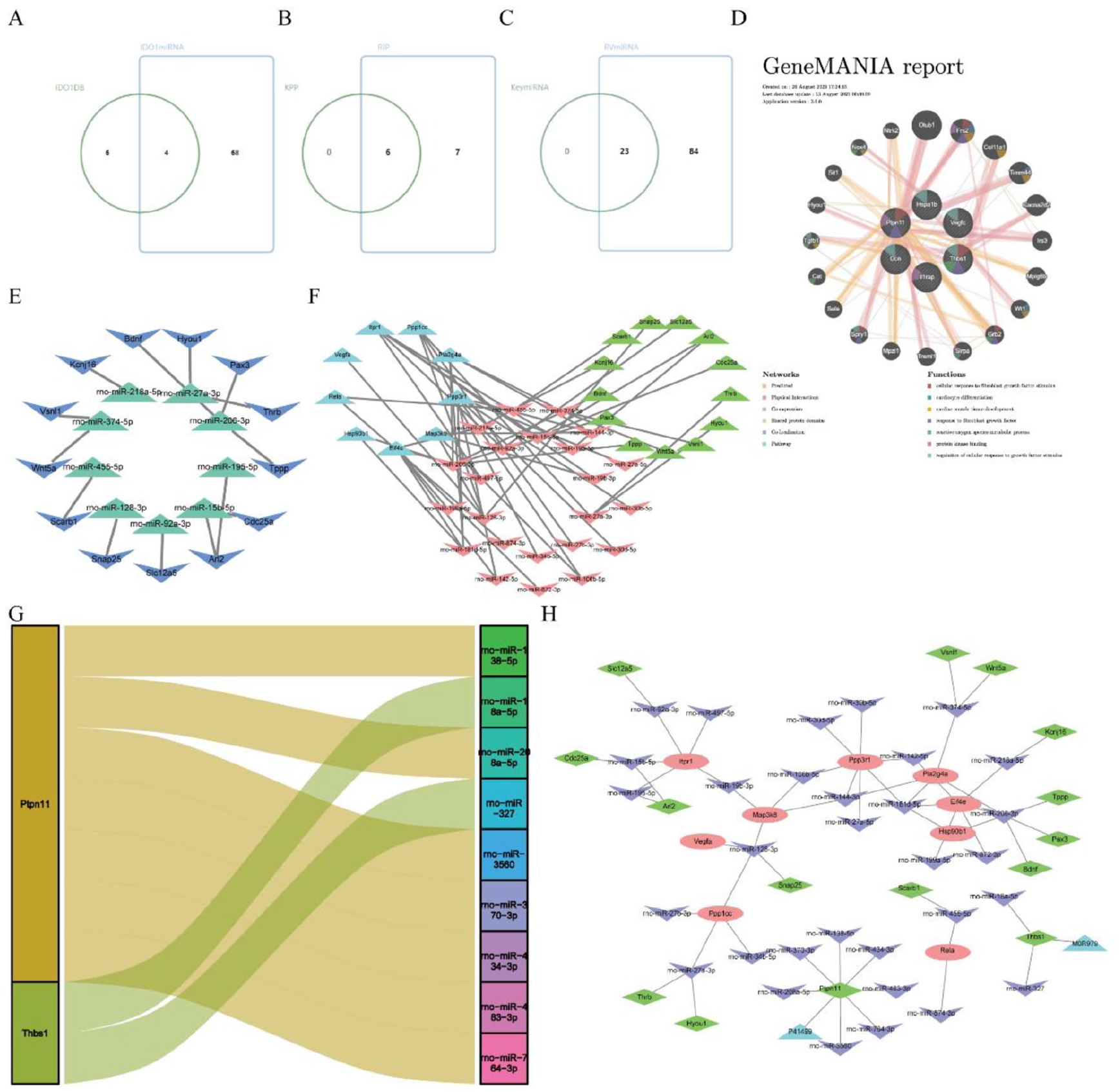
Analysis and identification of key miRNAs and key proteins. (A–C) Venn diagrams. (B) Venn diagram illustrating mRNA intersections of IDO1-related differential miRNAs. (D) IDO1-related key proteins GeneMANIA Network. (E) IDO1 TF-miRNA-mRNA network; green represents miRNA, and blue represents TF. (F) IDO1 TF-miRNA-mRNA network; red represents mRNA, green represents miRNA, and blue represents TF. (G) Sankey diagram of IDO1-related key protein networks. (H) Sankey diagram of IDO1-related key protein networks; red represents mRNA, purple represents miRNA, green represents TF, and blue represents protein.
Identification of the Expression Level of Key miRNAs and Key Proteins
Finally, we validated the expression level of key miRNAs and key proteins in the HTx rat tissues. Consistent with the aforementioned results, when compared to the BMSC group tissues, the expression of Anxa1, Anxa2, C3, Ctsb, Hp, Il1rap, Ntn1, Ptx3, and Thbs1 decreased, while Hspa1b, Vegfc, Dcn, and Ptpn11 increased in the IDO1 group tissues (Fig. 7A). In addition, the expression of 10 miRNAs decreased (Fig. 7B), including rno-let-7a-1-3p, rno-let-7b-5p, rno-let-7c-2-3p, rno-let-7f-2-3p, rno-let-7f-5p, rno-let-7i-5p, rno-miR-106b-5p, rno-miR-10a-3p, rno-miR-10a-5p, and rno-miR-10b-5p.
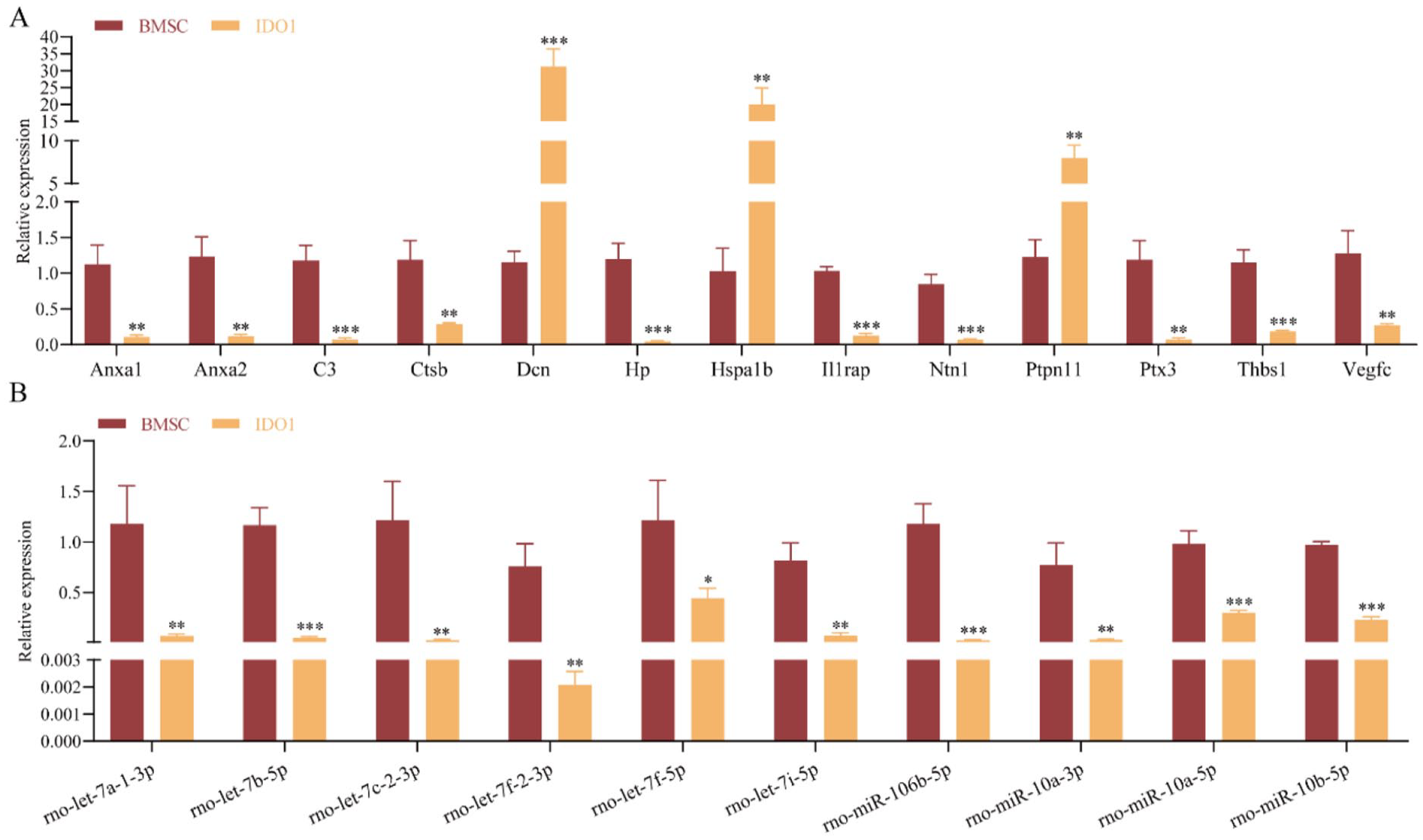
Identification of the expression level of the key miRNAs and key proteins. (A–B) RT-qPCR detected the expression level of the key proteins and key miRNAs. *
Discussion
IDO, an immunosuppressive enzyme, has been established to play a pivotal role in immunoregulatory mechanisms. IDO promotes apoptosis of T cells and natural killer (NK) cells by inhibiting the concentration of Trp 49 . Earlier studies have indicated that enhancing the expression of IDO can mitigate lung transplantation rejection and restore impaired T-cell activation 50 . Research by Li et al. 51 confirmed that IDO reduces rejection of heart-transplanted organs in mice, enhancing immune tolerance through interaction with Trp catabolic products (TC). IDO1 is crucial in various organ transplants. In transplant cells, IDO1 activated can inhibit transplant rejection and is the potential marker in rejection risk 52 . However, the mechanism of IDO1 in allogeneic HTx rejection is not fully understood yet. In our rat model of allogeneic HTx treated with Exos from transfected overexpression IDO1 BMSCs, we observed an improvement in the survival time and alleviation of transplantation rejection through the promotion of CD4 T-cell apoptosis. Consistent with our previous findings 22 . In this study, we delved into the immune-related functional mechanisms of Exos from overexpression IDO1 BMSCs in allogeneic HTx rejection.
Immunoregulation, a pivotal factor in allogeneic organ transplantation rejection, is crucial for enhancing immune tolerance and preventing rejection in organ transplantation recipients. Immunoregulation plays a key role in T-cell dysfunction, which often accompanies transplantation rejection 53 . In our investigation, we identified a strong correlation between Exos from overexpression IDO1 BMSCs and immune regulation in relieving HTx rejection in rats. We pinpointed 13 immune-related key proteins, such as C3, Ptx3, Hspa1b, Ntn1, Thbs1, Anxa2, Ctsb, Il1rap, Vegfc, Anxa1, Dcn, Ptpn11, and Hp. These proteins exhibit MFs like lipase inhibitor activity, calcium-dependent phospholipid binding, and phosphatidylserine binding. They are associated with the MAPK signaling pathway and the TGF-beta signaling pathway. Previous research has demonstrated that high IDO expression can inhibit T-cell receptor (TCR) activation and T-cell activity linked to calcium release and mitochondria 50 .
Exos, rich in proteins, mRNA, and miRNA, serve a crucial role in facilitating cell-to-cell communication 54 . Numerous studies have highlighted the significant role of miRNA derived from BMSC Exos in cardiac protection55,56. In this investigation, we identified DEmiRNAs in Exos from overexpression IDO1 BMSCs, including rno-let-7a-1-3p, rno-let-7b-5p, rno-let-7c-2-3p, rno-let-7f-2-3p, rno-let-7f-5p, rno-let-7i-5p, rno-miR-106b-5p, rno-miR-10a-3p, rno-miR-10a-5p, and rno-miR-10b-5p, with these 10 miRNAs exhibiting downregulation in IDO1 group tissues. Furthermore, we identified 34 immune mRNAs associated with IDO1-related differential miRNAs. MiRNAs, through complementary sequences, can bind to mRNA and play a pivotal role in cardiac remodeling and myocardial injury by regulating mRNA translation and degradation 57 . Novák et al. 58 confirmed the crucial role of miRNAs in the process of heart allograft transplantation rejection, demonstrating their potential diagnostic or predictive value in rejection outcomes. Research studies have reported that let-7b-5p and let-7c are abnormal expressions in acute cellular rejection (ACR) of liver transplantation patients 59 . Xu et al. 60 confirmed that miR-10a had a low expression of AR in lung transplant recipients (LTxR), the same as our results that rno-miR-10a-3p and rno-miR-10a-5p were downregulated. Subsequently, function enrichment analyses were performed on these immune mRNAs of IDO1-related differential miRNAs, finding that enrichment results mainly in the regulation of inflammatory response, lipid and atherosclerosis, TNF signaling pathway, MAPK signaling pathway, IL-17 signaling pathway, and so on. Previous research has discovered the relationship between organ transplantation and atherosclerosis, where organ transplantation leads the development 61 . TNF has been confirmed as a key factor that regulates cell inflammation response and mediates tumor rejection 62 . MAPK signaling pathway plays a crucial role in the activation of T cells and participates in the immune monitoring of kidney transplant patients 63 . The IL-17 signaling pathway is vital in the immune system. T helper cells produce IL-17, which is involved in the rejection of transplanted organs 64 . Therefore, immune mRNAs of IDO1-related differential miRNAs involved in regulating HTx rejection may be related to the aforementioned immunoregulation pathway, but the specific molecular mechanisms need to be further explored.
Through the identification of common enrichment pathways, we obtained 6 IDO1-related key immune proteins, namely Thbs1, Dcn, Ptpn11, Hspa1b, Il1rap, and Vegfc. Thbs1, a member of the thrombospondins (Thbs) family, has been linked to inducing lethal cardiac atrophy when highly expressed 65 . Previous studies have indicated that decorin (DCN) can alleviate cardiac dysfunction and inflammation, making it a potential treatment for diabetic cardiomyopathy 66 . Research by Sznajer et al. 67 established a connection between PTPN11 gene mutation and heart defects in 85% of patients. The study by Liu et al. 68 identified Hspa1b as a key gene associated with acute myocardial infarction (AMI). IL-1 receptor accessory protein (IL1RAP) is implicated in neutrophil activity and inflammation 69 . Vascular endothelial growth factor C (VEGFC) plays a vital role in cardioprotection, inhibiting proinflammatory macrophage activation and alleviating cardiac injury and inflammation 70 . Consequently, we hypothesize that Exos from overexpression of IDO1 BMSCs alleviates HTx rejection through the influence of these key immune proteins. Subsequently, we predicted the interaction and functional enrichment of IDO1-related key proteins in networks and pathways, predicted the TFs, and constructed the TF-miRNA-mRNA regulatory network and TF-miRNA-mRNA-proteins regulatory network.
In conclusion, our study demonstrates that Exos from overexpression IDO1 BMSCs play a crucial role in HTx rejection, particularly through their association with immune proteins. The exosome miRNA derived from overexpression IDO1 BMSCs may impact T-cell activation and regulate HTx rejection by interacting with mRNA. The 6 IDO1-related key immune proteins (Thbs1, Dcn, Ptpn11, Hspa1b, Il1rap, and Vegfc) emerge as pivotal players in transplantation rejection and hold promise as potential biomarkers in transplant rejection.
Supplemental Material
sj-docx-1-cll-10.1177_09636897241245796 – Supplemental material for Mechanism Exploration on the Immunoregulation of Allogeneic Heart Transplantation Rejection in Rats With Exosome miRNA and Proteins From Overexpressed IDO1 BMSCs
Supplemental material, sj-docx-1-cll-10.1177_09636897241245796 for Mechanism Exploration on the Immunoregulation of Allogeneic Heart Transplantation Rejection in Rats With Exosome miRNA and Proteins From Overexpressed IDO1 BMSCs by Rui Zheng, Xinxin Wu, Si Li, Xinhao Chen, Dan Yan and Jigang He in Cell Transplantation
Footnotes
Acknowledgements
The authors acknowledge ImmPort, InnateDB, miRDB, miRWalk, Uniprot, GSEA, GEO, GO, KEGG, STRING, MSigDB, miranda, and mirdb database for providing their platforms and contributors for uploading their meaningful datasets. The authors thank their colleagues for helpful suggestions.
Author Contributions
Conceived and designed the experiments: Jigang He and Rui Zheng; performed the experiments: Jigang He, Xinxin Wu, and Xinhao Chen; analyzed the data: Dan Yan; contributed reagents/materials/analysis tools: Xinhao Chen and Si Li; writing–original draft: Xinxin Wu and Jigang He; writing–review & editing: Jigang He and Rui Zheng.
Data Availability Statement
Data are applicable after the approval of co-authors.
Ethical Approval
With the assistance of Yunnan Labreal Biotech Co., Ltd., the animal experiment was conducted and received approval from the Experimental Animal Ethics Committee of Yunnan Labreal Biotech Co., Ltd (IACUC Issue No. PZ20221110).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: National Natural Science Foundation of China (no.: 82060299); Medical Discipline Leader Project of Yunnan Provincial Health Commission (no.: D-2019020); Yunnan Provincial Government Ten Thousand Person-Top Young Talents Project (no.: KH-SWR-QNBJ-2019-002); Clinical Medical Center of the First People’s Hospital of Yunnan Province (no.: 2021LCZXXF-XZ04); Kunming Medical Joint Special Project-Outstanding Youth Cultivation Project (no.: 202101AY070001-034); Clinical Medical Center of the First People’s Hospital of Yunnan Province (no.: 2022LCZXKF-HX05); Kunming Medical joint special project (no.: 202101AY070001-272); Famous Doctor Project of “Xingdian Talent Support Plan” of Yunnan Province (no.: XDYC-MY-2022-0037); Yunnan Province 2023 Undergraduate Education and Teaching Reform Research Project (2023BKXJJG-F04002).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
