Abstract
The tetrapod salamander species axolotl (Ambystoma mexicanum) is capable of regenerating injured brain. For better understanding the mechanisms of brain regeneration, it is very necessary to establish a rapid and efficient gain-of-function and loss-of-function approaches to study gene function in the axolotl brain. Here, we establish and optimize an electroporation-based method to overexpress or knockout/knockdown target gene in ependymal glial cells (EGCs) in the axolotl telencephalon. By orientating the electrodes, we were able to achieve specific expression of EGFP in EGCs located in dorsal, ventral, medial, or lateral ventricular zones. We then studied the role of Cdc42 in brain regeneration by introducing Cdc42 into EGCs through electroporation, followed by brain injury. Our findings showed that overexpression of Cdc42 in EGCs did not significantly affect EGC proliferation and production of newly born neurons, but it disrupted their apical polarity, as indicated by the loss of the ZO-1 tight junction marker. This disruption led to a ventricular accumulation of newly born neurons, which are failed to migrate into the neuronal layer where they could mature, thus resulted in a delayed brain regeneration phenotype. Furthermore, when electroporating CAS9-gRNA protein complexes against TnC (Tenascin-C) into EGCs of the brain, we achieved an efficient knockdown of TnC. In the electroporation-targeted area, TnC expression is dramatically reduced at both mRNA and protein levels. Overall, this study established a rapid and efficient electroporation-based gene manipulation approach allowing for investigation of gene function in the process of axolotl brain regeneration.
Introduction
The brain is established during embryonic development and maintained at a homeostasis state in adulthood. Once the adult mammalian brain including the human brain is challenged with injuries or suffers from diseases, such as Alzheimer’s disease, it is incapable of spontaneously restoring the lost cell types and damaged structures to recover function 1 . And a clinical therapeutic intervention for repairing brain injuries is missing, largely due to the lack of enough understanding of brain regeneration. Unlike mammals, salamanders have great power to regenerate injured brains2–4. Previous studies showed that, upon physical injuries or chemical-induced neuronal ablation, salamanders could rather precisely reproduce the lost cell types or structures2,5–7. In this process, a specific population of cells called ependymal glial cells (EGCs) plays a crucial role. EGCs are located within the ventricular zone of the brain, where they extend radially oriented processes 8 . They are characterized by certain molecular markers, such as SOX2, Nestin, and GFAP (glial fibrillary acidic protein)8,9. Upon injury, those EGCs that stay quiescent during homeostasis undergo activation and transit into a proliferative state, participating in the process of regeneration6,7,10. Moreover, using single-cell sequencing and spatial transcriptome techniques, both key EGCs and differentially expressed genes (DEGs) involved in axolotl brain regeneration have been identified6,7. However, key questions such as how EGCs are activated from a homeostatic to a regeneration state, what is the crucial molecule mediating such conversion, and what are the roles of DEGs in brain regeneration remain unresolved. Answering the above questions will facilitate an understanding of the mechanisms of brain regeneration and provide cues for improving brain regeneration in higher vertebrate species. In order to address these issues, establishing an efficient and rapid gene functional analysis approach in salamander, which is still missing, is very crucial to push the development of the field of brain regeneration molecular mechanism research.
Gain-of-function and loss-of-function studies are the mostly used method for gene functional analysis. Since the first transgenic line was established in axolotls 11 , Isopropyl-β-D-Thiogalactopyranoside (IPTG)-LacI-based and Cre-LoxP-based conditional gene overexpression systems were soon adapted to study the role of targeted genes in axolotl tissue regeneration12,13. By constitutive or conditional overexpressing targeted genes, the roles of thrombospondin-4 and p16INK4a were investigated in spinal cord and limb regeneration12,13. However, transgenesis-based conditional gene overexpression requires the generation of transgenic animals, which can be time-consuming and is limited by the lack of well-characterized tissue-specific promoters14,15. In terms of knockout, it has been reported that introducing TALEN or CRISPR/Cas9 into fertilized eggs could efficiently lead to constitutive knockout/knockdown of targeted genes in axolotls16,17. Yet complete loss of some genes, particularly the essential genes playing key roles at early developmental stages, potentially causes embryonic lethality and makes it difficult to study their function in regeneration.
To overcome the drawbacks of transgenesis, many transient gene delivery methods, such as virus transfection and plasmid/protein electroporation, have been developed for target gene manipulation in axolotls. There are two different types of viruses applied previously in axolotls, retrovirus and baculovirus. Retrovirus including Foamy virus and MLMV virus can integrate into the genome and then drive the target gene expression18,19, but baculoviruses maintain an episomal format to function 20 . Preparation of virus is relatively easy and fast, but the size of exogenous DNA for insertion is generally restricted. Plasmid electroporation offers an alternative method for gene overexpression in vivo. This approach involves introducing plasmids into the region of interest of an axolotl tissue, such as the central lumen of the spinal cord or the extracellular space of a solid tissue like limb, followed by electroporation21–24. The electric current from electroporation creates hole on the cell membrane of the adjacent cells and leads to the transport of the plasmid into these cells. Ultimately, the gene of interest located on the plasmid is expressed in the targeted cells 25 . Currently, virus transfection and plasmid electroporation have only been utilized for overexpressing target genes in axolotls, as there is a lack of appropriate RNA polymerase III promoters for driving gRNA expression 26 . As a result, these methods have not yet been applied for gene knockout or knockdown purposes. However, a recent study bypassed this bottleneck. By electroporation of protein, specifically CAS9-gRNA complexes to desired tissues, it has been shown that targeted genes could be successfully knocked out in the spinal cord and limb23,24.
Virus- and electroporation-mediated transient gene functional analysis has been broadly used in the spinal cord, the limb, and several other tissues in the axolotls27–29. However, for the brain of the salamanders, only labeling of EGCs by electroporation of a fluorescence protein coding plasmid has been reported, for the purpose of tracing the cell fate of EGCs7,10. Neither targeted gene knockout nor gene overexpression-based functional assay is established for transiently and rapidly investigating the role of defined genes during axolotl brain development or regeneration.
In this study, we optimized the electroporation condition, particularly the voltage, to achieve better brain transfection efficiency. Previous works showed that EGCs distributed in different regions express unique molecular markers and exhibit varied features, such as fast dividing, and quiescence3,6,7. In order to study the function of EGCs in diverse brain domains, we specifically targeted EGCs located in each domain of the brain by orientating the electrodes to specific angles. We then investigated the phenotype of brain regeneration upon overexpression of candidate gene. We chose Cdc42, a gene that has been previously reported to be involved in controlling the polarity of neural stem cells during brain development in other species30,31. We found that maturation and migration of newly born neurons, derived from Cdc42 overexpressing EGCs, were dramatically affected, which led to a delay of brain regeneration in axolotls. In addition, we also succeeded in knockout/knockdown of Tenascin-C (TnC) gene in the regenerating axolotl brain by electroporating CAS9 protein-gRNA complexes.
Materials and Methods
Animals
The d/d strain Ambystoma mexicanum used in this study was obtained from Elly M. Tanaka laboratory (Research Institute of Molecular Pathology, Vienna Biocenter, Vienna, Austria). Animals were housed and bred in dechlorinated tap water at 18°C–20°C with a 12-h light/12-h dark cycle 32 . All animals had been deeply anesthetized in 0.03% benzocaine (E1501, Sigma-Aldrich, St. Louis, MO, USA) before performing brain electroporation, brain injury, and harvest. Animal care was in accordance with the animal welfare legislation in China, and all animal experiments in this study were approved by the Biomedical Research Ethics Committee of Guangdong Provincial People Hospital (license number: KY2023-447-01).
Plasmids Construction
The reporter plasmid pCAGGs: EGFP was a kind gift from Elly M. Tanaka 11 . To construct pCAGGs: Cdc42 plasmid, Cdc42 coding sequence was PCR-amplified from axolotl brain cDNA with primer pair Cdc42-Fw (5′-ATGCAGACAATTAAATGTGTAGTTGTTGGG-3′) and Cdc42-Rev (5′-TCATAGCAGCACACACTTGCG-3′). Cdc42 PCR product was subcloned into pCAGGs vector. To construct pCAGGs: null plasmid for control, Cdc42 coding sequence was deleted from plasmid pCAGGs: Cdc42. All plasmids were verified by sequencing and purified with Endotoxin-free kit (12162, Qiagen, Hilden, Germany) following the manufacturer’s instructions.
gRNA
The TnC-gRNA1 (targeting site: 5′-GCACGAGCATCAGGCATGTG-3′), TnC-gRNA2 (targeting site: 5′-GCTCGTGCTGCTATGCCACC-3′), TnC-gRNA3 (targeting site: 5′-CACACCGCATCAACATCCCA-3′), TnC-gRNA4 (targeting site: 5′-TGATGAAGGCTTCACCGGCG-3′), TnC-gRNA5 (targeting site: 5′-CGATGAGGGATTCACCGGAG-3′), and Tyr-gRNA (targeting site: 5′- GGACTTCACTATCCCCTACT-3′, as control) 16 were synthesis from Genscript (Piscataway, NJ, USA).
Brain Electroporation
For the electroporation experiment, we diluted Gene-of-interest (GOI) overexpression plasmid’s concentration to 2 μg/μl and reporter plasmid’s concentration to 0.5 μg/μl in 0.8× phosphate-buffered saline (PBS). For knockout/knockdown electroporation, we mixed 1 μg/μl CAS9-NLS protein (5 mg/ml, MPI-CBG protein expression facility) and 2 μg/μl gRNA (in the case of TnC gRNA mix1, containing 1 μg/μl of TnC-gRNA 1, 2 respectively, and TnC gRNA mix2 containing 0.66 μg/μl of TnC-gRNA 3, 4, 5 respectively), followed by 5 min room temperature incubation for CAS9–gRNA complex formation. Fast Green (68724, Sigma-Aldrich, St. Louis, MO, USA) was added into the mixture before injection, serving as an injection accuracy indicator.
Electroporation was performed with the electroporator (NEPA21, Nepagene, Ichikawa, Japan). To carry out telencephalon electroporation, axolotls were anesthetized in 0.03% benzocaine. Using scalpels and spring scissors (15003-08, Fine Science Tools, Foster City, CA, USA), a rectangular scalp flap was created on the top of the head in each animal, in order to expose the left skull. Then, for animals less than 7 cm in body length from snout to tail tip, perforation was made on the rostral of exposed skull with syringe needles; for animals larger than 7 cm, a cranial flap was made on left skull to expose the left telencephalon. After loading plasmid mixture or CAS9-gRNA RNP mix into a microinjection glass capillary, which was connected to an aspirator tube assembly ready for injection, electroporation is performed as described in Fig. 2A. In brief, the capillary was inserted into the front tip of left lateral ventricle from the site of the olfactory bulb and approximately 2 μl of mixture was injected. Upon completion of the injection, animals were immersed into pre-chilled 1× PBS and electroporated using a pair of properly sized plate electrodes. Electroporation parameters were set according to animals’ size. (For example, for a 10-cm axolotl dorsal brain electroporation, we used electrodes 10 mm in diameter. Poring pulse was set to one unipolar pulse of 80 V for 5 ms duration and 50 ms interval. Transfer pulses were set to 5-time unipolar pulses of 60 V for 50 ms duration and 999 ms interval with 10% decay.) Afterward, cranial flap (for animals larger than 7 cm) and scalp flap were replaced and the animals were returned to dechlorinated tap water for recovery.
Brain Injury
Brain injury was executed in brain-electroporated animals as described previously 6 . In brief, anesthetized animal’s scalp flap and cranial flap were reopened to expose the left telencephalon. Assisted by fluorescence microscope (SZX16, Olympus, Shinjuku City, Japan), an approximately 0.25 mm × 0.25 mm square-shaped incision was made within the electroporation-positive region of the left telencephalon with spring scissors. The incision tissue was carefully removed with microsurgical forceps (11252-00, Fine Science Tools, Foster City, CA, USA). Afterward, cranial flap and scalp flap were replaced and the animals were returned to dechlorinated tap water for recovery.
Hematoxylin and Eosin Staining
Hematoxylin and eosin staining was performed on 16 μm brain coronal cryosections. Air-dried sections were rehydrated in ethanol (100%, 90%, 80%, 70%, 50%, ddH2O) for 5 min in each gradient. Then, the sections were immersed in hematoxylin to stain nuclei, with running tap water washed 5 times afterward. For differentiation, sections were dipped into 1% HCl in 70% ethanol and were immersed in tap water right away when the section turned red or pink. After bluing by ddH2O, sections were incubated in 60%, 70%, and 90% ethanol for 2 min consecutively. Following eosin staining, sections were incubated in 90% ethanol for 1 min, 100% ethanol twice for 1 min each, and xylene twice for 1 min each. Slides were mounted in neutral balsam and preserved at room temperature.
BrdU Injection
BrdU (B9285, Sigma-Aldrich, St. Louis, MO, USA) was dissolved in sterile 0.8× PBS to a stock concentration of 10 mg/ml. Anesthetized animals were performed intraperitoneal injection with a calculated BrdU volume (50 μg BrdU per gram of body weight).
Immunofluorescence Staining
Immunostaining analyses were carried out on 16 μm brain coronal cryosection. The harvested samples were immersed in MOPS/EGTA/Magnesium Sulfate/Formaldehyde Buffer (MEMFA) overnight at 4°C, then rinsed with 1× PBS and cryopreserved in 30% sucrose in 1× PBS at 4°C. Samples were snap-frozen in Tissue-Tek (4583, Sakura, Torrance, CA, USA) and stored at −80°C. Cryosections were preserved at −20°C.
For BrdU staining, sections were treated with 2M HCl at 37°C for 10 min to denature the DNA. They were then washed five times with 1× PBS for 20 minutes each and blocked with 5% donkey serum in 1× PBS/0.3% Triton X-100. Afterward, immunofluorescence staining was performed following the standard protocol. The following primary antibodies were used against the corresponding antigens: SOX2 (1:500, ab97959, Abcam, Cambridge, UK), CDC42(1:200, sc-8401, Santa cruz, Santa cruz, CA, USA), NEUN (1:500, MAB377, Sigma, St. Louis, MO, USA), TUJ1 (1:500, MAB1195, R&D system, Minneapolis, MN, USA), ZO-1 (1:500, 33-9100, Invitrogen, Waltham, MA, USA), BrdU (1:500, MA3-071, Thermo Fisher Scientific, Waltham, MA, USA), MCM2 (1:300, ab4461, Abcam, Cambridge, UK), and TNC (1:100, MT1, DSHB, Iowa, IA, USA).
gRNA Evaluation
In order to evaluate the gRNA efficacy for in vivo genome modification, TnC gRNAs were electroporated into spinal cord and followed by genotyping as previously described 24 . Briefly, CAS9-gRNA complexes were microinjected into the central canal of the spinal cord, followed by the electroporation procedure. Three days after electroporation, the electroporated segments of the spinal cord were dissected and genomic DNA was extracted using alkaline lysis method, followed by genotyping PCR and sequencing. The sequencing results were then analyzed using ICE Analysis (https://ice.synthego.com/) 33 .
Stereo-Seq Maps and In Situ Hybridization
The TnC spatial visualization stereo-seq maps were adapted from the interactive data portal at https://db.cngb.org/stomics/artista 6 .
Cdc42 CDS and TnC 3′-UTR sequence were PCR-amplified from axolotl brain tissues cDNA by the primer pairs, respectively: TnC-Fw (5′-GGCTACCACAGCTGTGGGATCTCCA-3′) and TnC-Rev (5′-GAAGGGCTTGGGAGCAGTCCTTGG-3′), Cdc42-Fw (5′-ATGCAGACAATTAAATGTGTAGTTGTTGGG-3′) and Cdc42-Rev (5′-TCATAGCAGCACACACTTGCG-3′). The PCR product was verified by sequencing and added T7 promoter by PCR amplification for the template of antisense RNA probe as previously described6,34.
In situ hybridization (ISH) was performed on 16 μm brain coronal cryosections as previously described6,34.
Statistical Analysis
For each sample, six to eight adjacent sections were selected for statistical analysis. The area adjacent to the lesion refers to the region within 500 µm of the wound. The ventricular zone was defined as the layered area where Sox2-positive cells are distributed along the ventricles, including the part protruding into the ventricle. To confirm the identity, condition, and location of EGFP-positive cells, we performed immunostaining with BrdU, MCM2, SOX2, TUJ1, and NEUN and labeled the cell nuclei with 4′6-diamdino-2-pheynlin-dole (DAPI). The exposure time for image acquisition was consistent between the experimental and control groups. It should be noted that cell was considered TUJ1-positive cell when the TUJ1 immunostaining clearly surrounded the DAPI-positive nucleus for counting. We counted the number of cells that co-expressed EGFP and these markers for statistical analysis. Statistical analysis was performed using GraphPad prism9.5 software (GraphPad Software, San Diego, CA, USA). All statistical results are represented as the mean ± standard error of the mean (SEM). Statistical significance was analyzed by the unpaired student’s t-tests. Values were considered statistically significant when P < 0.05. For all graphs, ** indicates P < 0.01, *** indicates P < 0.001, **** indicates P < 0.0001.
Results
Target EGCs at Different Ventricular Domain Via Orientated Electroporation
The axolotl telencephalon sits in between the olfactory bulb and the mesencephalon. It possesses a central luminal structure spanning the entire brain and the spinal cord (Fig. 1A). In the telencephalon region, the central lumen is surrounded by nuclei-enriched gray matter, followed by peripheral, TUJ1-stained white matter (Fig. 1B). Luminal-adjacent neural stem cells express transcription factor SOX2, and a proportion of which is capable of incorporating pulse-labeled BrdU (Fig. 1B), meaning in cycling status during homeostasis. Neurons in the telencephalon are positive for pan-neuronal markers TUJ1 and NEUN (Fig. 1B, C), markers labeling newly born and mature neurons, respectively. In addition, SOX2 expression is also detected in a few NEUN-positive neurons in the lateral brain area (Fig. 1C)7,35. However, SOX2 signals are predominantly detected in EGCs that are located adjacent to the lumen in the axolotl telencephalon, similar to the distribution observed in mammalian species. These EGCs are considered equivalent to neural stem cells (NSCs) in mammals and have the ability to contribute to neurogenesis during brain regeneration36,37. Such spatial distribution of EGCs in axolotl telencephalon makes it possible and easy to manipulate target gene expression by electroporation.

Morphology and cellular composition of axolotl’s telencephalon. (A) Schematic of axolotl brain (top panel) showing the location of the brain in the animal and the enlarged view of the brain showing the ventricular lumen (the yellow shape with orange outlines) inside the brain. HE staining (bottom panel) on cross-cryosections of the telencephalon in different regions from rostral to cauda (the cyan lines on the top panel indicate the section locations). The asterisks indicate the choroid plexus. Scale bar, 500 µm. (B) Immunofluorescence for SOX2 (red), TUJ1 (yellow), and BrdU (white) combined with DAPI (blue) on the axolotl left telencephalon cross-cryosections (top). The white dashed box indicates higher magnification (bottom). The white dotted line outlines the surface of telencephalon, and the cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. (C) Immunofluorescence for SOX2 (red) and NEUN (cyan) on the axolotl left telencephalon cross-cryosections (top) with DAPI (blue). The NEUN channel is shown in different detector gain values. The white dashed box indicates higher magnification (bottom). The yellow arrowheads indicate the double-positive cells of SOX2 and NEUN. The white arrows indicate the SOX2 single-positive cells located in the neuronal layer. The white dotted line outlines the surface of telencephalon, and the cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. HE: hematoxylin and eosin; VZ: ventricular zone.
To establish and optimize the electroporation condition (Supplemental Fig. S1), we used a plasmid pCAGGs: EGFP, in which a green fluorescent protein is driven by a constitutive CAGGs promoter (Fig. 2A), to target EGCs in the brain. The plasmid was injected into the lateral ventricle of left telencephalon, filling lumen with plasmid reagents (Fig. 2B). By placing the electrodes at different angles, we achieved specific expression of EGFP in EGCs located in the dorsal, ventral, medial, or lateral ventricular zone (VZ) (Fig. 2C). Based on the fact that (1) EGCs bear the apical membrane facing the lumen and (2) neighboring EGCs form tight junctions, expressing molecules such as ZO-1, which prevent passive penetration of large molecules from lumen to cortical tissue, SOX2-positive EGCs (Fig. 2D), but not NEUN-positive neurons, were the primary cell type receiving the plasmids from electroporation and expressing the EGFP (Fig. 2E).
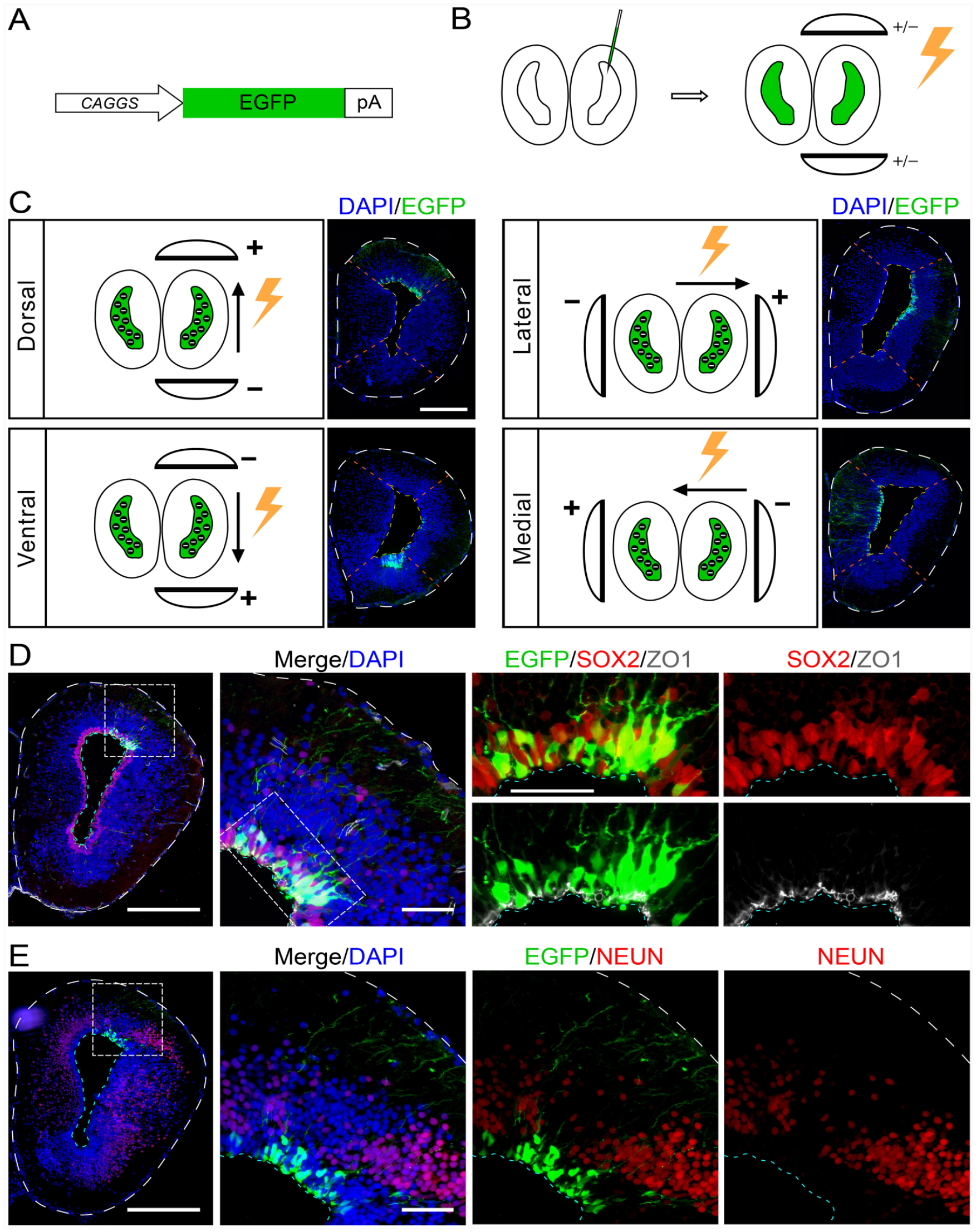
Electroporation of telencephalon EGCs in axolotl. (A) Schematic diagram showing the structure of reporter plasmid pCAGGs: EGFP. The construct contains ubiquitous chicken β-actin promoter with a CMV enhancer (CAGGs) driving EGFP expression, followed by rabbit β Globulin polyA signal sequence. (B) A schematic diagram of axolotl telencephalon electroporation procedure. In order to deliver specifically to EGCs in the VZ via electroporation, the mixture solution of plasmids was injected into the lateral ventricle of the telencephalon, following electroporation with tweezer-style plate electrodes. (C) The schemes and representative fluorescent images of electroporating reporter gene (pCAGGs: EGFP) toward specific regions of axolotl telencephalon VZ. For manipulating desired EGC region of telencephalon, the anode electrode is placed on the corresponding side. “+” indicates the anode electrode, “−” indicates the cathode electrode. The black circles with white “-” filled in the ventricle indicate the DNA plasmids which are negatively charged, and the black arrow aside the brain indicates the moving direction of plasmids. Noted that the negatively charged plasmids will always move toward the anode electrode under the electric field. EGFP fluorescence (unstained, green) combined with DAPI (blue) on cross-cryosections of electroporated telencephalon showing the corresponding electroporated area. Scale bar, 500 µm. (D and E) EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red in D), ZO-1 (white in D) and NEUN (red in E) combined with DAPI (blue) on cross-cryosections of electroporated telencephalon show the distribution and identity of EGFP-positive cells at 2 days after electroporated reporter plasmid pCAGGs: EGFP. The white dashed box indicates higher magnification as single channel or merged images. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. EGCs: ependymal glial cells; VZ: ventricular zone.
Cdc42 Overexpression in EGCs Does Not Affect Early Neurogenesis, But Hinders Maturation of Newly Born Neurons During Axolotl Telencephalon Regeneration
We next attempted to overexpress functional gene and examined its role in regulating EGC behavior during regeneration. We chose Cdc42 as a candidate gene for overexpression manipulation, since it has been reported that Cdc42 plays an essential role in controlling NSC activity during mouse brain development31,38. In the axolotl telencephalon, both mRNA ISH using an antisense Cdc42 probe and immunofluorescence using an antibody against CDC42 revealed that Cdc42 was expressed in the entire cortex, with lower expression level in the EGCs adjacent to the lumen, but higher in the neurons (Supplemental Fig. S2). We expressed Cdc42 in EGCs located at dorsal telencephalon, by electroporating the pCAGGs: EGFP plasmid mixed with either pCAGGs: Cdc42 or an empty pCAGGs: null plasmid as control (Fig. 3A). At 2 days post electroporation, we observed that exogenous CDC42 and EGFP were nearly co-expressed in ventricular zone (Fig. 3B, C), meanwhile, there was almost no detectable CDC42 signal in the control (Fig. 3B) under the same imaging condition, due to the lower expression level of endogenous Cdc42 (Supplemental Fig. S2). Notably, we, therefore, use EGFP-positive cells for the identification of Cdc42-overexpressed cells in this study. We then simultaneously examined the expression of ZO-1, TUJ1, and NEUN, in both the Cdc42 overexpressed and the control samples at 2 days post electroporation. Immunofluorescent results showed no significant differences in terms of the expression pattern of these markers in the Cdc42 overexpressed axolotl telencephalon, compared to the control (Supplemental Fig. S3).

Cdc42 overexpression combined with brain injury. (A) The combinate mixtures of plasmids used for overexpression electroporation. The left panel shows the structure of target gene construct pCAGGS: Cdc42 (top) and reporter construct pCAGGS: EGFP (bottom) which are mixed for injection in Cdc42 overexpression. The right panel shows the structure of two plasmids used for the control group, pCAGGS: null (top) and pCAGGS: EGFP (bottom). The red rectangle indicates full-length Cdc42 coding sequence, the green rectangle indicates EGFP coding sequence, the gray rectangle indicates multiple cloning sites, and pA stands for polyadenylation signal. (B) EGFP fluorescence (unstained, green) and immunofluorescence for CDC42 (red) combined with DAPI (blue) on brain cross-cryosections of 2 days post electroporation show the high expression level of CDC42 in Cdc42 overexpression group (upper panel) compared with the control (lower panel). The images of Cdc42 overexpression group show that the CDC42 signals nearly colocalize with EGFP. The white dash box indicates higher magnification of electroporated areas of dorsal telencephalon shown as merged and single channel images. The cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. (C) The graph shows that in transfected cells electroporated with Cdc42 and EGFP expressing plasmids, the majority (95.7%, counted from four animals) are CDC42 and EGFP double-positive. (D) The timeline for brain injury sample harvest after electroporation. Brain injury was performed 2 days post electroporation in both Cdc42 overexpression and control groups. Animals receive intraperitoneal BrdU injection and are harvested at 5, 10, 15, and 20 days post injury, respectively. For the animals harvested at 5 days, a single injection of BrdU is performed 6 h before harvest. For other time points, cumulative BrdU labeling is performed by injecting BrdU every 2 days from the day of injury until harvest. (E) EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red), BrdU (white) combined with DAPI (blue) on cross-cryosections of regenerating brains at 5 days post injury. In both Cdc42 overexpression brain (upper panel) and control brain (lower panel), there existed a part of EGFP-positive cells that are labeled by BrdU (yellow arrows showing one representative cell). The regenerating areas of dorsal telencephalon highlighted by white dash box are shown with higher magnification as merged or single channel images. The yellow dotted line outlines the surface of telencephalon, and the cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. (F) Quantification of the percentage of BrdU and EGFP double-positive cells over total EGFP-positive cells showing no significant difference between the overexpression group (triangles, n = 4) and the control (circles, n = 3). VZ: ventricular zone.
To evaluate the effect of Cdc42 overexpression on axolotl telencephalon regeneration, we next investigated EGC dynamics during regeneration, under Cdc42 overexpression condition. As we detect markers in Cdc42 overexpressed animals and found that the targeted brain area has no apparent abnormal expression pattern (Supplemental Fig. S3), we performed brain injury on both Cdc42 overexpression and the control animals at 2 days post electroporation. Subsequently, we harvested brain samples at 5, 10, 15, and 20 days post injury (Fig. 3D) and analyzed the effects of Cdc42 overexpression via various parameters of EGCs, including cell cycle, neurogenesis, cell polarity, and migration. For cell cycle kinetics analysis, we administered a single pulse of BrdU to 5-day brain regenerating axolotls prior to sample collection to reveal EGCs in S-phase. We quantified the ratio of BrdU and EGFP double-positive cells within EGFP-positive cells adjacent to the lesion and found that overexpression of Cdc42 did not cause significant change in the proportion of S-phase cells (Fig. 3E, F). Moreover, we analyzed the immunofluorescence of MCM2 at 5 days post injury, which is a marker of cells undergoing proliferation, combined with quantification of MCM2 and EGFP double-positive cells. They showed no significant difference between the Cdc42-overexpression group and the control (Supplemental Fig. S4), in accordance with the result of BrdU. It suggested that Cdc42 overexpression has no major impact on EGC cell cycle kinetics during brain regeneration, consistent with previous findings that perturbing Cdc42 expression does not affect NSC cell cycle during mouse neocortical development 31 .
We then analyzed the expression of TUJ1, a marker starting to express in newly born neurons, and observed that the percentage of TUJ1 and EGFP double-positive cells within EGFP-positive cells in the injured area was similar between the Cdc42 overexpression and the control group in 5-day regenerating brains (Fig. 4A, B). However, further quantification showed that most of the EGFP and TUJ1 double-positive cells in the Cdc42 overexpressed brains were distributed in the region close to the VZ where SOX2-positive NSCs reside, and in the lumen, likely extruded out from the VZ (Fig. 4A). It reached 78.2% in Cdc42 overexpression brains compared to the control, 23.6% (Fig. 4C). Moreover, at 20 days post injury, the proportion of TUJ1 and EGFP double-positive cells near the injury site was 78.6% in the Cdc42 overexpression group, which was significantly higher than the 48.1% observed in the control group (Supplemental Fig. S5A, S5B). Consistent with the observation at 5 days post injury, a majority of the TUJ1 and EGFP double-positive cells in the Cdc42 overexpression group (82.2%) were still located in the VZ at 20 days post injury, in comparison to the control (23.4%) (Supplemental Fig. S5C). However, when we detected NEUN at 20 days post injury, quantifying within the regenerating brain area adjoining the lesion, we found a dramatic decline in the proportion of EGFP and NEUN double-positive neurons over EGFP-positive cells in Cdc42 overexpression group (1.6%), compared to the control (12.7%) (Fig. 4D, E). Upon Cdc42 overexpression in EGCs, the ratio of EGC-derived TUJ1-positive newly born neurons was not affected at 5 days post injury, but increased at 20 days post injury, concomitant with a massive decline of NEUN-positive mature neurons. These results suggested that overexpression of Cdc42 has minor influence on early neurogenesis, instead, mainly blocking the maturation of newly born neurons.

Effect of Cdc42 overexpression on neurogenesis. (A) EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red), TUJ1 (yellow) combined with DAPI (blue) on cross-cryosections of regenerating brains at 5 days post injury. Noted that there are more TUJ1 and EGFP double-positive cells (cyan arrowhead indicated) that emerged at the VZ and inside the lumen in the overexpression group (upper panel) than the control (lower panel). The regenerating areas of dorsal telencephalon highlighted by white dash box are shown with two higher magnifications as merged or single channel images. The white dotted line outlines the surface of telencephalon, and the cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. (B) Quantification of the percentage of TUJ1 and EGFP double-positive cells to total EGFP-positive cells adjacent to the lesion of 5 days post injury shows no significant difference between Cdc42 overexpression group (triangles, n = 4) and the control (circles, n = 3). (C) Quantification of the percentage of TUJ1 and EGFP double-positive cells located at the VZ and inside the lumen over total TUJ1 and EGFP double-positive cells adjacent to the lesion at 5 days post injury. The proportion of double-positive cells in the VZ of overexpression group (78.2%, triangles, n = 4) is significantly higher than the control (23.6%, circles, n = 3). (D) EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red), NEUN (cyan) combined with DAPI (blue) on cross-cryosections of regenerating brains at 20 days post injury. In the Cdc42 overexpression group (upper panel), almost no cell is NEUN and EGFP double-positive while double-positive cells (yellow arrowhead indicated) emerge within the neuronal layer in the control (lower panel). The white dash box of dorsal telencephalon regenerating areas are shown in higher magnification as merged or single channel images. The white dotted line outlines the surface of telencephalon, and the yellow dotted line outlines the VZ surface facing the lumen. (E) Quantification of the percentage of NEUN and EGFP double-positive cells to EGFP-positive cells adjacent to the lesion of 20 days post injury. The proportion of double-positive cells in the overexpression group (1.6%, triangles, n = 4) is significantly lower than the control (12.6%, circles, n = 3). VZ: ventricular zone. Values are considered statistically significant when p < 0.05. *** indicates p < 0.001, **** indicates p < 0.0001.
Under the condition of Cdc42 overexpression, we observed the appearance of luminal SOX2-positive cells during brain regeneration (Fig. 4A as representative). Considering it has been reported that CDC42 is part of the Par-apical complex, and knockout of Cdc42 in NSCs in developing neocortex leads to polarity alteration of NSCs 31 , therefore, we next examined the expression of tight junction marker ZO-1 to indicate the changes of polarity, in both the Cdc42 overexpression and the control brains. Immunofluorescence performed on cryosections showed that the ZO-1 immunostaining signals, which are typically distributed apically, were almost completely lost in the Cdc42 overexpressing areas at both 5 and 20 days post injury, when compared to that in the controls (Fig. 5A, B). This result suggested that Cdc42 overexpression disrupted tight junctions and further interrupted cell polarity, which may explain the presence of neural cells in the lumen and TUJ1 and EGFP double-positive accumulated in the VZ upon Cdc42 overexpression. Based on this, we then analyzed the migration of the progeny of Cdc42 overexpressing EGCs (EGFP-positive cells) in more detail, at 5, 10, 15, and 20 days post injury. In the control, we found that EGFP-positive cells, which include EGCs initially labeled by electroporation and the derived progeny, were mainly distributed in the VZ at 5 days post injury, subsequently started to migrate out to the neuronal layer at 10 and 15 days post injury, and reached their final location at 20 days post injury. In contrast, we found that EGFP-positive cells failed to migrate to their proper position upon Cdc42 overexpression (Fig. 5C) and showed TUJ1 positivity (Supplemental Fig. S5). These migration defects started to appear at 5 days post injury and became obvious at 10, 15, and 20 days post injury (Supplemental Fig. S6). These results indicated that overexpression of Cdc42 attenuates the tight junction in EGCs and causes migratory defects of neurons derived from these EGCs.

Effect of Cdc42 overexpression on cellular polarity and migration. (A and B) EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red), ZO-1 (white) combined with DAPI (blue) on cross-cryosections of regenerating brains at 5 days and 20 days post injury, respectively. In both the 5-day and 20-day post-injury samples, the continuity of ZO-1 lining the VZ is observed to be interrupted at the electroporated area in the overexpression group (upper panel), compared to the control (lower panel). The yellow dotted line outlines the surface of telencephalon, and cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 100 μm. (C) Successive images of EGFP fluorescence (unstained, green) and immunofluorescence for SOX2 (red), combined with DAPI (blue) on telencephalon cross-cryosection for 5, 10, 15, and 20 days post injury (from left to right). The EGFP-positive cells reside basically in the VZ and in the lumen in Cdc42 overexpression animals (upper panel), while the EGFP-positive cells progressively migrate from the VZ into the neuronal cell layer in the control (lower panel). The yellow arrowheads indicated the EGFP-positive cells that migrate into the neuronal layer. The red dotted line outlines the edge of VZ facing the lumen, and the white dotted line divides the VZ area and neuronal cell layer. Scale bar, 100 μm. (D and E) Representative whole-brain fluorescent stereomicroscopy and immunofluorescent images of cross-cryosections are shown at 5 and 20 days post injury. At 5 days post injury (D), the incision at the dorsal part of left telencephalon is similar in both Cdc42 overexpression and control animals. In 20-day post-injury animals (E), the wound does not close in Cdc42 overexpression group, while the wound closes in the control. The dash boxes highlight the regenerating areas of dorsal telencephalon for higher magnification. The white dotted lines outline the wound edges of brains. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. VZ: ventricular zone. BF: bright field.
Furthermore, we checked the effect of Cdc42 overexpression on the overall brain regeneration phenotype. We found that Cdc42 overexpression caused a delayed brain regeneration defect when compared to the control (Fig. 5D, E). In the control, the opened brain injury was almost completely closed at 20 days post injury, where 9 out of 10 animals that we operated healed the wound. However, it was still maintained as a “mini-hole” in the Cdc42 overexpressed brain, where only 3 out of 10 healed the wound (Figs. 4D and 5E, Supplemental Table S1). Yet, the Cdc42 overexpressed brain showed closure of the wound until we observed at 30 days post injury (Supplemental Table S2). This delayed regeneration phenotype likely resulted from maturation and migration defects of newly born neurons, upon Cdc42 overexpression in EGCs.
The above results suggested that overexpression of target gene by electroporation is an effectual approach for studying the role of candidate gene during axolotl brain regeneration.
Target Gene Knockdown Via CAS9-gRNA Complex Electroporation
Loss of function is an important approach for studying gene function. We next proceeded to establish an electroporation-based, CRISPR/Cas9-mediated method to knockout/knockdown targeted gene in EGCs of the axolotl brains. It has been reported that electroporation of CAS9-gRNA could be applied to knockdown targeted gene in the spinal cord of the axolotls. In terms of target gene, we chose TnC, one of the genes detected to be up-regulated in EGCs adjacent to lesion during brain regeneration by spatial transcriptomics6,39,40 (Fig. 6A, top panel). In this experiment, we chose tyrosinase (Tyr) gene as the control, which is considered playing no or non-essential roles in the central nervous system, and has been used as a control for gene knockout experiments in previous publications16,41,42. mRNA ISH using an antisense probe indeed showed a similar expression regulation of TnC in brain regeneration (Fig. 6A, bottom panel). To knock down TnC, we first injected the pre-assembled CAS9-gRNA complex mixture against TnC into the telencephalic lumen, followed by electroporation orientated to the dorsal domain of the left telencephalon. We then carried out brain injury at 2 days post electroporation and collected regenerating brains at 5 days post brain injury (Fig. 6B, C). mRNA ISH showed that TnC expression was dramatically down-regulated in EGCs adjacent to the injury site upon electroporation of CAS9-gRNA mixture 2, but not so obvious with mixture 1, when compared to that in the control (Fig. 6D). Furthermore, when carried out immunofluorescence using an antibody against TnC, we observed similar phenotypes, except that the reduction of the protein level is more dramatic than mRNA upon electroporation of CAS9-gRNA mixture 2 (Fig. 6E), leading to more clear loss of protein than mRNA. To evaluate the efficacy of the gRNAs in inducing the TnC genome modification, we conducted genotyping after electroporating CAS9-gRNA complexes into the axolotl spinal cord. The sequencing analysis revealed that the gRNAs were capable of inducing indels in vivo at the targeted TnC loci (Supplemental Fig. S7). Among all animals that were electroporated with TnC gRNAs, 75% exhibited knockdown in electroporated brain area (n = 8), while none in the control group showed knockdown (n = 4) (Supplemental Table S3). It suggested that electroporation of CAS9-gRNA could efficiently knockdown targeted gene.
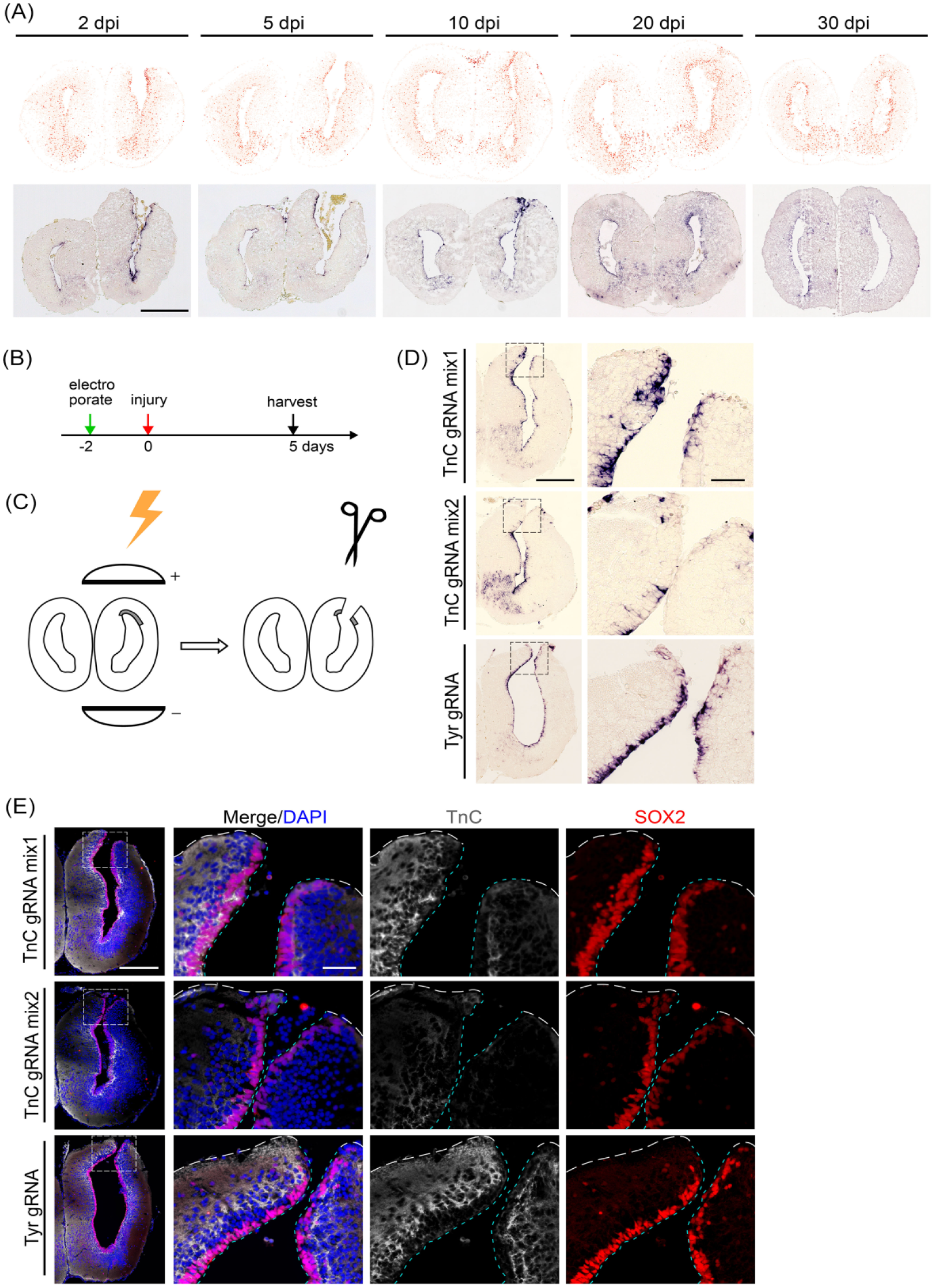
Knockdown of Tnc in axolotl telencephalon by CAS9 protein–gRNA complexes electroporation. (A) TnC expression at 2, 5, 10, 20, and 30 days post injury of the axolotl regenerating telencephalons, presented by spatial visualization Stereo-seq maps 6 (top panel, the color shade of dots denotes TnC expression level in cells, darker colors indicate higher expression level) and their corresponding in situ hybridization images (bottom). Scale bar, 500 µm. (B) The timeline of animal treatments involves surgical injury to the electroporated telencephalons at 2 days post electroporation, and harvested at 5 days post injury for Tnc expression detection. (C) A scheme is shown for electroporating CAS9 protein-gRNA complexes and performing an incision to the dorsal part of the electroporated telencephalons. (D) Representative in situ hybridization images on cross-cryosections show mRNA-level TnC expression at axolotl telencephalons incision site with electroporating different gRNA mixtures. For Tnc knockout, the mixture consisted of Tnc gRNAs (mix1 contains gRNA 1 and 2, mix2 contains gRNA 3, 4, and 5). For control, the mixture consists of Tyr gRNA. The stain of TnC signal in the VZ at incision site is significantly attenuated in Tnc gRNA mix2 group, compared to the control. The black dashed box indicates higher magnification. Scale bars, 500 μm at low magnification images and 100 μm at high magnification images. (E) Representative immunofluorescent images of TNC (white) and SOX2 (red) combined with DAPI (blue) on cross-cryosections show the loss of TNC expression at the incision site of a 5-day regenerating electroporated telencephalon compared with the control. The white dashed box indicates higher magnification as single channel or merged images. The white dotted line outlines the surface of telencephalon, and the cyan dotted line outlines the VZ surface facing the lumen. Scale bar, 500 μm at low magnification images and 100 μm at high magnification images. VZ: ventricular zone. TnC: Tenascin-C.
Discussion
Along with the development and application of multi-omic technologies in these years, a huge amount of bioinformatic data has been released and require further experimental analysis43,44. Recently, using single-cell sequencing and spatial transcriptomics technology, several studies analyzed DEGs and accessible open chromatin in EGCs during brain regeneration in axolotls. It was revealed that EGCs undergo a rejuvenation process, and switch back to an early embryonic-like EGC state, before participating in regeneration6,7. Applying gain-of-function and loss-of-function analyses to investigate the role of DEGs will disclose the essential molecules involved in the rejuvenation and brain regeneration process.
Here, we established rapid and robust functional gene manipulation approaches in the axolotl brain via electroporation of plasmid or CAS9 protein-gRNA complexes and demonstrated the role of Cdc42 on EGCs during brain regeneration. This study opened a great possibility for further mechanistic investigation of brain regeneration in axolotls via gain-of-function and loss-of-function approaches.
We observed that Cdc42 overexpression interrupted apical tight junctions, thus leading to delamination of EGCs. In addition, it does not play a major role in the onset of neurogenesis, but rather affects the maturation of newly born neurons and migration of neurons to their final location. During brain development, deletion of endogenous Cdc42 in mouse telencephalon yields similar defects, including disturbance of the apical adherent junctions followed by migration failure, and premature onset of neurogenesis, whereas the cell cycle of Cdc42-deficient neural progenitors is less affected. The phenotypes showed in rodent brain development after Cdc42 deletion is highly correlated to that observed in axolotl brain regeneration upon Cdc42 overexpression 31 . These findings suggest either up-regulation or down-regulation of Cdc42 leads to similar defects due to the interruption of proper Cdc42 expression in neural progenitors, as well as indicate a conserved Cdc42 function during evolution.
In this study, we achieved knockdown of TnC using electroporation based on CRISPR/Cas9 system. CAS9-gRNA complexes will recognize the coding sequence of TnC locus on genome and induce indels, which will disrupt the normal function of TnC. Interestingly, we not only observed the absence of TNC proteins in the electroporated area by immunofluorescent stain but also found that the TnC mRNA level was down-regulated, revealed by mRNA ISH. To address this, one of the potential explanations could be non-sense-mediated mRNA decay (NMD) 45 . NMD is a cellular process that selectively targets and degrades mRNAs containing premature stop codons, thereby preventing the production of truncated or non-functional proteins. In our case, the indels on the coding sequence of TnC locus might lead to the frameshift mutation, which may result in the occurrence of premature stop codons. Meanwhile, it is noteworthy that our current understanding does not rule out the possibility of abnormal initiation or regulation during the transcription process, which may cause the down-regulation of mRNA. Based on these mechanisms, it is feasible to evaluate the knockdown of target gene by detecting mRNA level using mRNA ISH.
As indicated recently, EGCs in adult axolotl brain can be categorized into three groups6,7. Besides common neural progenitor markers, each of them also expresses unique markers and is maintained at different states, such as slow dividing or quiescence. However, these EGC subtypes are mostly distinguished by the signature of combination of a group of genes, which makes it difficult to genetically label them using unique gene promoter by transgenics. In this study, we reached specific targeting of EGCs located in different domains by electroporation. It overcomes the limitations of genetic markers and allows for labeling and manipulating each EGC subtype to separately study their role during brain regeneration. Overall, electroporation-based gene functional manipulation method established in this study provides a powerful tool to facilitate the mechanistic investigation of brain regeneration in axolotls.
Supplemental Material
sj-docx-1-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-docx-1-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-2-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-2-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-3-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-3-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-4-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-4-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-5-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-5-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-6-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-6-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-7-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-7-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Supplemental Material
sj-jpg-8-cll-10.1177_09636897231200059 – Supplemental material for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain
Supplemental material, sj-jpg-8-cll-10.1177_09636897231200059 for Establishing an Efficient Electroporation-Based Method to Manipulate Target Gene Expression in the Axolotl Brain by Sulei Fu, Cheng Peng, Yan-Yun Zeng, Yuanhui Qiu, Yanmei Liu and Ji-Feng Fei in Cell Transplantation
Footnotes
Author Contributions
J-FF and YL conceived and planned the experiments. SF, CP, Y-YZ, and YQ performed the experiments. SF and CP analyzed the data and prepared the original figures. J-FF, SF, CP, and Y-YZ wrote the manuscript. All authors revised the manuscript and approved the final manuscript.
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
All animal experiments in this study were approved by the Biomedical Research Ethics Committee of Guangdong Provincial People Hospital (license number: KY2023-447-01), Guangzhou, China. This article does not contain any studies with human subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (31970782; 92268114; 32070819), the National Key R&D Program of China (2021YFA0805000; 2019YFE0106700), and BGI grant (BGIRSZ20210002).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
