Abstract
Gastric cancer is the third leading cause of cancer-related deaths worldwide, and research on gastric cancer pathogenesis is fundamental. Long intergenic non-coding RNAs (lincRNAs) control cancer initiation and progression through several mechanisms, with the competitive endogenous RNA (ceRNA) regulatory network being the most common. In this study, in situ hybridization revealed that long intergenic non-protein coding RNA-regulator of reprogramming (linc-ROR) was highly expressed in gastric cancer cells and was mainly cytoplasmic-positive. Cell counting kit-8 (CCK-8), plate colony formation, wound healing, and Transwell assay revealed that linc-ROR knockdown impedes the growth, proliferation, and migration of gastric cancer cells, while linc-ROR overexpression promoted gastric cancer cell growth, migration, and colony formation ability. Combined with previous studies, the molecular mechanism axis of linc-ROR/miR-145-5-5p/POU5F1/SOX2 was verified. The expression of linc-ROR knockdown significantly suppressed the protein expression of POU5F1 and SOX2. Co-transfection with linc-ROR siRNA reverses the carcinogenic effect of the miR-145-5p inhibitor on gastric cancer cell proliferation, cloning, and migration. These findings lay a foundation for developing novel targets for gastric cancer treatment.
Introduction
Gastric cancer is the third leading cause of cancer-related fatality worldwide 1 . Gastric cancer is generally diagnosed at an advanced stage and has a poor prognosis. Systemic therapy can relieve symptoms, prolong survival, and improve the quality of life of patients with locally advanced or metastatic diseases. Implementing biomarker testing, particularly analyzing HER2 status, microsatellite instability status, and the expression of programmed death ligand 1 (PD-L1), has remarkably influenced clinical practice and patient care. In clinical trials, targeted therapies using drugs such as trastuzumab, nivolumab, and pembrolizumab have yielded positive outcomes for the treatment of locally advanced or metastatic diseases 1 . Nevertheless, the 5-year survival rate of patients with gastric cancer is still less than 30%. In 2020, the number of deaths from gastric cancer worldwide was approximately 769,0002,3. Therefore, it is essential to actively conduct basic and clinical research on gastric cancer.
Since 2005, numerous studies have revealed that non-coding RNA is indispensable for life. The ENCODE project demonstrated that at least 80% of the sequences in the human genome have biological functions, most of which are non-coding RNA, which may have been transcribed from DNA regulatory elements 4 . The ENCODE project included approximately 9,640 human lncRNA loci 5 . More researchers are devoted to the field of long intergenic non-coding RNA (lincRNA) research. LincRNAs are involved in multiple essential pathways in cancer, including p53, NF-κB, PI3K/AKT, and Notch 6 . Furthermore, lincRNAs regulate the occurrence and development of gastric cancer through multiple mechanisms 7 . For example, the transcription of lincRNA THAP7-AS1 with m6A modification mediated by mettl3 is activated by the transcription factor SP1. Mechanically, THAP7-AS1 interrelates with the 1–50 amino acid region (nuclear localization sequence [NLS]) of CUL4B through its 1–442 nucleotide sequence, thereby stimulating the interaction between NLS and transcription factor α1 and facilitating CUL4B protein entry into the nucleus. This constrains the expression of miR-22-3p and miR-320a through CUL4B-catalyzed H2AK119ub1 and EZH2-mediated H3K27me3, thereby activating the PI3K/AKT signaling pathway to promote gastric cancer progression 8 . Nonetheless, several mechanisms regulating lncRNAs in gastric cancer remain unknown.
Long intergenic non-protein coding RNA-regulator of reprogramming (linc-ROR) controls the occurrence and development of cancers, including pancreatic cancer 9 , colorectal cancer 10 , breast cancer 11 , and hepatocellular carcinoma 12 . As linc-ROR can alter cell proliferation13,14, migration 15 , and invasion 16 , it may act as a biomarker and curative target for patients with malignant tumors. Linc-ROR exerts its oncogenic effects post-transcription through the competitive endogenous RNA (ceRNA) network in cancer pathogenesis17,18. Our laboratory has previously published articles reporting that linc-ROR is an excellent diagnostic marker for gastric cancer; however, its mechanism has not been explored 19 . The proliferative phenotype of cancer cells is a crucial cancerous hallmark 20 . Linc-ROR was thought to act as a sponge for hsa-miR-145-5p (miR-145-5p) to modulate osteoblast proliferation and apoptosis in osteoporosis 21 . miR-145-5p is a predictive biomarker for breast cancer stemness, targeting POU5F1 and SOX2 22 . In this study, we investigated the mechanism of linc-ROR which acts as a sponge for miR-145-5p to regulate POU5F1 and SOX2 and may affect cell growth, proliferation, and migration in gastric cancer.
Materials and Methods
Cell Culture and Passage
The normal gastric mucosal epithelial cell line, GES-1, was purchased from the Cancer Hospital, Chinese Academy of Medical Sciences. The gastric cancer cell lines, MGC-803 and SGC-7901, were purchased from the Shanghai Institute of Life Sciences, Chinese Academy of Sciences. The culture conditions were RPMI Medium 1640 (containing 10% fetal bovine serum [FBS]), 37°C, and 5% CO2. Cells in the logarithmic growth phase were used for experiments. A passage was formed when the cell confluence reached 90%.
Synthesis of Linc-ROR siRNA and miRNA Mimics
Linc-ROR small interfering RNA (siRNA), siRNA negative control (NC), miR-145-5p mimics, and microRNA (miRNA) NC were synthesized by GenePharma (Shanghai, China) (Table S1).
Construction of Linc-ROR Overexpression Plasmid
Plasmid vector (pEX-3) that overexpressed linc-ROR and empty vector were purchased from GenePharma (Shanghai, China) and were transfected into SGC-7901 and MGC-803 using a Lipo3000 (Thermo Fisher Scientific, Inc.) according to the provided protocol.
Transient Transfection
Cells in the logarithmic growth phase were inoculated into six-well plates and cultured in RPMI Medium 1640 medium (HyClone, Fairfield, CT, USA) containing 10% FBS (Bovogen, Keilor East, Australia) for 24 h. When the cell confluence reached 50%, the medium was discarded, washed twice with phosphate-buffered saline (PBS), and 1.8 ml of Opti-MEM I Reduced Serum Medium (Gibco, New York, NY, USA) was added to each well. miRNA mimics or linc-ROR siRNA (12 μl) was added to 88 μl of Opti-MEM I Reduced Serum Medium, and 2 μl of lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) was added to 98 μl of Opti-MEM I Reduced Serum Medium, gently mixed, and incubated at 25°C for 5 min. Diluted miRNA mimics or linc-ROR siRNA was mixed with liposomes, and a transfection solution was prepared at 25°C for 20 min. The dye solution (200 μl) was added to each well, gently shaken, and incubated at 37°C in a 5% CO2 incubator for 5 h. The medium with dye solution was then replaced with RPMI Medium 1640 medium containing 10% FBS. Subsequent experiments were conducted after 24–48 h.
RNA Extraction
Cells were collected, mixed with 1 ml of TRIzol Reagent (Invitrogen, Waltham, MA, USA), and incubated on ice for 3 min. Chloroform (200 μl) was added to the cell lysate, which was then oscillated for 15 s, incubated on ice for 3 min, and centrifuged at 12,000 × g and 4°C for 15 min. The upper aqueous phase was transferred to a 1.5-ml centrifuge tube without RNAase, and an equal volume of isopropanol (Solarbio, Beijing, China) was added and oscillated for 5 s. The sample was then incubated on ice for 10 min and centrifuged at 12,000 × g and 4°C for 10 min. The supernatant was discarded, and 1 ml of 75% ethanol (Solarbio) was added to wash RNA plates and centrifuged at 12,000 × g and 4°C for 3 min. The liquid supernatant was removed. The RNA plate was redissolved in RNA-free enzymatic water, and RNA concentration was measured using a Nanodrop UV Spectrophotometer (Denovix, Houston, TX, USA).
QRT-PCR Analysis
Linc-ROR, POU5F1, SOX2, and β-actin were reverse-transcribed using the ReverTra Ace qPCR RT Master Mix with genomic DNA (gDNA) Remover (FSQ-301; Toyobo Co., Ltd., Osaka, Japan) according to the manufacturer’s instructions. The total RNA amount added to the premixed reverse transcription system was 500 ng. Reverse transcription quantitative polymerase chain reaction (RT-qPCR) of these RNAs was performed according to the protocol of GoTaq 2-StepRT-qPCR System (Promega, Madison, WI, USA) in an Mx3005P QPCR System (Stratagene, La Jolla, CA, USA). After mixing and centrifuging, 5 μl of complementary DNA (cDNA) was added to the total volume of 25 μL qPCR reaction system. The qRT-PCR of mature miR-145-5p experiment was conducted according to the manufacturer’s instruction of Hairpin-itTM miRNA and U6 snRNA Normalization RT-PCR Quantitation Kit (E22001–E22010; GenePharma, Shanghai, China). The total RNA amount added to the volume of 20 μl reverse transcription system was 1,000 ng. After mixing and centrifuging, 2 μl of cDNA was added to the total volume of 20 μl qPCR reaction system. qRT-PCR primers in experiments are listed in Table S2.
Fluorescence In Situ Hybridization (FISH) Analysis
Cells were fixed in an in situ hybridization fixative (G1113, Servicebio, Wuhan, China) for 20 min, and the linc-ROR probe sequence was 5′- CTGTGATTGTAGTCATAGTGCCATTTTA-3×. The linc-ROR probe hybridization solution with a concentration of 500 nM was added and hybridized at 40°C for 12 h. The photographs were taken using a positive fluorescence microscope (Nikon, Tokyo, Japan). The nuclei stained with DAPI (G1012; Servicebio) were blue under ultraviolet excitation, and linc-ROR-positive expression was red, labeled by the corresponding fluorescein CY3 tyramide (G1223; Servicebio).
Cell Proliferation Analysis
After transfection, cells were seeded into 96-well plates at an intensity of 3,000 cells per well. After the cells adhered to the wall, 10 μl of cell counting kit-8 (CCK-8) detection reagent (Dojindo, Kumamoto, Japan) was added at different time points (24, 48, 72, and 96 h), and the cells were incubated at 37°C for 3 h in the dark. The optical density (OD) at 450 nm was measured using a microplate reader (LabSystems, Vantaa, Finland). Cell proliferation in rescue experiments was done in E-Plates 96 (ACEA Biosciences, San Diego, CA, USA) using a real-time cell analyzer and impedance-based xCELLigence System (RTCA DP Analyzer; Roche Applied Science, Mannheim, Germany). This appliance can monitor cell proliferation dynamically.
Clone Formation
After transfection, cells were seeded into six-well plates at a density of 500 cells/well. After 1 week, the old culture medium was replaced with a fresh medium. After 2 weeks, cells were fixed with 4% paraformaldehyde (Solarbio) for 30 min, stained with crystal violet (Solarbio) for 30 min, and washed with PBS to remove the floating color. The formation rate of the cell clones was observed.
Wound Healing Assay
Regular scratches were made in confluent monolayers using a 200-μl pipette tip, cells were incubated with a low-serum medium to reduce cell proliferation effects, and scratches were photographed at 0 and 24 h to observe the cell healing distance.
Transwell Migration Assay
After transfection, cells were suspended in a serum-free medium and seeded into the upper chamber. The upper chambers with cells were placed into 24-well plates containing a 20% FBS medium. After 24 h of incubation, cells that did not migrate to the underlying membrane were removed with a cotton swab. The cells on the lower membrane were fixed with 4% paraformaldehyde for 30 min, stained with crystal violet for 30 min, washed with water to remove the floating color, and photographed under a light microscope (Olympus, Tokyo, Japan) to record the number of cells.
Western Blotting Analysis
Gastric cancer cells (SGC-7901 and MGC-803) transfected with NC or small interfering (si)-linc-ROR were lysed using a radioimmunoprecipitation buffer (Solarbio). The total protein was extracted by centrifugation at 12,000 × g at 4°C for 25 min. The total protein was boiled and then separated using 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then blotted onto polyvinylidene fluoride (PVDF) membranes (Millipore, Bedford, MA, USA), which were blocked with 3% nonfat dry milk in TBST for 1 h. The membranes were incubated with POU5F1 rabbit polyclonal antibody (Abclonal, Wuhan, China) or SOX2 rabbit polyclonal antibody (Abclonal) at 4°C overnight, followed by incubation with secondary antibodies HRP goat anti-rabbit IgG (H+L) (Abclonal). POU5F1 and SOX2 proteins were then detected using an ECL Basic Kit (Abclonal). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (Abcam, Cambridge, UK) was used as an external control.
Data Statistics and Analysis
The experiment was repeated thrice. The experimental data were analyzed using SPSS Statistics 20.0 (IBM, Armonk, NY, USA). The statistics were analyzed using two independent-samples t tests. Statistical data are expressed as mean ± SD, and differences were considered statistically significant at P < 0.05 (*P < 0.05, **P < 0.01, ***P < 0.001).
Results
Linc-ROR Is Upregulated in Human Gastric Cancer Cells and Is Present in the Cytoplasm
To determine the influence of linc-ROR on cell biological functions in vitro, the expressional level of linc-ROR was detected by qRT-PCR. Compared with the human normal gastric epithelial cell line GES-1, linc-ROR was upregulated in SGC-7901 and MGC-803 cell lines. The expression level of linc-ROR in SGC-7901 was more than twice that in GES-1. The expression level of linc-ROR in MGC-803 was approximately three times higher than that in GES-1 (Fig. 1A). Therefore, linc-ROR may play a carcinogenic role in gastric cancer. FISH revealed that linc-ROR were present in the cytoplasm, indicating that linc-ROR plays a major biological role in the cytoplasm of gastric cancer cells (Fig. 1B).

The expression and subcellular localization of linc-ROR in gastric cancer cells. (A) Linc-ROR is upregulated in human gastric cancer cells. (B) Fluorescence in situ hybridization shows that linc-ROR is mainly cytoplasmic-positive.
Knockdown of Linc-ROR Impedes Cell Growth and Proliferation
We propose that linc-ROR is associated with the growth and proliferation of gastric cancer cells. To test this hypothesis, CCK-8 and plate colony formation assays were used to investigate the effect of linc-ROR on the proliferative ability of gastric cancer cells. siRNA was used to precisely and efficiently knock down linc-ROR expression. By transfecting linc-ROR siRNA into cells, we found that linc-ROR knockdown constrained the growth and proliferation of SGC-7901 and MGC-803 cells (Fig. 2). Cell growth was determined by measuring OD at 0, 24, 48, 72, and 96 h. After 24 h of transfection with linc-ROR siRNA, SGC-7901 began to display growth restriction. After 48 h of transfection, the OD of the linc-ROR siRNA group was lower than that of the NC group for both SGC-7901 and MGC-803. Cell growth inhibition by linc-ROR siRNA was more pronounced over time (Fig. 2A–C). Cell proliferation was identified based on colony formation efficiency. The colony formation efficiency of the linc-ROR siRNA group was lower than that of the NC group for both SGC-7901 and MGC-803 (Fig. 2D, E). Hence, linc-ROR siRNA impeded gastric cancer cell growth and proliferation.

Knockdown of linc-ROR impedes cell growth (A, B, C) and proliferation (D, E).
Knockdown of Linc-ROR Inhibits Cell Migration
Wound healing and Transwell migration assays were used to investigate the influence of linc-ROR on cell migration. After 24 h of transfection with linc-RoR siRNA, the horizontal migration speeds of SGC-7901 and MGC-803 cells decreased. The scratch width in the linc-ROR siRNA group was broader than that in the NC group (Fig. 3A, B). Similarly, fewer cells migrated vertically downward in the linc-ROR siRNA group than in the control group for both cell lines (Fig. 3C, D). Consequently, linc-ROR siRNA inhibited cell migration.

Knockdown of linc-ROR obstructs cell horizontal migration (A, B) and vertically downward migration (C, D).
Overexpression of Linc-ROR Promotes Gastric Cancer Cell Growth, Migration, and Colony Formation Ability
By transfecting the linc-ROR overexpression plasmid into gastric cancer cells, we found that linc-ROR overexpression promoted the growth of SGC-7901 and MGC-803 cells (Fig. 4A). After 24 h of transfection with linc-ROR overexpression plasmid, SGC-7901 began to display growth promotion. After 48 h of transfection, the OD of the linc-ROR overexpression group was higher than that of the NC group for both SGC-7901 and MGC-803 (Fig. 4A).

Overexpression of linc-ROR promotes gastric cancer cell growth (A), vertically downward migration (B), and colony formation ability (C).
Transwell migration assays were used to investigate the influence of linc-ROR overexpression on cell migration. After 24 h of transfection with linc-RoR overexpression plasmid, more cells migrated vertically downward in the linc-ROR overexpression group than in the control group for SGC-7901 and MGC-803 cells (Fig. 4B).
The colony formation efficiency of the linc-ROR overexpression group was higher than that of the NC group for both SGC-7901 and MGC-803 (Fig. 4C). Hence, linc-ROR overexpression promoted gastric cancer cell growth, migration, and colony formation ability.
Linc-ROR Shares miR-145-5p With POU5F1 and SOX2
To validate the theory that linc-ROR acts as a sponge for miR-145-5p to regulate POU5F1 and SOX2, the bioinformatics tool Miranda (http://www.bioinformatics.com.cn/local_miranda_miRNA_target_prediction_120) and miRcode (http://mircode.org/) were used to search for binding sites between miR-145-5p and other genes (POU5F1, SOX2m and linc-ROR) (Fig. 5).

Bioinformatic analysis to predict linc-ROR shares miR-145-5p with POU5F1 and SOX2.
The target validation through bioinformatics tools was consistent with dual-luciferase assay results in HEK293 cells from the published article (POU5F1 also named OCT4) 23 . Thus, linc-ROR, POU5F1, and SOX2 have miRNA response elements that bind to miR-145-5p. Next, miR-145-5p mimics were transfected into cells to overexpress miR-145-5p (Fig. 6A). Linc-ROR, POU5F1, and SOX2 were downregulated when miR-145-5p was upregulated (Fig. 6B–D). Similarly, linc-RoR siRNA was transfected into cells to knock down linc-ROR (Fig. 6E). When linc-ROR was downregulated, miR-145-5p was upregulated (Fig. 6F), whereas POU5F1 and SOX2 were downregulated (Fig. 6G, H). Linc-ROR was synergistic with POU5F1 and SOX2, and the variation trend of the three molecules is opposite to that of miR-145-5p. Based on the miRNA response elements and gene variation trends, we concluded that linc-ROR competes with POU5F1 and SOX2 to bind miR-145-5p, which is closely related to the growth, proliferation, and migration of gastric cancer cells.

Linc-ROR shares miR-145-5p with POU5F1 and SOX2 at the RNA levels. (A, B) Downregulation of linc-ROR after efficient overexpression of miR-145-5p in gastric cancer cells. miR-145-5p downregulates POU5F1 (C) and SOX2 (D). (E, F) Upregulation of miR-145-5p after efficient knockdown of linc-ROR in gastric cancer cells. POU5F1 (G) and SOX2 (H) were downregulated with knockdown of linc-ROR.
Linc-ROR Knockdown Suppressed the Protein Expression of POU5F1 and SOX2
Compared with transfection with siRNA NC in SGC-7901 and MGC-803 cells, results showed that the brightness of POU5F1 and SOX2 protein bands was remarkably reduced when transfected with linc-ROR siRNA (Fig. 7A). Moreover, three independent experiments showed that linc-ROR siRNA significantly suppressed the protein expression of POU5F1 and SOX2 in gastric cancer cells (Fig. 7B, C).
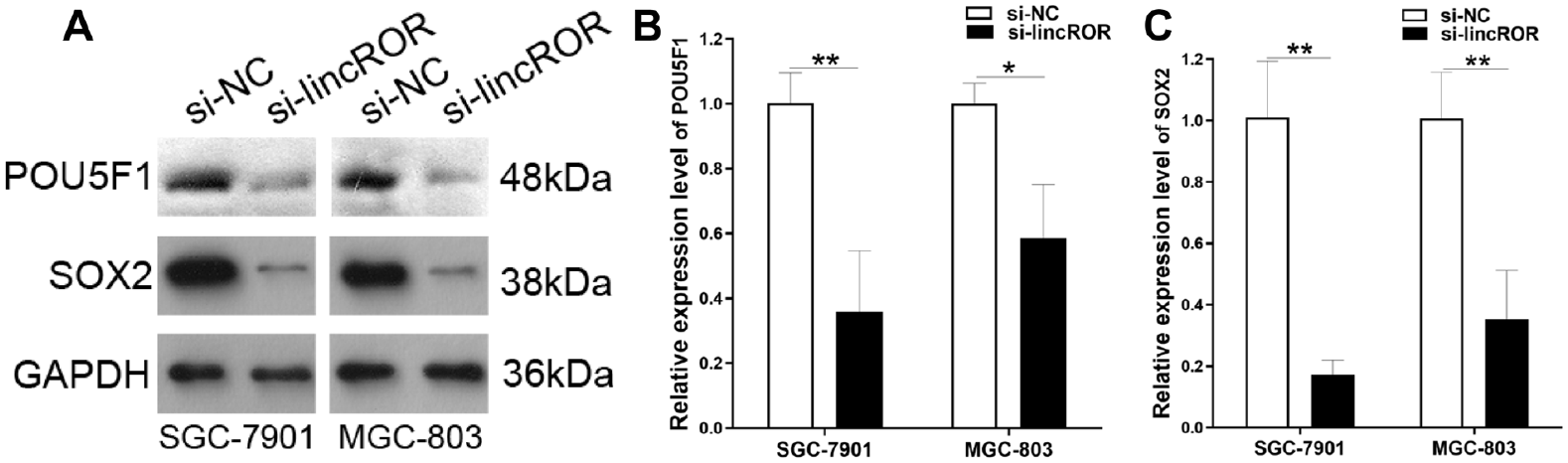
Western blotting analysis of protein expression changes of POU5F1 and SOX2 in gastric cancer cells. (A) A representative result. (B) Results from three independent experiments showed that si-linc-ROR significantly suppressed the protein expression of POU5F1 and SOX2. GAPDH was used as a loading control. Compared with si-NC, *P < 0.05, **P < 0.01, ***P < 0.001.
Linc-ROR siRNA Rescues the Carcinogenic Effect Induced by the miR-145-5p Inhibitor
Through co-transfection of linc-ROR siRNA and miR-145-5p into gastric cancer cells, we explored whether linc-ROR siRNA could rescue the carcinogenic effect induced by the miR-145-5p inhibitor. The knockdown of linc-ROR expression reversed the carcinogenic effect of the miR-145-5p inhibitor on gastric cancer cell proliferation (Fig. 8A, B), cloning (Fig. 8C, D), and migration (Fig. 8E, F). These results indicated that linc-ROR siRNA affects the biological function of gastric cancer cells by directly regulating miR-145-5p.

Linc-ROR siRNA rescues the carcinogenic effect induced by the miR-145-5p inhibitor. The knockdown of lincROR expression reversed the carcinogenic effect of the miR-145-5p inhibitor on the gastric cancer cell proliferation (A, B), cloning (C, D), and migration (E, F). *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
Several studies have reported that linc-ROR regulates the proliferation, metastasis, and differentiation of cancer cells through various pathways. It plays a crucial role in the incidence and development of tumors and is a conceivable biological marker for cancer screening and treatment 24 . For instance, linc-ROR is upregulated in hepatocellular carcinoma and promotes cancer metastasis 25 . Furthermore, linc-ROR is involved in the self-renewal and differentiation of glioma stem cells 26 . In addition, linc-ROR is an exceptional biological marker for breast cancer diagnosis and prognosis 27 .
With high molecular and phenotypic heterogeneity, gastric cancer has the highest morbidity and mortality rates worldwide 28 . The primary action is endoscopic resection of early gastric cancer. Patients with non-early gastric cancer can be treated with surgery29,30. Adjuvant chemotherapy and perioperative treatment enhance the survival rate of patients with advanced gastric cancer31,32. In first-line clinical treatment, patients with advanced gastric cancer undergo an incessant chemotherapy procedure for platinum and fluoropyrimidine dual chemotherapy drugs 33 . Targeted therapies have been approved from the first (trastuzumab) through the third (pembrolizumab) line, indicating significant progress1,34. However, the survival of patients with gastric cancer can be improved by diagnosing and treating the disease at an early stage. Moreover, the 5-year survival of advanced-stage patients is still under 10% 2 . However, patients with gastric cancer still face considerable challenges. Exploring the mechanism of linc-ROR in gastric cancer can lead to the discovery of new therapeutic targets.
In this study, qRT-PCR was used to quantitatively detect linc-ROR expression in gastric cancer cell lines. The results showed that linc-ROR was more highly expressed in gastric cancer cell lines than in normal gastric epithelial cells. Therefore, we speculate that linc-ROR plays a tumor-promoting role in gastric cancer cell. Subcellular localization FISH showed that linc-ROR might play a major biological role in the cytoplasm. Linc-ROR knockdown impedes cell growth, proliferation, horizontal migration, and vertical downward migration at the cellular phenotype level. On the contrary, overexpression of linc-ROR promoted gastric cancer cell growth, migration, and colony formation ability. At the machinery stage, combined with previous studies, we verified the existence of the ceRNA regulatory network of linc-ROR/miR-145-5p/POU5F1/SOX2 in gastric cancer, which is a potential target for gastric cancer therapy. In addition, linc-ROR siRNA significantly suppressed the protein expression of POU5F1 and SOX2. Through the co-transfection of linc-ROR siRNA and miR-145-5p inhibitor into gastric cancer cells, linc-ROR siRNA rescued the carcinogenic effect of the miR-145-5p inhibitor on gastric cancer cell proliferation, cloning, and migration. These results showed that linc-ROR regulated POU5F1 and SOX2 by competitively binding miR-145-5p to affect the proliferation and migration of gastric cancer cells. In this study, we found that linc-ROR functions as an oncogene in gastric cancer cell. Moreover, our previous published article reported that downregulated expression of linc-ROR is a promising biomarker for patients with gastric cancer 19 . The results indicated that linc-ROR may play dual role in gastric cancer cells and tissues. The limitation of this study is the translation of linc-ROR research into effective treatment and clinical decisions for patients with gastric cancer is also a significant challenge.
In conclusion, this is the first study that investigated that the mechanism of linc-ROR competes with POU5F1 and SOX2 to bind miR-145-5p in gastric cancer tumorigenesis. We believe that our study makes a significant contribution to the literature because our findings provide a basis for developing therapeutic targets for gastric cancer.
Supplemental Material
sj-rar-1-cll-10.1177_09636897231178902 – Supplemental material for Linc-ROR Regulates POU5F1 and SOX2 by Competitively Binding miR-145-5p to Affect the Proliferation and Migration of Gastric Cancer Cells
Supplemental material, sj-rar-1-cll-10.1177_09636897231178902 for Linc-ROR Regulates POU5F1 and SOX2 by Competitively Binding miR-145-5p to Affect the Proliferation and Migration of Gastric Cancer Cells by Xiuchong Yu, Xinyi Ma, Yangtao Jia, Xinxin Wu and Zhilong Yan in Cell Transplantation
Footnotes
Acknowledgements
Author Contributions
Xiuchong Yu analyzed the data and wrote the manuscript, Xinyi Ma, Yangtao Jia, and Xinxin Wu revised the manuscript, and Zhilong Yan designed the study. All authors have read and approved the final manuscript.
Availability of Data and Materials
The data used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Research Ethics and Patient Consent
This article does not contain any studies with human or animal subjects. Ethical Approval is not applicable for this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Medical and Health Science and Technology Project of Zhejiang Province (2021RC122 and 2022RC064), the Natural Science Foundation of Ningbo (2018A610371), the Natural Science Foundation of Zhejiang Province (LGF19H030007), The Foundation of Zhejiang Key Laboratory of Pathophysiology (202302), and the Natural Science Foundation of Ningbo (2023J167).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
