Abstract
Genomic loss of mismatched human leukocyte antigen (HLA loss) is one of the most vital immune escape mechanisms of leukemic cells after allogeneic hematopoietic stem cell transplantation (allo-HSCT). However, the methods currently used for HLA loss analysis have some shortcomings. Limited literature has been published, especially in lymphoid malignancies. This study aims to evaluate the incidences, risk factors of HLA loss, and clinical outcomes of HLA loss patients. In all, 160 patients undergoing partially mismatched related donor (MMRD) transplantation from 18 centers in China were selected for HLA loss analysis with the next-generation sequencing (NGS)-based method, which was validated by HLA-KMR. Variables of the prognostic risk factors for HLA loss or HLA loss–related relapse were identified with the logistic regression or the Fine and Gray regression model. An HLA loss detection system, HLA-CLN [HLA chimerism for loss of heterozygosity (LOH) analysis by NGS], was successfully developed. Forty (25.0%) patients with HLA loss were reported, including 27 with myeloid and 13 with lymphoid malignancies. Surprisingly, 6 of those 40 patients did not relapse. The 2-year cumulative incidences of HLA loss (22.7% vs 22.0%, P = 0.731) and HLA loss–related relapse (18.4% vs 20.0%, P = 0.616) were similar between patients with myeloid and lymphoid malignancies. The number of HLA mismatches (5/10 vs <5/10) was significantly associated with HLA loss in the whole cohort [odds ratio (OR): 3.15, P = 0.021] and patients with myeloid malignancies (OR: 3.94, P = 0.021). A higher refined-disease risk index (OR: 6.91, P = 0.033) and donor–recipient ABO incompatibility (OR: 4.58, P = 0.057) contributed to HLA loss in lymphoid malignancies. To sum up, HLA-CLN could overcome the limitations of HLA-KMR and achieve a better HLA coverage for more patients. The clinical characteristics and outcomes were similar in patients with HLA loss between myeloid and lymphoid malignancies. In addition, the results suggested that a patient with HLA loss might not always relapse.
Keywords
Introduction
Allogeneic hematopoietic stem cell transplantation (allo-HSCT) is one of the most important therapeutic strategies to improve the survival of patients with hematopoietic malignancies. The mainstay therapy for hematologic malignancies is high-dose chemotherapy and/or radiotherapy followed by transplantation of donor cells. The success of HSCT depends on the eradication of residual malignant cells by immune cells1–3. Currently, although several strategies including interferon (IFN) therapy, targeted drugs, and hypomethylating agents have been introduced for relapse intervention, post-transplantation relapse remains a major drawback of HSCT and carries a poor prognosis4–7. Cell antigen presentation by human leukocyte antigen (HLA) complex is a prerequisite for effective T-cell recognition and cytotoxicity8,9. Downregulation of HLA II molecule expression can impair donor T-cell recognition and mitigate graft-versus-leukemia (GVL) effect10,11. HLA loss through copy-neutral loss of heterozygosity (LOH) of chromosome 6p has also been considered as an important mechanism of relapse, especially in haploidentical donor transplantation12–14.
HLA loss was first reported in 1987 15 . The traditional detection method for HLA loss relied heavily on the sorting and purification of leukemic blasts to perform subsequent HLA typing or single-nucleotide polymorphism (SNP) array analysis, which was difficult to follow. HLA-KMR assay, developed by GenDx, has been widely used in Europe and America for rapid detection of HLA loss 16 . However, the polymerase chain reaction (PCR) panel in HLA-KMR was specific for only three HLA loci (HLA-A, HLA-C, and HLA-DPB1). Therefore, based on next-generation sequencing (NGS), we developed a novel HLA loss detection system named HLA-CLN (HLA chimerism for LOH analysis by NGS), which might overcome the limitations of HLA-KMR, and achieved a better HLA allele coverage (ie, HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1) for more patients.
Several retrospective studies have reported that the incidence of HLA loss was around 30% in relapsed patients with myeloid malignancies after haploidentical HSCT12,13,17. To date, the largest scale for HLA loss reported in literatures was 23 patients with an incidence of 33% 17 . However, few studies concerning HLA loss in lymphoid malignancies18,19 have been reported so far, as the vast majority of published works confined to myeloid malignancies such as acute myeloid leukemia (AML)20–22 and myelodysplastic syndrome (MDS)23,24.
HLA loss is thought to play a role in disease relapse for lacking T-cell-mediated alloreactivity12–14, yet its predictive value remains poorly defined. Until now, all studies for post-transplant HLA loss analysis were conducted for morphological relapsed patients14,17,22,25–27. Today, it is not clear whether patients with HLA loss will experience disease relapse or patients who achieved long-term complete remission (CR) are certainly free of HLA loss. In the present study, the chimerism of below 97% was the principal criterion to select patients for HLA loss analysis.
Materials and Methods
Patient Cohorts and Transplantation Procedures
A total of 3,542 patients with hematologic malignancies undergoing allo-HSCT between January 2006 and November 2020 from 18 transplantation centers in China were screened. Stem cells from a partially mismatched related donor (MMRD) were infused to a corresponding patient. According to the criteria of donor T-cell non-HLA chimerism below 97%, 181 patients were selected. Twenty-one patients were excluded due to early graft failure, leaving 160 eligible patients for this retrospective study. All patients or their families signed institutional review board–approved consent forms. HLA typing and compatibility among donor–recipient pair were performed by molecular typing on HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1 loci and haploidentical type was determined by three or more HLA mismatched loci between family donor and recipient. Graft-versus-host disease (GVHD) prophylaxis regimens included antithymocyte globulin (ATG)-based (Thymoglobin®; Genzyme Polyclonals S.A.S.) and post-transplant cyclophosphamide (PTCy)-based regimens. Bone marrow aspiration was performed at regular intervals of 1 month within the first year and 2 or 3 months thereafter or as clinically indicated for monitoring the disease status with flow cytometry and/or quantitative reverse transcriptase PCR. Non-HLA chimerism analysis on CD3+ T-cells for all patients was analyzed at the time of bone marrow sample collection by PCR-STR (short-term repeats) 28 . Relapse, including molecular and hematological relapse, was defined by the European Society for Blood and Marrow Transplantation (EBMT)29,30. Acute and chronic GVHD were diagnosed and graded according to the National Institutes of Health (NIH) criteria31,32. The study was approved by the Ethics Review Committee of Shanghai General Hospital (IRB# 2021KY005).
HLA Loss Bioinformatics Data Collection Based on NGS
Genomic DNA was extracted from bone marrow mononuclear cells using DNA Blood Kit (GenMag Bio, Changzhou, China) according to the manufacturer’s protocol. Specific DNA fragments for HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1 loci were separately amplified by PCR as previously reported 33 . DNA library for the PCR products was built as described before 34 . Sequencing reagent preparation and processes were performed as recommended by Illumina (NextSeq 500 High Output Kit V2—Reagent Preparation Guide). NGS-based HLA typing was selectively performed for the loci that encompassed the patient–donor HLA mismatches. The average read-depth for all samples was at least 5,000. DNA samples from the patient prior to transplantation and from the corresponding donor were analyzed at the same time to confirm typing information and to assess the level of nonspecific background reads for patient-specific alleles in 100% donor DNA. For HLA chimerism analysis, filters excluding allele calls with low read numbers were removed. HLA-CLN chimerism was determined as the proportion of read counts for patient-specific alleles to recipient (R)- and donor (D)-specific alleles, that is, R/(R + D) × 100%.
HLA Loss Determination and Validation
In the case of HLA loss, HLA-CLN chimerism is expected to be close to zero. When HLA-CLN chimerism (for HLA locus) is ≤0.5% and STR% (for non-HLA locus) is ≥3%, it demonstrates HLA loss has occurred. If HLA-CLN chimerism is ≥3%, it demonstrates a negative HLA loss. If 0.5% ≤ HLA-CLN chimerism <3%, then it could not be determined due to rare false positive at this low chimerism rate. In this study, chimerism on only HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1 was tested, as DPB1 matching was not a requirement for donor selection in China 35 . Although all the loci (HLA-A, HLA-B, HLA-C, HLA-DRB1, HLA-DQB1) have variations from the expected value due to preferential amplification, the average of five loci is close to the expected value. Thus, HLA loss in HLA-CLN system is determined by average chimerism of five loci instead of the specific ones (Supplemental Fig. S1). For limit of detection (LOD), serial dilution samples of five HLA gene segments were tested five times using HLA-CLN system. The simulated HLA chimerism ratio was 0%, 3%, 5%, 10%, 20%, 30%, 50%, and 100% based on PCR-STR.
To verity the consistency, the results of HLA loss analysis by HLA-CLN in 23 patients were compared with that by HLA-KMR (Supplemental Table S1). Mixed chimeric samples with different chimeric levels were produced with two transplantation models (haploidentical and full MMUDs) for sensitivity determination.
To clarify whether nonmalignant cells exposed to potent post-HSCT immune pressure could develop HLA loss, bone marrow samples were harvested from patients with HLA loss who remained in CR. HLA loss status was confirmed by HLA-CLN. The origin of cells with HLA loss was identified by PCR-STR. The cells developing HLA loss were confirmed to be not malignant with flow cytometry analysis, gene sequencing for mutations, and/or fluorescence in situ hybridization (FISH) for cytogenetics. Relapse was classified as HLA loss–related relapse or classical relapse depending on whether the relapse was accompanied with HLA loss.
Statistical Analysis
Differences in distribution were analyzed by the Chi-square test or two-sided Fisher’s exact test if needed. Continuous variables were compared using the nonparametric Mann–Whitney U test. Cumulative incidence of HLA loss and classical relapse was calculated using death without relapse or progression and other forms of relapse as competing events. Fine and Gray’s test was used for comparison to assess the statistical significance in subdistribution hazards model. Overall survival (OS) and relapse-free survival (RFS) were analyzed by the Kaplan–Meier method and compared by the log-rank test. Variables of the prognostic risk factors for HLA loss or HLA loss–related relapse were examined in the logistic regression and competing risk regression model, respectively. In the analysis of the effects of GVHD on HLA loss or HLA loss–related relapse, the occurrence of acute or chronic GVHD was treated as time-varying covariate. Factors in univariate analysis with a P value <0.10 were included in the following multivariate analysis. All analyses were performed using R package (version 3.6.3). Two-sided P < 0.05 was considered statistically significant.
Results
Establishment of HLA-CLN System
To determine the sensitivity of the HLA-CLN system, the simulated chimeric samples were analyzed with the HLA-CLN system. All the five HLA locus genes of HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1 could be detected in samples when the simulated chimeric ratio was >3%, while not all samples could be detected when <3%. The results indicated that the actual sensitivity of the HLA-CLN system was at least 3% (Fig. 1A); that is why the cutoff value of 97% for chimerism was chosen. Then, 50 samples with simulated HLA chimerism rate of 3% for each HLA gene were fully detected with the HLA-CLN system; each assay was repeated five times (Fig. 1B). Integrating the above results, the LOD for the HLA-CLN system was determined to be 3%. This indicated that HLA loss could be demonstrated when recipient-specific HLA chimerism was close to 0 (≤0.5%) and non-HLA chimerism was 3% or more (≥3%) in the HLA-CLN system (HLA-CLN ≤0.5% and STR ≥3%). We evaluated the consistency of the HLA-CLN and HLA-KMR assays (Supplemental Table S1). Specifically, 23 patient samples with different T-cell chimerisms were analyzed (Fig. 1C, D). We observed concordant results between HLA-KMR and HLA-CLN in all samples except two patients (CFL and MDQ), yielding an agreement rate of 91.7% (Supplemental Table S1).

HLA-CLN assay system. Positive samples of HLA loss detected by HLA-CLN were confirmed with the HLA-KMR assay, especially when the donor T-cell chimerism was between 95% and 97% and for special cases (ie, patients with ALL and nonrelapsed patients). (A) The simulated HLA chimerism ratio was 0%, 3%, 5%, 10%, 20%, 30%, 50%, and 100% based on PCR-STR, while the HLA chimerism ratio was detected by HLA-CLN, which is represented as mean ± SD for three independent tests. Serial dilution samples of five HLA genes were tested five times using the NGS detection system. (B) Fifty samples with 3% simulated HLA chimerism with PCR-STR were tested with the HLA-CLN system. (C, D) Host chimerism quantification with non-HLA markers (STR on T-cells); HLA-KMR and HLA-CLN were performed on 11 HLA loss–negative (C) and 12 HLA loss–positive patients (D) based on HLA-KMR. As expected, the two methods of HLA-KMR and HLA-CLN yielded concordant results in all HLA loss–negative and HLA loss–positive samples except two patients (CFL and MDQ). One patient named CFL with a host STR chimerism of 46.89% presented HLA loss with HLA-KMR (HLA chimerism of 0.724% for gene A), while with the HLA-CLN system, the patient did not present HLA loss because the HLA chimerism for gene loci A, B, C, DRB1, and DQB1 was 0.93%, 7.92%, 17.34%, 0.92 %, and 5.21%, respectively (average chimerism of five gene loci, 6.46%). Combined with the analysis of STR chimerism, it was speculated that the patient was likely to be in the process of HLA loss and has not been completely lost. For the other patient named MDQ, HLA loss was interpreted to be positive according to the HLA chimerism of 0.323% for gene A locus via HLA-KMR combined with the host STR chimerism of 3.12%, whereas the HLA chimerisms were 2.96%, 0%, match, 0.24%, and 0.51% for A, B, C, DRB1, and DQB1 loci, respectively (average chimerism rate of five gene loci, 1.23%) with the HLA-CLN system, so HLA loss for the patient was judged as negative. HLA-CLN, HLA chimerism for loss of heterozygosity analysis by next-generation sequencing; HLA: human leukocyte antigen; ALL: acute lymphoblastic leukemia; STR: short-term repeats; NGS: next-generation sequencing; PCR: polymerase chain reaction.
Patient Characteristics
Clinical data of 160 patients who underwent transplantation from partially MMRDs (146 patients from haploidentical donors) were retrospectively collected. High-resolution HLA typing revealed that 103 (64.4%) of 160 donor–recipient pairs had five HLA-A, HLA-B, HLA-C, HLA-DRB1, HLA-DQB1 locus mismatches, 27 (16.9%) had four mismatches, and 30 (18.8%) had three or less HLA mismatches at these loci. The demographics and disease- and transplant-related characteristics of patients are summarized in Table 1 and Supplemental Table S2. Most of the patients were diagnosed with myeloid malignancies, mainly AML (89/110, 80.9%) and MDS (17/110, 15.5%). Fifty patients were diagnosed with lymphoid malignancies, mainly acute lymphoblastic leukemia (ALL) (45/50, 90.0%). The median age was 32.0 (range, 1.0–64.0) years. The patients with lymphoid malignancies were younger than those with myeloid malignancies (P < 0.001), while the donors for patients with lymphoid malignancies were relatively older than those with myeloid malignancies (P = 0.006). At the time of transplantation, 104 (65.0%) of 160 patients had active disease, which included more patients with myeloid malignancies than lymphoid malignancies (40.9% vs 22.0%, P = 0.032). The proportions of haploidentical transplantation were similar between patients with myeloid and lymphoid malignancies (90.9% vs 92.0%, P = 0.940). No significant differences in the incidences of acute GVHD between patients with and without HLA loss were found (P = 0.095). A higher incidence of chronic GVHD was observed in patients with HLA loss than those without (P = 0.026). The follow-up time was longer for patients with HLA loss than those without HLA loss (P < 0.001), most likely due to the difference in relapse time. The other clinical characteristics were similar between patients with and without HLA loss (Table 1).
Patient Characteristics and Clinical Outcomes.

HLA: human leukocyte antigen; HSCT: hematopoietic stem cell transplantation; CR: complete remission; NR: non-remission; R-DRI: refined-disease risk index; HCT-CI: hematopoietic cell transplantation–specific comorbidity index; ECOG: performance status of Eastern Cooperative Oncology Group; GVHD: graft-versus-host disease; ATG: antithymocyte globulin; CTX: cyclophosphamide; PBSC: peripheral blood stem cell; UCB: umbilical cord blood; BM: bone marrow; CMV: cytomegalovirus; aGVHD: acute graft-versus-host disease; cGVHD: chronic graft-versus-host disease.
Incidences of HLA Loss and HLA Loss–Related Relapse
To match the sensitivity of the HLA-CLN system, the patients were enrolled into the study according to donor T-cell chimerism below 97%, rather than the disease status (relapse or not), so the incidences of HLA loss and HLA loss–related relapse were analyzed separately. At a median follow-up of 20.1 (range, 1.4–68.3) months, 40 (25.0%) of 160 patients were found to develop HLA loss. The 1- and 2-year cumulative incidences of HLA loss were 12.7% [95% confidence interval (CI), 6.4%–19.0%] and 22.7% (95% CI, 14.7%–30.7%) in myeloid malignancies, whereas 14.0% (95% CI, 4.2%–23.8%) and 22.0% (95% CI, 10.0%–34.0%) in lymphoid malignancies (P = 0.731) (Fig. 2A).

Cumulative incidences of HLA loss and HLA loss relapse in partially MMRD transplantation. (A) Cumulative incidences of HLA loss in patients with myeloid malignancies (red) or lymphoid malignancies (blue). (B) Cumulative incidences of classical relapse (blue) or HLA loss relapse (red) after transplantation. (C) Cumulative incidences of HLA loss relapse in patients with myeloid malignancies (red) or lymphoid malignancies (blue). HLA: human leukocyte antigen; MMRD: mismatched related donor; HSCT: hematopoietic stem cell transplantation.
The 2-year cumulative incidence of HLA loss–related relapse [18.9% (95% CI, 12.8%–25.0%)] was significantly lower than that of classical relapse [55.2% (95% CI, 47.5%–63.0%)] (P < 0.001; Fig. 2B). The 1- and 2-year cumulative incidences of HLA loss–related relapse were 10.1% (95% CI, 4.4%–15.8%) and 18.4% (95% CI, 11.1%–25.8%) for myeloid malignancies, whereas 18.0% (95% CI, 7.3%–28.8%) and 20.0% (95% CI, 8.8%–31.2%) for lymphoid malignancies. The cumulative incidences of HLA loss–related relapse were similar between myeloid and lymphoid malignancies (P = 0.616) (Fig. 2C). The median relapse time was 10.0 (range, 2.0–46.0) months and 4.6 (range, 1.3–24.6) months for HLA loss and classical relapse, respectively (P < 0.001).
Characteristics of Patients With HLA Loss
The characteristics and clinical outcomes of patients with HLA loss are shown in Table 2. The genomic loss of both HLA class I and class II loci occurred in 33 (82.5%) of 40 patients, and either HLA-I (n = 6) or HLA-II (n = 1) locus-specific loss occurred in 7 (17.5%) of 40 patients. The median time of HLA loss was 10.1 (range, 1.0–49.3) months after transplantation for all HLA loss patients. For myeloid malignancies, the median time is 11.2 (range, 1.5–49.3) months, whereas it is 9.1 (range, 1.0–46.6) months for lymphoid malignancies. In all, 31 of 40 patients with HLA loss experienced hematological relapse, whereas 2 of 40 patients with molecular relapse (Nos 37, 75) and 1 of 40 ALL patient died too early to be evaluated (No. 21). It was worth noting that 6 of 40 patients with HLA loss did not relapse, with a median follow-up time of 46.7 months (range, 25.0–66.0) months, including 5 with myeloid malignancies (3 with AML and 2 with MDS) and 1 with ALL. Among these six patients, five cases (Nos 20, 28, 31, 36, 161) developed both HLA-I and HLA-II gene locus loss and one case (No. 4) developed a single HLA-DQB1 locus loss. To confirm that the patients with HLA loss did not relapse, bone marrow samples from two patients (at 36 and 66 months after HSCT for No. 28 and No. 20, respectively) were further analyzed for HLA loss with HLA-CLN and HLA-KMR, and the results were positive for HLA loss. In the meantime, no abnormal cells were detected by flow cytometry.
Characteristics and Clinical Outcomes of Patients With HLA Loss.
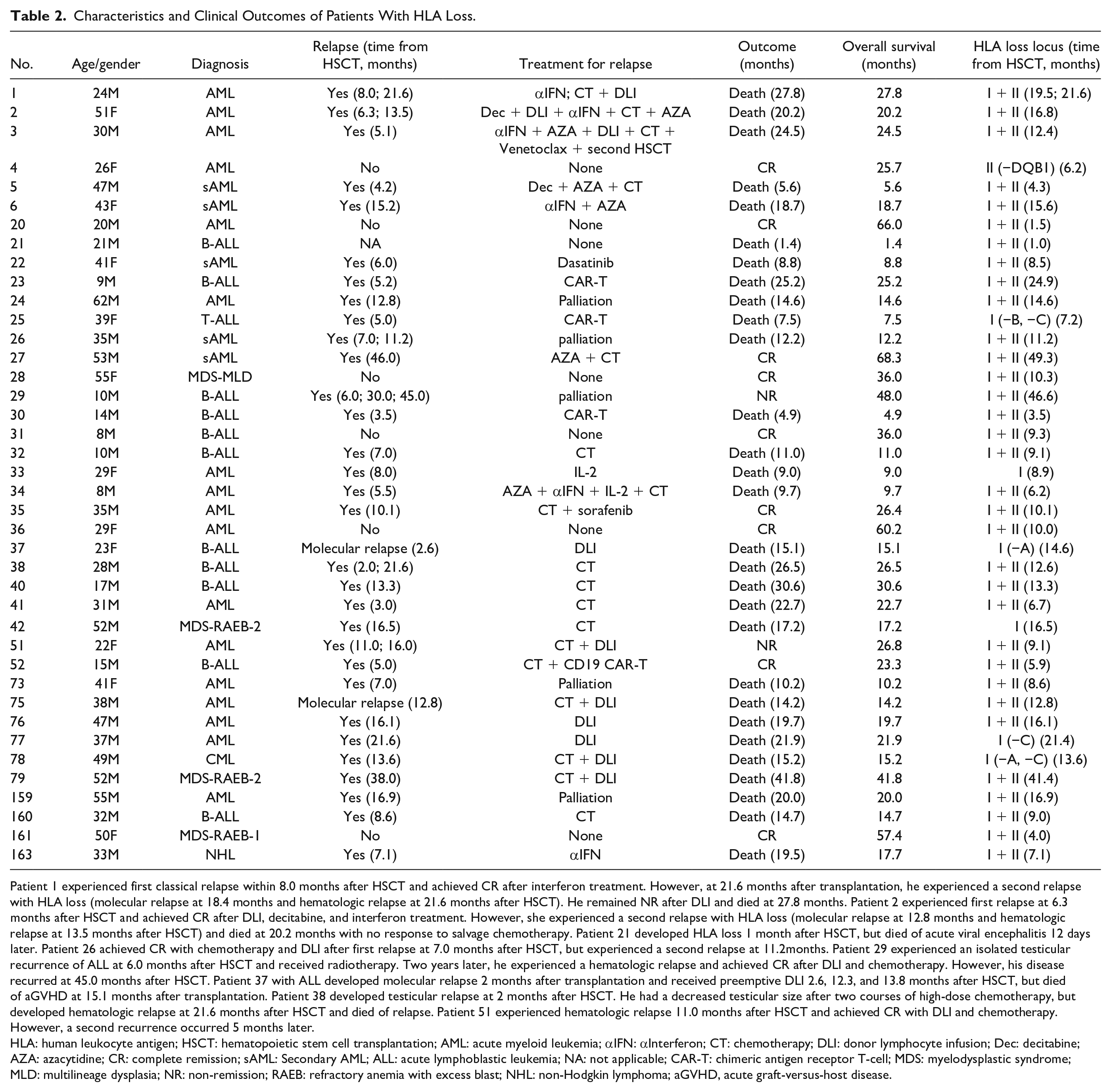
Patient 1 experienced first classical relapse within 8.0 months after HSCT and achieved CR after interferon treatment. However, at 21.6 months after transplantation, he experienced a second relapse with HLA loss (molecular relapse at 18.4 months and hematologic relapse at 21.6 months after HSCT). He remained NR after DLI and died at 27.8 months. Patient 2 experienced first relapse at 6.3 months after HSCT and achieved CR after DLI, decitabine, and interferon treatment. However, she experienced a second relapse with HLA loss (molecular relapse at 12.8 months and hematologic relapse at 13.5 months after HSCT) and died at 20.2 months with no response to salvage chemotherapy. Patient 21 developed HLA loss 1 month after HSCT, but died of acute viral encephalitis 12 days later. Patient 26 achieved CR with chemotherapy and DLI after first relapse at 7.0 months after HSCT, but experienced a second relapse at 11.2months. Patient 29 experienced an isolated testicular recurrence of ALL at 6.0 months after HSCT and received radiotherapy. Two years later, he experienced a hematologic relapse and achieved CR after DLI and chemotherapy. However, his disease recurred at 45.0 months after HSCT. Patient 37 with ALL developed molecular relapse 2 months after transplantation and received preemptive DLI 2.6, 12.3, and 13.8 months after HSCT, but died of aGVHD at 15.1 months after transplantation. Patient 38 developed testicular relapse at 2 months after HSCT. He had a decreased testicular size after two courses of high-dose chemotherapy, but developed hematologic relapse at 21.6 months after HSCT and died of relapse. Patient 51 experienced hematologic relapse 11.0 months after HSCT and achieved CR with DLI and chemotherapy. However, a second recurrence occurred 5 months later.
HLA: human leukocyte antigen; HSCT: hematopoietic stem cell transplantation; AML: acute myeloid leukemia; αIFN: αInterferon; CT: chemotherapy; DLI: donor lymphocyte infusion; Dec: decitabine; AZA: azacytidine; CR: complete remission; sAML: Secondary AML; ALL: acute lymphoblastic leukemia; NA: not applicable; CAR-T: chimeric antigen receptor T-cell; MDS: myelodysplastic syndrome; MLD: multilineage dysplasia; NR: non-remission; RAEB: refractory anemia with excess blast; NHL: non-Hodgkin lymphoma; aGVHD, acute graft-versus-host disease.
The prognosis of patients with HLA loss was poor. The 2-year OS (Fig. 3A) and RFS (Fig. 3B) were 42.5% (95% CI, 29.6%–60.9%) and 20.0% (95% CI, 10.8%–37.2%), respectively. The 2-year nonrelapse mortality (NRM) was significantly lower in patients with HLA loss than those without (P = 0.035) (Fig. 3C). The cumulative incidences of relapse (CIRs) were similar between patients with [77.5% (95% CI, 64.1%–90.9%)] and without [73.8% (95% CI, 65.8%–81.8%)] HLA loss (P = 0.610) (Fig. 3D). The 2-year OS was 40.7% (95%C I, 25.9%–64.2%) in patients with myeloid malignancies and 46.2% (95% CI, 25.7%–83.0%) in patients with lymphoid malignancies (P = 0.600). The 2-year CIRs were similar between patients with myeloid [74.1% (95% CI, 56.8%–91.4%)] and lymphoid [76.9% (95% CI, 51.1%–100%)] malignancies (P = 0.119). A higher RFS was observed in patients with myeloid malignancies (25.9%, 95% CI, 13.7%–49.0%) compared with patients with lymphoid malignancies (7.7%, 95% CI, 1.2%–50.6%) (P = 0.004).

Clinical outcomes of patients with and without HLA loss. (A) Overall survival of patients with and without HLA loss (P = 0.100). (B) Relapse-free survival of patients with and without HLA loss (P = 0.020). (C) Cumulative incidences of NRM of patients with and without HLA loss (P = 0.035). (D) Cumulative incidences of relapse of patients with and without HLA loss (P = 0.610). HLA: human leukocyte antigen; NRM: nonrelapse mortality; HSCT: hematopoietic stem cell transplantation.
Characteristics of Patients With HLA Loss–Related Relapse
Thirty-three patients with HLA loss relapsed, which included 22 patients with myeloid malignancies and 11 with lymphoid malignancies. The median time of relapse for myeloid malignancies was 12.4 (3.0–46.0) months, whereas 5.2 (2.0–45.0) months for lymphoid malignancies. Nine of 22 patients with myeloid malignancies developed chronic graft-versus-host disease (cGVHD) (2 mild, 5 moderate, and 2 severe). Three patients (Nos 37, 76, 77) were only administered with donor lymphocyte infusion (DLI), two of whom (Nos 37, 77) died due to disease progression; the other developed severe post-treatment aGVHD and died. Two patients (Nos 1, 163) were treated with IFN-α. Patient 1 presented an initial response (without loss at first relapse), but subsequently developed a second relapse. Then the patient was administered with salvage chemotherapy plus DLI, but remained NR and died. Patient 163 did not respond to IFN-α and died from relapse. Five patients received palliative care and died shortly due to disease progression. Twenty patients received various salvage therapies including DLI, targeted therapies, chemotherapy, and chimeric antigen receptor T (CART) cells, alone or in combination, but showed disappointing results. Three patients (Nos 27, 35, 52) achieved long-term CR with a median OS of 26.4 months. They received chemotherapy in combination with azacitidine, sorafenib, or CD19-CART cells, respectively (Table 2). The response rates of salvage therapies for late-relapsed patients (6 months after transplantation) and early-relapsed patients (within 6 months after transplantation) were 12.5%, and 8.3%, respectively. The 2-year OS of late-relapsed patients was 35.0% (95% CI, 19.3%–63.6%), which was similar to that of early-relapsed patients [37.5% (95% CI, 15.3%–91.7%)] (P = 0.300).
Eleven of 13 ALL patients with HLA loss relapsed. Seven (64%) patients developed cGVHD (five mild, one moderate, and one severe) when relapsed. Five patients with active disease and five in CR2 or CR3 at transplantation experienced early relapse (within 8.6 months after transplant), while one patient in CR1 relapsed later (1 year after transplantation).
The 2-year OS of patients with HLA loss–related relapse was 33.3% (95% CI, 20.6%–54.0%), which was concordant with classical relapse [29.5% (95% CI, 20.2%–43.0%), P = 0.300]. The 2-year OS was 27.3% (95% CI, 13.8%–54.0%) for myeloid malignancies and 45.5% (95% CI, 23.8%–86.8%) for lymphoid malignancies (P = 0.700).
Risk Factors for HLA Loss and HLA Loss–Related Relapse
In the univariate analysis for HLA loss in the whole cohort (160 patients, Supplemental Table S3), the number of HLA-mismatched loci between donor–recipient pair (5/10 vs <5/10), ABO incompatibility, and cGVHD were associated with an increased risk for HLA loss. In the multivariate analysis (Table 3), 5 of 10 HLA mismatches [odds ratio (OR): 3.15, 95% CI, 1.26–9.06; P = 0.021] and cGVHD (OR: 2.69; 95% CI, 1.21–6.00; P=0.015) showed a significant association with HLA loss. For patients with myeloid malignancies, 5 of 10 HLA mismatches (OR: 3.94; 95% CI, 1.34–14.50; P = 0.021) were the only independent risk factor, whereas for patients with lymphoid malignancies, high- or very-high-risk R-DRI (OR: 6.91; 95% CI, 1.33–54.45; P = 0.033) was an independent risk factor for HLA loss. A weaker correlation was observed between donor–recipient ABO incompatibility and HLA loss in lymphoid malignancies (OR: 4.58; 95% CI, 1.00–24.38, P = 0.057).
Multivariate Analysis for Factors Associated With HLA Loss or HLA Loss Relapse.

HLA: human leukocyte antigen; OR: odds ratio; HR: hazard ratio; CI: confidence interval; cGVHD: chronic graft-versus-host disease; R-DRI: refined-disease risk index.
*P < 0.05.
In the whole cohort for HLA loss–related relapse analysis, a higher R-DRI, higher number of HLA mismatch (five loci), ABO incompatibility, and cGVHD were putative variables in univariate analysis (Supplemental Table S4). In multivariate analysis (Table 3), occurrence of cGVHD increased the risk of HLA loss–related relapse [hazard ratio (HR): 2.17; 95% CI, 1.08–4.36; P = 0.029]. Five of 10 HLA mismatches had a strong trend as an independent risk factor (HR: 2.61; 95% CI, 0.97–7.05; P = 0.058). For patients with myeloid malignancies, the number of HLA mismatches (5/10) was an independent risk factor (HR: 3.72; 95% CI, 1.09–12.68; P = 0.036). For lymphoid malignancies, a higher R-DRI significantly increased the risk of HLA loss–related relapse (HR: 8.51; 95% CI, 1.13–64.32; P = 0.038).
ATG was administered in most patients (80%), PTCy in 10% of patients, and ATG in combination with PTCy in 10% for GVHD prophylaxis. The 1-year cumulative incidences of HLA loss and HLA loss–related relapse in patients with different regimens of GVHD prophylaxis had no significant differences (P = 0.146 for HLA loss, P = 0.491 for HLA loss–related relapse).
Discussion
In this study, an NGS-based HLA loss detection system named HLA-CLN was developed and utilized to analyze HLA loss for patients after partially MMRD transplantation from 18 centers in China. Forty patients with HLA loss were documented, including 27 with myeloid malignancies and 13 with lymphoid malignancies. The cumulative incidence of HLA loss and other clinical characteristics of patients with myeloid malignancies were concordant with lymphoid malignancies. To our knowledge, it was first reported in our study that the patients with HLA loss might not definitely experience disease relapse.
HLA-KMR is a rapid and economic quantitative PCR (qPCR)-based assay for the detection of HLA loss. However, the qPCR reactions were only specific for HLA-A, HLA-C, and HLA-DPB1 alleles and could not cover all HLA loci 16 , while HLA-CLN might overcome the limitations of HLA-KMR and achieve a better coverage for more patients. First, HLA-CLN might cover more HLA alleles for HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1. Second, HLA haplotype matching was defined according to HLA-A, HLA-B, HLA-C, HLA-DRB1, and HLA-DQB1 alleles, not including HLA-DPB1 in most reports about haploidentical transplantation36–39, which could lower the coverage for patients with HLA-KMR. Despite several advantages, limitations of HLA-CLN should also be acknowledged. It does not target HLA II DPB1 locus, which would limit its usage in some patients with HLA-DPB1 locus–mismatched matched unrelated donor (MUD). In addition, imbalanced amplification of HLA locus genes in HLA-CLN might lead to false positives, especially when the non-HLA chimerism rate for a host was relatively low. A positive result of HLA loss should be carefully interpreted when a donor cell chimerism was of 95%–97%. Moreover, during the validation of HLA-CLN assay, we found an agreement rate of 91.7% with HLA-KMR. The results with HLA-CLN and HLA-KMR for the two patients (CFL and MDQ as shown in Fig. 1D) were discordant, but unfortunately were unable to be verified because of the limitation of sample availability. The discrepancy between the two methods might be caused by different platforms (NGS vs qPCR) and different algorithms of HLA-CLN and HLA-KMR. In consequence, the HLA-CLN system needs to be validated and verified with a larger sample size.
Forty patients were reported in our study, which was the largest scale about HLA loss in the partially MMRD HSCT setting to date. Although HLA loss could develop at diagnosis or pretransplantation23,40,41, it is rare in AML at diagnosis or relapse after intervention with chemotherapy, with a rate of at least 10 times less frequent than that in patients at relapse after HSCT from large series analysis by SNP array42–44. In addition, bone marrow samples at diagnosis from two patients with ALL were analyzed and no evidence of HLA loss in our study was found (data not shown). Although HLA loss occurring at pretransplantation could not be excluded in our study, it did not obviously affect the results. Vago and colleagues 45 recently presented the first global collaborative study about the incidence of HLA loss for post-transplant relapsed patients using HLA typing for sorted blasts, HLA-KMR, or a novel NGS method covering HLA-A, HLA-B, HLA-C, HLA-DRB1, HLA-DQB1, and HLA-DPB1 loci. The results showed that the incidences of HLA loss were 22.6% for haploidentical HSCT, followed by unrelated donor HSCT (11.9% in MMUD and 4.3% in MUD) and umbilical cord blood transplantation (0%). The incidence of HLA loss for haploidentical HSCT was concordant with our data. For myeloid malignancies, the incidence of HLA loss in our study was consistent with previous reports12,13. For lymphoid malignancies (mainly ALL), we first reported that the incidence of HLA loss was concordant with that for myeloid malignancies in partially MMRD transplantation.
Loss of the unshared HLA haplotype may circumvent allogeneic antigen recognition and render donor T-cells (ie, DLI or second HSCT from the same donor) to be ineffective for relapse intervention. Other treatments such as chemotherapy, novel targeted therapies, and second HSCT from an alternative donor have the potential to improve survival and achieve long-term responses. In our study, three patients were only administered with DLI, all of whom died of relapse or severe aGVHD, while three patients treated with chemotherapy in combination with hypomethylating agents or adoptive T-cell therapy achieved long-term survival. In addition, one AML patient (No. 4) with HLA DQB1 locus loss responded to azacitidine plus IFN-α and achieved a good prognosis, which indicated that a single HLA II locus loss might not affect the antigen presentation by HLA-I molecules.
GVL effects are critical for eradicating the residual leukemia cells post-transplantation to prevent relapse. Measurable residual disease is the most important relapse risk factor for patients with ALL46–49. In our study, 9 of 10 ALL patients with HLA loss, including 5 in NR and 5 in CR2 or more at transplantation, relapsed early after transplantation. This suggested that the preconditioning intensity is more important than GVL for early relapsed patients with HLA loss, while GVL is more crucial for late relapsed patients. The results indicated that a different haplotype donor compared with the haplotype donor of the first transplantation should be selected for a second transplantation to a relapsed ALL patient, especially for a late relapsed patient, as suggested for a relapsed patient with myeloid malignancy 50 . HLA loss should also be routinely monitored for a relapsed ALL patient to select a better salvage therapeutic modality.
Notably, an interesting phenomenon was found in our study. Six patients with HLA loss did not relapse. The results suggested that normal cells rather than malignant cells could develop HLA loss51,52. In that case, we assumed that the GVL effects from non-T-cells might play a critical role in preventing leukemia relapse. The importance of natural killer (NK) cell alloreactivity is emphasized to achieve the GVL effect in partially MMRD, especially in haploidentical transplantation 53 . HLA loss may enhance the NK cell alloreactivity to kill mutant leukemic cells54–56. The alloreactivity of NK cells in nonrelapsed patients with HLA loss needs to be further investigated.
The incidence of HLA loss was the highest of 30% in haploidentical transplantation, second by about 10% in MUD transplantation for patients with myeloid malignancies17,22. Only one patient with extramedullary myeloid sarcoma was reported to develop HLA loss in matched related donor transplantation 57 . These data suggested that HLA loss was associated with the intensity of HLA mismatches because the greater number of HLA locus mismatches could result in stronger immune pressure against patient-specific HLA. Our data showed that the number of HLA mismatches was an independent risk factor for HLA loss in the whole cohort and for patients with myeloid malignancies, although not with lymphoid malignancies. HLA mismatched intensity (five locus mismatches) also increases the risk of HLA loss–related relapse (P = 0.058in the whole cohort). According to Crucitti and coworkers’ report 17 , 23 relapsed myeloid malignant patients were documented with HLA loss. They found that the number of HLA mismatches was not associated with HLA loss–related relapse. However, active disease at HSCT and older patient were strongly correlated to HLA loss occurrence, while cGVHD and higher T-cell dose in graft mildly correlated. The differences in the effects of HLA-mismatched locus number on HLA loss between our study and Crucitti’s report might be associated with the different size of samples. In theory, the occurrence of cGVHD could trigger stronger immune pressure on patient-specific HLA and result in the development of HLA loss. In our study, the occurrence of cGVHD (total 19 cases) only increased the risk of HLA loss (OR: 2.69, P = 0.015) and HLA loss–related relapse (HR: 2.17, P = 0.029) in the whole cohort, but not in the myeloid or lymphoid malignancy subgroups. In a previous report 17 regarding a total of nine cases with cGVHD, cGVHD was weakly associated with HLA loss (P = 0.082). In our study, the limited number of patients in the subgroups of myeloid (nine cases) or lymphoid (seven cases) malignancies might be the reason for cGVHD not showing significant impacts on HLA loss.
High- or very-high-risk R-DRI was significantly associated with HLA loss and loss relapse only in patients with lymphoid malignancies, but not with myeloid malignancies and in the whole cohort, which suggested that the risk factors for HLA loss and loss relapse were different between lymphoid and myeloid malignancies. To date, few studies with good magnitude of HLA loss in patients with lymphoid malignancies were reported. There are several potential reasons for the differences in risk factors for HLA loss and HLA loss–related relapse between myeloid and lymphoid malignancies. First, myeloid and lymphoid malignancies are a diverse group of hematologic neoplasms derived from different hematologic lineage clones. Second, apart from heterogeneous disease type, the variations could also be related to the differences in disease status (CR vs active disease) before HSCT, although no statistical difference between the two groups was found in R-DRI. Third, there were differences in sample sizes of two groups (ie, myeloid malignancies: 110 patients; lymphoid malignancies: 50 patients). These findings must be interpreted cautiously and further study with a larger cohort is required for validation, especially for the risk factor analysis.
As expected, the survival of patients with HLA loss or HLA loss–related relapse was poor. No significant difference was shown in the survival of patients with or without HLA loss. One possible reason was that a fraction of patients with HLA loss did not relapse, and the NRM of patients with HLA loss was lower than those without HLA loss. The relatively small sample size of patients might limit the statistical power to detect associations between survival, response to salvage therapies, and the timing of HLA loss. However, the analysis of HLA loss has already been implemented in the routine management in our center for post-HSCT follow-up, which may allow us to carry out a further larger study to confirm the results.
In conclusion, HLA-CLN, a new detection system for HLA loss based on NGS, was developed and could cover more patients who require HLA loss analysis. In the present study, we documented 40 patients with HLA loss by HLA-CLN. The incidences of HLA loss in patients with lymphoid and myeloid malignancies were concordant. Furthermore, the results suggested that a patient with HLA loss might not always relapse, although the mechanisms are not fully elucidated and need to be further explored.
Supplemental Material
sj-docx-1-cll-10.1177_09636897221102902 – Supplemental material for Clinical Characteristics and Outcome Analysis for HLA Loss Patients Following Partially Mismatched Related Donor Transplantation Using HLA Chimerism for Loss of Heterozygosity Analysis by Next-Generation Sequencing
Supplemental material, sj-docx-1-cll-10.1177_09636897221102902 for Clinical Characteristics and Outcome Analysis for HLA Loss Patients Following Partially Mismatched Related Donor Transplantation Using HLA Chimerism for Loss of Heterozygosity Analysis by Next-Generation Sequencing by Andi Wang, Wenjun Li, Fei Zhao, Zhongzheng Zheng, Ting Yang, Sanbin Wang, Jinsong Yan, Jianpin Lan, Shengjin Fan, Mingfeng Zhao, Jianpin Shen, Xin Li, Tonghua Yang, Quanyi Lu, Ying Lu, Hai Bai, Haiyan Zhang, Dali Cai, Ling Wang, Zhiyang Yuan, Erlie Jiang, Fang Zhou and Xianmin Song in Cell Transplantation
Footnotes
Acknowledgements
The authors would like to express their gratitude to all the medical staff at different centers for their contribution and all the patients and their families for participation in this study.
Author Contributions
Xianmin Song, Fang Zhou, and Erlie Jiang designed research studies; Andi Wang, Wenjun Li, and Fei Zhao collected the data and wrote the manuscript. Xianmin Song, Zhongzheng Zheng, and Zhiyang Yuan designed the detection method based on NGS and Zhongzheng Zheng detected HLA loss for all the samples. Ting Yang, Sanbin Wang, Jinsong Yan, Jianpin Lan, Shengjin Fan, Mingfeng Zhao, Jianpin Shen, Xin Li, Tonghua Yang, Quanyi Lu, Ying Lu, Hai Bai, Haiyan Zhang, Dali Cai, and Ling Wang contributed to treating patients. All authors read and approved the final manuscript.
Data Availability
All the authors confirm the availability of data and materials. The datasets generated during and/or analyzed during the current study are available from the corresponding author(s) upon request.
Consent to Publish
All the authors confirm that the manuscript represents our honest work and agree to consent for publication.
Ethical Approval
The collection of HLA typing data and DNA samples for HLA loss analysis was approved by the Ethics Review Committee of Shanghai General Hospital (2021KY005).
Statement of Human and Animal Rights
This study was approved by the Ethics Review Committee of Shanghai General Hospital (2021KY005).
Statement of Informed Consent
All patients or their families signed institutional review board–approved consent forms. Written informed consent was obtained from the patient(s) for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Three-Year Development Project from Shanghai Shenkang Hospital Development Center (grant numbers SHDC2020CR1012B and 16CR1010A) and Translational Research Grant of NCRCH (grant number 2020ZKPC02).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
