Abstract
This study aimed to explore the characteristics of TGFBR1–epidermal growth factor receptor (EGFR)–CTNNB1–CDH1 axis in regulating the invasion and migration in lung cancer. Using the small interfering RNA technology, EGFR was silenced in H2170 and H1299 cells. Then, the colony formation, migration, and invasion abilities were detected using colony-forming assay and transwell assay. Moreover, the mRNA expression of smad2, smad3, CTNNB1, and CDH1, and the protein expression of TGFBR1, CDH1, and TCF were determined using the real-time polymerase chain reaction and western blotting. The results showed that silencing EGFR could significantly decrease the colony-forming ability in H2170 and H1299. Knocking down EGFR could significantly inhibit the invasion and migration ability of H2179 and H1299. Inhibiting the expression of EGFR could significantly decrease the expression of smad2, smad3, CDH1, and CTNNB1, with all P-values <0.05. In addition, silencing EGFR could markedly decrease the expression of TGFBR1 and CDH1 in H1299 and H2170, with all P-values <0.05. In conclusion, silencing EGFR could significantly regulate the progression of lung cancer via TGFBR1–EGFR–CTNNB1–CDH1 axis in Wnt signaling pathway.
Introduction
Lung cancer is an important health problem and responsible for a great part of cancer-associated mortality worldwide 1 . In China, lung cancer is the first leading cause of cancer-related mortality, which accounts for more than 569,000 deaths per year 2 . Non-small-cell lung cancer (NSCLC) is the main type of lung cancer, accounting for 87% of lung cancer 3 . Despite the development of cancer treatment, the outcomes for lung cancer patients are still unsatisfied. Therefore, it is important to explore the mechanism of NSCLC improving the therapy outcomes of patients suffering from lung cancer.
Wnt signaling pathway is a crucial pathway involved in the development of cancer, including lung cancer 4,5 . Activation of Wnt signaling pathway accelerates the progression of lung cancer, leading to a poor prognosis 6 . Rab37 could suppress the Wnt signaling to inhibit the stemness of lung cancer cells via mediating exocytosis of secreted firzzled-relayed protein 1 7 . SOX30 could suppress the metastasis of lung cancer via Wnt/β-catenin signaling pathway 8 . More and more studies have revealed that transforming growth factor β (TGF-β) signaling pathway could crosstalk with Wnt signaling in regulating the development of cancer. Wang et al. have reported that TGF-β-induced factor homeobox1 increases the development of colorectal cancer via activating Wnt/β-catenin signaling pathway 9 . Peng et al. have demonstrated that FOXO1 mediates the signaling pathway crosstalk between TGF-β and Wnt signaling pathways to regulate the progression of colorectal cancer 10 . However, the crosstalk between TGF-β and Wnt signaling pathways is still unclear in the progression of lung cancer.
In this study, epidermal growth factor receptor (EGFR), a link between TGF-β and Wnt signaling pathways, was silenced followed by the detection of the downstream targets to explore the underlying mechanism of the crosstalk between TGF-β and Wnt signaling pathways in regulating the development of lung cancer. According to these investigations, we hope to provide some glances on the crosstalk between TGF-β and Wnt signaling pathways in lung cancer.
Materials and Methods
Cell Culture
Human NSCLC cell lines H2170 and H1299 were purchased from the Chinese Academy of Science cell bank (Shanghai, China). Both cells lines were maintained in RPMI-1640 (Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS, Gibco), 100 IU/ml penicillin, and 100 μg/ml streptomycin in a humidity incubator with 5% CO2 in atmosphere at 37°C.
Small Interfering RNA Silencing EGFR
The sequences for si-EGFR and si-negative control (NC) sequences and their corresponding lentivirus were purchased from Genpharm (Shanghai, China). First, H1299 and H2170 were seeded in a six-well plate overnight with 70% confluency. Then, the cells were transfected with lentivirus using lipofectamine 2000 according to the manufacturer’s protocol. After 48-h transfection, cells were harvested and verified for the following instrument. The model group had no treatment.
Colony-Forming Assay
A total of 5 × 103 cells in each group were seeded in 6-cm plates and suppled with RPMI-1640 medium containing 10% FBS for 3 weeks. Subsequently, colonies in plates were fixed with methanol and stained with 0.1% crystal violet (Sagon, Shanghai, China) dissolved in 20% methanol at room temperature for 0.5 h. After rinsing, the colony was pictured and calculated.
Transwell Invasion Assay
Cell invasion ability was assessed using transwell plates (Corning Life Sciences, Tewksbury, MA, USA). First, the transwell plates were coated with Matrigel Basement Membrane Matrix according to the manufacturer’s protocol. Following this, a total of 2 × 104 cells in 200 µl medium supplemented with 2% FBS were seeded in the upper chamber and 600 µl medium containing 10% FBS was filled into the lower chamber. After incubation for 48 h, cells in the upper chamber were removed by a cotton swab. Then, the chambers were fixed in methanol at room temperature for 10 min and stained with 0.1% crystal violet (Sagon) dissolved in 20% methanol at room temperature for 0.5 h.
Quantitative Real-Time Polymerase Chain Reaction
Total RNA was extracted from cells using TRIzol reagent (Takara, Dalian, China) according to the manufacturer’s protocol. Then, a total of 3 µg total RNA was used to synthesize cDNA using a PrimerScript RT Reagent Kit (Takara, Shiga, Japan). Following this, real-time polymerase chain reaction was conducted using SYBR Green (Sigma, St. Louis, Missouri, USA) on an ABI 7500 instrument (Applied Biosystems, Foster City, CA, USA). Glyceraldehyde 3-phosphate dehydrogenase was used as the internal control. Relative gene expression was analyzed using 2- ΔΔCt method.
Western Blotting
After culturing, cells were lysed using RIPA lysis buffer (Pierce, Rockford, IL, USA) supplemented with a protease inhibitor cocktail (Sigma). After centrifugation at 4°C for 10 min, the supernatant was harvested and qualified using bicinchoninic acid assay method. Following this, 30 µg of protein was subjected for sodium dodecyl sulfate polyacrylamide gel electrophoresis and electrophoretically transferred onto polyvinylidene difluoride membrane. Then, membranes were blocked with 5% nonfat milk at room temperature for 1 h. After rinsing with TBST, membranes were incubated with anti-TGFBR1 (1:1,000; ab31013, Abcam), EGFR (1:1,000, ab36039, Abcam), TCF (1:1,000, ab134275, Abcam), CDH1 (1:50, GTX125890), and actin antibodies at 4°C overnight. Membranes were washed with TBST three times and incubated with the secondary antibodies at room temperature for 1 h. Following by washing three times, protein bands on membranes were visualized by the electrochemiluminescence method (Pierce).
Statistical Analyses
In this study, data analysis was performed using the GraphPad Prism 6.0. All experiments were conducted at least in triplicate. Data were presented with mean ± standard deviation. Comparisons were estimated using student’s t-test (two groups) or analysis of variance (more than two groups). P-value <0.05 was considered statistically significant. All groups complete posttest measures in order to allow for better control of pretesting effects.
Results
Silencing EGFR Significantly Inhibited the Proliferation of Lung Cancer Cells
First, EGFR was knocked down in lung cancer cell lines H2170 and H1299 and their proliferation was detected by colony-forming assay. The result showed that si-EGFR could obviously decrease the proliferation ability of H2170 compared with the NC group (all P-values <0.05, Fig. 1A). In addition, compared with the NC group, si-EGFR could also markedly reduce the proliferation of H1299 (all P-values <0.05, Fig. 1B).

Silencing EGFR inhibits the proliferation of H2170 (A) and H1299 (B) determined using colony-forming assay. *P < 0.05.
Silencing EGFR Suppressed the Invasion Ability of H2170 and H1299 Cells
For further exploration, the effect of EGFR on the invasion of H2170 and H1299 cells was also determined. The results showed that inhibiting the expression of EGFR could markedly reduce the invasion ability of H2170 and H1299 cells compared with the model and NC group (all P-values <0.05, Fig. 2A, B).

Silencing EGFR inhibits the invasion of H2170 (A) and H1299 (B) determined by Matrigel transwell assay. *P < 0.05.
Knocking Down EGFR Inhibited the Migration of H2170 and H1299 Cells
Moreover, the effect of EGFR on the migration of H2170 and H1299 was also detected in this study. The result presented that silencing EGFR could significantly decrease the migration ability of H2170 (all P-values <0.05, Fig. 3A) and H1299 (all P-values <0.05, Fig. 3B). These findings suggested that EGFR could serve as a tumor suppressor gene in the development of lung cancer.

Silencing EGFR inhibits the migration of H2170 (A) and H1299 (B) determined by transwell assay. *P < 0.05.
Silencing EGFR Reduced the RNA Expression of smad2, smad3, CTNNB1, and CDH1
Bioinformatic method had revealed that EGFR could interact with CTNNB1 and CDH1. Therefore, the relative expression of smad2, smad3, CTNNB1, and CDH1 was also determined in EGFR-silenced cells. The results showed that silencing EGFR could significantly decrease the relative mRNA expression of smad2, smad3, CTNNB1, and CDH1 in both H2170 (all P-values <0.05, Fig. 4A) and H1299 (all P-values <0.05, Fig. 4B).
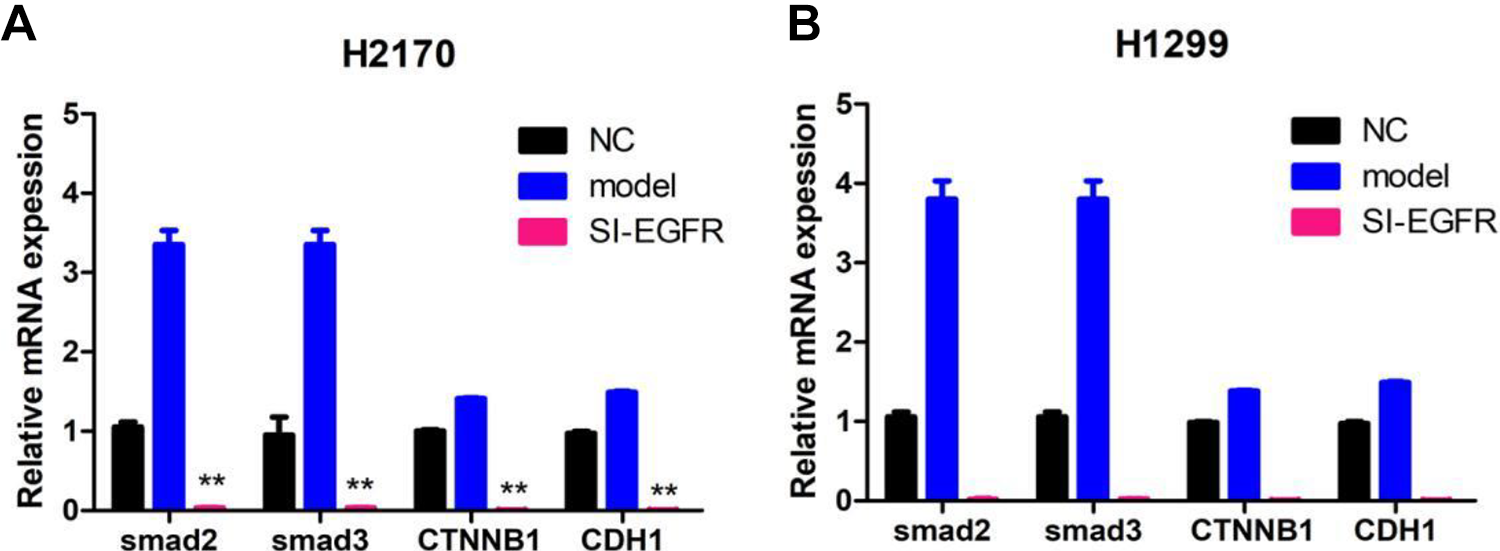
Silencing EGFR inhibits the expression of CTNNB1, Smad2, Smad3, and CDH1 in H2170 (A) and H1299 (B) determined by real-time polymerase chain reaction. **P < 0.01.
Silencing EGFR Reduced the Protein Expression of TGFBR1, TCF, and CDH1
Previous study reported that TGFBR1 could significantly regulate the transcription of CTNNB1 and TCF via smad2/smad3-dependent pathway. Therefore, the protein expression of TGFBR1, TCF, and CDH1 was detected in EGFR-silenced cell lines. The results showed that si-EGFR could markedly decrease the expression of TGFBR1 and CDH1 in H2170 and H1299 cells (all P-values <0.05, Fig. 5A, B). However, knocking down the expression of EGFR had no significant effect on the expression of TCF in both H2170 and H1299 cells (all P-values <0.05, Fig. 5A, B).
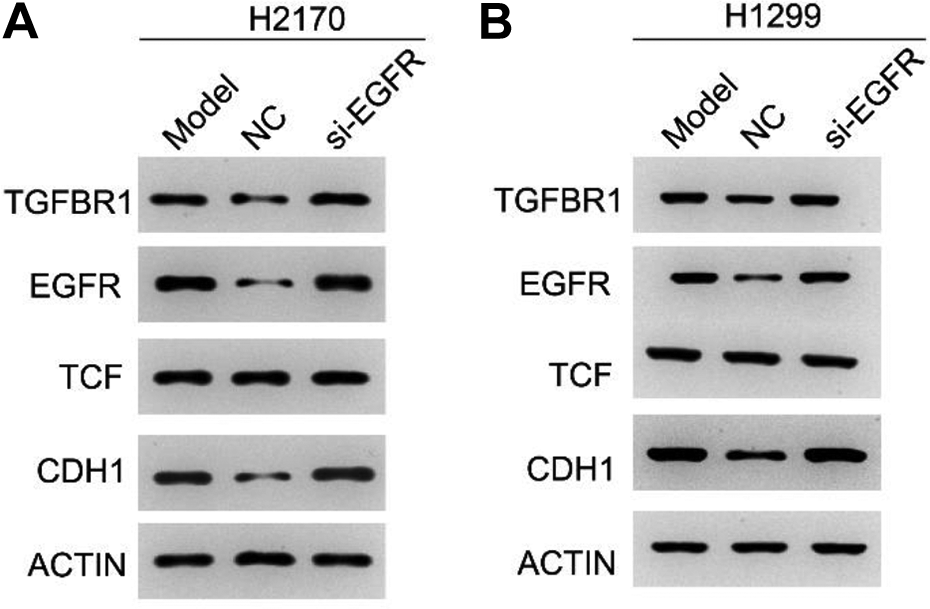
Expression of TGFBR1, TCF, and CDH1 in EGFR-silenced H2170 (A) and H1299 (B) determined by real-time polymerase chain reaction. *P < 0.05.
Discussion
It is notable that there are some crucial pathways controlling the growth, death, and differentiation of cells during the growth and development of individuals, such as Wnt, Notch, Hedgehog, and TGF-β signaling pathways 11 . However, dysfunctions of these pathways also serve a critical role in multiple diseases and cancers 12 . As a tyrosine kinase receptor, EGFR is one of the hallmarks of NSCLC. Multiple clinical studies have suggested EGFR tyrosine kinase inhibitor as the standard first-line therapy in treating advanced NSCLC 13,14 . In this study, silencing EGFR could significantly decrease the proliferation and metastasis of H2170 and H1299 cells. Hence, it is significant to explore EGFR-associated molecules and pathways in regulating NSCLC, improving the understanding and treatment of NSCLC.
In this study, bioinformatics method found that EGFR could interact with CDH1 and CTNNB1, which are involved in the Wnt signaling pathway 15,16 . CHD1 (E-cadherin) is an important tumor metastasis suppressor in multiple cancers, and it is also a crucial epithelial–mesenchymal transition (EMT) marker, as well as the migration and invasion of several cancers 17 . Deficiency of CDH1 improves the expression of EGFR in NSCLC cells via phosphorylation of YBX1 18 . Ma et al. have revealed that miR-23a could target CDH1 in activating Wnt/CTNNB1 signaling pathway to promote TGF-β1-induced EMT and metastasis of breast cancer cells 19 . miR-708-5p targeting DNMT3A inhibits the stemness of lung cancer cells via suppressing Wnt/β-catenin signaling pathway 20 . In this study, silencing EGFR could significantly decrease the expression of CDH1 and CTNNB1 in both H2170 and H1299 cells. These findings suggested that there are some underlying regulatory mechanisms among EGFR, CDH1, and CTNNB1, and EGFR–CDH1–CTNNB1 axis in Wnt signaling pathway plays a crucial role in the migration and invasion of NSCLC.
In 2015, Funa et al. have revealed that β-catenin (CTNNB1) collaborative interacted with Smad2/Smad3 to modulate promotive streak induction 21 . In the same year, Antrodin C could suppress the Smad2/Smad3 and β-catenin signaling pathway to inhibit the EMT transition and metastasis of breast cancer cells 22 . TGFBR1 is an important component of TGF-β/smad signaling pathway, which plays a critical role in the proliferation and metastasis of several cancers 23 . Yang et al. have reported that miR-769-5p could suppress the proliferation, migration, and invasion of NSCLC cells via inhibiting the expression of TGFBR1 24 . Therefore, the expressions of TGFBR1, Smad2, and Smad3 were determined in si-EGFR H1299 and H2170 cells. The results showed that knocking down EGFR could significantly decrease the expressions of TGFBR1, Smad2, and Smad3. These findings suggested that EGFR–TGFBR1–Smad2/Smad3–CTNNB1 axis might play a crucial role in the migration and invasion of NSCLC. EGFR mutations arise in many cases of NSCLC, which in turn comprise ∼85% of all lung cancer cases, highlighting the potential of EGFR mutations as biomarkers. The activation of canonical Wnt signaling is associated with a higher rate of cancer recurrence in patients with Stage I NSCLC, wherein activation of canonical Wnt signaling reflects a more aggressive tumor phenotype suggesting the need for a more aggressive therapy. NSCLCs with KRAS mutations are resistant to EGFR-ligand TKIs: oncogenic activation of KRAS obviates the need for signaling via ligand-dependent activation and phosphorylation of EGFR. Exogenous expression of mutant KRAS in human bronchial epithelial cells reduced the phosphotyrosine immunoprecipitates of EGFR in the phosphoproteomic studies. For KRAS and NRAS mutations, we will further study cell lines to explore the mechanism.
Conclusion
There is some underlying crosstalk between the TGF-β/smad signaling pathway and Wnt/β-catenin signaling pathway in regulating the migration and invasion of NSCLC. Specifically, TGFBR1–EGFR–CTNNB1–CDH1 axis in Wnt signaling plays crucial roles in this crosstalk. However, the exact mechanism of this crosstalk in regulating the progression of NSCLC is still unclarified and needs to be further explored in the future.
Footnotes
Authors Note
Rong Liu and Yusui Zhang are joint first authors.
Authors Contribution
Long Pan and Shuai Zhang wrote the manuscript; Rong Liu, Yusui Zhang, and Yuan Ding conducted most of the experiments; Yusui Zhang and Yuan Ding conducted the experiments and collected the data; Yusui Zhang and Yuan Ding collected and analyzed the data; and Long Pan and Shuai Zhang designed the study and all authors approved the submission.
Availability of Data and Material
Data and material are available on request.
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
