Abstract
Secreted frizzled-related proteins (SFRPs) are a group of five secreted glycoproteins—SFRP1, SFRP2, SFRP3 (frizzled related protein, FRZB), SFRP4, and SFRP5—which contain a frizzled-related cysteine-rich domain and a netrin module. We analyzed SFRPs’ expression levels, mutations, regulation, functional networks, and correlation with immune infiltration in breast cancer (BC) patients using data from multiple open databases. SFRP1/3/4/5 were downregulated and SFRP2 was upregulated in BC patients compared to healthy controls. Furthermore, higher levels of SFRP1/3/4 were significantly associated with favorable prognosis. In addition, the prognostic significance of the infiltrating B cells was correlated to the SFRPs. Based on these findings, we hypothesize that SFRPs play a synergistic role in BC progression, and are, therefore, promising prognostic biomarkers as well as therapeutic targets.
Introduction
Breast cancer (BC) is the most prevalent cancer and the second leading cause of cancer-related death in women 1 . It is the result of a complex interplay of genetic and environmental factors 2 , and frequently involves lymph node and distant metastasis 3 . Therefore, despite the encouraging outcomes of locoregional, endocrine, chemo and molecular-targeted therapies, the mortality rate of BC is still considerably high 4 . The therapeutic outcomes of BC depend on its molecular subtype—luminal A, luminal B, erb-b2 receptor tyrosine kinase 2 (HER2) enriched, basal-like, claudin-low or normal breast-like—as well as on its histological stratification based on the expression of estrogen receptor (ER), progesterone receptor, and HER2 5 . More recently, genomic and transcriptomic analyses have identified 10 BC subtypes, including 4 ER-negative and 6 triple-negative subtypes 6 .
Secreted frizzled-related proteins (SFRPs) are a family of glycoproteins, including SFRP1, SFRP2, SFRP3 (FRZB), SFRP4, and SFRP5, which are characterized by a frizzled-related cysteine-rich domain (CRD) and a netrin module. They are known repressors of the pro-oncogenic Wnt/β-catenin pathway and inhibit signaling by binding CRDs to Wnts 7,8 . The Wnt/β-catenin pathway is constitutively active in multiple cancers, and closely associated with BC initiation and metastasis 9,10 . Studies show a diverse functional profile of the SFRPs in BC with respect to Wnt signaling. For instance, SFRP1 acts a Wnt antagonist in the breast carcinoma cells, and blocks thrombospondin-induced adhesion and migration, whereas SFRP2 has a pro-tumorigenic role 11 . SFRP4 is also a Wnt antagonist that mediates apoptosis in breast cancer stem cells, and inhibits proliferation, metastasis, and epithelial mesenchymal transition 12 . We explored the mutation signatures and expression levels of various SFRPs in BC datasets through integrated bioinformatics analysis in order to provide new insights into their potential functions, molecular mechanisms, and prognostic significance.
Materials and Methods
ONCOMINE Database
The expression data of SFRP mRNAs in BC and normal samples were downloaded from the Oncomine 4.5 (www.oncomine.org) 13 database that currently includes 715 gene expression data sets and 86,733 samples. P < 0.05 was used as the cutoff criterion.
GEPIA Dataset
GEPIA (Gene Expression Profiling Interactive Analysis) 14 is a newly created interactive online web server tool that provides with key interactive and customizable functions including differential expression analysis, profiling plotting, correlation analysis, patient survival analysis, similar gene detection, and dimensionality reduction analysis. It is used to analyze RNA sequencing data of BC/normal tissues from The Cancer Genome Atlas (TCGA).
UALCAN Analysis
UALCAN (http://ualcan.path.uab.edu) 15 , a portal for the analysis, integration, and discovery of cancer transcriptomic data, was used to compare the expression levels of SFRPs across different subgroups using TCGA data.
Kaplan–Meier Plotter
Kaplan–Meier plotter (http://kmplot.com/analysis/, K-M) was used to analyze the prognostic relevance of the SFRPs 16 . The patients were divided into respective low and high expression groups according to the median values of each SFRP, and their survival was assessed using a K-M survival plot.
cBioPortal
The cBioPortal portal (http://cbioportal.org) 17 encompassing data sets of 245 cancer studies was used to analyze SFRP alterations in TCGA BC samples and construct an interaction network of the SFRPs and their coexpressed genes, including neighboring genes. The functions of SFRP mutations and neighboring genes that were significantly related to SFRP mutations were performed using Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) with the Database for Annotation, Visualization, and Integrated Discovery (Enrichr) online tool. P < 0.05 was considered statistically significant.
TIMER Analysis
Tumor IMmune Estimation Resource (TIMER) (https://cistrome.shinyapps.io/timer/) 18 was used to evaluate tumor infiltration of CD8+ T cells, B cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells, and their correlation with SFRPs expression in BC. The survival module was used to determine the prognostic relevance. P < 0.05 was considered statistically significant.
Enrichr
GO and KEGG pathway enrichment analyses were performed on the most frequently modified neighbor genes using Enrichr (http://amp.pharm.mssm.edu/Enrichr.) 19 , which currently contains 180,184 annotated gene sets from 102 gene set libraries. P < 0.05 was considered statistically significant.
Results
SFRPs Are Aberrantly Expressed in BC
The transcriptional levels of the SFRPs were analyzed in BC and normal breast tissues using the ONCOMINE databases. According to the results of Ma et al 20 , SFRP2 expression was 4.89-fold higher in the ductal breast carcinoma tissues compared to normal tissues (P = 4.81E-7), and 4.202-fold higher in the invasive ductal breast carcinoma versus normal breast tissues (P = 7.66E-5). In addition, the expression levels of SFRP4 mRNA increased 11.053-fold in ductal BC (P = 7.98E-9) and 19.752-fold in the invasive ductal BC tissues (P = 7.82E-5) compared to the normal samples (Fig. 1). TCGA data also indicated a 2.747-fold increase in SFRP2 mRNA expression in male BC tissues (P = 2.17E-14, Table 1). Furthermore, Finak et al 21 showed a 19.435-fold, 12.763-fold, and 79.315-fold increase in SFRP2 (P = 1.82E-23), SFRP3 (P = 4.45E-16), and SFRP4 (P = 9.43E-28) levels, respectively, in the invasive BC tissues compared to normal tissues.

The transcriptional levels of SFRPs in breast cancer and normal samples from multiple ONCOMINE datasets, redder means higher expression and bluer means lower expression, the analysis that can give us an understanding of the expression of SFRPs family in different cancer types.
The Significant Changes of SFRP Expression in Transcription Level Between Different Types of BC and Normal Tissues (Oncomine Database).

BC: breast cancer; NA: not available; TCGA: The Cancer Genome Atlas.
SFRPs Are Independent of the Clinicopathological Features of BC
According to the GEPIA dataset, the expressions of SFRP1/3/4/5 were significantly lower in BC tissues and that of SFRP2/4/5 differed across the tumor stages (Figs. 2 –4). Subgroup analysis on the basis of clinical stage (I to X) further showed that SFRP2 was upregulated in BC patients of all stages compared to healthy individuals. In contrast, other SFRPs were downregulated in the tumor versus normal tissues regardless of clinical stage. Subgroup analysis by histological type (luminal, HER2 positive and triple negative), gender, and age (21 to 40, 41 to 60, 61 to 80, and 81 to 100 years) also showed that the expression of SFRP2 was significantly higher and that of SFRP1/3/4/5 significantly lower in BC patients across all subgroups (Fig. 5).

Box plot showing expression of SFRPs in breast cancer. The box color of red indicates tumor and blue indicates normal. We used log2(TPM + 1) for y-axis log-scale, the method for differential analysis is one-way analysis of variance, using disease state (tumor or normal) as variable for calculating differential expression and asterisk means statistically significant, with each dot representing a distinct tumor or normal sample (GEPIA Database, TPM: transcripts per million).
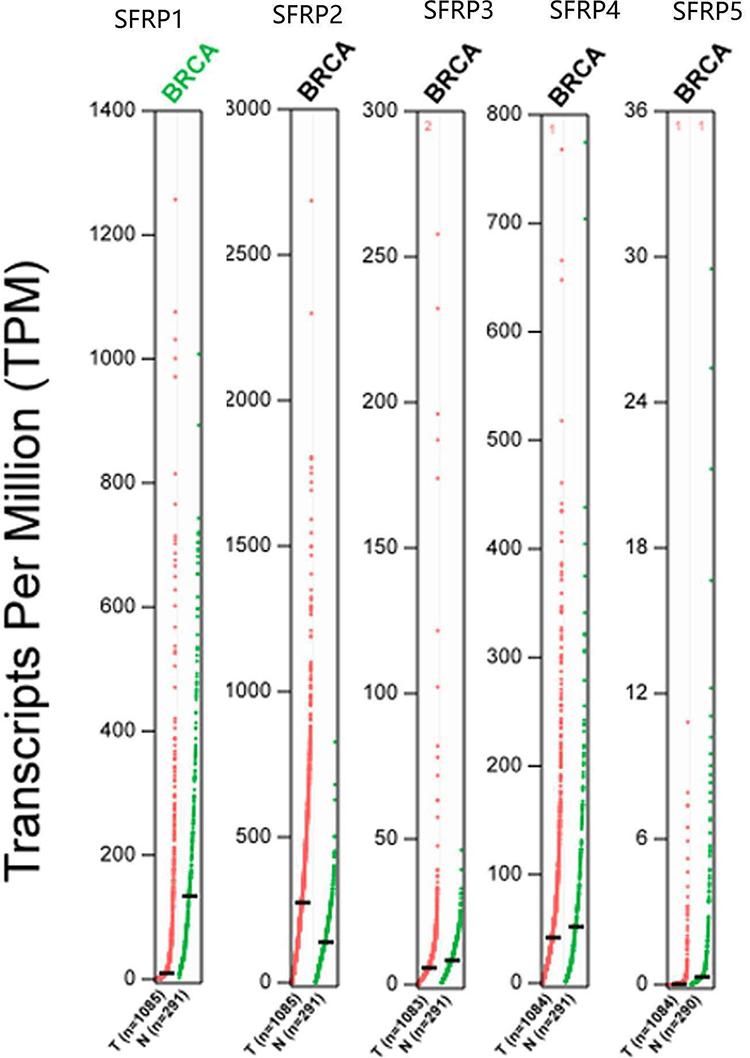
Expression profile of SFRPs in breast cancer, from the results we can see that SFRP1 is highly expressed in normal samples and SFRP2/3/4/5 highly expressed in tumor samples (GEPIA Database).

The expression of SFRPs across different cancer stages. We used log2(TPM + 1) for y-axis log-scale, the method for differential gene expression analysis is one-way analysis of variance, using pathological stage as variable for calculating differential expression. The larger F, the better the study fit, and P-value <0.05 as statistically significant (GEPIA Database).
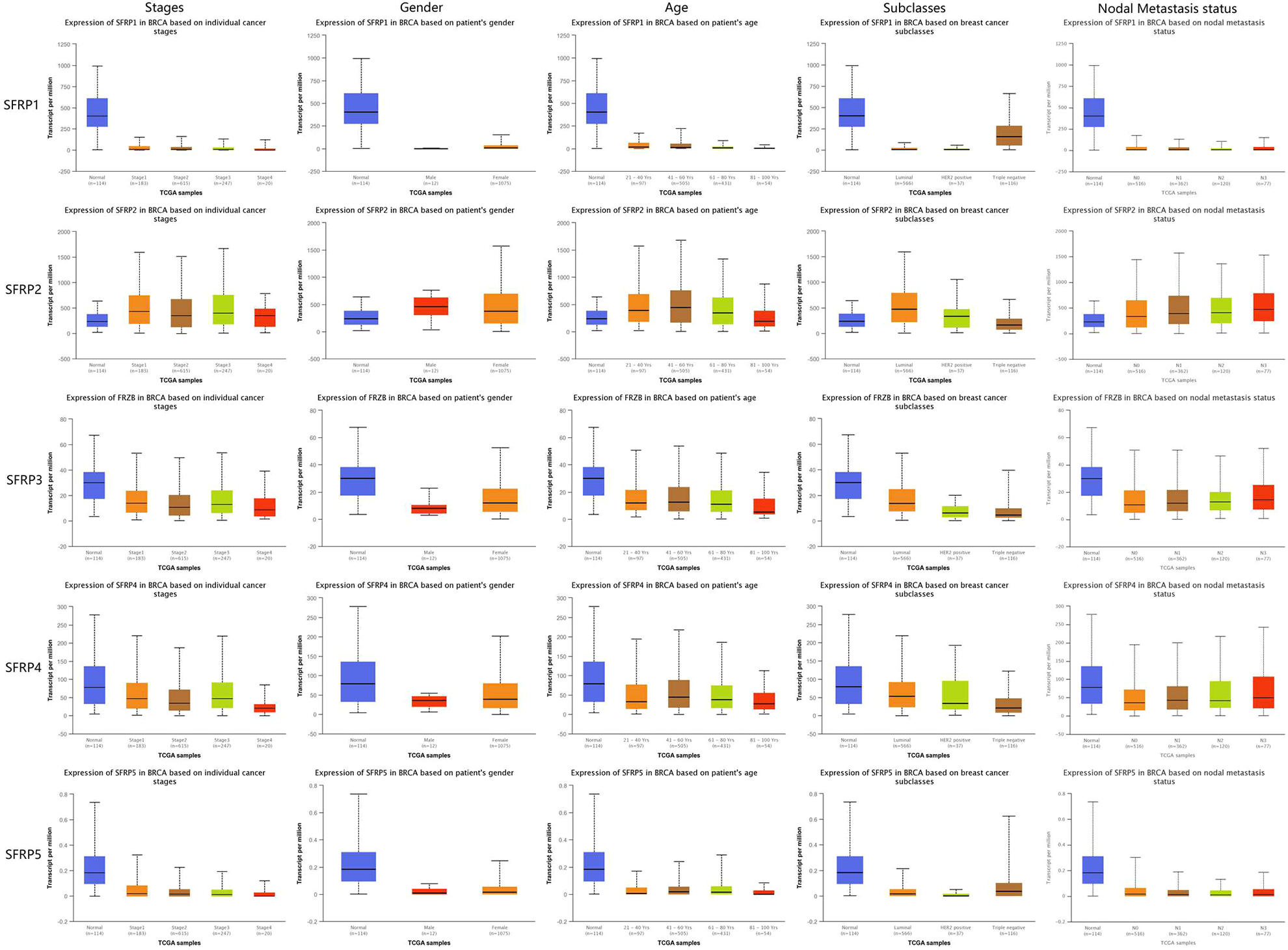
Boxplot showing relative expression of SFRPs in breast cancer patients stratified on the basis of gender, age, stages, subclasses, and nodal metastasis (UALCAN). P < 0.05 was statistically significant.
SFRPs Have Prognostic Significance in BC
As shown in Fig. 6, SFRP1 (hazard ratio [HR] = 0.72, 95% confidence interval [CI]: 0.58 to 0.9, P = 0.0031), SFRP3 (HR = 0.63, 95% CI: 0.51 to 0.78, P = 2.2e-5), and SFRP4 (HR = 0.69, 95% CI: 0.56 to 0.86, P = 0.00075) overexpression were significantly associated with longer overall survival of BC patients, while lower levels of all SFRPs correlated with shorter relapse-free survival. Interestingly, lower expression of SFRP3 (HR = 0.78, 95% CI: 0.61 to 0.99, P = 0.042) and SFRP4 (HR = 0.78, 95% CI: 0.61 to 1, P = 0.046) were significantly associated with worse post-progression survival. Taken together, SFRP1/3/4 levels are prognostically relevant in BC, and are promising biomarkers for predicting survival.

The prognostic value of SFRPs in breast cancer. The patient samples are split into two groups according to various quantile expressions of the proposed biomarker and gene expression data (Kaplan–Meier Plotter Database).
Predicted Functions and Pathways of SFRPs in BC
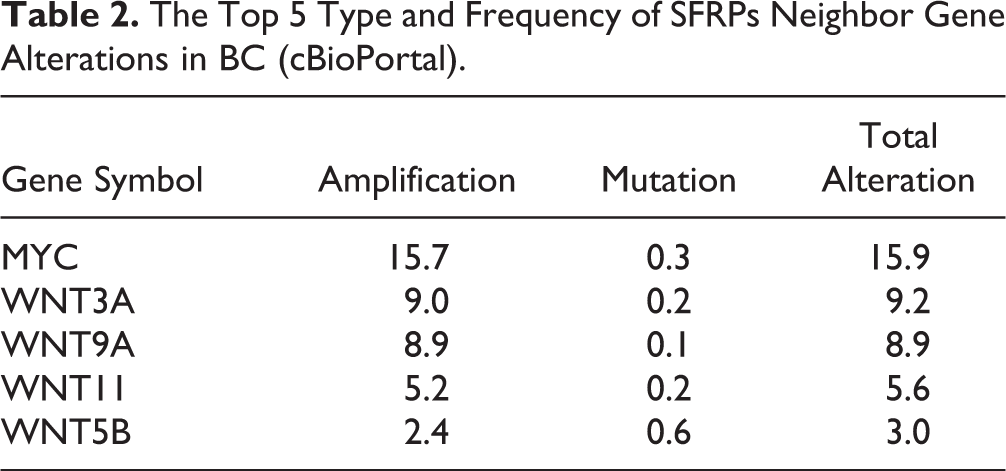
To further explore the biological role of SFRPs in BC, we analyzed their mutation patterns and functional networks using the cBioPortal online tool. Of the 996 BC tumor samples that were sequenced, genetic alterations were detected in 125 samples with a mutation rate of 13%. SFRP1 showed the highest mutation rate of 10%. The network of SFRPs and the most frequently altered neighboring genes was also constructed (Fig. 7), of which myelocytomatosis oncogene (MYC), wingless-type integration site family (Wnt), member 3A (WNT3A), Wnt family member 9A (WNT9A), Wnt family member 11 (WNT11), and Wnt family member 5B (WNT5B) ranked among the top five mutated genes (Table 2). GO analysis of the SFRPs and their neighboring genes further indicated a significant enrichment in biological processes including Wnt signaling pathway, canonical Wnt signaling pathway, generation of neurons, neuron differentiation, response to retinoic acid, cellular response to retinoic acid, cellular response to acid chemical, cellular response to lipid, cellular response to oxygen-containing compound, and regulation of canonical Wnt signaling pathway. In addition, the major molecular function terms were frizzled binding, G-protein-coupled receptor binding, Wnt-activated receptor activity, G-protein-coupled receptor activity, transcription regulatory region DNA binding, E-box binding, cytokine activity, regulatory region DNA binding, and kinase activator activity. Finally, the significantly enriched cell component terms included Golgi lumen, endoplasmic reticulum lumen, clathrin-coated endocytic vesicle membrane, and nuclear chromatin. KEGG pathway enrichment showed that these genes were mainly enriched in Wnt signaling pathway, basal cell carcinoma, melanogenesis, signaling pathways relating pluripotency of stem cells, BC, gastric cancer, hippo signaling pathway, hepatocellular carcinoma, proteoglycans in cancer, and mechanistic target of rapamycin signaling pathway (Fig. 8). Taken together, the SFRPs are associated with multiple cancer-related functions and pathways.

SFRPs gene expression and mutation analysis in breast cancer (cBioPortal Database). A. SFRPs gene expression and mutation analysis; B. Network of SFRPs and the 50 most frequently altered neighbor genes.
The Top 5 Type and Frequency of SFRPs Neighbor Gene Alterations in BC (cBioPortal).

Predicted functions and pathways of SFRPs and their frequently altered neighbor genes in breast cancer patients (cBioPortal and Enrichr). GO enrichment analysis predicted the functional roles of target host genes based on BP, CC, and MF. KEGG pathway analysis identified the enriched pathways. The lighter the red, the more enriched it is.
Immune Infiltrates in Correlation With SFRPs in BC
We detected a significant correlation between SFRPs expression in BC tissues and immune infiltration (P < 0.05, Fig. 9). Compared to the basal and luminal tumors, the HER2-enriched and normal tumors showed significantly higher infiltration of B cells (P < 0.05), which was positively correlated to SFRPs expression. Therefore, we hypothesize that infiltration of immune cells in BC tumors is associated with the expression of SFRPs and affects prognosis.

Correlation between SFRPs expression and infiltration of B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells in BRCA, BRCA-Basal, BRCA-Luminal, and BRCA-Her2 (TIMER).
Discussion
Previous studies have established key roles of SFRPs in cancer, although their exact function in BC remains to be elucidated. We explored the expression levels and mutational landscape of various SFRPs in multiple BC datasets, and found that the expression levels of specific SFRPs correlated with prognosis and immune infiltration.
SFRPs are aberrantly expressed in tumors, and regulate tumor cell cycle, proliferation, and apoptosis by targeting the WNT signaling pathway. In addition, the expression levels of SFRPs have been associated with the prognosis of various cancers. SFRP1 is a Wnt antagonist that blocks BC cell adhesion and migration 11 . Ling-Yu Kong et al 22 showed that SFRP1 knockdown activated the Wnt/β-catenin signaling pathway, which in turn promoted the proliferation, migration, and invasion of BC cells. Ba et al 23 further showed that miR-27a increased the proliferation and invasiveness of colon cancer cells by targeting SFRP1 through the Wnt/β-catenin pathway. Likewise, Li et al 24 found that miR-1254 promoted lung cancer cell proliferation by targeting SFRP1. SFRP3 contains a CRD that regulates Wnt signaling. Bernascone et al 25 found that loss of SFRP3 predisposed mice to invasive BC. Schlensog et al 26 also reported a tumor-suppressive role of SFRP3 in lung adenocarcinoma likely via the regulation of canonical Wnt signaling. However, another study established an oncogenic function of SFRP3 in paired box 3-forkhead box O1 (PAX3-FOXO1)-positive alveolar rhabdomyosarcoma 27 . SFRP4 is another Wnt antagonist that inhibits the pathway by binding to Wnt ligands and frizzled receptors. Abhijeet et al 28 found that SFRP4 regulated the basal glucose uptake in the cancer stem cells derived from breast and prostate tumor cell lines, indicating a crucial role in tumor cell proliferation and survival. Consistent with this, KEGG analysis of the genes significantly correlated to SFRP mutations in our study was enriched in pathways regulating stem cells’ pluripotency. Sandsmark et al 29 found that high SFRP4 expression is associated with postsurgery recurrence of prostate cancer. Chen et al 30 found that SFRP5 was markedly downregulated in melanoma tissues and cell lines, which is in line with the enrichment of melanogenic genes observed in this study. Veeck et al 31 showed that SFRP5 was epigenetically inactivated in human BC, indicating its potential as a DNA-based biomarker.
We detected low levels of SFRP1/3/4/5 in BC patients that correlated with poor prognosis. Contradictory to previous studies, we found that SFRP2 is highly expressed in BC tissues, thereby warranting further exploration of its functions. GO and KEGG analyses showed that the SFRPs and their frequently altered neighboring genes were significantly enriched in Wnt signaling pathway, frizzled binding, G-protein-coupled receptor binding, Golgi lumen, endoplasmic reticulum lumen, basal cell carcinoma, and melanogenesis. The Wnt signaling pathway is a key player in the initiation and progression of many cancers, as well as normal tissue development and stem cell function 32 . There is evidence indicating a role of SFRPs and tumor-infiltrating immune cells. We found a significantly higher infiltration of B cells in some BC subtypes, which correlated positively to SFRPs expression. This indicates that immune cells may have a significant effect on BC prognosis, and the potential role of SFRPs ought to be investigated further.
There were several limitations in our study that should be addressed. Since we only analyzed data retrieved from online databases, our findings will have to be validated on larger cohorts. Furthermore, we did not evaluate the diagnostic and therapeutic potential of SFRPs due to the considerable heterogeneity in histological types and anatomical sites. Finally, the specific functions and mechanisms of action of the SFRPs, especially SFRP2, need further investigation.
Footnotes
Author Contributions
WZH and ZT designed and analyzed the research study; WZH and ZYJ wrote and revised the manuscript, ZYJ, YJX, and WZH collected the data and all authors contributed to and approved the final version of manuscript.
Availability of Data and Materials
Ethical Approval
As the work is a bioinformatics analysis article, ethical approval was not necessary and all the data were retrieved from the free online databases.
Statement of Human and Animals Rights
This article does not contain studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Nature Science Foundation of China (No: 81771005, 81400462) and Hubei provincial Natural Foundation of China (2017CFB623).
