Abstract
Accumulating evidence suggests that long noncoding RNA (lncRNA) functions as a critical regulator in cancer biology. Here, we characterized the role of lncRNA PCED1B antisense RNA 1 (PCED1B-AS1) in glioblastoma (GBM). PCED1B-AS1 was notably upregulated in GBM tissues and cell lines and closely associated with larger tumor size and higher grade. Patients with high PCED1B-AS1 had shorter survival time than those with low PCED1B-AS1. Functional experiments showed that depletion of PCED1B-AS1 significantly inhibited, while overexpression of PCED1B-AS1 promoted cell proliferation, glucose uptake, and lactate release. Mechanistically, PCED1B-AS1 was able to directly bind to the 5′-UTR of HIF-1α mRNA and potentiate HIF-1α translation, leading to increased HIF-1α protein level, thereby promoting the Warburg effect and tumorigenesis. Importantly, PCED1B-AS1 lost the carcinogenic properties in the absence of HIF-1α. In addition, we also confirmed the existence of the PCED1B-AS1/HIF-1α regulatory axis in vivo. Taken together, our findings demonstrate that PCED1B-AS1 is a novel oncogenic lncRNA in GBM and functions in a HIF-1α-dependent manner, which provides a promising prognostic biomarker and druggable target for GBM.
Introduction
Glioblastoma (GBM) is the most common and fatal malignant primary brain tumor in adults, accounting for 12% to 15% of all intracranial tumors and 50% to 60% of astrocytic tumors 1 . It is able to rapidly invade and infiltrate normal peripheral brain tissue, making complete resection impossible 2 . Therefore, the standard treatment at present is surgery combined with radiotherapy and chemotherapy 3 . The 5-year survival rate of patients with GBM is very unfavorable, less than 5%, and this indicator drops to 2% among patients aged 65 years or older 4 . A better understanding of the pathogenesis of GBM will provide new approaches for treatment and improve the outcomes of patients.
The occurrence and progression of cancer is an extremely complex process, accompanied by a variety of pathophysiological changes. Emerging evidence suggests that abnormal glucose metabolism is a hallmark of cancer 5 . In normal cells, the final product of glucose can be lactic acid, or it can be completely oxidized by respiration in the mitochondria to produce CO2, depending on the oxygen content 6 . Cancer cells, however, prefer glycolysis, even under aerobic conditions, leading to dramatically increasing glucose uptake and lactate production, a phenomenon known as aerobic glycolysis, also called the Warburg effect 7 . Enhanced Warburg effect helps cancer cells to minimize oxidative stress, thereby promoting tumorigenesis and aggressive progression 8 .
Long noncoding RNA (lncRNA) is a class of endogenous noncoding RNA longer than 200 nt in length. Multiple lines of evidence show that lncRNA functions as a pivotal regulator in the Warburg effect. Fox example, lincRNA-p21 9 , IDH1-AS1 10 , and MALAT1 11 were proposed to govern the Warburg effect through regulating some key oncogenes, such as c-Myc and HIF-1α. In this study, we identified a glycolysis-related lncRNA, PCED1B antisense RNA 1 (PCED1B-AS1), which was remarkably elevated in GBM tissues and cell lines. Furthermore, the underlying mechanism by which it facilitated the Warburg effect and tumorigenesis was also deciphered.
Materials and Methods
GBM Tissues and Cell Lines
We collected 76 pairs of primary GBM and adjacent normal tissues from The First Affiliated Hospital of Zhengzhou University. All specimens were histologically confirmed by two independent experienced pathologists. The detailed clinical information of patients is summarized in Table 1. Patients were followed up with routinely after surgery, and each patient provided informed consent. This study was approved by the Ethics Committee of The First Affiliated Hospital of Zhengzhou University. A total of six GBM cell lines (U87MG, U251MG, SHG44, Ln229, T98, and A172) and one normal human astrocyte cell line NHA were obtained from Chinese Academy of Sciences (Shanghai, China) and cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) (Gibco, Invitrogen, Carlsbad, CA, USA) complete medium containing 10% fetal bovine serum (FBS) (Gibco, Invitrogen, Carlsbad, CA, USA), penicillin, and streptomycin.
Correlation Between PCED1B-AS1 Expression and Clinicopathological Features in GBM Patients (

GBM: glioblastoma; PCED I B-AS1: PCED1B antisense RNA 1.
The Primer Sequences Used in This Study.

Quantitative Reverse Transcription PCR Analysis
GBM cells and tissues were treated with Trizol reagent (Invitrogen, Carlsbad, CA, USA), followed by extraction by chloroform, precipitation by isopropanol, and dissolution by diethyl pyrocarbonate water. Then, 1 μg RNA was reverse transcribed into cDNA using Molony Murine Leukemia Virus (MMLV) transcriptase (Promega, Madison, WI, USA). Lastly, qPCR was performed using SYBR Green SuperMix (Roche, Basel, Switzerland) on a CFX96 real-time PCR detection system (Bio-Rad, Richmond, CA, USA). The 2−ΔΔCt method was employed to calculate RNA expression and the results were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Table 2).
Cell Transfection and Generation of Stably Engineered Cell Lines
Cell transfection was performed using Lipofectamine 3000 reagent (Invitrogen) according to the manufacturer’s instructions. To generate stable PCED1B-AS1-depleted (sh-RNA#1: 5′-GGCCUGAAAUACAUUUAAA-3′; sh-RNA#1: 5′-GGUGACUCAUGCCUAUAAU-3′) or overexpressed GBM cells, the pCDH-CMV-MCS-EF1-copRFP lentiviral vector was used and infected into cells in the presence of 8 μg/ml polybrene. Then, the stable cell lines were selected using puromycin for 2 wk.
Cell Counting Kit-8 (CCK-8) and EdU ( 5-ethynyl-2’-deoxyuridine) Assays
For CCK-8 assay, GBM cells were plated on 96-well plates and grown for 24, 48, and 72 h, followed by treatment with 10 μl CCK-8 solution (Dojindo, Kumamoto, Japan) for 3.5 h at 37°C. Then, the absorbance value at 450 nm in each well was recorded with a microplate reader. For EdU assay, the Cell-Light EdU Apollo488
Glucose Consumption and Lactate Production
The analysis of glucose consumption and lactate production was carried out using Glucose Assay Kit (ab65333, Abcam, Cambridge, UK) and Lactate Assay Kit (ab65330, Abcam) based on the manufacturer’s instructions. The absorbance at 570 nm was detected using a microplate reader.
Western Blot and Immunohistochemistry (IHC)
Total protein was extracted using lysis buffer supplemented with protease and phosphatase inhibitors. Then, the protein was loaded in SDS–PAGE gel, transferred onto polyvinylidene fluoride (PVDF) member, and blocked by 5% skimmed milk. Finally, the member was incubated with corresponding primary and secondary antibodies and exposed using immobilon western chemiluminescent HRP substrate (Millipore, Schwalbach, Germany). The primary antibodies used in this study are as follows: anti-HIF-1α (#ab16066, Abcam), anti-c-Myc (#ab32072, Abcam), and anti-β-actin (#ab8226, Abcam). For IHC staining, GBM tissues were fixed and embedded into tissue microarray in the Department of Pathology. Then, the paraffin-embedded sections were dewaxed and rehydrated, followed by antigen retrieval, blockade, and incubation with anti-HIF-1α (#ab16066, Abcam) antibody at 4°C overnight. After incubation with ready-to-use DAKO secondary antibody for 1 h at room temperature, the sections were visualized using diaminobenzidine reagent. The results were analyzed with an optical microscope in 10 random fields by using H-score method 12 .
Assessment of Translation Level
For polysome profile analysis, the lysates of U87MG and U251MG cells were collected with polysome lysis buffer and loaded onto different sucrose density gradients. After high-speed centrifugation, the RNA in polysome fraction was isolated using Trizol reagent and the expression of HIF-1α mRNA was detected by quantitative reverse transcription PCR (qRT-PCR). For the luciferase reporter assay, the HIF-1α 5′-UTR containing PCED1B-AS1 binding site was embedded into bicistronic reporter vector; then, the reporter was transfected into stable PCED1B-AS1-depleted U87MG and U251MG cells for 48 h. Finally, the luciferase activity was recorded using dual luciferase reporter system (Promega). For detecting newly synthesized HIF-1α protein, the lysates of U87MG and U251MG cells were collected after incubation with
RNA Pull-Down Assay
The biotinylated probes were
In Vivo Xenograft Assay
This animal experiment was carried out with the approval of the Animal Welfare Committee of The First Affiliated Hospital of Zhengzhou University. A total of 15 BALB/c nude mice were obtained from Shanghai Experimental Animal Center, Chinese Academy of Sciences. They were randomly divided into three groups and subcutaneously injected with 1 × 107 control or PCED1B-AS1-depleted U87MG cells, followed by growth under specific-pathogen-free condition for 5 weeks. At the end of the observation, all mice were euthanized by cervical dislocation and the tumors were weighed and collected for western blot and qRT-PCR analysis.
Statistical Analysis
The difference between the two groups was statistically analyzed by Student’s
Results
Upregulation of PCED1B-AS1 Predicts Poor Prognosis for Patients with GBM
By analyzing the Gene Expression Profiling Interactive Analysis (GEPIA) database, we found that PCED1B-AS1 was the most upregulated lncRNA in GBM as compared with normal tissues (Fig. 1A). Then, we collected 76 paired GBM and paracancerous normal tissues to test the above result. As shown in Fig. 1B, PCED1B-AS1 was lowly expressed only in 7 (9%) cases. Also, PCED1B-AS1 was notably elevated in six GBM cell lines in comparison to normal NHA astrocyte cells (Fig. 1C). Next, we evaluated the relationship between PCED1B-AS1 expression and clinicopathological features of GBM patients. The results showed that high PCED1B-AS1 was positively correlated with larger tumor size and higher glioma grade (Table 1). More importantly, patients with high PCED1B-AS1 displayed shorter survival time than those with low PCED1B-AS1 (Fig. 1D). These data suggest that PCED1B-AS1 is upregulated in GBM and may be used as a potential prognostic indicator.

PCED1B-AS1 is frequently overexpressed in GBM. (A) The relative expression of PCED1B-AS1 in GEPIA database. (B) qRT-PCR analysis of PCED1B-AS1 expression in 76 GBM and adjacent normal tissues. (C) qRT-PCR analysis of PCED1B-AS1 expression in GBM cell lines. (D) The survival curve of GBM patients with low or high PCED1B-AS1 based on median PCED1B-AS1 value in 76 GBM tissues. *
Depletion of PCED1B-AS1 Represses the Warburg Effect and Proliferation of GBM Cells
To explore the biological function of PCED1B-AS1 in GBM, we established stable PCED1B-AS1 knockdown U87MG and U251MG cell lines by using the pCDH-CMV-MCS-EF1-copRFP lentiviral vector (Fig. 2A). As shown in Fig. 2B, the viability of U87MG and U251MG cells was drastically reduced after depletion of PCED1B-AS1. Similarly, less DNA synthesis was observed in PCED1B-AS1-silenced cells compared to control cells (Fig. 2C, D). Given that abnormal glucose metabolism is responsible for uncontrolled proliferation of cancer cells, we then tested the levels of glucose uptake and lactate production. The results showed that knockdown of PCED1B-AS1 significantly decreased the levels of glucose uptake (Fig. 2E) and lactate production (Fig. 2F) in both U87MG and U251MG cells. These functional assays indicate that PCED1B-AS1 knockdown inhibits GBM cell growth via blocking the Warburg effect.
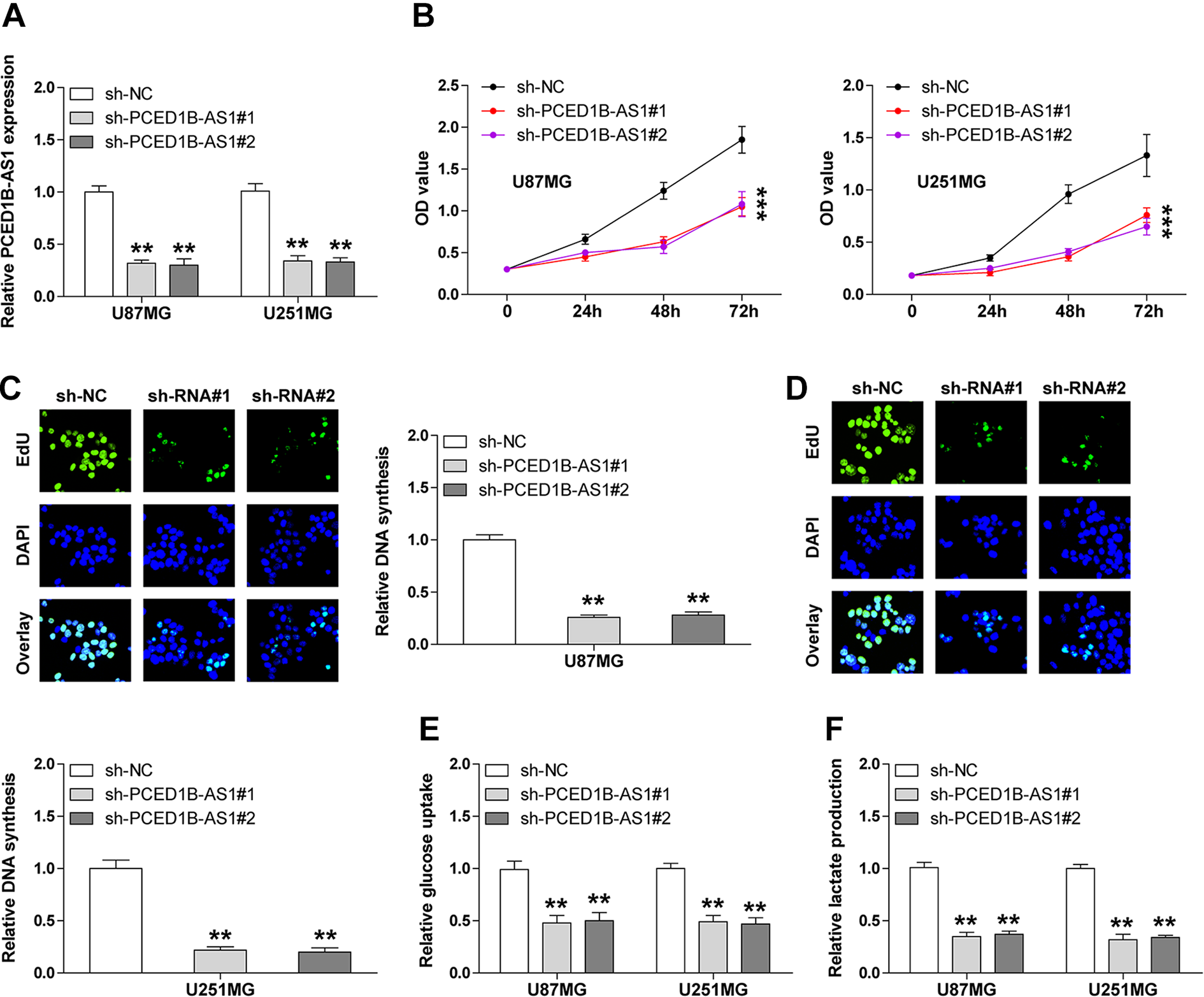
PCED1B-AS1 knockdown inhibits GBM cell proliferation, glucose uptake, and lactate release. (A) qRT-PCR analysis for verification of PCED1B-AS1 knockdown in U87MG and U251MG cell lines. (B) Cell viability in PCED1B-AS1-depleted U87MG and U251MG cell lines tested by CCK-8 assay. (C and D) DNA synthesis rate in PCED1B-AS1-depleted U87MG and U251MG cell lines detected by EdU assay. (E and F) The levels of glucose uptake and lactate release in U87MG and U251MG cell lines with or without stable PCED1B-AS1 knockdown. **
Overexpression of PCED1B-AS1 Potentiates the Warburg Effect and Proliferation of GBM Cells
We also established the PCED1B-AS1-overexpressing A172 and Ln229 cell lines. The overexpression efficiency was verified by qRT-PCR (Fig. 3A). In contrast, exogenous PCED1B-AS1 expression remarkably increased cell viability and DNA synthesis rate (Fig. 3B, C), as illustrated by CCK-8 and EdU assays, respectively. Likewise, more glucose uptake and lactate production were observed after enforced expression of PCED1B-AS1 in both A172 and Ln229 cells (Fig. 3D, E). These data further confirm the oncogenic role of PCED1B-AS1 in GBM.

PCED1B-AS1 overexpression promotes GBM cell proliferation, glucose uptake, and lactate release. (A) qRT-PCR analysis for verification of PCED1B-AS1 overexpression in A172 and Ln229 cell lines. (B) Cell viability in PCED1B-AS1-overexpressing A172 and Ln229 cell lines tested by CCK-8 assay. (C) DNA synthesis rate in PCED1B-AS1-overexpressing A172 and Ln229 cell lines detected by EdU assay. (D and E) The levels of glucose uptake and lactate release in A172 and Ln229 cell lines with or without PCED1B-AS1 overexpression. **
HIF-1α Is a Downstream Target of PCED1B-AS1
In light of the pivotal regulatory role of HIF-1α and c-Myc in the Warburg effect, we then tested whether PCED1B-AS1 functioned via them. As shown in Fig. 4A, knockdown of PCED1B-AS1 significantly reduced the protein expression levels of HIF-1α, but not c-Myc. And HIF-1α protein was uniformly overexpressed in GBM cells in comparison with normal NHA cells (supplemental Figure S1). However, PCED1B-AS1 did not affect HIF-1α mRNA level (Fig. 4B), implying that PCED1B-AS1 controlled HIF-1α expression at the posttranscriptional level. Subsequently, less HIF-1α mRNA was observed in the heavy polysome fraction after PCED1B-AS1 knockdown (Fig. 4C), suggesting that PCED1B-AS1 promoted HIF-1α mRNA translation. Consistently, depletion of PCED1B-AS1 resulted in a dramatic reduction in newly synthesized HIF-1α protein in both U87MG and U251MG cells (Fig. 4D). In addition, we tested the expression of HIF-1α protein in GBM tissues by IHC staining. The results showed that HIF-1α protein expression was notably upregulated in GBM as compared with adjacent normal tissues (Fig. 4E), and its expression was strongly positively correlated with PCED1B-AS1 expression (

PCED1B-AS1 upregulates HIF-1α expression via promoting its translation process. (A) The protein expression of HIF-1α and c-Myc in PCED1B-AS1-depleted U87MG and U251MG cell lines detected by western blot. (B) qRT-PCR analysis of HIF-1α mRNA expression in PCED1B-AS1-depleted U87MG and U251MG cell lines. (C) qRT-PCR analysis of HIF-1α mRNA level in the indicated polysome fractions of PCED1B-AS1-depleted U87MG and U251MG cell lines. (D) Western blot analysis of the newly synthesized HIF-1α protein in PCED1B-AS1-depleted U87MG and U251MG cell lines. (E) IHC staining of HIF-1α protein in GBM and normal tissues. (F) The correlation between PCED1B-AS1 and HIF-1α protein in GBM tissues. *
PCED1B-AS1 Directly Binds to the 5′-UTR of HIF-1α mRNA
To determine how PCED1B-AS1 promotes HIF-1α mRNA translation, we first tested the subcellular localization of PCED1B-AS1. As shown in Fig. 5A, PCED1B-AS1 was predominantly located in the cytoplasm, which was consistent with its role in posttranscriptional regulation of HIF-1α. It has been well documented that lncRNA is able to govern gene expression by directly binding to DNA, RNA, or protein. Through sequence alignment, we found that PCED1B-AS1 might interact with the 5′-UTR of HIF-1α mRNA (energy = −15.25480 kcal/mol; Fig. 5B). To verify this possibility, we performed RNA pull-down assay coupled with qRT-PCR analysis. The results showed that more HIF-1α 5′-UTR/PCED1B-AS1 was enriched by PCED1B-AS1/HIF-1α 5′-UTR probes in comparison to control probes, respectively (Fig. 5C, D). In addition, the luciferase activity of bicistronic reporter containing HIF-1α 5′-UTR was dramatically decreased in PCED1B-AS1-depleted U87MG and U251MG cells compared with control cells (Fig. 5E). These data indicate that PCED1B-AS1 accelerates HIF-1α translation by physically interacting with the 5′-UTR of HIF-1α mRNA.

PCED1B-AS1 directly binds to the 5′-UTR of HIF-1α mRNA. (A) qRT-PCR analysis of the location of PCED1B-AS1 in U87MG and U251MG cell lines. GAPDH and U1 were used as cytoplasmic and nuclear control references, respectively. (B) The interaction sequence of PCED1B-AS1 and 5′-UTR of HIF-1α mRNA. (C and D) RNA pull-down coupled with qRT-PCR analysis in U87MG and U251MG cell lines using the indicated biotinylated probes. (E) The translation activity of 5′-UTR of HIF-1α mRNA in U87MG and U251MG cell lines detected by luciferase reporter assay. **
PCED1B-AS1 Functions in a HIF-1α-Dependent Manner
We then tested the expression levels of HIF-1α downstream targets after manipulating PCED1B-AS1 expression. The qRT-PCR results showed that ectopic expression of PCED1B-AS1 significantly elevated the expression of EGR1, ANGPTL4, IGFBP3, PKM2, and LDHA in both A172 and Ln229 cells, whereas these effects were evidently blocked by HIF-1α silencing (Fig. 6A, supplemental Figure S2). Besides, exogenous PCED1B-AS1 expression enhanced cell viability, DNA synthesis, glucose uptake, and lactate production only in control cells, but not in HIF-1α-depleted cells (Fig. 6B–E), implying that HIF-1α is indispensable for the oncogenic role of PCED1B-AS1 (Fig. 6F).

HIF-1α is required for the pro-oncogenic role of PCED1B-AS1. (A) qRT-PCR analysis of the indicated gene expression in PCED1B-AS1-overexpressing A172 and Ln229 cell lines transfected with HIF-1α siRNA. (B) Cell viability in PCED1B-AS1-overexpressing A172 and Ln229 cell lines transfected with HIF-1α siRNA detected by CCK-8 assay. (C) DNA synthesis rate in PCED1B-AS1-overexpressing A172 and Ln229 cell lines transfected with HIF-1α siRNA detected by EdU assay. (D and E) The levels of glucose uptake and lactate release in PCED1B-AS1-overexpressing A172 and Ln229 cell lines transfected with HIF-1α siRNA. (F) The proposed model of the pro-oncogenic role of PCED1B-AS1 via regulation of HIF-1α translation in GBM. **
Knockdown of PCED1B-AS1 Inhibits Tumor Growth
To test whether PCED1B-AS1 also functioned in vivo, we established the xenograft tumor model by subcutaneous injection of control or PCED1B-AS1-depleted cells into nude mice (

Depletion of PCED1B-AS1 retards tumor growth. (A) The tumor images of nude mice in the indicated three groups (
Discussion
In the present study, we described a novel GBM-related lncRNA, PCED1B-AS1, which was markedly overexpressed in GBM and closely correlated with aggressive clinical features and adverse prognosis. Loss- and gain-of-function assays indicated that knockdown of PCED1B-AS1 inhibited the Warburg effect and cell proliferation, while overexpression of PCED1B-AS1 resulted in opposite effects. Further mechanistic study revealed that HIF-1α was a direct downstream target of PCED1B-AS1, in which PCED1B-AS1 was able to directly bind to the 5′-UTR of HIF-1α mRNA, resulting in accelerated HIF-1α translation, thereby elevating HIF-1α protein expression and promoting the Warburg effect and tumorigenesis of GBM. In addition, we also observed a strong correlation between PCED1B-AS1 and HIF-1α protein expression in GBM tissues and in the xenograft tumor model. Therefore, our data highlight the importance of PCED1B-AS1 in the progression of GBM and also shed new light on the regulation of lncRNA on protein translation.
lncRNA is emerging as a crucial player in the occurrence, development, and progression of cancer 13 . It can act as a tumor suppressor or an oncogene depending on the different contexts 14 . The mechanism by which lncRNA works is extremely complex, but accumulating evidence suggests that this is closely related to its subcellular localization. Cytoplasmic lncRNA mainly functions via sponging miRNAs, binding to proteins and even translating peptides 15 . On the other hand, nuclear localized lncRNA is able to directly interact with epigenetic modification complex, influence chromatin architecture, and control gene expression at the transcriptional level 16 . Herein, we found that PCED1B-AS1 was mainly located in the cytoplasm, and it could directly bind to the 5′-UTR of HIF-1α mRNA to potentiate HIF-1α translation. To the best of our knowledge, only a few studies have focused on the direct binding of lncRNA to mRNA to affect its translation. For instance, lncRNA PXN-AS1 was reported to physically interact with the coding sequences of PXN mRNA, leading to dissociating the translation elongation factors from PXN mRNA and inhibiting its translation 17 . lncRNA NPCCAT1 was proposed as a translation promoter of YY1 by directly binding to the 5′-UTR of YY1 mRNA 18 . In this study, we hypothesize that PCED1B-AS1 may recruit some translation triggers (such as YB1 19 ) by binding to HIF-1α 5′-UTR, thus promoting HIF-1α translation, which requires further study.
HIF-1α has been widely recognized as a key trigger of the Warburg effect that increases the expression of glycolysis genes by binding to hypoxia-responsive elements (HRE) on their promoters 20 . Dysfunctions of the regulatory system for HIF-1α activity are critical for tumorigenesis 21 . A growing body of evidence has shown that HIF-1α is significantly upregulated in various human cancers and predicted poor prognosis 22 . Up to now, several lncRNAs have been reported to be responsible for the dysregulation of HIF-1α, such as TUG1 23 , MEG3 24 , PVT1 25 , and MTA2TR 26 . They controlled HIF-1α expression by altering HIF-1α mRNA levels, modulating the HIF-1α translation process, or affecting HIF-1α protein stability. Here, we confirmed that HIF-1α protein was elevated in GBM tissues and found that PCED1B-AS1 promoted HIF-1α protein expression, but not mRNA expression, by directly binding to the 5′-UTR of HIF-1α mRNA and accelerating its translation process. Importantly, PCED1B-AS1 had no tumor-promoting effect in the absence of HIF-1α, suggesting that PCED1B-AS1 functions in a HIF-1α-dependent manner in GBM. Therefore, our study advances the understanding of the regulation of HIF-1α, and meanwhile underlines the essential relevance of lncRNA in gene regulation. Intriguingly, we also found some HRE motifs on the promoter PCED1B-AS1. Whether a regulatory feed-forward loop was formed between PCED1B-AS1 and HIF-1α to amplify the tumor-promoting effect of PCED1B-AS1 is worthy of in-depth investigation.
Of note, by analyzing the GEPIA database, we found that PCED1B-AS1 was also upregulated in several human cancers, such as diffuse large B cell lymphoma and kidney renal clear cell carcinoma, as shown in supplemental Figure S3. Further study is needed to confirm this observation to expand the clinical implication of PCED1B-AS1.
Collectively, our findings for the first time demonstrate that PCED1B-AS1 is a pro-oncogenic lncRNA in GBM through regulation of HIF-1α mRNA translation. Targeting this regulatory axis of PCED1B-AS1/HIF-1α may be a promising approach for the treatment of GBM patients.
Supplemental Material
Supplementary_Figure_1 - Long Noncoding RNA PCED1B-AS1 Promotes the Warburg Effect and Tumorigenesis by Upregulating HIF-1α in Glioblastoma
Supplementary_Figure_1 for Long Noncoding RNA PCED1B-AS1 Promotes the Warburg Effect and Tumorigenesis by Upregulating HIF-1α in Glioblastoma by Zhiqiang Yao, Qiansheng Zhang, Fuyou Guo, Shewei Guo, Bo Yang, Binghui Liu, Panxing Li, Jinyi Li, Sheng Guan and Xianzhi Liu in Cell Transplantation
Footnotes
Ethical Approval
Ethical approval to report this case series was obtained from the Ethics Committee of The First Affiliated Hospital of Zhengzhou University.
Statement of Human and Animal Rights
All procedures involving the care and use of laboratory animals were approved by the Animal Policy and Welfare Committee of The First Affiliated Hospital of Zhengzhou University, and all efforts were made to minimize use of animals as well as to minimize their pain or discomfort during the course of the study.
Statement of Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grant from Post-doctoral Science Foundation Project (No. 61).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
