Abstract
Long noncoding RNA (lncRNA) is emerging as an essential regulator in the development and progression of cancer, including cervical cancer (CC). In this study, we found a CC-related lncRNA, KCNMB2-AS1, which was significantly overexpressed in CC and linked to poor outcomes. Depletion of KCNMB2-AS1 remarkably inhibited CC cell proliferation and induced apoptosis.
Introduction
Cervical cancer (CC) is the fourth most frequently diagnosed cancer and the fourth leading cause of cancer-associated death among women worldwide, which has become a huge economic burden, especially in many underdeveloped countries 1 . Over the last few decades, despite many drugs or vaccines have been developed to antagonize CC, such as human papillomavirus (HPV) vaccination programs 2,3 , the 5-year survival rate is still not optimistic, especially for high-risk cases 4 . Therefore, continued dissection into mechanisms governing CC progression may yield promising therapeutic insights.
It is estimated that more than 90% of mammalian genomes are transcribed into noncoding RNA. Long noncoding RNA (lncRNA), as a special kind of noncoding RNA, has a wide range of biological functions with a length of over 200 nucleotides
5
. Emerging evidence shows that lncRNA plays important role in the initiation, development, and progression of various human diseases, especially in cancer
6
. The lncRNA has been proposed to harbor diverse functions, including the organization of nuclear architecture, transcriptional regulation in
Up to now, extensive studies have shown that lncRNA is frequently deregulated in cancer and functions as a tumor suppressor or oncogene through sponging miRNAs. For instance, linc00173 was recently proposed as a promoter of small cell lung cancer, and it could abundantly absorb miR-218 and elevate GSKIP and NDRG1, leading to the nuclear translocation of β-catenin and subsequent transcriptional upregulation of oncogenic Etk 11 . HLA-F-AS1 was shown to be significantly overexpressed in colorectal cancer and promoted colorectal cancer cell proliferation, migration, and invasion by increasing PFN1 expression via sponging and repressing miR-330-3p 12 .
In the present study, we characterized a previously unconcerned lncRNA, KCNMB2-AS1. We found that KCNMB2-AS1 was dysregulated in CC tissues and played a pivotal role in CC tumorigenesis. Moreover, we also deciphered the underlying mechanism of KCNMB2-AS1 in CC.
Materials and Methods
CC Tissues and Cell Lines
The current study was conducted on 82 pairs of CC and adjacent normal specimens, which was histopathologically and clinically diagnosed at Shengjing Hospital of China Medical University. We collected the clinicopathological data of these patients and analyzed the correlations between them and KCNMB2-AS1 expression (Table 1). Patients who received antitumor therapy before the operation were excluded. All patients provided informed consent, and this study was approved by the ethics committee of Shengjing Hospital of China Medical University (No. SHK168509). Two CC cell lines, including SiHa and HeLa, were purchased from ATCC (Manassas, VA, USA) and cultured in high-glucose Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum and 100 U/ml penicillin, 100 g/ml streptomycin at 37 °C in humidified air containing 5% carbon dioxide. Mycoplasma contamination was tested by the EZ-PCR Mycoplasma Test Kit (#20-700-10, BIOIND, Guangzhou, China) before the cells were used.
The Correlations Between KCNMB2-AS1 Expression and Clinicopathological Characteristics of Patients with Cervical Cancer.
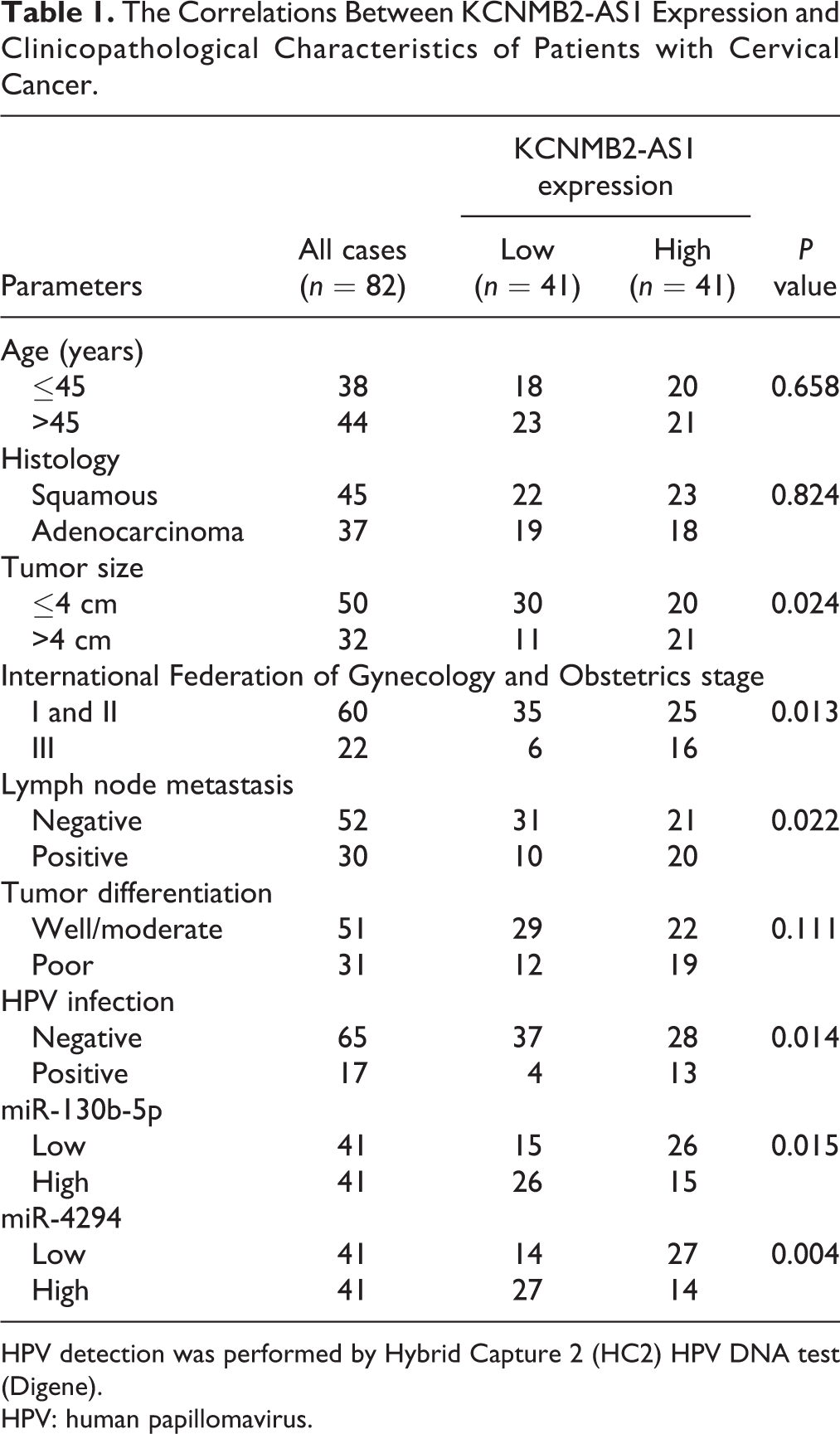
HPV detection was performed by Hybrid Capture 2 (HC2) HPV DNA test (Digene).
HPV: human papillomavirus.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated by using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) as per the standard protocols, followed by reverse transcription into cDNA by using GoScript Reverse Transcription System with AMV Reverse Transcriptase (Promega, Madison, WI, USA). Then, cDNA was amplified and quantified by using GoTaq qPCR Master Mix (Promega) with 7500 FAST Real-Time PCR Detection System (Applied Biosystems, Cambridge, UK). 2−△△CT formula was applied to calculate relative gene expression.
Subcellular Fractionation and Fluorescence In Situ Hybridization (FISH)
Nuclear and cytoplasmic fragments were extracted by the nuclear/cytoplasmic fractionation PARIS Kit (Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instruction. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 were used as endogenous references for cytoplasm and nucleus, respectively. FISH assay was carried out by using the lncRNA FISH detection kit provided by RiboBio (Guangzhou, China) according to the manufacturer’s instruction.
Generation of Stably Engineered Cell Lines
Two KCNMB2-AS1 shRNA plasmids were transfected into the PT67, an NIH3T3-derived packaging cell line (Clontech, San Francisco, CA, USA) by using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instruction. Subsequently, the supernatants were collected and infected into SiHa and HeLa cells for 24 h in the presence of 8 μg/ml polybrene. After infection, stably transduced cells were obtained by adding 1.5 μg/ml puromycin into the DMEM.
Functional Assays
For cell counting kit-8 (CCK-8) assay, SiHa and HeLa cells were plated onto 96-well plates and cultured for 1 to 3 days. Then, each well was added with CCK-8 solution (Dojindo, Kumamoto, Japan) and incubated for 2 h at 37 °C. The absorbance at 450 nm was detected by using an automatic microplate spectrophotometer (SpectraMax, San Francisco, CA, USA). For colony formation assay, cells were plated onto six-well plates and cultured for 14 days with the medium changing every 3 days, followed by staining with crystal violet. For measuring the DNA synthesis rate, the Cell-Light EdU Detection Kit (RiboBio, Guangzhou, China) was used, and the EdU-positive cells were counted and photographed in eight random fields. Besides, the apoptotic cells were detected by the Annexin V-PE/7-AAD Apoptosis Detection Kit (BD Biosciences, San Jose, CA, USA) following the manufacturer’s manual. The results were analyzed by a flow cytometer (FACSCanto II, BD Biosciences) with 20,000 acquired events. Cells negative for Annexin V-PE and negative for 7-AAD are living cells; cells positive for Annexin V-PE and negative for 7-AAD are cells in early apoptosis; cells positive for Annexin V-PE and positive for 7-AAD are cells in late apoptosis, and cells negative for Annexin V-PE and positive for 7-AAD are cells in necrosis. For the representation of apoptosis, the sum of early and late apoptosis was performed.
Tumor Xenografts In Vivo and Immunohistochemistry (IHC)
A total of 1 × 107 control or KCNMB2-AS1-depleted SiHa cells were resuspended in 0.1 ml phosphate-buffered saline and inoculated into the armpit of 5-week-old male BALB/c nude mice. Tumor volume was measured once a week by using a Vernier caliper. In the fourth week, all mice were sacrificed, and the tumor tissues were carefully isolated. Then, all tumor tissues were paraffin embedded and used for IHC staining with anti-Ki-67 (#ab15580, Abcam, Cambridge, UK) and anti-IGF2BP3 antibodies (#ab177942, Abcam). The positive cells were counted and photographed in eight random fields with an inverted optical microscope (CX22, Olympus, Tokyo, Japan). This study was approved by the Institutional Ethical Review Board and Animal Welfare and Research Ethics Committee of Shengjing Hospital of China Medical University (No. SHK168509).
RNA Pull-Down Assay
The biotin-labeled control and KCNMB2-AS1 probes were
Luciferase Reporter Assay
The full-length wild-type or mutant KCNMB2-AS1 and IGF2BP3 3′-untranslated region (UTR) sequences were synthesized and cloned into pmiGLO luciferase vector (Promega) at SacI and SalI restriction sites. Then, control or miR-130b-5p/miR-4294 mimics (RiboBio) were co-transfected with above luciferase vectors into SiHa and HeLa cells by using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instruction. After 48 h, the dual-luciferase reporter assay was performed by using a commercial kit (Promega) as per the standard protocols.
Western Blot
Western blot assay was performed according to a previously described standard method 13 . In brief, total protein was collected and boiled in sodium dodecyl sulfate buffer for 5 min at 100 °C, followed by electrophoresis, transfer, and blocking. Then, the blot was incubated with anti-IGF2BP3 (#ab177942, Abcam), anti-GAPDH antibodies (#ab9485, Abcam), and a goat-derived secondary antibody. The protein signal intensity was assessed by using the chemiluminescence detection system (Bio-Rad, Richmond, CA, USA).
RNA Immunoprecipitation (RIP)
We used anti-IGF2BP3 (#ab177942, Abcam) antibody to pull down KCNMB2-AS1. The protein A/G beads (Santa Cruz Biotechnology, Dallas, Texas, USA) were applied to recover the IGF2BP3 antibody, and then, the RNA level of KCNMB2-AS1 in the precipitates was determined by qRT-PCR analysis. For assessing the m6A level of KCNMB2-AS1, the methylated RIP (MeRIP) assay was performed by using Magna MeRIP m6A Kit (Merck Millipore, Schwalbach, Germany) according to the manufacturer’s instruction, followed by RNA extraction and qRT-PCR analysis of KCNMB2-AS1 expression.
Statistical Analysis
Unless otherwise stated, continuous variables are summarized as mean ± SD indicating at least three independent experiments carried out in triplicate. Differences between groups were compared as appropriate using Student’s
Results
KCNMB2-AS1 is Frequently Overexpressed in CC
By analyzing the Gene Expression Profiling Interactive Analysis online database, we found that KCNMB2-AS1 was significantly upregulated in CC tissues (Fig. 1A). Then, we collected 82 pairs of CC and adjacent normal tissues and performed qRT-PCR assay; the results confirmed the upregulation of KCNMB2-AS1 in CC (Fig. 1B). Moreover, high KCNMB2-AS1 expression was positively correlated with tumor size (

The upregulation of KCNMB2-AS1 is identified in CC. (A, B) The expression level of KCNMB2-AS1 in CC and normal tissues from the Gene Expression Profiling Interactive Analysis online tool and our own cohort. (C) The survival curve of CC patients is based on the median KCNMB2-AS1 expression level. (D, E) qRT-PCR analysis of the location of KCNMB2-AS1 in SiHa and HeLa cells (three independent experiments carried out in triplicate). (F) FISH assay showing the cytoplasmic localization of KCNMB2-AS1. The nucleus was stained with 4′,6-diamidino-2-phenylindole. ***
Depletion of KCNMB2-AS1 Represses the Malignant Phenotype of CC Cells In Vitro
To explore the functional roles of KCNMB2-AS1 in CC, we first generated the stable KCNMB2-AS1 knockdown SiHa and HeLa cell lines (Fig. 2A). The CCK-8 results showed that cell viability was dramatically weakened after the KCNMB2-AS1 knockdown (Fig. 2B, C). Likewise, less clone-forming cells were observed in KCNMB2-AS1-silenced CC cells in comparison to control cells (Fig. 2D, E). And depletion of KCNMB2-AS1 slowed down the DNA synthesis rate of CC cells, as shown by the EdU staining assay (Fig. 2F, G). Besides, the number of apoptotic cells was notably increased after the knockdown of KCNMB2-AS1 (Fig. 2H, I). The above functional assays demonstrate that KCNMB2-AS1 is a promoter of CC aggressive phenotype.

Knockdown of KCNMB2-AS1 inhibits CC cell growth and promotes apoptosis. (A) qRT-PCR analysis verifying the stable KCNMB2-AS1 knockdown effect in SiHa and HeLa cells. (B–G) CCK-8, colony formation, and EdU assays detecting the viability, colony formation ability, and DNA synthesis rate of SiHa and HeLa cells with KCNMB2-AS1 knockdown. (H, I) Annexin V-PE and 7-AAD double staining testing the apoptotic number of SiHa and HeLa cells with KCNMB2-AS1 knockdown. **
KCNMB2-AS1 Knockdown Retards Tumor Growth In Vivo
To test whether KCNMB2-AS1 also functions
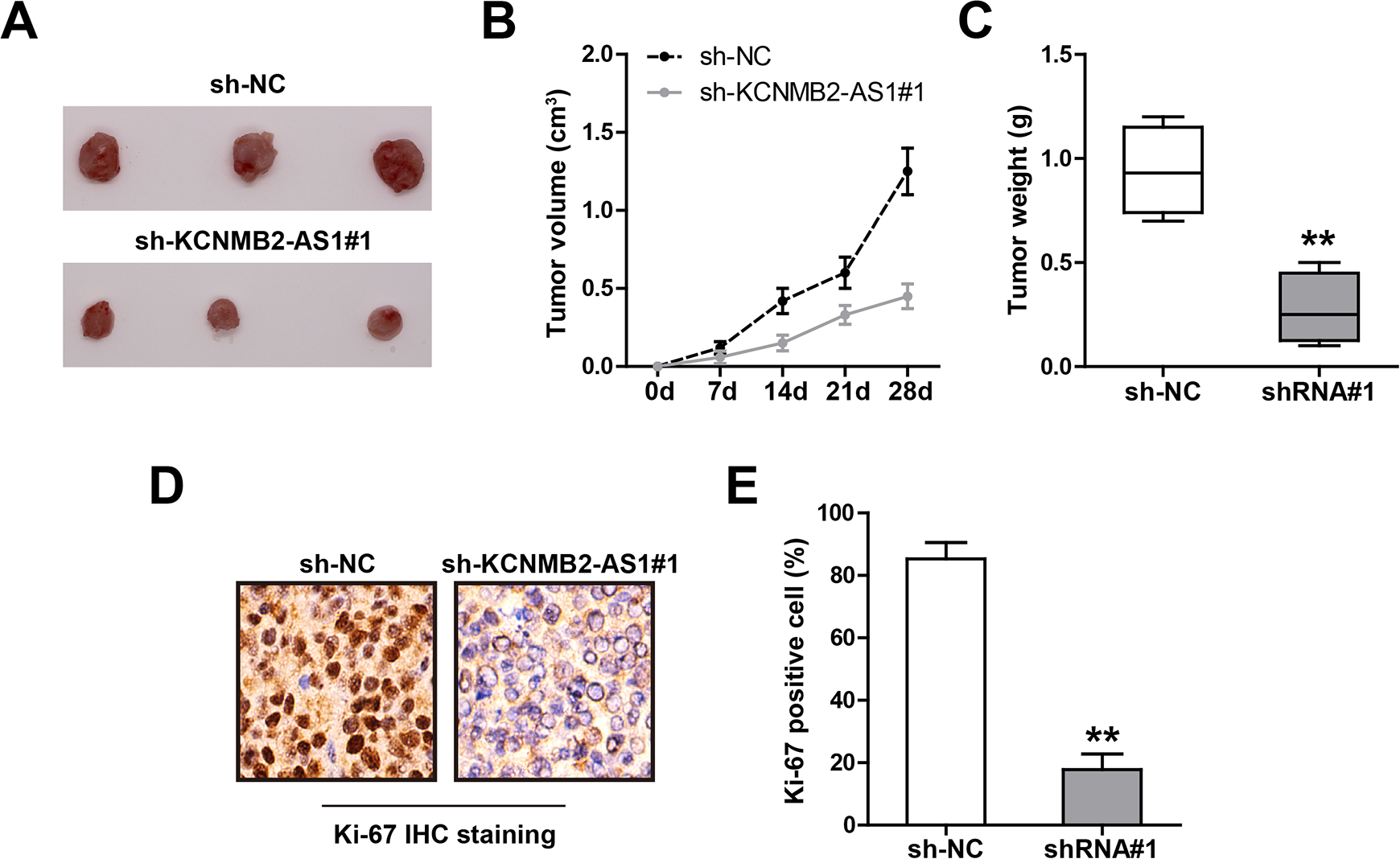
Knockdown of KCNMB2-AS1 inhibits CC
KCNMB2-AS1 Sponges miR-130b-5p and miR-4294 in CC Cells
Given that the cytoplasmic location of KCNMB2-AS1 in CC cells, we inferred that KCNMB2-AS1 might function as a ceRNA to sponge miRNAs. Through using the lncRNASNP2 online tool, a large number of miRNAs were predicted to be complemented with KCNMB2-AS1; we then selected the top five miRNAs to verify according to binding score. The results of RNA pull-down assay showed that miR-130b-5p and miR-4294, not the other three miRNAs, were abundantly pull down by KCNMB2-AS1 in both SiHa and HeLa cells (Fig. 4A, B). The complementary sequences of KCNMB2-AS1 and miR-130b-5p or miR-4294 were shown in Fig. 4C; we mutated them to perform luciferase reporter assay. The results showed that overexpression of miR-130b-5p or miR-4294 significantly reduced the luciferase activity of the wild-type vector, whereas had no effect on of the mutant one (Fig. 4D, E). Moreover, the expression levels of miR-130b-5p and miR-4294 were elevated in KCNMB2-AS1-depleted CC cells (Fig. 4F, G) and xenografts (Fig. 4 H), and KCNMB2-AS1 expression was significantly negatively correlated with miR-130b-5p or miR-4294 in CC tissues (Table 1). Functionally, the diminished cell proliferation caused by KCNMB2-AS1 knockdown was effectively rescued by silencing of miR-130b-5p or miR-4294 (Fig. 4I). Likewise, overexpression of KCNMB2-AS1 significantly enhanced cell proliferation, whereas this effect was effectively abolished after mutation of miR-130b-5p and miR-4294 binding sequences or after overexpression of miR-130b-5p/miR-4294 (Figure S1). These data suggest that miR-130b-5p and miR-4294 are the downstream functional targets of KCNMB2-AS1 in CC cells.

KCNMB2-AS1 functions as a ceRNA. (A, B) RNA pull down in SiHa and HeLa cells using biotin-labeled control or KCNMB2-AS1 probe, followed by qRT-PCR analysis. (C–E) The wild-type or mutant KCNMB2-AS1 luciferase reporter transfected with control or miR-130b-5p/miR-4294 mimics into SiHa and HeLa cells, followed by the analysis of the luciferase activity. (F–H) qRT-PCR analysis of miR-130b-5p/miR-4294 expression in KCNMB2-AS1-depleted CC cells and xenografts. (I) miR-130b-5p/miR-4294 mimics were transfected into KCNMB2-AS1-depleted CC cells, followed by the assessment of cell proliferation. *
KCNMB2-AS1 Regulates the miR-130b-5p/miR-4294/IGF2BP3 Axis in CC Cells
Through analyzing the miRWalk database, we found that IGF2BP3, the well-known oncogene in CC, might be the common downstream target of miR-130b-5p and miR-4294 (Fig. 5A). The results of the luciferase reporter assay showed that ectopic expression of miR-130b-5p or miR-4294 dramatically decreased the luciferase activity of wild-type IGF2BP3 3′-UTR vector, while the above effects were disappeared after mutation of the binding sites (Fig. 5B, C). Importantly, IGF2BP3 mRNA and protein levels were both downregulated in KCNMB2-AS1-depleted SiHa and HeLa cells, whereas silencing of miR-130b-5p or miR-4294 could partially rescue IGF2BP3 expression levels (Fig. 5D–F). Consistently, the results of IHC staining in xenografts showed that KCNMB2-AS knockdown reduced the number of IGF2BP3 positive cells (Fig. 5G, H). Functionally, IGF2BP3 overexpression almost completely blocked the diminished cell proliferation caused by KCNMB2-AS depletion (Fig. 5I). These data indicate that IGF2BP3 is the downstream functional target of the KCNMB2-AS/miR-130b-5p/miR-4294 axis in CC.

KCNMB2-AS1 regulates IGF2BP3 via sponging miR-130b-5p/miR-4294. (A–C) The wild-type or mutant IGF2BP3 3′-UTR luciferase reporter was transfected with control or miR-130b-5p/miR-4294 mimics into SiHa and HeLa cells, followed by the analysis of luciferase activity. (D–F) qRT-PCR and Western blot analysis of IGF2BP3 mRNA and protein levels in KCNMB2-AS1-depleted CC cells transfected with miR-130b-5p/miR-4294 inhibitors. (G, H) The representative image showing the IHC staining of IGF2BP3 in control and KCNMB2-AS1-depleted groups. (I) IGF2BP3-expressing vector was transfected into KCNMB2-AS1-depleted CC cells, followed by the assessment of cell proliferation. *
KCNMB2-AS is Stabilized by IGF2BP3 Via N6-Methyladenosine (m6A) Modification
Recently, many lncRNAs have been found to be modified and regulated by m6A 14 , and we then tested whether KCNMB2-AS also appeared in m6A modification. The MeRIP assay results showed that KCNMB2-AS was significantly enriched by the anti-m6A antibody in comparison to the immunoglobulin G antibody (Fig. 6A). Given that IGF2BP3 is an RNA-binding protein (RBP) that functions as an m6A “reader” and stabilizes RNA through its KH3-4 domain 15 , we then wondered whether KCNMB2-AS was affected by IGF2BP3. The qRT-PCR results showed that KCNMB2-AS1 expression was notably increased by overexpression of wild-type IGF2BP3, but not by overexpression of IGF2BP3 with KH3-4 domain mutation (Fig. 6B), and this upregulation was abrogated after treatment with 3-deazaadenosine (DAA), the global methylation inhibitor (Fig. 6B). Congruously, exogenous expression of IGF2BP3 significantly extended the half-life of KCNMB2-AS1 in both SiHa and HeLa cells (Fig. 6C, D). The RIP results showed that KCNMB2-AS1 was abundantly enriched by the IGF2BP3 antibody, and this effect was completely diminished by DAA treatment (Fig. 6E). Through sequence alignment, we found three IGF2BP3-binding motifs (TGGAC) on KCNMB2-AS1, and then, we replaced the adenosine base in m6A consensus sequence with thymine to abolish m6A modification (Fig. 6F) and constructed the corresponding luciferase reporter vectors (Fig. 6G). As shown in Fig. 6H, I, the luciferase activity of the wild-type vector, not of the mutant one, was substantially increased by overexpression of IGF2BP3. These results clearly suggest that IGF2BP3 prevents the degradation of KCNMB2-AS1 via m6A modification (Fig. 7).

KCNMB2-AS1 is stabilized by IGF2BP3 via m6A modification. (A) MeRIP assay in CC cells using the anti-m6A antibody, followed by the qRT-PCR analysis of KCNMB2-AS1 enrichment. (B) qRT-PCR analysis of KCNMB2-AS1 expression in CC cells transfected with the indicated vectors or treated with DAA. (C, D) qRT-PCR analysis of KCNMB2-AS1 expression in control or IGF2BP3-overexpressing CC cells after treatment with actinomycin D. (E) RIP assay in mock or DAA-treated CC cells using the anti-IGF2BP3 antibody, followed by the qRT-PCR analysis of KCNMB2-AS1 enrichment. (F, G) Three IGF2BP3-binding motifs were found on KCNMB2-AS1, and they were mutated to conduct luciferase reporter assay using the indicated vector. (H, I) The above luciferase reporters were co-transfected with control or IGF2BP3-overexpressing into SiHa and HeLa cells, followed by the analysis of the luciferase activity. **

The proposed model showing m6A-stabilized KCNMB2-AS1 promoting cervical cancer (CC) growth through regulating the miR-130b-5p/miR-4294/IGF2BP3 axis. In normal cells, oncogenic IGF2BP3 is lowly expressed; m6A is not recognized by IGF2BP3, and then KCNMB2-AS1 is destabilized and loses its ability to adsorb miR-130b-5p and miR-4294, thereby IGF2BP3 expression is decreased, forming a regulatory loop. In CC cells, the situation is reversed; IGF2BP3 is increased, ultimately promoting CC tumorigenesis.
Discussion
Recent studies have shown that lncRNA plays a pivotal role in cancer biology. Up to now, although a small portion of lncRNAs has been functionally characterized, many members in the class remain uncharacterized. Herein, we identified a new oncogenic lncRNA in CC (KCNMB2-AS1). KCNMB2-AS1 was remarkably upregulated in CC and modified by m6A. KCNMB2-AS1 promoted CC tumorigenesis both
Therefore, our data highlight the essential role of KCNMB2-AS1 in CC and also provide new evidence for the biological relevance of m6A modification.
The ceRNA network of lncRNA/miRNA/mRNA has been proposed as an important and extensive mode of action in cancer, provided that lncRNA is predominantly localized in the cytoplasm 16,17 . In this study, we used two assays, including qRT-PCR and FISH, and found that KCNMB2-AS1 is a cytoplasmic lncRNA in CC cells; thus, we speculated that KCNMB2-AS1 might function as a ceRNA. A series of subsequent experiments confirmed our hypothesis and showed that KCNMB2-AS1 could directly bind miR-130b-5p and miR-4294, leading to increasing the expression of their common target, IGF2BP3. IGF2BP3 is a well-documented oncogene that is shown to be frequently upregulated in various human cancers, including CC, which is linked to aggressive clinical features and unfavorable prognosis 18,19 . Consistently, silencing of miR-130b-5p/miR-4294 or overexpression of IGF2BP3 could effectively rescue the attenuated malignant phenotype of CC cells induced by KCNMB2-AS1, indicating that the ceRNA network of KCNMB2-AS1/miR-130b-5p/miR-4294/IGF2BP3 does exist and is functional. Whether KCNMB2-AS1 is also an oncogene as well as a ceRNA in other malignant tumors may be worthy of further investigation.
Recently, RNA modifications are gaining great interest of biologists worldwide, especially m6A modification. m6A modification is the most prevalent internal RNA modification that mainly occurs at a consensus sequence of RRACH (R=G or A; H=A, C, or U) 20 . It is involved in RNA splicing, stabilization, translocation, and even translation, and this process includes “writers,” “readers,” and “erasers,” and they are delicately balanced to support normal cellular functions 21 . Specifically, “readers” are some RBPs (YTH domain family proteins and IGF2BPs) that can recognize m6A modification and affect gene expression and cancer biology 22 . IGF2BPs consists of IGF2BP1, IGF2BP2, and IGF2BP3, which have early been proven to preferentially bind to the “UGGAC” consensus sequence containing the “GGAC” m6A core motif and increase the stability and storage of their targets 15 . In agreement with the recent report showing that IGF2BPs acted as a “reader” for m6A modified lncRNA 23 , we found that IGF2BP3 could physically bind to m6A modified KCNMB2-AS1 and decrease its decay, resulting in elevating KCNMB2-AS1 expression. Of note, mutation of the KH3-4 domain, the critical region for IGF2BP3 m6A “reader” function, evidently blocked the upregulation effect of KCNMB2-AS1, suggesting that IGF2BP3 increases KCNMB2-AS1 level in an m6A-dependent manner. Therefore, a regulatory feed-forward loop mediated by m6A modification is formed between KCNMB2-AS1 and IGF2BP3. It will be of great interest to explore whether this phenomenon also exists in other malignancies.
Taken together, our study for the first time demonstrates that m6A modified KCNMB2-AS1 functions as an oncogenic lncRNA in CC by regulating the miR-130b-5p/miR-4294/IGF2BP3 signal axis, which provides a promising prognostic biomarker and druggable target for CC patients.
Supplemental Material
Supplemental Material, Figure_S1 - Long Noncoding RNA KCNMB2-AS1 Stabilized by N6-Methyladenosine Modification Promotes Cervical Cancer Growth Through Acting as a Competing Endogenous RNA
Supplemental Material, Figure_S1 for Long Noncoding RNA KCNMB2-AS1 Stabilized by N6-Methyladenosine Modification Promotes Cervical Cancer Growth Through Acting as a Competing Endogenous RNA by Yao Zhang, Dian Wang, Dan Wu, Donghong Zhang and Ming Sun in Cell Transplantation
Footnotes
Authors’ Contributions
YZ designed the study, conducted the experiments, and drafted and revised the manuscript. DW and DWu collected clinical information. DHZ and MS analyzed the data. All authors read and approved the final manuscript.
Ethical Approval
Ethical approval to report this case series was obtained from the ethics committee of Shengjing Hospital of China Medical University.
Statement of Human and Animal Rights
All procedures involving the care and use of laboratory animals were approved by the Animal Policy and Welfare Committee of Shengjing Hospital of China Medical University, and all efforts were made to minimize the use of animals as well as to minimize their pain or discomfort during the course of the study.
Statement of Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: this work was supported by the grant from Special Research Fund for Doctoral Discipline of Universities (No. 20122104120012).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
