Abstract
Purpose
This study investigated the mechanism by which Rosmarinic acid (RA) may alleviate doxorubicin (DOX)- induced cardiomyocyte apoptosis.
Methods
The target genes of RA, DOX-related differentially expressed genes, and GEO database related genes were retrieved by bioinformatics analyses. The results of these analyses were further intersected to identify candidate genes. The protein-protein interaction network was constructed to develop the pharmacophore model. The molecular docking was simulated to determine the core target B-cell lymphoma 2-like 1 (BCL2L1) for subsequent molecular mechanism investigation in vitro. The effects of DOX and RA on the apoptosis of H9c2 cells were assessed using the CCK8 assay. The present study investigated the effect of RA on DOX-induced oxidative stress in cardiomyocytes. This investigation was conducted using an ELISA test and a DCFH-DA probe. The JC-1 probe was utilized to assess the effect of RA on DOX-induced cardiomyocyte mitochondrial membrane permeability. A Western blot assay was conducted to ascertain the activation of multiple signaling molecules, including those belonging to the BCL-2 and caspase-3 families, within the apoptosis pathway.
Results
A total of 17 differentially expressed genes (DEGs) were screened, and five genes were selected as hub DEGs. A subsequent KEGG enrichment analysis revealed that these DEGs were significantly enriched in various biological processes and pathways, including the MAPK signaling pathway, autophagy, apoptosis, and the TNF signaling pathway. The pharmacophore model and molecular docking of five candidate targets with RA were successfully established. It is noteworthy that DOX treatment led to a suppression of SOD and GSH levels, an exacerbation of oxidative stress, and a promotion of cardiomyocyte apoptosis. Furthermore, it has been demonstrated to suppress mitochondrial membrane permeability. Subsequent RT-qPCR analysis of the hub genes revealed that only BCL2L1 exhibited significant alterations. Treatment with DOX altered the expression levels of apoptosis-associated proteins, BCL-2 family members, and caspase-3 family members. However, the administration of RA mitigated the deleterious effects of DOX on cardiomyocytes.
Conclusions
The protective effects of RA may against myocardial cell apoptosis are likely mediated through its activation of BCL2L1 and inhibition of caspase cascade protein expression in myocardial cells.
Introduction
Despite the demonstrated efficacy of anthracyclines in enhancing the long-term survival rates of cancer patients, their accumulation beyond a certain threshold within the body has been shown to result in substantial cardiac damage.
1
In 1971, Middleman identified a correlation between the dosage of anthracyclines and the extent of cardiac injury, highlighting a dose-dependent relationship.
2
Due to the lack of differentiation ability of myocardial cells, they are particularly susceptible to adverse effects of anthracycline antibiotics, which can accelerate heart damage.
3
Currently, there is a paucity of effective strategies for the prevention and management of anthracycline-induced cardiotoxicity, representing a critical challenge in the realm of oncology and cardiology research globally. In the context of experimental studies on anthracycline-induced cardiac injury, the model that utilizes doxorubicin (DOX, Figure 1a) is distinguished by its preeminent status as the most established.4,5 This includes a variety of DOX-induced models such as animal models (e.g., rabbit,
6
rat,
7
and mouse
8
cardiac injury models) and cellular models (e.g., rat cardiac H9c2 cell injury model
9
and mouse myocardial HL-1 cell injury model9,10). Differentially expressed genes in Control and DOX-induced cardiotoxicity. (a) The chemical 2D and 3D structures of 2 sesquiterpenes (Rosmarinic acid and doxorubicin); (b) Normalized expression matrices of GSE12260, PCA diagrams of the GSE12260 and Volcano plot of differentially expressed genes in GSE12260; (c) Normalized expression matrices of GSE59672, PCA diagrams of the GSE59672 and Volcano plot of differentially expressed genes in GSE59672; (d) Normalized expression matrices of GSE40289, PCA diagrams of the GSE40289 and Volcano plot of differentially expressed genes in GSE40289.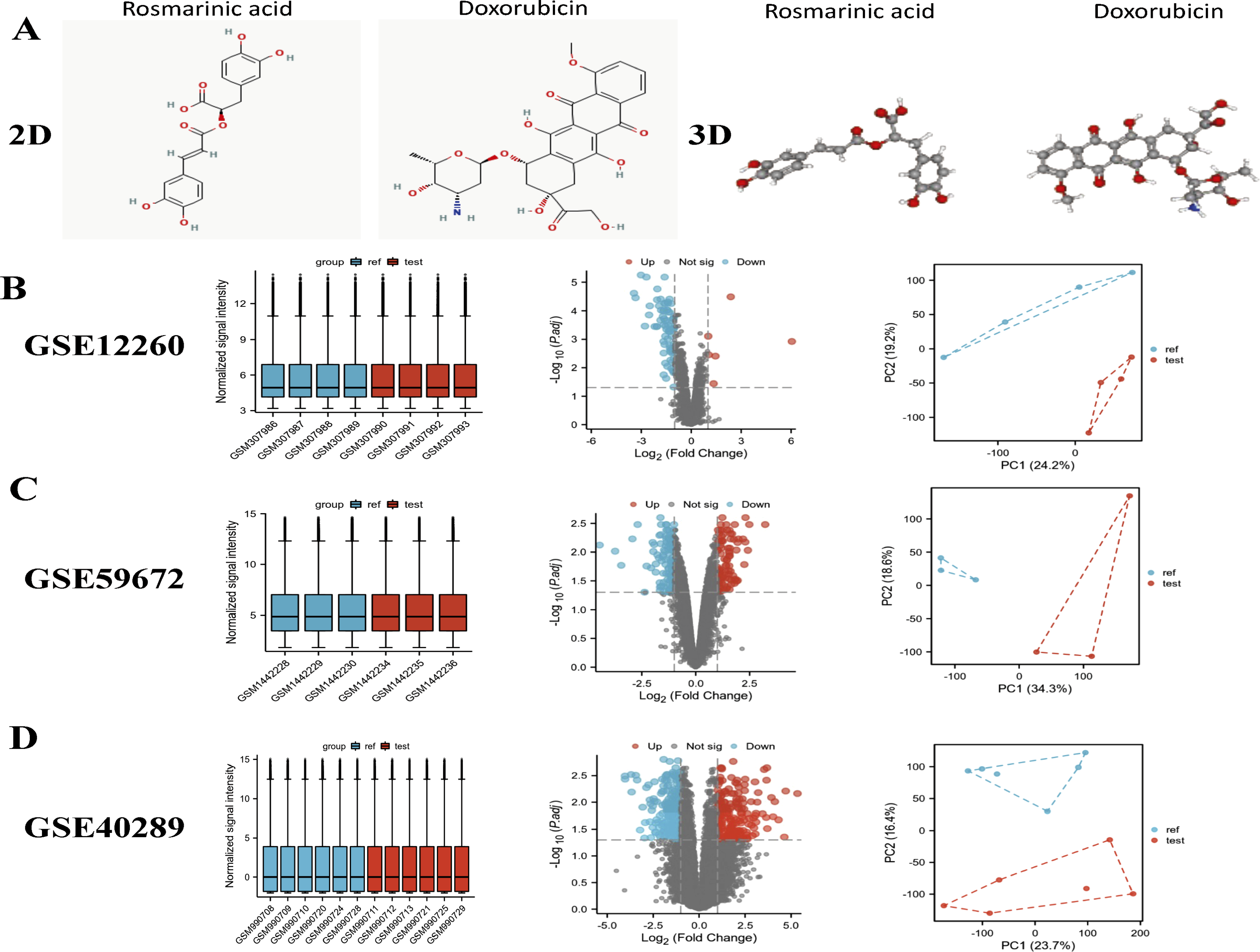
Rosmarinic acid (alpha-o-caffeoyl-3,4-dihydroxyphenyllactic acid, RA, Figure 1a), a phenolic acid compound prevalent in the plant kingdom, is predominantly found in various medicinal plants belonging to the Labiatae and Comfrey families, including rosemary, perilla leaf, sage, spearmint, bee balm, and tansy.11,12 RA can be obtained through direct extraction from plants or biosynthesized from L-phenylalanine and L-tyrosine via enzymatic reactions involving enzymes such as phenylalanine deaminase and tyrosine aminotransferase. 13 Additionally, it can be chemically synthesized through the esterification of caffeic acid and 3,4-dihydroxyphenyl lactic acid. 14 RA demonstrates a broad spectrum of biological activities, including anti-oxidant, 15 anti-inflammatory, 16 anti-microbial, 17 anti-viral, 18 and immune-modulatory effects. 19 It also shows therapeutic potential in treating tumors, 20 liver fibrosis, 21 and other conditions, as well as protective effects on the cardiovascular 22 and nervous systems. 23
Previous studies have shown that RA has a protective effect against DOX-induced cardiotoxicity, but the exact mechanism remains unclear. Earlier research directly selected a specific pathway for related experiments without further justification for the choice of pathway. The present study is designed to investigate the underlying mechanisms of RA’s cardioprotective effects against DOX-induced toxicity. A combination of network pharmacology, bioinformatics, and in vitro experimental approaches will be employed to this end.
Materials and methods
Reagents
RA and methanol were obtained from Shanghai Aladdin Biochemical Technology Co., Ltd (Shanghai, China). DOX was sourced from Sigma (Santa Clara, CA, USA). Assay kits for glutathione (GSH), superoxide dismutase (SOD), and malondialdehyde (MDA) were procured from Nanjing Jiancheng Bioengineering Institute (Nanjing, China). The 2,7-dichlorofluorescein diacetate (DCFH-DA) fluorescent probe was acquired from Beyotime Biotech Inc (Shanghai, China). The 5,5′,6,6′-tetrachloro-1,1′,3,3′-tetraethyl-imidacarbocyanine iodide (JC-1) fluorescent probe, horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG antibody, and ECL chemiluminescence substrates A and B were purchased from Yeasen Biotech Co., Ltd (Shanghai, China). Antibodies targeting cleaved-caspase-3, pro-caspase-3, BCL-2, BAX, and BCL2L1 were supplied by Abclonal Technology Co., Ltd (Wuhan, China). Antibodies against BAX were obtained from Abcam Technology Co., Ltd (UK). Antibodies specific to GAPDH were provided by Affinity Technology Co., Ltd (USA).
Bioinformatics analyses for differentially expressed genes (DEGs)
Drug Target Gene Screening:The two-dimensional (2D) and three-dimensional (3D) chemical structures of rosmarinic acid (RA) (Compound CID: 5281792) and doxorubicin (DOX) (Compound CID: 31703) were retrieved from the PubChem database. The 3D chemical structure of RA was saved in the “SDF” format and uploaded to the PharmMapper server (https://www.lilab-ecust.cn/pharmmapper/). PharmMapper uses Cavity to detect potential binding sites on the surface of a given protein structure, and sorts them based on their corresponding pharmacological efficacy scores. Then use the receptor-based pharmacophore modeling program Pocket to extract pharmacophore features from the cavity. Screen out pharmacophores with pKd values higher than 6.0. Molecular docking simulations between RA and potential target proteins were performed using the reverse docking function of PharmMapper. The target range was set to “Human Protein Targets Only,” while other parameters remained at their default settings.
Additionally, target genes associated with DOX were downloaded from the Comparative Toxicogenomics Database (CTD) (https://ctdbase.org/).
GEO Microarray Data Analysis: Expression profile datasets related to DOX-induced cardiotoxicity, including GSE12260, GSE59672, and GSE40289, were obtained from the Gene Expression Omnibus (GEO) database.
GSE12260 is a dataset based on the GPL1355 platform (Affymetrix Rat Genome 230 2.0 Array) and includes eight rat myocardium samples: four control samples (males), four DCM samples (males), and four DOX-treated samples (males). (https://www.ncbi.nlm.nih.gov/geo/download/?acc=GSE12260)
GSE59762 is a dataset based on the GPL1261 platform (Affymetrix Mouse Genome 430 2.0 Array) and includes six mouse myocardium samples: three control samples (males) and three DOX-treated samples (males). (https://www.ncbi.nlm.nih.gov/geo/download/?acc=GSE59672)
GSE40289 is a dataset based on the GPL13912 platform (Agilent-028005 SurePrint G3 Mouse GE 8 × 60 K Microarray) and includes 12 mouse myocardium samples: six control samples (males) and six DOX-treated samples (males). (https://www.ncbi.nlm.nih.gov/geo/download/?acc=GSE40289)
These datasets comprised 13 samples of cardiac tissue with DOX-induced cardiotoxicity and 13 normal cardiac tissue samples from healthy volunteers. Differential expression analysis was conducted using the R “limma” package, with normal cardiac tissues serving as controls. A threshold of | log2 (FoldChange)| > 1 and p < 0.05 was applied to identify significantly differentially expressed genes. The Sangerbox tool was utilized to screen for target genes. Candidate targets were identified by performing Venn analysis on the potential targets of RA, DOX, and the results from the GEO microarray analysis.
Disease ontology enrichment analysis
A thorough Disease Ontology (DO) enrichment analysis was performed using the DO package (version 3.30.1) in R, following methodologies outlined by Yu et al. and other pertinent studies in bioinformatics and systems biology. The gene sets associated with the molecules were derived from Section 2.2. The DO package combines semantic similarity measures and enrichment analysis to investigate the relationships between genes and diseases. This enrichment analysis identifies disease terms that are overrepresented in a given gene set, offering valuable insights into potential disease-related pathways. The analysis employs a hypergeometric distribution model to compute p-values, which are used to determine statistically significant disease terms. Furthermore, semantic similarity scores are calculated to evaluate the relatedness between the identified diseases. This ensures a precise analysis of gene sets within the framework of disease ontology. 24
Protein-protein interaction (PPI) network construction
By querying the STRING database with “Homo sapiens” as the specified species, a protein-protein interaction (PPI) network of the candidate targets was established. The regulatory network was subsequently constructed and imported into Cytoscape (v3.8.2) for detailed analysis and visualization. The software was employed to evaluate and rank the network nodes based on key topological metrics, such as Maximum Clique Centrality (MCC), Betweenness, Bottle Neck, Degree, Density of Maximum Neighborhood Component (DMNC), Edge Percolated Component (EPC), and Maximum Neighborhood Component (MNC). These metrics, along with the combined score, were visualized using distinct colors to emphasize their importance. The candidate targets were then ranked according to these values to identify the most critical nodes within the network.
Functional enrichment analysis
Gene Ontology (GO) enrichment analysis of the candidate targets was performed using the R language “ClusterProfiler” package, encompassing biological process (BP) and cellular component (CC) categories. A significance threshold of p < 0.05 was applied to identify enriched terms. Additionally, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis was conducted using the online tool available at https://sangerbox.com/Tool. The results were visualized through bar and loop graphs to illustrate the enriched pathways and their relationships.
Molecular docking simulation
The five core proteins-HMOX1 (PDB ID: 3CZY), STAT1 (PDB ID: 7NUF), SELE (PDB ID: 1G1T), BCL2L1 (PDB ID: 7JGW), and LCN2 (PDB ID: 6Z6Z)-were retrieved and filtered from the Protein Data Bank. The chemical structure of rosmarinic acid (RA) was obtained from the PubChem database and converted into a 3D structure using Chem3DUltra 14.0 software. The structure was energy-minimized using the MM2 algorithm. The target protein receptors were prepared by removing water molecules and organic substances with PyMOL software, followed by hydrogenation and charge calculation using AutoDockTools 1.5.6. Appropriate box center and lattice parameters were defined, and both the compounds and target protein receptors were converted into “pdbqt” files. Molecular docking was performed using the Vina 1.1.2 tool, and docking energy values were calculated to evaluate the binding interactions.
Cell culture
Rat cardiac myocytes (H9c2) were acquired from the American Type Culture Collection. The H9c2 cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM) with a glucose concentration of 4.5 mg/mL, enriched with 10% fetal bovine serum. The cells were incubated in a humidified atmosphere containing 5% CO2 at 37°C, and the culture medium was refreshed every 2–3 days to ensure optimal growth conditions. In addition, this experiment has been approved by the Ethics Committee of Nanjing University of Traditional Chinese Medicine for in vitro study of H9c2 cells.
Cytotoxicity of RA and the dose of RA inhibiting DOX induced cell damage
The cells were plated into 96-well plates at a density of 5 × 104 cells/mL prior to treatment, with varying concentrations of rosmarinic acid (RA) (0, 12.5, 25, 50, 100, 200, and 400 μM). Cells in the Control group (CON) were treated with equivalent volumes of serum-free medium. After 24 h of incubation, cell viability was assessed using the CCK8 assay. In a separate experiment, H9c2 cells were pretreated with different concentrations of RA (5, 10, 30, 50, 100, and 200 μM) for 30 min, followed by exposure to doxorubicin (DOX) at a concentration of 1 μM. Control cells (CON) received equal amounts of serum-free medium. Cell viability was subsequently measured using the CCK8 assay.
Cell treatments
H9c2 cells were categorized into three experimental groups. The control group (CON) consisted of cells cultured in serum-free DMEM high-glucose medium. The MOD group (MOD) involved cells cultured in serum-free DMEM high-glucose medium supplemented with 1 μM DOX. The RA group (RA) included cells pretreated with 50 μM RA for 30 min prior to the addition of 1 μM DOX. Cells from all groups were incubated for 24 h under the specified conditions.
CCK-8 assay
The CCK-8 assay (C0039, Beyotime Biotechnology) was employed to evaluate cell viability. H9c2 cells were plated into 96-well plates at a density of 1 × 103 cells per well in 100 μl of medium devoid of 10% fetal bovine serum (FBS). After allowing the cells to adhere completely, the culture medium alone was designated as the blank group. Following a 24-h incubation period, 10 μl of CCK-8 solution was added to each well, and the cells were further incubated for 4 h at 37°C. The optical density (OD) at 450 nm was then measured to determine cell viability. 25
ELISA
Cell supernatant was collected where the levels of SOD, MDA and GSH. The OD450 value of each well was measured and the protein concentration was calculated according to a standard curve. 26
Intracellular ROS levels
H9c2 cells were plated into a 6-well culture plate and exposed to DOX for 24 h. The intracellular level of reactive oxygen species (ROS) was measured using a fluorescent probe, adhering to the manufacturer’s guidelines. The fluorescence intensity for each group was captured and analyzed using an inverted fluorescence microscope (Olympus, Tokyo, Japan).
JC-1
H9c2 cells were plated into a 6-well culture plate and treated with DOX for 24 h. Changes in mitochondrial membrane potential (MMP) were assessed using the JC-1 fluorescent probe, following the manufacturer’s protocol. 27 The fluorescence intensity for each group was measured and visualized using an inverted fluorescence microscope (Olympus, Tokyo, Japan).
RT-qPCR
Primer sequences for real-time PCR.

Western blot
Western blot analysis was employed to assess protein expression levels in H9c2 cells. Briefly, H9c2 cells were lysed in 100 μL of RIPA lysis buffer, and protein concentrations were quantified using the BCA assay. Equal amounts of protein samples were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred onto nitrocellulose membranes. The membranes were blocked with 5% non-fat milk in Tris-buffered saline (TBS) and then incubated with primary antibodies, including anti-BAX (1:1000 dilution), anti-BCL2L1 (1:500 dilution), anti-cleaved-caspase-3 (1:500 dilution), anti-pro-caspase-3 (1:500 dilution), anti-BCL-2 (1:500 dilution), anti-BAX (1:500 dilution), anti-BCL2L1 (1:500 dilution), and anti-GAPDH (1:1000 dilution). The membranes were then subjected to an overnight incubation with primary antibodies at 4°C, followed by three washes with TBS-T. Subsequently, the membranes were incubated with appropriate horseradish peroxidase (HRP)-conjugated secondary antibodies (1:5000 dilution) for 60 min. Protein bands were visualized using enhanced chemiluminescence (ECL) reagents, and protein expression levels were analyzed using a digital gel imaging system (Alpha Imager2200, Alpha Innotech Corporation, San Leandro, CA, USA). 28 Uncut original Western blot images are provided in Supplemental 1.
Statistical analysis
The data are presented as mean ± standard deviation and were analyzed using SPSS 21.0 (IBM, Armonk, NY). Comparisons between two groups were performed using the unpaired t-test, while comparisons among multiple groups were conducted using one-way ANOVA or repeated measures ANOVA, followed by Tukey’s post-hoc test for further analysis. Univariate analysis was carried out using the Log-rank test. A p-value of less than 0.05 (p < 0.05) was considered statistically significant.
Results
The differentially expressed genes of GSE12260, GSE59672 and GSE40289 were screened by Limma package
RA, a naturally occurring polyphenolic compound predominantly found in rosemary, mint, perilla frutescens, and sarcandra glabra, has been documented to exhibit a wide array of biological activities. These include anti-inflammatory, antioxidant, antibacterial, and HIV-1 inhibitory properties, among others. 29
We analyzed three datasets to evaluate gene expression differences between cardiac toxicity groups induced by anthracycline drugs and control groups. The expression matrices of the datasets GSE12260, GSE59672, and GSE40289 were normalized, and the box plots demonstrated consistent distribution trends, appearing as straight lines. In order to assess the reproducibility of data from the same group, principal component analysis (PCA) was conducted on the three datasets, revealing adequate reproducibility. Using the threshold criteria of an adjusted |log2(FC)| > 1 and p < 0.05, we identified 86 differentially expressed genes (DEGs) in the GSE12260 dataset, 434 DEGs in the GSE40289 dataset, and 210 DEGs in the GSE59672 dataset. Volcano plots illustrating the DEGs for the three datasets are presented in Figure 1(b)–(d).
RA is an active ingredient in the treatment of anthracyclines induced cardiac toxicity
Figure 2(a) displays the partial gene heatmap expression patterns for the three datasets. Additionally, genes associated with anthracyclines were identified through the CTD, resulting in a total of 8111 genes. Screening target genes for RA to improve cardiac toxicity of anthracycline drugs. (a) Part of hierarchical clustering heatmap of the 3 datasets differentially expressed genes. (b) Venn analysis was performed between the target genes of RA and GEO database and CTD database. RA: Rosmarinic acid.
Next, target genes of RA were screened, identifying 401 related genes. Venn analysis was performed between the target genes of RA and those from the GEO and CTD databases, revealing 3, 12, and 15 target genes associated with anthracycline-induced cardiac toxicity, respectively (Figure 2(b)). These genes were combined, resulting in a total of 24 genes: HSPA1A, HMOX1, BCL2L1, GSTM1, MAPK10, MMP3, S100A9, CFB, ERBB4, SULT1A1, CDK5, DCXR, GLRX, PLEKHA4, TAP1, TTR, LCN2, CTSL, SELE, STAT1, AHCY, GATM, KIT, and QPCT.
Identification of hub genes and GO, DO and KEGG analysis
The 24 genes were imported into the STRING database for protein-protein interaction (PPI) analysis. The results were then imported into Cytoscape software to construct a drug-gene interaction network. By retaining genes that interact with each other, a total of 17 genes were identified: HSPA1A, HMOX1, BCL2L1, GSTM1, MMP3, S100A9, SULT1A1, GLRX, PLEKHA4, TAP1, TTR, LCN2, CTSL, SELE, STAT1, AHCY, and KIT (Figure 3(a)). GO/KEGG and DO functional analysis and PPI analysis of the five candidate targets. (a) Construction of drug and gene interaction network diagram in Cytoscape software; (b) DO function analysis of candidate RA targets genes; (c) GO function analysis of candidate RA targets regarding the BP, CC, and MF and KEGG pathway enrichment analysis; (d) five hub genes of using Cytoscape analysis. RA: Rosmarinic acid.
The 17 candidate targets were further analyzed using GO and KEGG enrichment analyses. The GO functional analysis revealed that these targets are involved in biological processes (BP) such as the regulation of apoptotic signaling pathways, intrinsic apoptotic signaling pathways, extrinsic apoptotic signaling pathways, and cellular responses to toxic substances. In terms of cell component (CC), they were primarily enriched in vesicle lumen, cytoplasmic vesicle lumen, and secretory granule lumen. KEGG pathway analysis indicated that these targets participate in key pathways, including the MAPK signaling pathway, autophagy, apoptosis, and the TNF signaling pathway (Figure 3(c)). These findings underscore the involvement of the 17 candidate targets in the mechanisms underlying cardiac toxicity, supporting their potential role as therapeutic targets against anthracycline-induced cardiac toxicity. To further refine the analysis, the proteins encoded by the 17 candidate genes were subjected to PPI analysis using the STRING database. The top five targets, ranked based on their Maximum Clique Centrality (MCC), Betweenness, Bottle Neck, Degree, Density of Maximum Neighborhood Component (DMNC), Edge Percolated Component (EPC), and Maximum Neighborhood Component (MNC) values, were identified as HMOX1, BCL2L1, LCN2, SELE, and STAT1 (Figure 3(d)). These five targets were selected for subsequent in-depth analysis.
Construction of pharmacophore models of RA and candidate targets
The pharmacophore structure of the drug molecule RA and the top five receptor targets involved in anthracyclines induced cardiac toxicity regulation according to MCC, Betweenness, Bottle Neck, Degree, DMNC, EPC, MNC values were analyzed with the help of the PharmMapper database. The results showed that in the RA molecule, 5 Hydrophobic, 1 donor, and 2 Acceptor interact with HMOX1; 1 Hydrophobic, 1 Positive, 2 Donor and 4 Acceptor interact with the STAT1; 1 Negative, 1 Donor and 7 Acceptor interact with SELE; 1 Hydrophobic, 1 Negative, 1 Donor and 1 Acceptor interact with the LCN2 and 6 Hydrophobic, 1 Positive, 1 Donor and 1 Acceptor regions interact with BCL2L1 (Figure 4(a)–(f)). Pharmacophore model of interactions between candidate targets and RA molecule. (a-e) Pharmacophore model diagrams of RA molecule with HMOX1, BCL2L1, LCN2, SELE and STAT1 protein molecule, respectively; (f) The binding situation of each pharmacophore model (g) The number of binding elements in each pharmacophore; higher Normalized Fit Score, Z’-score, and Normalized Fit Score reflect a greater probability of being the targets of the RA molecule. RA: Rosmarinic acid.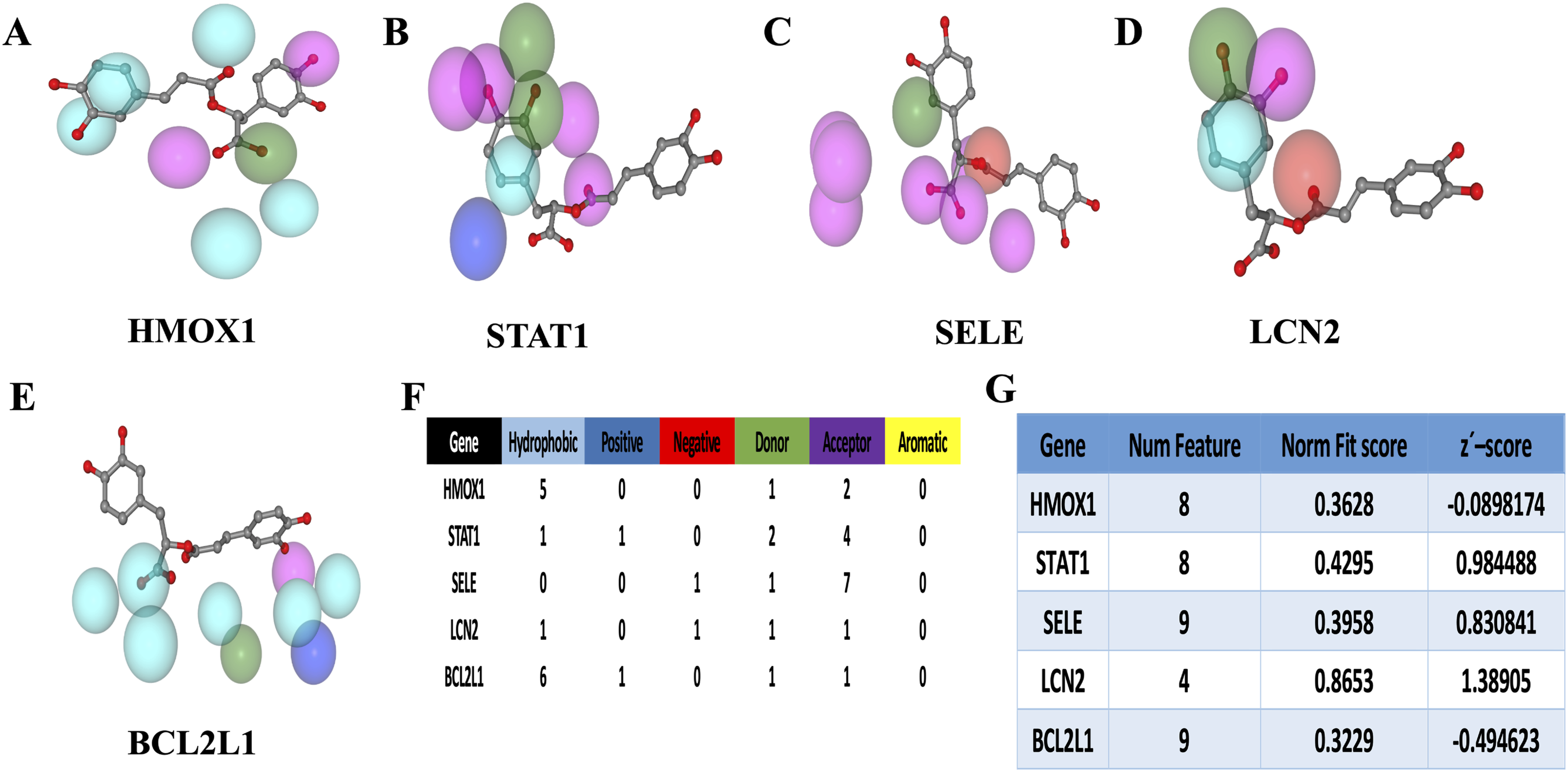
Based on the corrected scores, the receptor targets were ranked as LCN2, STAT1, SELE, HMOX1, and BCL2L1. The Normalized Fit Score indicates that higher scores correspond to a greater likelihood of the target being a molecular target of RA (Figure 4(g)). Utilizing these results, a pharmacophore model was constructed to illustrate the interactions between the candidate targets and RA molecules, thereby establishing a foundation for subsequent analyses.
Docking validation of candidate targets with RA molecules
We conducted molecular docking analysis between the five candidate proteins (HMOX1, STAT1, SELE, LCN2, and BCL2L1) and the RA molecule using AutoDockTools 1.5.6 and Vina 1.1.2 software. Each protein-ligand pair was docked three times, and the minimum binding free energy values were averaged to ensure accuracy. The 3D diagrams visually illustrate the binding modes between the target protein receptors and the RA molecule, as well as their interactions with surrounding amino acid residues (Figure 5(a)–(e)). These results provide detailed insights into the molecular interactions and binding affinities, supporting further functional analysis. Molecular docking between candidate targets and RA. (a-e) Molecular docking between five candidate targets and RA; (f) Bar chart of the binding energy of molecular docking between each candidate target and RA. RA: Rosmarinic acid.
In instances where the binding energy is less than 0 kJ/mol, it is indicative of the spontaneous binding and interaction of molecular proteins with one another. Lower binding energy values are indicative of more stable molecular conformations. The analysis revealed that the five proteins ranked in terms of their binding affinity from low to high as follows: BCL2L1, HMOX1, SELE, STAT1 and LCN2 (Figure 5(f)). This ranking suggests that BCL2L1 has the strongest binding affinity with RA, while LCN2 has the weakest.
Cytotoxicity of RA on H9c2 cells and the optimal protective concentration for cardioprotection
RA did not exhibit cytotoxicity toward H9c2 cells within a specific concentration range, as evidenced by the unchanged cell survival rates, indicating its non-toxic nature (Figure 6(a)). Treatment with RA significantly enhanced the survival rates of H9c2 cells in a dose-dependent manner within the range of 5–50 μM (Figure 6(b)). The highest survival rate was observed at 50 μM RA (p < 0.05). However, further increases in RA concentration beyond 50 μM did not result in additional changes in cell survival rates. In light of the findings, 50 μM RA was identified as the optimal concentration for subsequent experiments in this study. Chen et al.
30
also chose 50 µM RA and found that RA inhibits vascular remodeling and exerts a cardioprotective effect. Cytotoxicity of RA on H9c2 Cells and the Optimal Protective Concentration for Cardioprotection. (a) Effect of RA on the survival rate of H9c2 cells. *p < 0.05 compared with the control group; (b) effects of RA on the cell viability of H9c2 cells induced by DOX; (c) Superoxide dismutase (SOD) levels; (d) malondialdehyde (MDA) levels; (e) glutathione (GSH) levels. *p < 0.05 compared with the control group, #p < 0.05 compared with the DOX group. Data are expressed as mean ± SEM (n = 6). CON: Control, MOD: DOX-induced model, RA: Rosmarinic acid.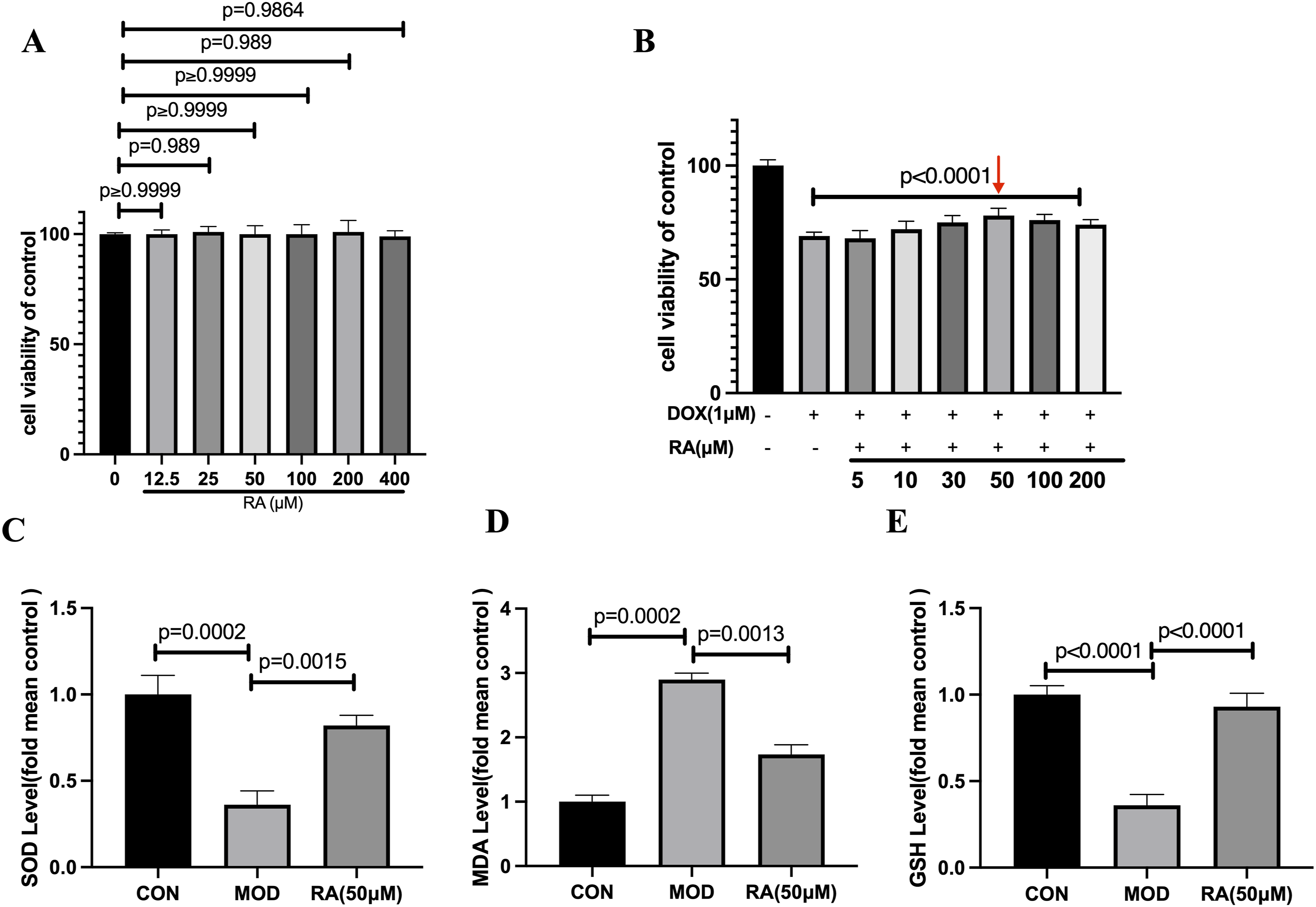
Effects of RA on DOX-induced oxidative stress
The results demonstrated that SOD and GSH levels were significantly lower, while MDA levels were significantly higher in the DOX group compared to the control group (p < 0.05) (Figure 6(c)–(e)). Further analysis revealed that in the RA group, SOD and GSH levels were significantly elevated, whereas MDA levels were significantly reduced compared to the DOX group (p < 0.05). Additionally, the results of DCFH-DA staining indicated that DOX treatment markedly increased intracellular ROS levels. However, RA significantly attenuated DOX-induced ROS generation in the cells (Figure 7(a)). RA inhibited DOX-induced ROS and Cell mitochondrial membrane. (a) ROS levels in H9c2 cells (n = 3); (b) Cell mitochondrial membrane potential using JC-1 (n = 3). CON: Control, MOD: DOX-induced model, RA: Rosmarinic acid.
Effects of RA on the mitochondria in cells treated with DOX
Cells treated with DOX exhibited a significant reduction in mitochondrial membrane potential (MMP) compared to those in the control group (Figure 7(b)). In contrast, the RA group demonstrated a significantly higher mitochondrial membrane potential than the DOX group, indicating that RA effectively mitigated the DOX-induced loss of MMP.
qRT-PCR detection of the expression of hub genes in H9c2 cells
Quantitative real-time PCR (qRT-PCR) was employed to assess the expression levels of HMOX1, BCL2L1, LCN2, SELE, and STAT1 in the different experimental groups (Figure 8(a)–(e)). The results revealed that only BCL2L1 exhibited significant changes. Specifically, compared to the control group, the expression of BCL2L1 in the DOX group was significantly reduced (p < 0.05). Conversely, in the RA group, the expression of BCL2L1 was significantly increased compared to the DOX group (p < 0.05). These findings suggest that RA may exert its protective effects by modulating the expression of BCL2L1. qRT-PCR detection of the expression of hub genes in H9c2 Cells. (a) Representative HMOX1 mRNA expression levels; (b) representative STAT1 mRNA expression levels; (c) representative SELE mRNA expression levels; (d) representative BCL2L1 mRNA expression levels; (e) representative LCN2 mRNA expression levels; *p < 0.05 compared with the control group, #p < 0.05 compared with the DOX group. Data are expressed as mean ± SEM (n = 3). CON: Control, MOD: DOX-induced model, RA: Rosmarinic acid.
RA reduces DOX induced cardiac toxicity in H9c2 cells by inhibiting the apoptotic signaling pathway
The expression levels of pro-caspase 3, BCL2L1, and the BCL-2/BAX ratio in the DOX group were significantly lower compared to the control group (CON), while the level of cleaved-caspase 3 was significantly higher (p < 0.05). In contrast, the RA group exhibited significantly higher expression levels of pro-caspase 3, BCL2L1, and the BCL-2/BAX ratio compared to the DOX group, along with a significantly lower level of cleaved-caspase 3 (p < 0.05) (Figure 9). These results suggest that RA mitigates DOX-induced apoptosis by modulating the expression of key apoptotic regulators. RA ameliorates the expression of BCL2L1, BCL-2, BAX and Caspase-3 proteins in the H9c2 cells. (a) Representative BCL2L1, BCL-2, BAX and Caspase-3 proteins expression levels; GAPDH was used as an internal control; (b–e) representative BCL2L1, Caspase-3 and GAPDH protein ratios; BCL-2 and BAX ratio. *p < 0.05 compared with the control group, #p < 0.05 compared with the DOX group. Data are expressed as mean ± SEM (n = 3). CON: Control, MOD: DOX-induced model, RA: Rosmarinic acid.
Discussion
Doxorubicin (DOX), an anthracycline drug, is widely used in the treatment of various cancers, including breast cancer, bladder cancer, lymphoma, and acute lymphoblastic leukemia. Despite its high efficacy, DOX is associated with dose-dependent acute, subacute, and chronic irreversible cardiotoxicity, which can progress to heart failure. 2 Recent studies have further demonstrated that RA possesses potent anti-apoptotic capabilities and can protect cells against hydrogen peroxide-induced DNA damage and apoptosis. 31
A substantial body of research has been dedicated to investigating the pharmacological effects of RA on DOX-induced cardiac toxicity. However, the underlying mechanisms that underpin these effects remain to be fully elucidated.32,33 To date, only two studies have examined the potential mechanism of RA in relation to DOX-induced cardiac toxicity. Zhang et al. reported that RA inhibits the expression and release of FasL in cardiac fibroblasts through paracrine mechanisms, thereby reducing cardiomyocyte apoptosis. However, the authors did not provide a clear theoretical basis for selecting this signaling pathway. 29 Similarly, Kim et al. demonstrated that RA inhibits ADR induced apoptosis in H9C2 cardiomyocytes by reducing ROS production and suppressing JNK and ERK activation, but also did not explain the selection of apoptotic signaling pathways. 34 In contrast, the present study employed bioinformatics and network pharmacology to identify pivotal targets for the enhancement of RA. This study constitutes a significant addition to the extant body of research in this field. The findings of the present study offer supplementary mechanistic evidence that lends further credence to the therapeutic potential of RA in the context of DOX-induced cardiac injury.
Employing bioinformatics methodologies, we identified potential DEGs associated with anthracycline-induced cardiac toxicity. This identification was made by intersecting DEGs from the GSE12260, GSE59672, and GSE40289 datasets with cardiotoxicity-related genes from the CTD and genes regulated by RA. This analysis yielded a total of 17 DEGs. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis further indicated that these targets participate in key pathways, including the apoptosis signaling pathway. Chen et al. demonstrated that resveratrol effectively alleviates Dox-induced cardiotoxicity by suppressing ferroptosis possibly through modulating the MAPK signaling pathway. 35 Mohammed et al. suggest that azilsartan effectively reduced doxorubicin-induced cardiotoxicity, likely by mitigating apoptosis, inflammation, and oxidative stress in cardiac tissues. 36 To further refine our analysis, we conducted protein-protein interaction (PPI) analysis using the STRING database for the proteins encoded by the 17 candidate genes. The results highlighted the top five action targets based on their interaction scores and functional relevance. A summary of these findings identified five common hub genes: HMOX1, BCL2L1, LCN2, SELE, and STAT1. These genes are likely central to the molecular mechanisms underlying anthracycline-induced cardiac toxicity and represent potential therapeutic targets for intervention.
The presence of elevated oxidative stress and compromised antioxidant mechanisms has been identified as a primary contributing factor to the development of cardiotoxicity induced by DOX. 37 DOX has been shown to reduce the levels of various antioxidants in H9c2 cells. Previous studies, such as those by Wu et al. have demonstrated that DOX increases MDA and LDH levels while suppressing SOD activity, leading to oxidative stress in primary cardiomyocytes. 38 In this study, we observed that treatment with RA significantly increased SOD and GSH levels while reducing MDA levels in H9c2 cells. These findings suggest that RA enhances the body’s antioxidant capacity, thereby providing protection against DOX-induced cardiotoxicity. Apoptotic signals reduce mitochondrial membrane potential (MMP), triggering the release of apoptotic signaling molecules that activate caspase proteases and initiate cell apoptosis. 39 In a study by Jiang et al. the JC-1 probe was used to assess changes in MMP in DOX-induced H9c2 cells. 40 They observed that DOX increased green fluorescence intensity (indicative of depolarized mitochondria) and decreased red fluorescence intensity (indicative of healthy mitochondria). In the present study, DOX demonstrated a substantial inhibitory effect on MMP, while pretreatment with RA led to a significant restoration of MMP.
BCL2L1, an anti-apoptotic protein and a key regulator of apoptotic genes, primarily modulates apoptosis through mitochondrial pathways. 41 Shan et al. suggest that Hsp10 and Hsp60 modulated post-translational modification of Bcl-xl. Antisense Hsp60 reduced the abundance of endogenous Hsp60 in cardiomyocytes and amplified the cytotoxicity of doxorubicin. 42 In our study, the RA group exhibited significantly elevated expression levels of pro-caspase 3, BCL2L1, and the BCL-2/BAX ratio compared to the DOX group, along with a significantly reduced level of cleaved-caspase 3. These results suggest that RA may plays a protective role in DOX-induced apoptosis by enhancing anti-apoptotic mechanisms and suppressing caspase activation, thereby preventing cell death. Divya et al. also provided evidence demonstrating the efficacy of RA in combating UVB induced apoptosis related proteins, including BCL-2 and BAX. Mdivi-1 was used as a positive control in these experiments. 43 These experiments indicate that the RA have the anti-apoptotic effect.
The present study demonstrates that RA effectively protects against DOX-induced cardiomyocyte apoptosis, may through BCL2L1 activation and caspase cascade inhibition. The results of this study underscore the RA’s dualistic mechanism of enhancing anti-apoptotic pathways while concomitantly reducing oxidative stress. This dual mechanism positions RA as a promising candidate for the treatment of anthracycline-induced cardiotoxicity. While these findings provide strong preclinical evidence, future studies involving in vivo models and BCL2L1 functional experiments will be crucial to fully validate RA’s cardioprotective mechanisms and therapeutic potential.
Supplemental Material
Supplemental material - Network pharmacology and experimental verification: Rosmarinic acid alleviates doxorubicin-induced cardiomyocyte apoptosis by regulating BCL2L1
Supplemental material for Network pharmacology and experimental verification: Rosmarinic acid alleviates doxorubicin-induced cardiomyocyte apoptosis by regulating BCL2L1 Sicong Xie, Cheng Chang, Rongxing Jiang, Lifeng Wang, Yunli Yang, Zongjin Li and Yang Zhang in Human & Experimental Toxicology
Footnotes
Ethical consideration
Author contributions
Yang Zhang, Sicong Xie, Rongxing Jiang, Lifeng Wang, Yunli Yang, Zongjin Li and Cheng Chang designed and coordinated the study; Sicong Xie, Cheng Chang and Rongxing Jiang performed the experiments, acquired and analyzed data; Yang Zhang, Zongjin Li, Cheng Chang, Rongxing Jiang and Lifeng Wang interpreted the data; Yang Zhang, Zongjin Li and Yunli Yang wrote the manuscript; all authors approved the final version of the article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project supported by the National Natural Science Foundation of China (No. 82302847), Jiangsu Province Basic Research Program (No. BK20241907) and Nanjing University of Traditional Chinese Medicine National Natural Science Foundation Youth Science Fund Supporting Project (No. 012062005001-26).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Raw data (e.g., qPCR and Western blot quantifications) are available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
