Abstract
The use of pesticides in farmland has increased considerably to protect crops against pests, weeds, and diseases. However, pesticides and/or their residues in ecosystems may affect non-target organisms. Indaziflam is a widely used herbicide in agricultural areas in the southern region of Turkey. Therefore, this study aimed to investigate the possible genotoxic and cytotoxic effects of indaziflam on HepG2 cells using comet assay, micronucleus assay, and xCELLigence. The HepG2 cells were treated with various concentrations of indaziflam for different duration of time based on xCELLigence results. Accordingly, the cells were incubated with indaziflam at final concentrations of 1, 5, 10, 20, 40, and 80 μg/mL for 96 h for cytotoxicity assay. To assess genotoxicity, cells were treated with indaziflam at final concentrations of 10, 40, and 100 μg/mL for 4 and 24 h. Ethanol was used as a solvent for indaziflam. Hydrogen peroxide (40 μM) was used as a positive control. Studies have revealed that indaziflam did not show a statistically cytotoxic effect at the tested doses. Nevertheless, genotoxicity studies showed that indaziflam induced both DNA strand breaks and micronucleus numbers depending on the exposure time and dose.
Introduction
Soil, one of the essential components of ecosystems, has been contaminated consistently by various agents, including pesticides, heavy metals, or Polycyclic Aromatic Hydrocarbon (PAHs).1,2 Among contaminants, pesticides are the most worrisome. 2 Pesticides can reach the soil directly (agricultural applications) and indirectly (pesticide production sites, occupational accidents, surface runoff, etc.). 3 Ideally, a pesticide should have some properties. It should only affect the target organism. In addition, it should not remain in the ecosystem for a long time and have no harmful environmental effects. 4 However, many pesticides do not have these properties.
Pesticides may lead to various toxic effects on target and non-target organisms. 5 Because of the chemical and physical properties of many pesticides, they may migrate between ecosystems. Thus, many living organisms may exposure to increasing pesticide doses through food chains. Many organisms deal with it through antioxidant systems to prevent and minimize the adverse effects of pesticides. However, antioxidant systems and other regulatory processes may be inefficient in some organisms. For this reason, determining pesticide toxicity in the target and non-target organisms is crucial worldwide. 5
Pesticides are classified as herbicides, fungicides, and insecticides according to targeted organisms. Indaziflam is a herbicide that belongs to the chemical class of alkylazines and is currently used as a cellulose biosynthesis inhibiting. 6 It has broad-spectrum activity at low application rates. 7 Indaziflam is commonly used in crops (e.g. citrus, grapes, pome, and stone fruit) and commercial areas (e.g. turfgrass, landscape ornamentals, and Christmas trees). 8 Indaziflam can reach non-target organisms, including animals and humans, through the food cycle as a result of its widespread use. This phenomenon raises concerns for animal and human health safety. When non-target organisms are constantly exposed to indaziflam, adverse effects such as cytotoxicity and genotoxicity can occur on non-target organisms. However, it has not been sufficient data on genotoxic and cytotoxic effects on non-target organisms.
The toxicity mechanism of many pesticides is not well known. Some of them may cause oxidative stress, endocrine disruption, and deplete antioxidant defences. Moreover, pesticides can increase the levels of reactive oxygen species in the cell and thus affect genome integrity. The genome integrity should be preserved in living things. Otherwise, DNA damage in the genome can cause cancer, infertility, and extinction.9,10 DNA damages are measured using the comet and micronucleus assays.11,12 These methods detect DNA strand breaks and chromosome damage in almost any cell type.
Various metabolic enzymes involved in xenobiotic biotransformation in normal cells may lose their functions during the proliferation of cells. However, HepG2 cells retain cytochrome P450-dependent monooxygenase secretion and glucuronic-and-sulfate-conjugation abilities, which play an essential role in the detoxification process of cells.13–16 Therefore, HepG2 cells are generally chosen as an in vitro model to investigate the genotoxicity of different chemical compounds like plant extract, pyrrolizidine alkaloids, and herbicides.14,15,17 In addition, their excellent ability to re-establish their cellular organization makes them a good in vitro model for cytotoxicity tests that require long-term repeated dose exposure. In this context, the present study aimed to detect the genotoxic and cytotoxic effects of indaziflam on the HepG2 cell lines.
Materials and methods
Chemicals
The analytical standard of indaziflam was purchased from Sigma Aldrich CAS-No 950782-86-2. Eagle’s minimum essential medium, heat-inactivated fetal bovine serum, penicillin, and streptomycin were obtained from Biowest. All other chemicals used in experiments were purchased from Sigma (Darmstadt, Germany) and Merck (Darmstadt, Germany).
Cell culture
HepG2 [ATCC® HB8065] cell line was acquired from the American Type Culture Collection. HepG2 cells were cultured in an EMEM medium with 10% heat-inactivated fetal bovine serum and 1% penicillin-streptomycin. Cells were incubated in an incubator with a humidified atmosphere containing 5% CO2 at 37°C. The medium was changed 2-3 times a week. 1M stock solution of indaziflam was prepared using pure ethanol. The stock solution was diluted by culture medium to the other concentrations (1, 5, 10, 20, 40, and 80 μg/mL). The final volume of ethanol represents under 1% of the total medium volume. Cells were seeded on 24 wells culture plates at 25 × 104 cells/well for the comet and cytokinesis-block micronucleus (CBMN) assays. Following seed, the cells were incubated for 24 h and then treated with studied concentrations (1, 5, 10, 20, 40, and 80 μg/mL indaziflam, 40 μM hydrogen peroxide (H2O2), 200 μg/mL cyclophosphamide).
Cell viability and density
Cells were washed twice with PBS and incubated in trypsin-EDTA solution (0.25%) at 37°C for 5 min. The cell suspension was centrifuged at 130 × g for 5 min. The pellet was resuspended in a 1 mL complete medium. Then, 10 μL of cell suspension was mixed with 10 μL of trypan blue. Finally, the mixture was subjected to Cedex XS Analyzer to determine cell viability and density.
Measurement of cytotoxicity as cell index
The Real-time cell analysis (RTCA) systems including xCELLigence™, have been used to examine proliferation, viability, invasion, and cytotoxicity in cancer cells. The xCELLigence system measures cell number in each well of microtiter plates through gold microelectrode biosensors. Increasing the number of adherent cells in each well leads to altering in the impedance. The impedance difference detects various biological events including, proliferation, invasion, and apoptosis. The Increasing proliferation of cells at different time intervals is recorded as cell index. As a result, the test system is reliable for evaluating interactions between agents and cancer cells. 18
The impedance-based cytotoxicity assay was performed using the xCELLigence (Real-Time Cell Analysis (RTCA) Dual Plate (DP)) (Roche Diagnostics GmbH, Penzberg) instrument at 37°C with 5% CO2. Firstly, 100 μL of the medium was added to each well of 16 wells plates. After measuring background impedance, 90 μL of the cell suspension (5 × 103 cell/well) were seeded into the 16 wells E-plates. E-plates were incubated for 24 h. Afterward, different concentrations of indaziflam and ethanol were added to each well for treatment. Cell index was recorded at 15 min intervals until the end of the treatment (96 h).
Comet assay
The comet assay protocol described by Singh et al. (1988) was used with minor modifications in the study. 19 HepG2 cells were treated with indaziflam (10, 40, and 100 μg/mL) and hydrogen peroxide (40 μM) for 4 and 24 h. After treatment, cells were washed with phosphate-buffered saline (PBS) and mixed to a low melting point agarose (0.65%). The mixture was then rapidly shed onto microscope slides covered with standard melting point agarose (1%). Slides were kept in the refrigerator for 15 min at 4°C to allow the mixture to freeze. Following freezing, the slides were transferred to a fresh lysis buffer (2.5 M NaCl, 100 mM Na2-EDTA, 10 mM Tris, pH 10, with 10% dimethyl sulfoxide and 1% Triton X-100 added fresh) and incubated for 1 h at 4°C. The slides were placed on horizontal electrophoresis and submerged in an alkaline solution (1 mM Na EDTA, 300 mM NaOH and pH 13) for 20 min. All procedures were practiced to prevent DNA damage in dark conditions. Electrophoresis was performed at 25 V (300 mA) for 20 min. After then, the slides were removed to a cold neutralizing buffer for 10 min. Ethidium bromide dye was added to each slide to stain the DNA. DNA damage was measured for the 100 cells at each slide. The damage type was calculated as the tail length in the comet assay.
Cytokinesis-block micronucleus assay
The micronucleus assay was performed according to Darroudi and Natarajan (1994). 20 The HepG2 cell line was seeded in 24 wells plates at a 2 × 105 cell/well density. After 24 h attachment, the medium was removed and cells were exposed to indaziflam, cyclophosphamide (200 μg/mL), and ethanol for 4 and 24 h. Then, cells were washed twice with PBS. To acquire binucleated cells, cells were incubated with a medium containing cytochalasin B (4 μg/mL) for 28 h. After incubating, cells were harvested, treated with cold hypotonic solution (KCl, 0.075 M), and fixed with Carnoy fixative (3 methanol:1 acetic acid). The cells were spread to coded slides and stained with a 5% aqueous Giemsa solution for 15 min. Slides were scored at 40X magnification using a light microscope to measure micronucleus formation.
Statistical analysis
Statistical analyses were performed using SPSS 11.5. After assessing the normality of the distribution of the data, all data are expressed as mean ± standard deviation. The mean values among different doses were compared using the Tukey test and Paired t test (p < 0.05).
Results
Measurement of cytotoxicity as cell index
Cell index values on HepG2 cells after treatment with 1–80 μg/mL of the indaziflam for 4–72 h.

Data are shown as mean ± SD.
* Statistical difference p ˂ 0.05.
NC: negative control; SC: solvent control (ethanol).
Comet assay
Determination of DNA damage by comet assay on the HepG2 cells in the presence of indaziflam, ethanol (SC), hydrogen peroxide (PC).

NC: negative control; SC: solvent control (ethanol); PC: positive control (H2O2); DCP: damage cell percent; GDI: genetic damage index.
*Statistical difference p ˂ 0.05.
Genetic damage index (GDI) = (Type I + 2 Type II + 3 Type III +4 Type IV)/(Type 0 + I + II + III + IV). 21
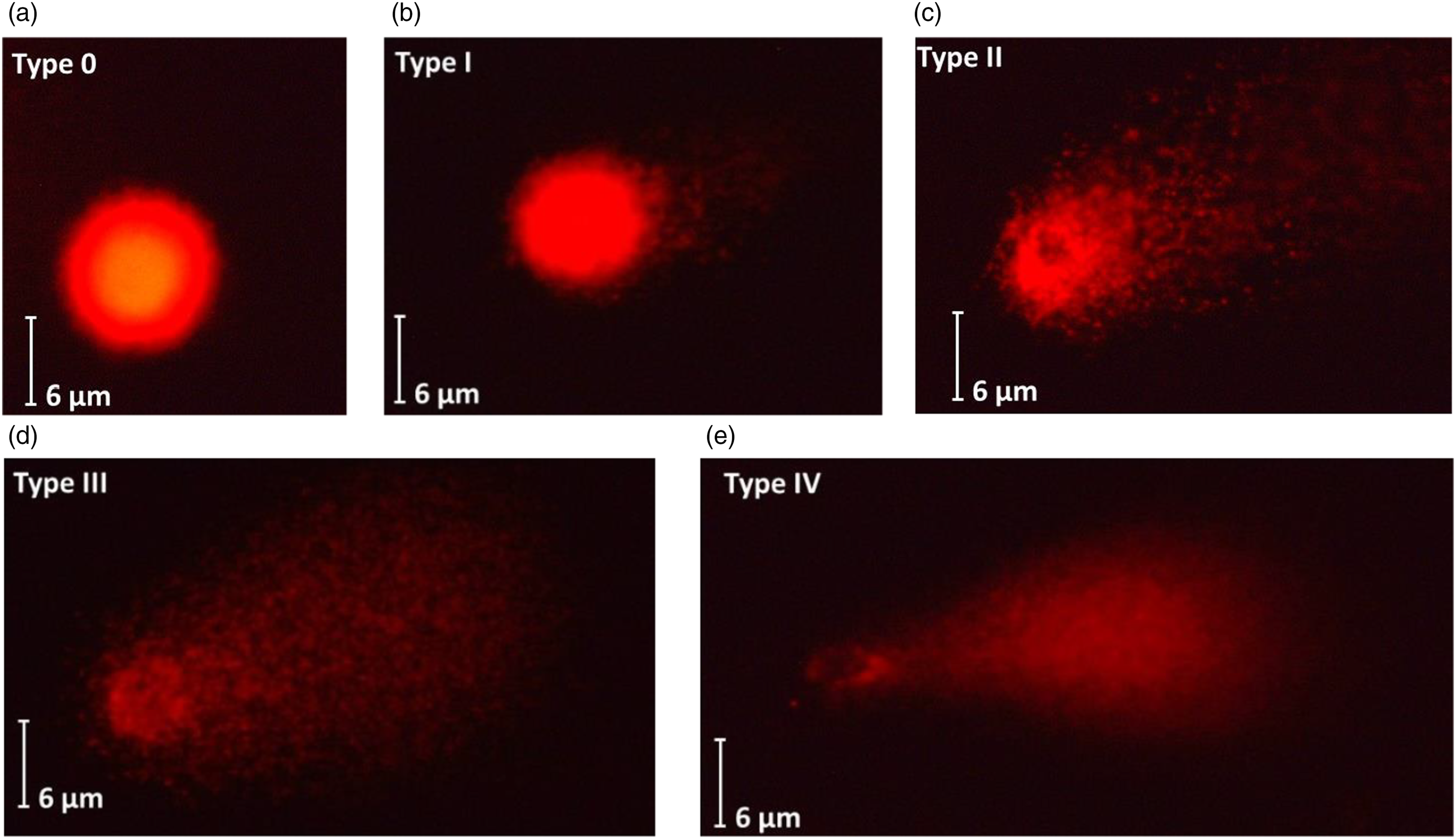
Levels of DNA damage on HepG2 cells after exposure to indaziflam evaluated by comet assay. Undamaged cell: Type 0
Cytokinesis-block micronucleus assay
Frequency of micronuclei formation on HepG2 cells exposed to the indaziflam for 4 and 24 h.

*Statistical difference p ˂ 0.05.
NC: negative control; SC: solvent control (ethanol); PC: positive control (cyclophosphamide).
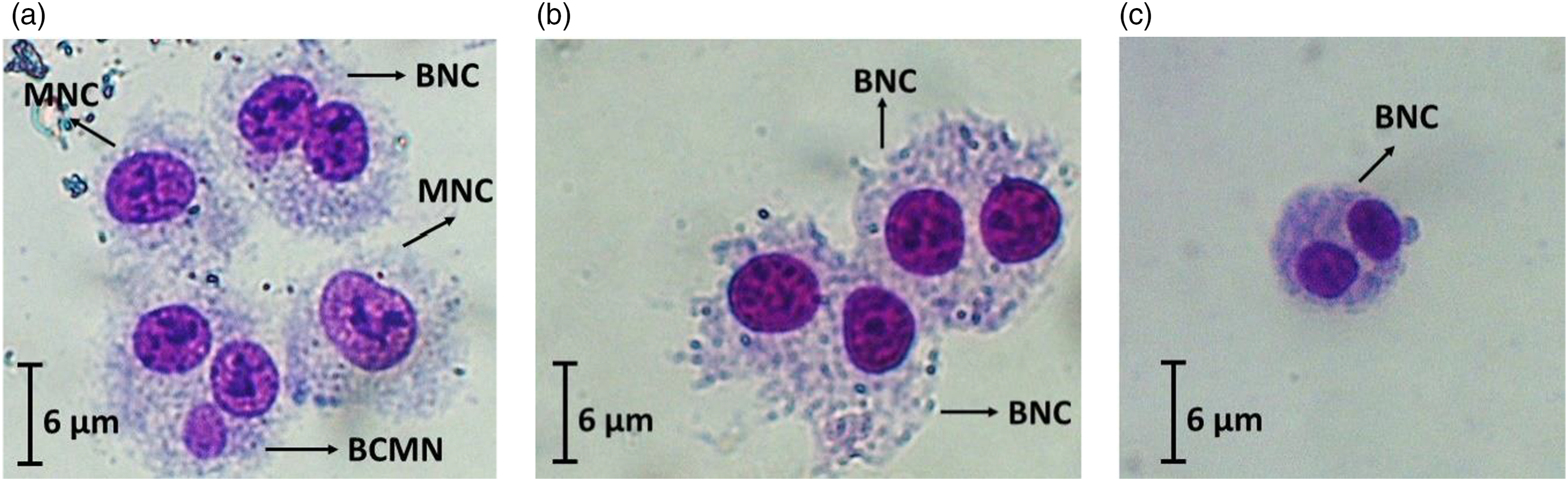
HepG2 cells exposed to the indaziflam (a), HepG2 cells in solvent control group (b), and HepG2 cells in negative control group (c). BNC, MNC, and BCMN means binucleated cell, mononucleated cell and binucleated cell including micronucleus, respectively.
Discussion
Pesticides are chemicals widely used in agriculture to protect crops. 23 Due to the intensive pesticide application in modern agriculture, pesticide exposure is constantly increasing worldwide. Moreover, many pesticides can cause adverse effects on non-targeted organisms, including animals and humans. 24 Therefore, environmental, or occupational exposure to pesticides has been related to health risks, including genetic damage in cells and cancerogenesis in various organs such as the lung, brain, prostate, testis, cervix, bladder, kidney, and rectum. 25 For this reason, investigating adverse effects caused by pesticides is a growing need nowadays.
The utilization of only one genotoxicity assay does not give enough information to determine the reliability of pesticides. Therefore, different biomarkers are used to evaluate the genotoxicity of pesticides on organisms. 26 Previous researchers have preferred micronucleus assay, sister chromatids exchange, chromosomal aberration, comet assay, and gamma H2AX assay to assess genotoxicity in vitro and in vivo.27–29 Micronucleus and comet assays are commonly used to evaluate pesticide genotoxicity on different models.30–33 Moreover, although tested pesticide has shown negative results on a selected model organism, it can lead to positive results on other organisms. Consequently, different test systems and model organisms should be selected to observe the genotoxicity of pesticides.
Indaziflam is a herbicide belonging to the alkylazine chemical group and inhibits cellulose-biosynthesis in plants. Also, it has broad-spectrum activity at low application rates between 25 and 150 g of active ingredient (ai) per hectare in agricultural and nonagricultural areas. 6 Indaziflam is commonly used on both monocotyledons and dicotyledons. The half-life of indaziflam in soil has longer than 150 days. 34 Therefore, it is essential to evaluate the effect of indaziflam on living things.
There is no information about the cytotoxicity and genotoxicity of indaziflam on HepG2 cells. Previous studies related to indaziflam focused on its metabolites, its effect on the diversity of microorganisms in the soil, and its effects on target organisms. Tissot et al. (2022) have examined the toxic effects of indaziflam individually and in combination on aquatic bivalves (Mya arenaria) as filter-feeding organisms. 35 They have quantified indaziflam concentration in the clam tissue and water. They also showed that the indaziflam treatment caused strong mortality. Scully-Engelmeyer et al. (2021) have measured 12 pesticides including herbicide, fungicide, and insecticide, in bivalve samples. 36 They reported that they could detect indaziflam even in the forest. 36 In another study, Sun et al. (2018) reported that indaziflam caused root mass changes in hybrid Bermuda grass and decreased accumulation of macro/micro-nutrients in roots. 37 As a result, the present study focused on filling a gap related to cytotoxicity and genotoxicity of indaziflam on HepG2 cells. In this study, our results showed that indaziflam is non-cytotoxic on HepG2 cells. In contrast, increasing concentrations of indaziflam lead to a genotoxic effect (DNA strand breaks and micronucleus frequency) on HepG2 cells. Moreover, pesticides are usually transformed into two or more metabolites after use in crops and soils. These metabolites might have more hazardous than the parent pesticides. 38 A previous study showed that indaziflam has four metabolites, including triazine-indanone, 1-fluoroethyl triazinediamine, indaziflam-olefin, indaziflam-carboxylic acid on fruits and vegetables. 38 Therefore, the genotoxicity observed at increasing doses in our study may be due to the metabolites of indaziflam.
Conclusion
Indaziflam is one of the most widely used herbicides in agriculture for protecting crops from unwanted plants. This is the first study involving the cytotoxic and genotoxic effects of indaziflam on the HepG2 cells. Tested doses of indaziflam did not cause a cytotoxic effect on HepG2 cells. However, it has been observed that increasing concentrations of indaziflam lead to a genotoxic effect on HepG2 cells. The toxicity mechanism of indaziflam can be clarified with additional studies involving various pathway mechanisms. Moreover, this study showed that HepG2 cells are a proper model for studying the cytotoxic and genotoxic effects of pesticides.
Footnotes
Author contributions
The corresponding author, Serpil Könen Adıgüzel, designed the study. Material preparation, data collection, analysis, and the draft of the manuscript were performed by Serpil Könen Adıgüzel.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Research Fund of Mersin University in Turkey with Project Number: 2018-2-AP3-2969.
