Abstract
Background
Long intergenic non-protein-coding RNA 173 (LINC00173) executes vital functions in various cancers. Nevertheless, its role and expression in nasopharyngeal carcinoma (NPC) have yet to be investigated. Here, we investigated its effects on the malignancy characteristics of NPC and elucidated the potential molecular mechanism of LINC00173 in NPC progression.
Methods
Quantitative real-time reverse transcription-PCR (qRT-PCR) and immunoblotting were conducted to estimate the LINC00173, microRNA-765 (miR-765), and Gremlin 1 (GREM1) expressions in NPC cells and tissues. Cell counting kit-8 (CCK8), colony formation, and wound healing experiments were done to evaluate the proliferation, growth, and migration of NPC cells, respectively. The tumorous growth of NPC cells in vivo was assessed through the xenograft tumor experiment. Furthermore, the interactions among miR-765, LINC00173, and GREM1 were investigated through bioinformatics analyses, luciferase reporter and RNA immunoprecipitation chip assays.
Results
An upregulated LINC00173 expression was found in NPC cell lines and tissues. The functional experiments uncovered that its downregulation repressed NPC cell proliferation, growth, and migration. In addition, LINC00173 knockdown hampered the NPC cells’ tumorous growth in vivo. These effects could partially be reversed by downregulating miR-765. GREM1 is a downstream target of miR-765. GREM1 knockdown could repress the proliferation, growth, and migration of NPC cells. Nonetheless, these anti-tumor effects could be abolished by miR-765 downregulation. Mechanistically, LINC00173 increased the expression of GREM1 by binding with miR-765.
Conclusions
LINC00173 functions as an oncogenic factor by binding with miR-765 to promote the progression of NPC via GREM1 upregulation. This study provides a novel insight into the molecular mechanisms involved in NPC progression.
Keywords
Introduction
Nasopharyngeal carcinoma (NPC) is a type of cancer that arises from the nasopharyngeal epithelium with fossa of Rosenmüller being the tumor epicenter in most cases.1,2 NPC has been reported to be mostly prevalent in East and Southeast Asia, thus showing a unique pattern of racial and geographical distribution. The rate of occurrence of NPC varies widely—from 30 in 100,000 individuals in endemic regions to <1 in 100,000 individuals in non-endemic areas with mainly white population. 3 The risk factors associated with its pathogenesis include genetics, viral infections such as Epstein–Barr virus and human papillomavirus, and other environmental factors.4–8 Over the last two decades, the outcomes for NPC have certainly gotten better with the improvements in radiotherapy techniques and combination therapy strategies, including radiotherapy and chemotherapy. 9 However, control of advanced stages of NPC remains unsatisfactory. 9 In addition, toxicity and recurrence remain major challenges after a successful anti-cancer treatment. 3 Hence, it is necessary to explicate the mechanisms underlying NPC progression and oncogenesis, which will eventually help in identifying a novel therapeutic approach.
The development of high-throughput technologies over the past decade has led to a rigorous examination of the noncoding genome, majority of which are transcribed under some conditions. 10 Among the various types of non-protein coding RNAs, long noncoding RNAs (lncRNAs) have gained the attention of several researchers. They are RNAs that have over 200 nucleotides and no protein-coding capability. 10 Several reports have shown that lncRNAs induce various hallmarks of cancer cells by interacting with other biomolecules such as proteins, RNA, and DNA. 11 Many lncRNAs have been found to be dysregulated in association with NPC progression. For instance, lncRNA LOC401317 was found be upregulated by p53 and exerted anti-tumor effects in NPC cells. 12 The lncRNAs AFAP1-AS1, XIST, and MALAT1 were also upregulated in NPC cell lines and tissues.13–15 Moreover, they functioned as oncogenes in NPC progression by targeting different miRNAs.13–15 On the contrary, the expressions of the two lncRNAs, LINC00312 16 and LET, 17 were downregulated and exhibited anti-tumor effects in NPC. The previous studies indicate that different lncRNAs play different roles in NPC. Long intergenic nonprotein coding RNA 173 (LINC00173) is situated at chromosome 12q24.22, and it was first functionally identified in non-small-cell lung cancer (NSCLC) in 2019, in which its poor expression was detected in the cancer tissues. The LINC00173 deficiency led to the accumulation of miR-182 resulting in accelerated NSCLC cell migration and proliferation while inhibiting apoptosis via the Advanced Glycosylation End-Product Specific Receptor (AGER)/NF-κB axis. 18 Nonetheless, it remains unknown whether LINC00173 is dysregulated in NPC and plays any role in its progression. In addition, the mechanism by which it regulates tumor function is still obscure.
In the present study, we investigated how LINC00173 affected NPC’s malignant characteristics in vivo and in vitro. Next, we investigated the potential molecular mechanism of LINC00173 that may have an involvement in the mediation of NPC progression. We determined that LINC00173 served as an oncogenic factor by binding with microRNA (miR-765) to boost NPC progression through the upregulation of Gremlin 1 (GREM1). These outcomes contribute to a novel understanding of the molecular mechanism involved in NPC progression. This study has been the first to provide clinical evidence of the upregulation of LINC00173 in NPC cells and tissues. In view of the findings above, LINC00173 may possibly be adopted as a novel diagnostic and prognostic biomarker for NPC patients.
Materials and methods
Clinical specimens
The Fifth Hospital of Wuhan granted approval to this study. The Ethics Committee of the hospital approved all the experiments. A total of 38 pairs of human NPC and adjacent healthy tissues were acquired from the Fifth Hospital of Wuhan. Patients that had other types of malignancies were excluded from this study as well as those who underwent chemotherapy, radiotherapy, or any other therapies. The tissue samples were promptly flash-frozen with liquid nitrogen once excised. They were then kept at −80°C for future usage. All patients submitted their informed/written consents.
Cell culture
Human NPC cell lines C666-1 and NPC/HK1 and the human nasopharyngeal epithelial cell line NP69 (control) were all bought from the American Type Culture Collection (ATCC, USA). The cell lines were cultivated in Roswell Park Memorial Institute (RPMI) 1640 medium (Thermo Fisher Scientifc, USA) that also had 10% fetal bovine serum (Thermo Fisher Scientifc, USA). The cultures were kept at 37°C in an incubator that was filled with 5% CO2. The cells were passaged when they reached a confluency of 70–90%.
Cell transfection
Cells were transiently transfected with: (i) validated small-interfering RNA (siRNAs, GenePharma Technology) for LINC00173 (si-lnc) and GREM1 (si-GREM1), with a scrambled siRNA (si-NC) oligonucleotide as the corresponding negative control; and (ii) miR-765 mimic (a type of synthetic RNA molecule to overexpress miR-765, 5′-UGGAGGAGAAGGAAGGUGAUG-3′), miR-765 inhibitor (a type of synthetic RNA molecule to inhibit miR-765, 5′-CAUCACCUUCCUUCUCCUCCA-3′), and their negative controls (GenePharma Technology). Lipofectamine™ 2000 (Invitrogen, USA) was utilized, as described in its protocol, for the transfection of the 50 nM oligonucleotides into NPC cells. Finally, transfection efficiency was ascertained via quantitative real-time reverse transcription-PCR (qRT-PCR) at 24 h post-transfection.
RNA isolation and quantitative real-time reverse transcription-PCR
Real-time PCR primer synthesis list.

Cell counting kit-8 assay
The transfected C666-1 and NPC/HK1 cells were inoculated on 96-well plates (2000 cells/well) and maintained at 37°C in an incubator that contained 5% CO2. At specific periods following the inoculation (0, 24, 48, and 72 h), the cells in each well were supplied with 10 μL of CCK-8 solution (Sigma, USA) and then left to incubate for two more hours. Finally, each well was analyzed with a microplate reader (Applied Biosystems, USA) to determine their absorbance at 450 nm.
Colony formation assay
Two hundred transfected NPC/HK1 and C666-1 cells were plated on each of the wells of 6-well plates and then incubated with 5% CO2 for 2 weeks at 37°C. After incubation, the cells were rinsed three times with phosphate-buffered saline. Thereafter, the cells were fixed with 4% paraformaldehyde and then stained with 0.5% crystal violet. Both steps were conducted for 30 min each at room temperature. Finally, the colonies were photographed with the light microscope (Olympus, Japan). Colony formation was assessed by calculating the number of colonies per well.
Wound healing assay
The transfected NPC cells were inoculated on 6-well plates (5 × 105 cells/well). The cells were then cultivated at 37°C with 5% CO2 for 36 h to allow the cells to spread completely and achieve >95% monolayer confluency. Gentle cross scratching was done in each well using a sterile 100 μL pipette tip. The wound was rinsed to remove non-adherent cells, and then it was photographed under an Olympus light microscope. The cells were subsequently cultured in a serum-free medium and were photographed again after 24 h. The migration of cells was evaluated using the following formula: cell migration rate = (wound width at 0 h-wound width at 24 h)/wound width at 0 h × 100%.
Tumor xenograft experiments
The Ethics Committee of our hospital granted approval to all procedures implemented in this animal study. The xenograft tumor models were established using 3-4-week-old male BALB/c nude mice bought from Beijing HFK Bioscience (Beijing, China). The mice were all kept in pathogen free condition at 23 ± 3°C under a relative humidity of 55 ± 15% and with a 14 h/10 h light/dark cycle. A LINC00173-specific short hairpin RNA (sh-LINC00173) as well as its corresponding negative control (GenePharma Technology) were delivered into lentiviral pLKO vectors to produce pLKO-sh-LINC00173 and pLKO-sh-NC. A total of 5 mice each were distributed into sh-control (sh-NC) and sh-LINC00173 (sh-lnc) groups. Following the stable transfections with sh-LINC0031 or sh-NC vectors, 4 × 106 C666-1 cells were subcutaneously injected into the right flank of each mouse. The growth of the tumors was plotted by measuring the size of the xenograft tumor. After 5 weeks, the mice were euthanized before their tumors were excised for weighing.
Bioinformatics analyses and dual luciferase assay
MiRDB (http://www.mirdb.org/) was utilized in predicting the LINC00173–miRNA interactions while TargetScan (https://www.targetscan.org/vert_80/) was for the prediction of miRNA-GREM1 interactions. LINC00173 and GREM1 wild-type (WT) and mutant (Mut) sequences that harbor miR-765 complementary or mutated sequences, respectively, were cloned into pmirGLO vectors (Promega, USA). Each vector was co-transfected into C666-1 and NPC/HK1 cells with a miR-765 mimic/NC. Following the 24-h transfection, the luciferase activities in the cells were tested with the help of a Dual-Luciferase® Reporter (DLR™) Assay Kit (Promega, USA).
RNA immunoprecipitation assay
The RIP experiment was performed by means of a Magna RIP Kit (Millipore, USA) to test the interaction of LINC00173 with miR-765 in NPC cells. The cell extract was obtained from treating the cells with the RIP lysis buffer. It was then incubated at 4°C overnight with magnetic beads with attached anti-Argonaute 2 (Ago2) antibody or control IgG (both from Millipore, USA). On the following day, the magnetic beads were gathered, rinsed, then treated with proteinase-K. The immunoprecipitated RNA was subsequently isolated and then subjected to RT-qPCR for the estimation of the miR-765 and LINC00173 enrichments on the Ago2 beads.
Western blotting
RIPA lysis buffer (Thermo Fisher Scientifc, USA) was utilized to dissolve NPC cells to extract their total proteins. The BCA Protein Assay Kit (Thermo Fisher Scientifc, USA) was then used for the determination of the protein concentration. Protein (40 μg) was separated via 12% sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) before being moved onto a polyvinylidene difluoride (PVDF) membrane (Thermo Fisher Scientifc, USA). The membrane was subsequently blocked with 5% fat-free milk for 2 h at room temperature. It was then maintained at 4°C overnight with primary antibodies targeting GREM1 (Abcam, USA: ab231065; 1:1000 dilution) and GAPDH (Abcam, USA: ab9485; 1:2000 dilution). On the next day, a horseradish peroxidase (HRP)–conjugated secondary antibody (ab205718; 1:5000; Abcam, USA) was utilized to probe the membrane for 2 h at room temperature. Finally, an enhanced chemiluminescence kit (Beyotime) was employed to visualize the protein signals. GAPDH was adopted as the loading control.
Statistical analysis
Each assay was independently carried out thrice, and the results were expressed as the mean ± standard deviation (SD) from the three independent experiments. GraphPad Prism 7.0 (GraphPad, USA) was utilized in the processing of data. Student’s t test was utilized in comparing data between groups. Meanwhile, One-way or two-way analysis of variance (ANOVA) together with Tukey’s or Bonferroni’s post hoc test was applied to compare data from more than two groups. The correlation between the expressions of two groups was tested through Pearson’s correlation analysis. A p-value smaller than 0.05 was deemed to be statistically significant.
Results
NPC cell lines and tissues exhibit LINC00173 upregulation
We first analyzed lncRNA LINC00173 expression, via qRT-PCR, in an expanded sample set that consisted of 38 NPC and 38 normal nasopharyngeal tissues. The outcomes demonstrated that LINC00173 levels were significantly higher in NPC than in normal tissue samples (p < .0001, Figure 1(a)). A similar pattern was observed in the NPC cell lines as they exhibited considerably higher LINC00173 levels than the NP69 (control) cells (p < .001, Figure 1(b)). Furthermore, we also observed that LINC00173 was mostly found within the C666-1 and NPC/HK1 cytoplasm, thereby implying its possible involvement in NPC progression through a competing endogenous RNA (ceRNA) network (Figure 1(c)). Collectively, these results are indicative of an important role executed by LINC00173 in NPC progression. Relative expression of LINC00173 in NPC cell lines and tissues. (a) LINC00173 expression was detected via qRT-PCR in NPC tissues and matched normal tissues (n = 38). Tumor, NPC tissues. Normal, matched normal tissues. (b) qRT-PCR analysis of LINC00173 expression in nasopharyngeal epithelial cells (NP69) and NPC cell lines (NPC/HK1 and C666-1). (c) Subcellular analysis of LINC00173 in NPC cells. U6 and GAPDH were utilized as loading controls. Bar graphs: The data show the mean ± SD from three independent experiments. **p < .001 vs. NP69.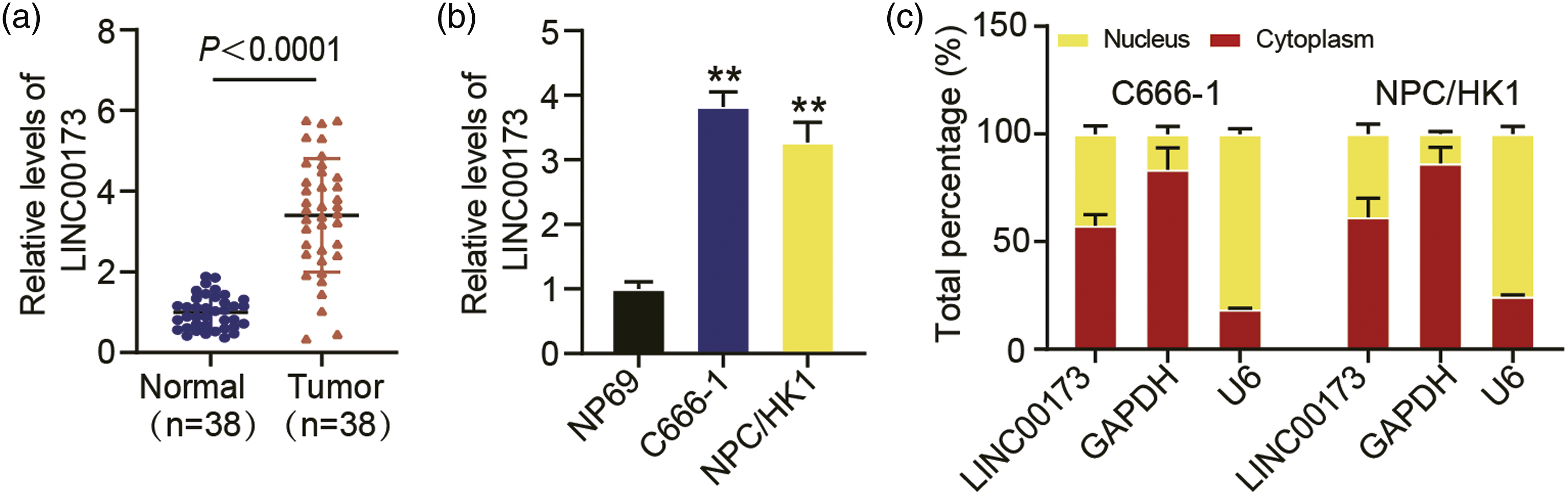
LINC00173 knockdown suppresses the proliferation, growth, and migration of NPC cells
To identify LINC00173’s exact functional role in the progression of NPC, it was knocked down by introducing a siRNA into the C666-1 and NPC/HK1 cell lines, which had relatively high relatively high LINC00173 expression. The outcomes of the qRT-PCR indicated that LINC00173 expression remarkably declined in the NPC cell lines, following the introduction of the siRNA (p < .001, Figure 2(a)). The CCK-8 assay revealed that the proliferative abilities of the C666-1 and NPC/HK1 cells considerably decreased upon the knockdown of LINC00173 (p < .05 at 48 h and p < .001 at 72 h, Figure 2(b)). The NPC cell lines transfected with si-lnc manifested a significant reduction in colony numbers relative to the controls (p < .001, Figure 2(c)). The wound healing experiment further demonstrated that LINC00173 silencing impaired the migrative ability of NPC cells (p < .001, Figure 2(d)). Taken together, the abovementioned results show that LINC00173 is possibly an oncogene in NPC and stimulates the proliferation, migration, and growth of NPC cells. LINC00173 knockdown restricts the proliferation, growth, and migration of NPC/HK1 and C666-1 cells. (a) LINC00173 expression was gauged in NPC/HK1 and C666-1 cells via qRT-PCR at 24 h post-transfection with si-NC and si-lnc. (b) Cell proliferation was evaluated by CCK-8 assay in si-NC and si-lnc treated C666-1 and NPC/HK1 cells. (c) Colony formation experiments were done to assess the growth of si-NC- and si-lnc-transfected C666-1 and NPC/HK1 cells. (d) Wound healing assays were conducted to investigate the effect of si-NC and si-lnc treatment on cell migration. Bar graphs: The data show the mean ± SD from three independent experiments. *p < .05 and **p < .001 vs. si-NC. si-NC, the negative control of small interfering RNA. si-lnc, small interfering RNA targeting LINC00173.
LINC00173 knockdown suppresses NPC growth in vivo
To figure out how LINC00173 affects tumor growth in vivo, we subcutaneously transplanted C666-1 cells with diminished LINC00173 expression into nude mice. The xenograft tumors that grew from the LINC00173-knocked-down C666-1 cells had smaller sizes, mean volumes, and weights than the tumors that developed from the control cells (p < .001, Figure 3). These results suggest that the depletion of LINC00173 hampers NPC cell tumorigenesis in vivo. LINC00173 knockdown suppresses the growth of NPC cells in vivo. Sh-NC or sh-lnc was stably transfected into C666-1 cells. Then, each group of cells was injected into mice (n = 5 mice for each group) before they were monitored for 5 weeks. Thereafter, each mouse was euthanatized, and their tumors were excised. (a) A representative image of the excised tumors from both groups (sh-NC and sh-lnc) is shown. (b) The plot of tumor growth was established by measuring the volume of the xenograft tumors every week. (c) The weight of the tumors as determined after 5 weeks. Bar graphs: The data indicate the mean ± SD (n = 5 biological replicates). *p < .05 and **p < .001 vs. sh-NC. sh-NC, the negative control of short hairpin RNA. sh-lnc, short hairpin RNA targeting LINC00173.
miR-765 is a direct target of LINC00173
We explored the miRDB (http://www.mirdb.org/) database and identified miR-765 as the most potential target miRNA of LINC00173 as it scored the highest and showed two binding sites (Figure 4(a)). Furthermore, we performed a dual luciferase experiment to substantiate the direct binding of miR-765 with LINC00173 in NPC cells. The miR-765 upregulation, mediated by the miR-765 mimic, effectively diminished the luciferase activity of the LINC00173-WT plasmid in the C666-1 and NPC/HK1 cells (p < .001), but did not have notable effects on that of the LINC00173-Co-Mut plasmid since all binding sites on LINC00173 were mutated (Figure 4(b)). The RIP assay further demonstrated that both miR-765 and LINC00173 were highly enriched on the magnetic beads with anti-AGO2 antibody (p < .001, Figure 4(c)). The experimental findings substantiate the interaction of miR-765 with LINC00173 in NPC cells. We also observed a notably poor miR-765 expression in NPC tissues (p < .0001) as well in cell lines (p < .001), relative to their respective controls (Figure 4(d) and (e)). Moreover, the miR-765 and LINC00173 expressions in NPC tissue samples were revealed to have an inverse relationship with each other (Figure 4(f)). These findings suggest that LINC00173 possibly acts as a ceRNA in NPC cells via miR-765 sponging. LINC00173 acts as a ceRNA to sponge miR-765. (a) miRDB database analysis predicted the miR-765 and LINC00173 binding sites. Mut1, LINC00173 with binding site 1 mutation. Mut2, LINC00173 with binding site 2 mutation. (b) The luciferase reporter plasmid harboring the wild-type or mutant LINC00173 was co-transfected into NPC cells with either a miR-765 mimic/NC; the association between LINC00173 and miR-765 was validated through the luciferase reporter experiment. *p < .05 and **p < .001 vs. WT + NC. WT, wild-type LINC00173. Mut1, LINC00173 with binding site 1 mutation. Mut2, LINC00173 with binding site 2 mutation. Co-Mut, Mut1 + Mut2. NC, the negative control of mimic. mimic, miR-765 mimic. (c) RIP assay was accomplished using anti-Ago2 in transfected NPC cells (either with LINC00173 or miR-765 mimics), with anti-IgG as the negative control. The miR-765 and LINC00173 enrichments in NPC/HK1 and C666-1 cells were evaluated through qRT-PCR. **p < .001 vs. Anti-IgG. (d) MiR-765 expression in NPC tissues (n = 38) as well as in paired normal tissues was detected via qRT-PCR. Tumor, NPC tissues. Normal, paired normal tissues. (e) MiR-765 expression in NPC cell lines (NPC/HK1 and C666-1) and their respective control was estimated through qRT-PCR. **p < .001 vs. NP69. (f) The correlation of LINC00173 expression in tumor tissues with that of miR-765 as ascertained by Pearson correlation coefficient (R2 = 0.6519, p < .0001). Bar graphs: Data indicate mean ± SD from three independent experiments.
miR-765 inhibitor reversed the functional effects of LINC00173 knockdown on NPC cells
We performed rescue assays in order to determine whether LINC00173 was involved in NPC by regulating miR-765. Si-lnc, miR-765 inhibitor, and si-lnc + miR-765 inhibitor were individually transfected into the NPC cells. MiR-765 expression significantly rose upon the solo transfection of si-lnc but declined in those with miR-765 inhibitor alone (p < .001). Furthermore, the additional transfection of si-lnc + miR-765 inhibitor partially restored miR-765 expression in the NPC cells (Figure 5(a)). The CCK-8 assay revealed a reduced cell proliferation resulting from the si-lnc-transfection-mediated enhancement of miR-765 (p < .001). Meanwhile, a boost in cell proliferation was observed upon the solo transfection of the miR-765 inhibitor (p < .001). Interestingly, the proliferative ability of the NPC cells was partially restored by the combined transfection of si-lnc + miR-765 inhibitor (Figure 5(b)). Similarly, the colony formation and wound healing assays demonstrated partial restoration of colony numbers and migratory capacity of NPC cell lines upon the combined transfection of si-lnc + miR-765 inhibitor (p < .001, Figure 5(c) and (d)). These results show that downregulating miR-765 partially reversed the functional effects of LINC00173 knockdown. Thus, it may be implied that LINC00173 modulates NPC characteristics by sponging miR-765. miR-765 inhibitor reversed the effects of LINC00173 on NPC cells. Rescue experiments were conducted on NPC/HK1 and C666-1 cell lines transfected with si-NC, inhibitor-NC, si-lnc, miR-765 inhibitor, and si-lnc + miR-765 inhibitor (a) Relative levels of miR-765 expression were evaluated through qRT-PCR. (b) Cell proliferation was assessed via the CCK-8 experiment. (c) Colony formation assays were carried outs to assess the growth of cells. (d) Wound healing experiments were done to scrutinize how the transfections affected the migration of cells. Bar graphs: The data indicate the mean ± SD from three independent experiments. *p < .05, **p < .001 vs. si-NC; #p < .05, ##p < .001 vs. inhibitor-NC; &p < .05, &&p < .001 vs. to si-lnc + inhibitor. si-NC, the negative control of small interfering RNA. si-lnc, small interfering RNA targeting LINC00173. inhibitor-NC, the negative control of inhibitor. inhibitor, miR-765 inhibitor.
GREM1 is a miR-765 target gene
The bioinformatics analysis with TargetScan revealed GREM1 to have the most potential as miR-765 target as it has three potential binding sites, each of them showing a percentile score higher than 97% (Figure 6(a)). To substantiate this, we constructed luciferase reporters for the dual luciferase assay. The outcomes of the assay revealed that the luciferase activity of GREM1-WT declined after being treated with the miR-765 mimic (p < .001), and the luciferase activity of GREM1-Co-Mut did not reduced since all binding sites on GREM1 3′UTR were mutated (Figure 6(b)). This suggests the interplay of miR-765 with GREM1 in both the NPC cell lines. The cohort of clinical tissues as well as the C666-1 and NPC/HK1 cells manifested higher GREM1 expressions than their respective controls (p < .0001 in tissues and p < .001 in cells, Figure 6(c) and (d)). Moreover, miR-765 expression was revealed to have an inverse association with GREM1 expression in NPC tissues (Figure 6(e)). These findings indicate that GREM1was a target effector for miR-765. MiR-765 directly binds to GREM1. (a) Targetscan predicted three miR-765 and GREM1 3′UTR binding sites. Mut1, GREM1 3′UTR with binding site 1 mutation. Mut2, GREM1 3′UTR with binding site 2 mutation. Mut3, GREM1 3′UTR with binding site 3 mutation. (b) The luciferase reporter plasmid with wild-type or mutant GREM1 was co-transfected into NPC cells along with miR-765 mimics/NC. The association between GREM1 and miR-765 was then validated through the luciferase reporter experiment. *p < .05 and **p < .001 vs. WT + NC. WT, wild-type GREM1 3′UTR. Mut1, GREM1 3′UTR with binding site 1 mutation. Mut2, GREM1 3′UTR with binding site 2 mutation. Mut3, GREM1 3′UTR with binding site 3 mutation. Co-Mut, Mut1+Mut2+Mut3. NC, the negative control of mimic. mimic, miR-765 mimic. (c) GREM1 levels in NPC tissues (n = 38) and paired normal tissues were detected through qRT-PCR. Tumor, NPC tissues. Normal, paired normal tissues. (d) The GREM1 levels in NPC cell lines (NPC/HK1 and C666-1) and their respective control were detected through qRT-PCR. **p < .001 vs. NP69. (e) The correlation between the miR-765 and GREM1 expressions in tumor tissues was ascertained through Pearson correlation analysis (R2 = 0.6224, p < .0001). Bar graphs: Data indicate mean ± SD from three independent experiments.
MiR-765 downregulation reversed the anti-tumor function of GREM1 knockdown in NPC cells
We investigated the miR-765-mediated molecular pathway in the progression of NPC by performing rescue experiments to define the interplay between miR-765 and GREM1. Immunoblotting revealed that, in both NPC cells, the solo transfection of the miR-765 inhibitor resulted in higher GREM1 levels than the inhibitor-NC (p < .001). This upregulation of GREM1 was partially abolished when si-GREM1 was added along with the miR-765 inhibitor during transfection (Figure 7(a)). Furthermore, the CCK-8 assays uncovered that the proliferative capacity of the cells substantially improved upon the inhibition of miR-765 but got impaired by the silencing of GREM1 expression (p < .001). Moreover, reintroducing miR-765 inhibitor partly restored the inhibitory influence of si-GREM1 transfection on the proliferative capacity of the cells (Figure 7(b)). As expected, the number of colonies decreased after knocking down GREM1, increased upon inhibiting miR-765, and was partially alleviated when miR-765 was downregulated while GREM1 was knocked down (p < .001, Figure 7(c)). Furthermore, the wound healing assays demonstrated that the enhanced migration of NPC cell lines, which was triggered by miR-765 repression, was abolished by downregulating GREM1 (p < .001). This combined transfection of si-GREM1 + miR-765 inhibitor resulted in the partial restoration of the migratory capacity of the NPC cell lines (Figure 7(d)). These findings indicate that GREM1 knockdown impaired the ability of NPC cells to proliferate, grow, and migrate. Nevertheless, these effects may be reversed when NPC were transfected with miR-765 inhibitor. MiR-765 downregulation reversed the effects of GREM1 knockdown on NPC cells. Rescue experiments were conducted on NPC/HK1 and C666-1 cell lines transfected with si-NC, inhibitor-NC, si-GREM1, miR-765 inhibitor, and si-GREM1 + miR-765 inhibitor. (a) Western blotting was accomplished to detect the GREM1 protein bands. GAPDH was adopted as the internal control for the immunoblot experiments. (b) Cell proliferation was evaluated via the CCK-8 experiment. (c) Colony formation experiments were done to assess the growth of the cells. (d) Wound healing experiments were conducted to investigate how the transfections affected the migration of the cells. Bar graphs: The data indicate the mean ± SD from three independent experiments. *p < .05, **p < .001 vs. si-NC; #p < .05, ##p < .001 vs. inhibitor-NC; &p < .05, &&p < .001 vs. to si-G + inhibitor. si-NC, the negative control of small interfering RNA. si-GREM1 or si-G, small interfering RNA targeting GREM1. inhibitor-NC, the negative control of inhibitor. inhibitor, miR-765 inhibitor.
Discussion
In the past decade, the roles and functions of several lncRNAs in NPC have been studied. 20 In the present research, we first assessed the LINC00173 levels in NPC cell lines and tissues and found LINC00173 upregulation in both. Silencing LINC00173 remarkably suppressed NPC cell proliferation, growth, and migration. Moreover, knocking down LINC00173 impeded the NPC’s tumorous growth in vivo. By means of in-silico approaches, we identified miR-765 as a potential LINC00173 target and also found GREM1 to be a potential miR-765 target. These were corroborated by the results of the luciferase reporter assay. We observed a significant downregulation of miR-765 as well as an upregulation of GREM1 in NPC cells and tissues, relative to their respective controls. Our findings indicate that GREM1 may be crucial mediator for miR-765-induced effects on NPC cell proliferation, growth, and migration.
Recent reports and ongoing studies have found that lncRNAs are novel regulators of tumorigenesis and tumor progression.21–23 Contemporary accumulated evidence confirm that LINC00173 is an oncogenic lncRNA. LINC00173 has been found to be upregulated in small cell lung cancer, where it was also closely linked to chemo resistance and more advanced stages among the cancer patients. 24 Similarly, in prostate cancer, the experimental data indicated that LINC00173 may serve as an oncogene and exert its biological effects by modulating the miR-338-3p/Rab25 axis. 25 Triple-negative breast cancer tissues manifested an elevated LINC00173 expression that was also linked to advanced TNM stages and lymph node metastasis. 26 In addition, patients with high expression of LINC00173 in their cancer tissues had shorter overall survival. 26 LINC00173 expression was markedly elevated in glioma and colorectal cancer cells and tissues. An elevated LINC00173 expression has been associated with poor prognosis in those two cancers.27,28 Interestingly, in both of those studies, LINC00173 interacted with miR-765 to enhance distinct target genes. In our study, we also found miR-765 to be the potential target of LINC00173. Our data also confirms that LINC00173 is an oncogenic lncRNA in NPC. The subcellular localization of lncRNAs plays a vital role in the determination of lncRNAs’ functions. The ceRNA hypothesis suggests that, when present mainly in the cytoplasm, lncRNAs act as molecular sponges by binding to target miRNAs, thereby consequently regulating the expression of related genes that have shared miRNAs. 29 In our study, LINC00173 was highly expressed in NPC and was found to be localized predominately within the NPC cell cytoplasm. That suggests that LINC00173 may operate as a ceRNA.
Numerous studies suggest that miR-765 is a vital regulator in the progression and prognosis of numerous tumors. In fact, miR-765 may have anti-oncogenic or oncogenic effects in different types of tumor cells. For example, one group reported a miR-765 upregulation in both hepatocellular carcinoma cell lines and tissues, which resulted in increased cell proliferation and tumorigenicity via the inhibition INPP4B. 30 Another group demonstrated significant upregulation of miR-765 in non-small cell lung cancer cell lines and tissues, which had a correlation with the TNM stages of patients. 31 In osteosarcoma (OS) tissues, miR-765 was discovered to be upregulated. 32 Moreover, miR-765 stimulated the invasion, migration, and proliferation of OS cells by targeting MTUS1. 32 On the contrary, miR-765 overexpression/upregulation has been observed to repress the proliferation and invasion of SK-BR-3 and oral squamous cancer cell lines through the regulation of EMP3.33,34 Another recent report suggested that miR-765 may operate as a tumor suppressor as its overexpression inhibited clear cell renal cell cancer development through the suppression of PLP2 expression. 35 We also observed the anti-oncogenic role of miR-765 in NPC, as revealed by its downregulation in the NPC tissues. Our experimental assays provided strong evidence that supported the interaction of LINC00173 with miR-765. Furthermore, the correlation analysis revealed that LINC00173 had an inverse association with miR-765. Taken together, our findings suggest that LINC00173 facilitates NPC cell growth by sponging miR-765.
As far as GREM1 is concerned, numerous reports suggest that it executes a crucial function in the initiation and progression of numerous cancer types. GREM1 is among the bone morphogenetic protein (BMP) antagonists, and it bind directly to the BMPs. 36 Furthermore, GREM1 has an involvement in diabetic nephropathy, organogenesis, organ fibrosis, and tissue differentiation. 36 Recently, a number of studies have reported GREM1’s oncogenic role in cancer. A high GREM1 expression has been observed in cervical and pancreatic cancer tissues and was found to promote the progression of these cancers.37,38 Similarly, in NSCLC tissues and cells, its levels are high, and knocking down GREM1 hampered the proliferation of NSCLC cells and increased the expression of BMP7 protein. 39 GREM1 has been reported to have an important role in breast cancer progression. GREM1 knockdown diminished the breast cancer cell proliferation while its overexpression boosted their growth, invasion, and migration. 40 Several other studies have reported the upregulation of GREM1 and its association with metastasis in ER-negative breast cancer cells.41,42 Consistent with the other cancer types, we also observed elevated levels of GREM1 in NPC cell lines and tissues. Our findings suggest that GREM1 is upregulated in NPC via the LINC00173/miR-765 axis. This could be through multiple pathways, including the BMP dependent and independent pathways like the Vascular Endothelial Growth Factor Receptor (VEGFR) signaling pathway. Our findings are indicative of GREM1’s promising potential as a therapeutic target and prognostic biomarker for NPC. Still, GREM1’s exact underlying mechanisms as well as its interactions with other cellular components in NPC progression need further exploration.
The present work and its scope certainly have left some questions which will hopefully be addressed in future research. More experiments with in vivo assays are necessary to supplement more robust evidence for our results. Understanding of the detailed molecular mechanisms of GREM1 in the modulation of NPC progression would require further studies. Studying the downstream effectors or signaling pathways linked with the LINC00173/miR-765/GREM1 axis should also be very interesting and may give novel leads.
Conclusion
Our findings demonstrated that LINC00173 executes an important oncogenic role in NPC by modulating the miR-765/GREM1 axis. Hence, a new LINC00173/miR-765/GREM1 pathway, which appears to regulate tumorigenesis and tumor progression, has been identified NPC. In addition, LINC00173 has shown its potential as a therapeutic target and prognostic biomarker for NPC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
