Abstract
Background
Colorectal carcinoma (CRC) ranks the third most frequent malignancy worldwide. Makorin RING zinc finger-2 (MKRN2) has been identified as a tumor suppressor in CRC, and the bioinformatics prediction indicated that some non-coding RNAs (ncRNAs) that directly or indirectly regulate MKRN2 might play critical roles in CRC progression. This study aimed to analyze the regulatory effect of LINC00294 on CRC progression, and to explore the underlying mechanisms by assessing miR-620 and MKRN2. The potential prognostic value of the ncRNAs and MKRN2 was also investigated.
Methods
The expression of LINC00294, MKRN2, miR-620 was examined by qRT-PCR. Cell counting kit-8 assay was used to assess the proliferation of CRC cells. Transwell assay was used to evaluate the migration, invasion of CRC cells. Kaplan-Meier method and log-rank test were used to perform comparative analysis of overall survival in CRC patients.
Results
Lower expression of LINC00294 was observed in both CRC tissues and cell lines. In CRC cells, LINC00294 overexpression inhibited cell proliferation, migration and invasion, but these effects were directly reversed by the overexpression of miR-620, which was demonstrated as a target of LINC00294. Additionally, MKRN2 was found to be a target gene of miR-620, and might mediate the regulatory function of LINC00294 in CRC progression. In CRC patients, low LINC00294, MKRN2 and high miR-620 expression was associated poor overall survival of CRC.
Conclusions
LINC00294/miR-620/MKRN2 axis had the potential to provide prognostic biomarkers for CRC patients, and negatively regulated the malignant progression of CRC cells, including proliferation, migration and invasion.
Keywords
Introduction
Colorectal carcinoma (CRC) is the third most common malignancy in adults, preceded by lung and breast cancer. 1 Its presentation is usually confused with other diseases, leading to significant delays in diagnosis. 2 However, it is the most common primary gastrointestinal malignancy, after liver tumors. 2 Thus, timely diagnosis and treatment can improve the survival prognosis in patients with CRC. Due to its slow progression from detectable precancerous lesions and to better prognosis of CRC patients diagnosed at early stages, thus the potential for reducing the disease burden in patients with CRC through early detection is significant. 3 Although screening significantly reduced the risk of CRC associated mortality,4,5 the achievement of its effectiveness still required assistance from other biological diagnostic factors. 3
Long noncoding RNA LINC00294, as a functional long noncoding RNA (lncRNA), functioned to regulate lipopolysaccharide (LPS)-induced human umbilical vein endothelial cells (HUVECs) inflammation injury. 6 Moreover, LINC00294 has been reported to be involved in the tumor progression process of glioma and cervical cancer as a functional lncRNA,7,8 implying the potential of LINC00294 as a tumor regulator. In terms of the bioinformatics analysis results of LINC00294, we found that the expression of LINC00294 had a significantly high expression level in colon normal tissues according to NONCODE online database (http://www.noncode.org/), while the GEPIA (http://gepia.cancer-pku.cn/) analysis platform found that LINC00294 expression was significantly lower in CRC tumor tissues compared with normal control tissues. Therefore, it was reasonable to speculate that LINC00294 might have a potential association with the occurrence and development of CRC.
Makorin RING zinc finger-2 (MKRN2) was reported to be a member of the Makorin RING zinc finger family, which encoded multiple C3H1 zinc finger domains, a characteristic Makorin zinc finger and an interspersed RING domain. 9 Several physiological functions for MKRN2 have recently been specified. For instance, a previous study reported that MKRN2 was a novel ubiquitin E3 ligase targeting the p65 subunit of NF-κB to negatively regulate inflammatory responses. 10 In addition, MKRN2 has been implicated as a negative regulator of diverse cellular and physiological pathways, including tumor cells.11–13 In the field of malignancy, a study from Jiang et al. pointed out that MKRN2 inhibited migration and invasion of non-small-cell lung cancer. 12 Subsequent study by Chang et al. found that MKRN2 was to be potentially associated with CRC tumor progression. 14 In consideration of both LINC00294 and MKRN2 were associated with CRC progression, this study was intended to analyze the relationship between LINC00294 and MKRN2 and the roles of both in CRC tumor progression.
miRNA-620 (miR-620) was disclosed to be aberrantly expressed and exert regulatory functions in the progression of some cancers. For instance, the expression of miR-620 was lower in glioma tissues, and the inhibition of miR-620 showed opposite effects on glioma cell cycle arrest and apoptosis. 15 miR-620 was found to be up-regulated in osteosarcoma cells. 16 In this study, on account of the results of bioinformatics prediction using starBase 3.0 (https://starbase.sysu.edu.cn/), 17 we noted that LINC00294 and MKRN2 sequences had complementary sequences to miR-620 sequence, speculated that miR-620 might be have association with CRC progression. We have known that miR-620 could be aberrantly expressed and exert regulatory functions in the progression of some cancers, but the expression level and function in CRC were unknown.
Therefore, in this study, the regulatory effect of LINC00294 on cell proliferation, migration and invasion was analyzed in CRC cells. The relationship between miR-620, MKRN2 and LINC00294 in CRC cells, as well as the mechanism by which both participated in the function exertion of LINC00294, were investigated. Moreover, this study further validated the relationship between LINC00294, miR-620 and MKRN2 in CRC tissues, and analyzed its clinical significance in predicting the survival prognosis of CRC. We expect that the findings will further enrich the pathological mechanism of CRC and provide novel biomarkers and therapeutic targets for the treatment of CRC.
Materials and methods
Cell culture and transfection
A human colonic epithelial cell line NCM460, two CRC cell lines SW480 and SW620, were used to perform relevant experimental analyses in this study. All cell lines used for experimental analyses were purchased from Shanghai Cell Bank (Shanghai, China). All cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, NY, USA) supplemented with 10% fetal bovine serum (FBS, Invitrogen), 100 U/mL penicillin and 100 μg/mL streptomycin. All cells were maintained at 37°C with 5% CO2 in a saturated humidified atmosphere.
SW480 and SW620 cells were seeded in 6-well plates for cell transfection. pcDNA3.1-LINC00294 was used to obtain overexpression LINC00294 CRC cells, and pcDNA3.1 was transfected simultaneously as controls. In addition, miR-620 mimic (AUGGAGAUAGAUAUAGAAAU), miR-620 inhibitor (AUUUCUAUAUCUAUCUCCAU), mimic negative controls (NC) (UUCUCCGAACGUGUCACGU), inhibitor NC (CAGUACUUUUGUGUAGUACAA) were transfected into SW480 and SW620 cell lines to regulate the miR-620 expression in CRC cells. All the transfection was achieved using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocols. After 6 h of incubation at 37°C, the transfection reagent was removed, and fresh DMEM was added for further incubation.
CRC tissue collection
According to the pathological diagnostic criteria for CRC patients,4,18 77 CRC patients between 2018 and 2021 at Qinghai Provincial People’s Hospital were included in this study. Tumor tissues and adjacent normal control mucosal tissues (more than 5 cm from the edge of the tumor tissues) were collected from 77 CRC patients, all collected tissue samples were verified via histopathology and frozen in liquid nitrogen for further use. A signed informed consent was obtained from the patients before sampling. This study did not include patients that had received radiotherapy and immunotherapy prior to or following surgical treatment. Furthermore, follow-up surveys were conducted by telephone communication or outpatient visits with a follow-up, and survival information of all patients were recorded at fixed times per month for further survival analysis. All protocols were all in accordance with the guideline of the Ethics Committee and has approved by the Ethics Committee of Qinghai Provincial People’s Hospital (0017365).
Reverse transcription quantitative PCR (RT-qPCR)
Primer sequences for RT-qPCR.

RT-qPCR, reverse transcription quantitative PCR; LINC00294, long noncoding RNA LINC00294; MKRN2, Makorin RING zinc finger-2; miR-620, miRNA-620; GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Cell counting kit (CCK)-8 assay
Cell counting kit (CCK)-8 assay was used to assess the proliferation abilities of SW480 and SW620 cells. Transfected SW480 and SW620 cells were seeded at the density of 3 × 103 cells/well in 96-well plates and incubated for 0, 24, 48, 72 h. Then, CCK-8 reagents were added to each well and incubating at 37°C with 5% CO2. The absorbance at 450 nm was measured and recorded, and the change of optical density (OD) value could reflect the cell proliferation abilities.
Transwell assay
Transwell assay was used to evaluate the migration and invasion abilities of SW480 and SW620 cells. Transwell chambers (Corning, USA) with Matrigel precoating and without Matrigel coating were used to analyze the migration and invasion abilities of the SW480 and SW620 cells. CRC cells with serum-free culture medium were seeded into the upper chambers with the density of 3 × 104 cells/well, and the lower chambers included medium contained 10% FBS. After 48 h incubation at 37°C, the CRC cells in the lower chambers were stained with 0.2% crystal violet for 10 min and inverted microscope (Olympus Corporation, Tokyo, Japan) was used to count the cell number. All experiments were repeated at least three times.
Dual-luciferase reporter assay
Based on the results predicted using starBase 3.0 (https://starbase.sysu.edu.cn/), 17 it was known that the base sequence of LINC00294 and MKRN2 had complementary sequences with miR-620, respectively. Wild type (Wt) sequences containing the binding sequences of miR-620and mutant type (Mut) sequences were synthesized, including: LINC00294-Wt, LINC00294-Mut, MKRN2-Wt, MKRN2-Mut. Subsequent experiments were also conducted in 2 main sections.
In part, LINC00294-Wt and LINC00294-Mut sequence were cloned into the pmirGLO vector (Promega, Madison, WI, USA). LINC00294-Wt or LINC00294-Mut with miR-620 mimic and mimic NC respectively, co-transfected into CRC cell lines using Lipofectamine 3000 (Invitrogen, CA, USA). Post-transfected 48h, relative luciferase activity was detected by dual-luciferase reporter assay (Promega, WI, USA) following the instructions of manufacturer. pcDNA3.1-LINC00294 and pcDNA3.1, miR-620 mimic and mimic NC, LINC00294 + miR-620 mimic transfected into CRC cell lines using Lipofectamine 3000 (Invitrogen, CA, USA), respectively. The relationship between LINC00294 and miR-620 and the effect on CRC progression were assessed by examining the miR-620 expression levels, cell proliferation, migration and invasion abilities of CRC cells.
On the other part, MKRN2-Wt, MKRN2-Mut sequence were cloned into the pmirGLO vector (Promega, Madison, WI, USA). MKRN2-Wt, MKRN2-Mut with miR-620 mimic and mimic NC respectively, co-transfected into CRC cell lines using Lipofectamine 3000 (Invitrogen, CA, USA). Post-transfected 48h, relative luciferase activity was detected by dual-luciferase reporter assay (Promega, WI, USA) following the instructions of manufacturer. pcDNA3.1-LINC00294 and pcDNA3.1, miR-620 mimic and mimic NC, LINC00294 + miR-620 mimic transfected into CRC cell lines using Lipofectamine 3000 (Invitrogen, CA, USA), respectively. The relationship between LINC00294, MKRN2 and miR-620 were assessed by detecting the MKRN2 expression levels of CRC cells.
Statistical analysis
All data was analyzed using SPSS 21.0 (SPSS, Inc, Chicago, Illinois) and GraphPad 7.0 (GraphPad Software, Inc, USA). Student’s t test was used to compare the differences between two groups. One-way ANOVA was used for comparing the differences between multiple groups. Pearson correlation coefficient was used to evaluate the correlation between LINC00294, MKRN2 and miR-620 expression in CRC. Kaplan-Meier method and log-rank test were used to perform comparative analysis of prognostic survival in CRC patients. p < 0.05 was considered to indicate statistically significant difference.
Results
LINC00294 expression was reduced in CRC cell lines
The LINC00294 expression levels in CRC cells and normal cells were compared by culturing two CRC cell lines SW480 and SW620 and a human colonic epithelial cell line NCM460. The analysis results revealed that the expression levels of LINC00294 were markedly decreased in two CRC cell lines SW480 and SW620 compared with NCM460 cell line, and SW480 cell line had the lowest LINC00294 expression levels (Figure 1, all p < 0.001). Expression of LINC00294 in CRC cell lines. (***p < 0.001 vs. NCM460).
LINC00294 overexpression inhibited CRC cell proliferation, migration and invasion
Through cell transfection using Lipofectamine 3000, pcDNA3.1-LINC00294 was transfected into SW480 and SW620 cell lines, and pcDNA3.1 was transfected simultaneously as controls. The transfection results displayed that pcDNA3.1-LINC00294 significantly increased the expression level of LINC00294 in SW480 and SW620 cell lines compared with pcDNA3.1 (Figure 2(a), all p < 0.001), with the effect of overexpressing LINC00294. Effects of LINC00294 on the proliferation, migration and invasion of SW480 and SW620 cells. (a) LINC00294 expression was promoted by pcDNA3.1-LINC00294 in CRC cells. (b) and (c) The overexpression of LINC00294 inhibited cell proliferation in both SW480 and SW620 cells. (d) CRC cell migration was suppressed by LINC00294 overexpression. (e) LINC00294 upregulation inhibited CRC invasion. (*p < 0.05, **p < 0.01, ***p < 0.001 vs. pcDNA3.1 group).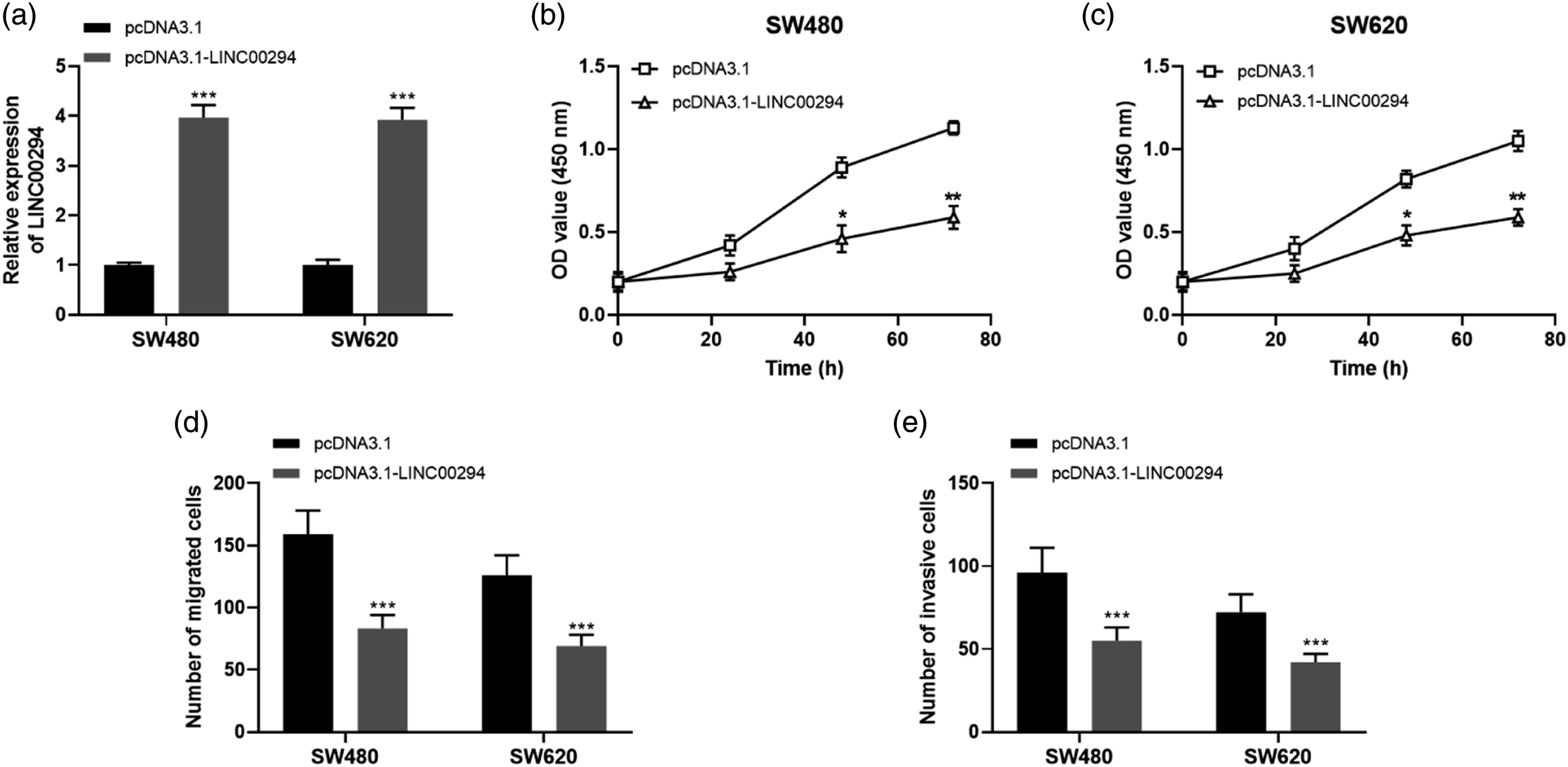
Subsequent cell proliferation experiments revealed that overexpressed LINC00294 inhibited the proliferation abilities of SW480 and SW620 cells, and the inhibitory effect were enhanced over time (Figure 2(b) and (c), 48h: all p < 0.05, 72h: all p < 0.01). Subsequent migration and invasion assays showed that overexpression of LINC00294 both significantly inhibited the migration abilities (Figure 2(d), all p < 0.001) and invasion abilities (Figure 2(e), all p < 0.001) of SW480 and SW620 cells.
LINC00294 suppressed CRC progression by sequestering miR-620
According to the results predicted using starBase 3.0, the complementary sequences of the base sequences of LINC00294 and miR-620 were demonstrated in Figure 3(a). Dual-luciferase reporter assay results demonstrated that relative luciferase activities were suppressed by miR-620 mimic compared with mimic NC in CRC cells transfected with LINC00294-Wt (Figure 3(b), p < 0.01). Whereas no significant changes were observed in CRC cells transfected with LINC00294-Mut (Figure 3(b)), implying that LINC00294 indeed was complementary targeting with miR-620. miR-620 mediated the regulation of LINC00294 in CRC cell biological behaviors. (a) The complementary sequences of the base sequences of LINC00294 and miR-620. (b) Results of dual-luciferase reporter assay. (c) Expression of miR-620 in CRC cells with dysregulation of LINC00294. (d) The overexpression of miR-620 promoted CRC cell proliferation, and abolished the inhibitory effects of LINC00294 overexpression. (e) and (f). miR-620 overexpression led to enhanced CRC cell migration and invasion abilities, and reversed the regulatory effects of LINC00294 on cell migration and invasion. (For Figure 3(b): **p < 0.01 vs. mimic NC; For Figure 3(c)–(f): *p < 0.05, **p < 0.01, ***p < 0.001 vs. pcDNA3.1; #p < 0.05, ##p < 0.01, ###p < 0.001 vs. mimic NC; △p < 0.05, △△p < 0.01, △△△p < 0.001 vs. pcDNA3.1-LINC00294).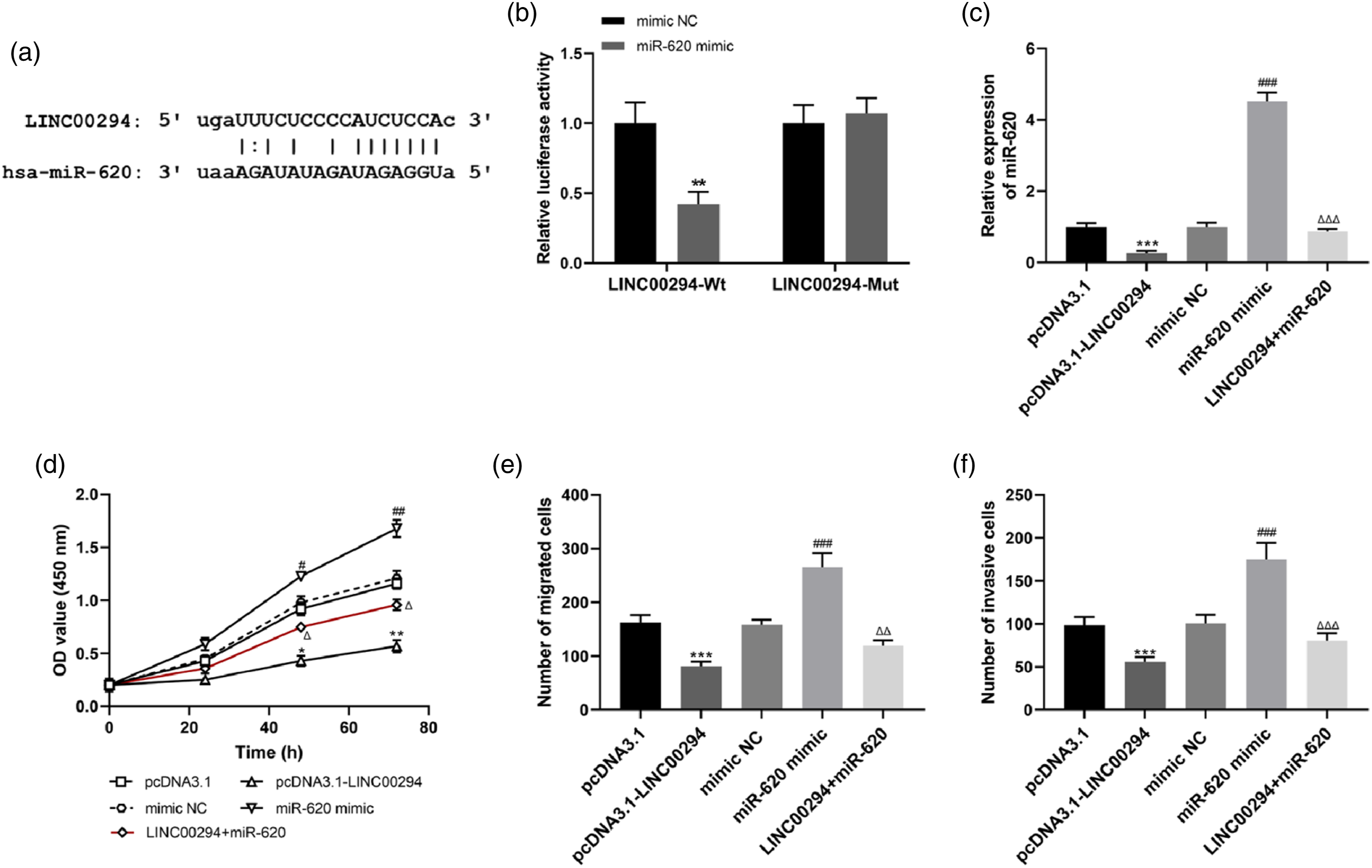
In addition, the relationship between relative expression of LINC00294 and miR-620 in CRC cells was analyzed in this study. The analysis found that overexpression of LINC00294 significantly inhibited the relative expression of miR-620 compared with pcDNA3.1 (Figure 3(c), p < 0.001), miR-620 mimic significantly promoted the relative expression of miR-620 compared mimic NC (Figure 3(c), p < 0.001), whereas the overexpression of miR-620 could be significantly suppressed by the effects of LINC00294 by sequestering miR-620 compared with miR-620 mimic, and to restore it to approximately normal levels (Figure 3(c), p < 0.001). The results of further cell proliferation experiments exhibited that overexpression of LINC00294 inhibited CRC cell proliferation (Figure 3(d), 48h: p < 0.05, 72 h: p < 0.01) and miR-620 mimic promoted CRC cell proliferation (Figure 3(d), 48 h: p < 0.05, 72 h: p < 0.01), and the effects of LINC00294 by sequestering miR-620 could alleviate the promotion of CRC cell proliferation by overexpression of miR-620 (Figure 3(d), 48 h: p < 0.05, 72 h: p < 0.05). The results of Transwell assays indicated that overexpression of LINC00294 markedly inhibited CRC cells migration and invasion (Figure 3(e) and (f), all p < 0.001), miR-620 mimic markedly promoted CRC cells migration and invasion (Figure 3(e) and (f), all p < 0.001), and the effects of LINC00294 by sequestering miR-620 alleviated the promotion of CRC cell migration (Figure 3(e), p < 0.01) and invasion (Figure 3(f), p < 0.001) by miR-620 mimic.
MKRN2 expression was enhanced by LINC00294 through miR-620
Based on the results predicted using starBase 3.0, the complementary sequences of the base sequences of MKRN2 and miR-620 were demonstrated in Figure 4(a). Dual-luciferase reporter assay results revealed that relative luciferase activities were suppressed by miR-620 mimic compared with mimic NC in CRC cells transfected with MKRN2-Wt (Figure 4(b), p < 0.001). There were no significant changes revealed in CRC cells transfected with MKRN2-Mut (Figure 4(b)), suggesting that MKRN2 was complementary targeting with miR-620. MKRN2 served as a target of miR-620. (a) The complementary sequences of the base sequences of MKRN2 and miR-620. (***p < 0.001 vs. mimic NC) (b) Results of dual-luciferase reporter assay. (c) MKRN2 expression was enhanced by LINC00294 overexpression, but was inhibited by miR-620 overexpression. (***p < 0.001 vs. pcDNA3.1; ###p < 0.001 vs. mimic NC; △△△p < 0.001 vs. pcDNA3.1-LINC00294).
The further analysis was observed that overexpression of LINC00294 significantly promoted the relative expression of MKRN2 compared with pcDNA3.1 (Figure 4(c), p < 0.001), miR-620 mimic significantly decreased the relative expression of MKRN2 compared mimic NC (Figure4(c), p < 0.001), the inhibitory effect of miR-620 mimic on MKRN2 expression was significantly reversed by LINC00294 through miR-620 (Figure 4(c), p < 0.001).
Expression of the LINC00294/miR-620/MKRN2 axis in the tissues of CRC patients
The expression levels of LINC00294, MKRN2 and miR-620 were examined by RT-qPCR. The results of RT-qPCR assay indicated that the expression of LINC00294 and MKRN2 were remarkable downregulated in CRC tissues compared with normal tissues (Figure 5(a) and (c), all p < 0.001). The expression level of miR-620 was significantly higher in CRC tissues compared with normal tissues (Figure 5(b), p < 0.001). Expression of the LINC00294/miR-620/MKRN2 axis in the tissues of CRC patients. (a)–(c) The expression of LINC00294 and MKRN2 was downregulated, but miR-620 expression was upregulated in the tissues of CRC patients. (***p < 0.001 vs. normal tissues). (d)–(f) LINC00294 was negatively correlated with miR-620, but positively correlated with MKRN2 in CRC. A negative correlation was observed between miR-620 and MKRN2.
Subsequent correlation analysis revealed significant negative correlation between relative LINC00294 expression and relative miR-620 expression (Figure 5(d), r = −0.642, p < 0.001). Correlation analysis shown a marked positive correlation between relative LINC00294 expression and relative MKRN2 expression (Figure 5(e), r = 0.471, p < 0.001). There was also significant negative correlation between relative miR-620 expression and relative MKRN2 expression (Figure 5(f), r = −0.684, p < 0.001).
Association of the LINC00294/miR-620/MKRN2 axis with CRC survival
The association of LINC00294/miR-620/MKRN2 axis with the prognostic survival of CRC patients was analyzed using the Kaplan Meier method and log-rank test. Total of the 77 patients participated the follow-up, and 5 (6.50%) patients lost contact during the survey (censored cases). The data of expression levels were presented as mean ± standard deviation (SD), and divided into low LINC00294 expression group (n = 39) and high LINC00294 expression group (n = 38), low miR-620 expression group (n = 38) and high miR-620 expression group (n = 39), low MKRN2 expression group (n = 39) and high MKRN2 expression group (n = 38), respectively. Survival analysis results indicated that low LINC00294 expression was associated with worse survival prognosis in CRC patients (Figure 6(a), Log-rank p = 0.011), high miR-620 expression was associated with worse overall survival in CRC patients (Figure 6(b), Log-rank p = 0.002), low MKRN2 expression was associated with poor survival prognosis in CRC patients (Figure 6(c), Log-rank p = 0.022). Association of the LINC00294/miR-620/MKRN2 axis with CRC survival. (a) Low LINC00294 expression was associated with worse survival prognosis in CRC patients (log-rank p = 0.011). (b) High miR-620 expression was associated with worse overall survival in CRC patients (log-rank p = 0.002). (c) Low MKRN2 expression was associated with poor survival prognosis in CRC patients (log-rank p = 0.022).
Discussion
Multiple studies have reported that diversified biological factors has directly or indirectly participated in CRC progression and aberrantly expressed in CRC.19-21 For instance, in the article by Ni et al. to explore the function pathways of CRC mentioned lncRNA and believed that long noncoding RNA GAS5 (GAS5) inhibited progression of CRC and overexpression of GAS5 significantly impaired tumor growth with a concomitant decline in cell viability. 19 LncRNA GLCC1 was revealed to be upregulated in CRC expression and promote CRC carcinogenesis by stabilizing c-Myc. 20 In addition, in a review reporting the relationship between CRC and miRNAs, investigators summarized that miR-17 promoted the malignant progression of CRC by downregulating SIK1 expression.21,22 In this study, we aimed at exploring the functional role of LINC00294 in CRC progression, as well as its related mechanisms. In addition, the expression of LINC00294 and its target was investigated in CRC tissues with the purpose to evaluate their clinical value in the prognosis of CRC. According to RT-qPCR analysis, it is demonstrated that the expression levels of LINC00294 were significantly reduced in CRC tissues and cells. Moreover, the expression of MKRN2 was remarkable downregulated in CRC tissues compared with normal tissues, the expression level of miR-620 was significantly higher in CRC tissues compared with normal tissues. Generally, the results of the aberrant expression of LINC00294, miR-620 and MKRN2 in CRC suggested that they might be involved in CRC progression.
The former mentioned that LINC00294 was involved in the process of tumor progression in glioma and cervical cancer and has the potential to be a tumor regulator,7,8 and the specific regulatory role of LINC00294 in CRC tumor progression required further analytical studies. Regarding the effects of LINC00294 on apoptosis, Dong et al. have proposed that silencing LINC00294 could inhibit the apoptosis of glioma cells under hypoxia via the miR-21-5p/CASKIN1/cAMP axis. 23 In the present study, we explored the effects of LINC00294 on cell proliferation, migration and invasion. pcDNA3.1-LINC00294 was transfected into CRC cells to obtain LINC00294 overexpressed CRC cells. Subsequent CCK-8 assay and Transwell assay demonstrated that LINC00294 overexpression inhibited CRC cell proliferation, migration and invasion, implying that LINC00294 overexpression suppressed the malignant progression of CRC.
With deeper exploration in the molecular field, lncRNA-mediated competing endogenous RNA networks including lncRNA, miRNA and mRNA, which continuously regulated CRC cell proliferation, invasion, tumor growth and metastasis. 24 Hence, the regulatory mechanisms underlying the development and progression of CRC need to be deeper explored and understood. The study conducted by Yu et al. revealed that lncRNA PVT1 promoted CRC tumorigenesis via the PVT1/miR-30d-5p/RUNX2 axis). 25 Similarly, previous investigation summarized that lncRNA H19 promoted CRC cell migration and invasion, as well as CRC tumor growth and liver metastasis, by competitively sponging miR-138 to derepress the oncoprotein HMGA1.24,26 Silencing LINC00294 has also been revealed to regulate apoptosis of glioma cells via the miR-21-5p/CASKIN1/cAMP Axis. 23 According to the results predicted using starBase 3.0, the complementary sequences of the base sequences of LINC00294 and miR-620 was obtained, dual-luciferase reporter assay results implied that LINC00294 indeed was complementary targeting with miR-620. Subsequent correlation analysis revealed significant negative correlation between relative LINC00294 expression and relative miR-620 expression. Next cell proliferation, migration and invasion assays suggested that LINC00294 might be suppress the malignant progression of CRC by sequestering miR-620. In addition, bioinformatics analysis results also predicted complementary sequences between MKRN2 and miR-620. MKRN2 functioned as a tumor suppressor11,12 and the relationship with LINC00294 and miR-620 in CRC needed to be further explored. Dual-luciferase reporter assay results suggested that MKRN2 was complementary targeting with miR-620. Correlation analysis shown that LINC00294 expression was positively correlated with MKRN2 expression, and miR-620 expression was negatively correlated with MKRN2 expression. Collectively, the above analysis results indicated that MKRN2 expression might be enhance by LINC00294 through inhibiting miR-620.
CRC diagnosis and treatment have been greatly improved in recent years, however, distant metastasis, especially liver metastasis, and the stem cell properties of cancer cells contribute to the poor prognosis and high mortality rate of CRC patients.24,27 The research of CRC prognostic biomarkers has been on ongoing. There were also lots of excellent previous works that have been recognized, for example, a research pointed out that lncRNA LINRIS was an independent prognostic biomarker for CRC, meanwhile the LINRIS-IGF2BP2-MYC axis promoted the progression of CRC and might be a promising therapeutic target. 28 Besides, lncRNA GLCC was also noted to be associated with poor prognosis in CRC. 20 The association of LINC00294/miR-620/MKRN2 axis with the prognostic survival of CRC patients was analyzed using the Kaplan Meier method and log-rank test in this study. Survival analysis results indicated that low LINC00294 expression, high miR-620 expression, low MKRN2 expression were associated with worse survival in CRC patients. Therefore, we speculate that LINC00294/miR-620/MKRN2 axis might serve as novel biomarker of poor survival in CRC patients and might provide a novel target for CRC treatment.
Conclusion
Collectively, the expression levels of LINC00294 was reduced in CRC cell lines and tissues, and LINC00294 overexpression inhibited CRC cell proliferation, migration and invasion. Moreover, combined with bioinformatics analysis we found that LINC00294 suppressed CRC progression by sequestering miR-620, MKRN2 expression was enhanced by LINC00294 through miR-620. Furthermore, survival analysis results indicated that low LINC00294, MKRN2 expression and high miR-620 expression were associated with worse survival in CRC patients, indicating that LINC00294/miR-620/MKRN2 axis might serve as novel prognostic biomarkers of poor survival in CRC patients, and might provide novel targets for CRC treatment.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
The experimental procedures were all in accordance with the guideline of the Ethics Committee of Qinghai Provincial People’s Hospital and has approved by the Ethics Committee of Qinghai Provincial People’s Hospital. This study complies with the Declaration of Helsinki.A signed written informed consent was obtained from each patient.
Availability of data and materials
The data used and analyzed can be obtained from the corresponding author under a reasonable request.
